Project
GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
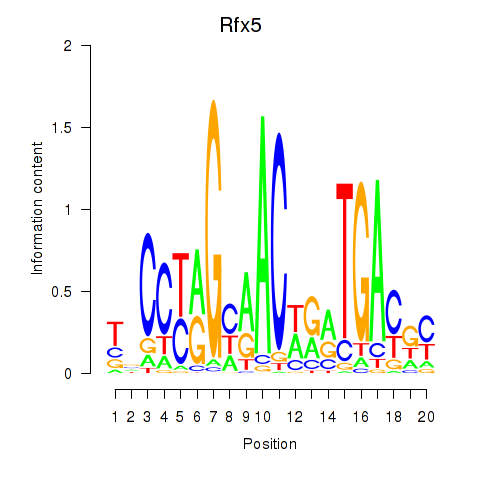
Results for Rfx5
Z-value: 0.78
Transcription factors associated with Rfx5
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Rfx5
|
ENSMUSG00000005774.13 | regulatory factor X, 5 (influences HLA class II expression) |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Rfx5 | mm39_v1_chr3_+_94861386_94861537 | 0.24 | 1.7e-01 | Click! |
Activity profile of Rfx5 motif
Sorted Z-values of Rfx5 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_-_28453374 | 6.89 |
ENSMUST00000028161.6
|
Cel
|
carboxyl ester lipase |
| chr17_+_34524841 | 4.17 |
ENSMUST00000235530.2
|
H2-Eb1
|
histocompatibility 2, class II antigen E beta |
| chr18_+_60936910 | 4.15 |
ENSMUST00000097563.9
ENSMUST00000050487.16 ENSMUST00000167610.2 |
Cd74
|
CD74 antigen (invariant polypeptide of major histocompatibility complex, class II antigen-associated) |
| chr17_+_34524884 | 3.98 |
ENSMUST00000074557.11
|
H2-Eb1
|
histocompatibility 2, class II antigen E beta |
| chr17_+_34482183 | 3.65 |
ENSMUST00000040828.7
ENSMUST00000237342.2 ENSMUST00000237866.2 |
H2-Ab1
|
histocompatibility 2, class II antigen A, beta 1 |
| chr5_-_145816774 | 3.58 |
ENSMUST00000035918.8
|
Cyp3a11
|
cytochrome P450, family 3, subfamily a, polypeptide 11 |
| chr1_+_88139678 | 3.51 |
ENSMUST00000073049.7
|
Ugt1a1
|
UDP glucuronosyltransferase 1 family, polypeptide A1 |
| chr17_+_34416707 | 2.99 |
ENSMUST00000025196.9
|
Psmb8
|
proteasome (prosome, macropain) subunit, beta type 8 (large multifunctional peptidase 7) |
| chr17_+_34372046 | 2.80 |
ENSMUST00000114232.4
|
H2-DMb1
|
histocompatibility 2, class II, locus Mb1 |
| chr17_+_34406523 | 2.71 |
ENSMUST00000170086.8
|
Tap1
|
transporter 1, ATP-binding cassette, sub-family B (MDR/TAP) |
| chr17_+_34406762 | 2.70 |
ENSMUST00000041633.15
|
Tap1
|
transporter 1, ATP-binding cassette, sub-family B (MDR/TAP) |
| chr17_+_34416689 | 2.68 |
ENSMUST00000173441.9
|
Psmb8
|
proteasome (prosome, macropain) subunit, beta type 8 (large multifunctional peptidase 7) |
| chr17_-_34406193 | 2.22 |
ENSMUST00000173831.3
|
Psmb9
|
proteasome (prosome, macropain) subunit, beta type 9 (large multifunctional peptidase 2) |
| chr17_+_34354556 | 1.78 |
ENSMUST00000236322.2
ENSMUST00000237402.2 |
H2-DMa
|
histocompatibility 2, class II, locus DMa |
| chr17_+_35598583 | 1.70 |
ENSMUST00000081435.5
|
H2-Q4
|
histocompatibility 2, Q region locus 4 |
| chr17_-_34219225 | 1.45 |
ENSMUST00000238098.2
ENSMUST00000087189.7 ENSMUST00000173075.3 ENSMUST00000172912.8 ENSMUST00000236740.2 ENSMUST00000025181.18 |
H2-K1
|
histocompatibility 2, K1, K region |
| chr11_+_115054157 | 1.28 |
ENSMUST00000021077.4
|
Slc9a3r1
|
solute carrier family 9 (sodium/hydrogen exchanger), member 3 regulator 1 |
| chr17_+_37581103 | 1.27 |
ENSMUST00000038580.7
|
H2-M3
|
histocompatibility 2, M region locus 3 |
| chr9_+_44684285 | 1.19 |
ENSMUST00000239014.2
|
Ift46
|
intraflagellar transport 46 |
| chr5_-_125371162 | 1.16 |
ENSMUST00000127148.2
|
Scarb1
|
scavenger receptor class B, member 1 |
| chr9_+_44684206 | 1.10 |
ENSMUST00000002099.11
|
Ift46
|
intraflagellar transport 46 |
| chr17_+_34457868 | 1.09 |
ENSMUST00000095342.11
ENSMUST00000167280.8 ENSMUST00000236838.2 |
H2-Ob
|
histocompatibility 2, O region beta locus |
| chr9_+_44684248 | 1.01 |
ENSMUST00000128150.8
|
Ift46
|
intraflagellar transport 46 |
| chr9_+_44684450 | 1.00 |
ENSMUST00000238800.2
ENSMUST00000147559.8 |
Ift46
|
intraflagellar transport 46 |
| chr10_-_78143384 | 1.00 |
ENSMUST00000150828.2
ENSMUST00000239481.2 |
Agpat3
ENSMUSG00000118646.2
|
1-acylglycerol-3-phosphate O-acyltransferase 3 |
| chr17_+_35658131 | 0.99 |
ENSMUST00000071951.14
ENSMUST00000116598.10 ENSMUST00000078205.14 ENSMUST00000076256.8 |
H2-Q7
|
histocompatibility 2, Q region locus 7 |
| chr17_+_25105617 | 0.91 |
ENSMUST00000117890.8
ENSMUST00000168265.8 ENSMUST00000120943.8 ENSMUST00000068508.13 ENSMUST00000119829.8 |
Spsb3
|
splA/ryanodine receptor domain and SOCS box containing 3 |
| chr9_+_44684324 | 0.87 |
ENSMUST00000214854.3
ENSMUST00000125877.8 |
Ift46
|
intraflagellar transport 46 |
| chr17_+_34364206 | 0.72 |
ENSMUST00000041982.9
ENSMUST00000171231.8 |
H2-DMb2
|
histocompatibility 2, class II, locus Mb2 |
| chr2_-_92264374 | 0.68 |
ENSMUST00000111278.2
ENSMUST00000090559.12 |
Cry2
|
cryptochrome 2 (photolyase-like) |
| chr6_+_94477294 | 0.63 |
ENSMUST00000061118.11
|
Slc25a26
|
solute carrier family 25 (mitochondrial carrier, phosphate carrier), member 26 |
| chr7_+_44926925 | 0.61 |
ENSMUST00000210861.2
|
Slc6a21
|
solute carrier family 6 member 21 |
| chr7_+_101027390 | 0.59 |
ENSMUST00000084895.12
|
Arap1
|
ArfGAP with RhoGAP domain, ankyrin repeat and PH domain 1 |
| chr5_-_140346132 | 0.55 |
ENSMUST00000196130.5
|
Snx8
|
sorting nexin 8 |
| chr8_+_27532583 | 0.55 |
ENSMUST00000033875.10
ENSMUST00000209525.2 |
Plpbp
|
pyridoxal phosphate binding protein |
| chr8_+_27532623 | 0.47 |
ENSMUST00000209856.2
ENSMUST00000098851.12 ENSMUST00000211393.2 ENSMUST00000211518.2 |
Plpbp
|
pyridoxal phosphate binding protein |
| chr2_+_121978156 | 0.46 |
ENSMUST00000102476.5
|
B2m
|
beta-2 microglobulin |
| chr5_-_36987917 | 0.44 |
ENSMUST00000031002.10
|
Man2b2
|
mannosidase 2, alpha B2 |
| chr7_-_44741138 | 0.43 |
ENSMUST00000210527.2
|
Rcn3
|
reticulocalbin 3, EF-hand calcium binding domain |
| chr17_+_36172210 | 0.36 |
ENSMUST00000074259.15
ENSMUST00000174873.2 |
Nrm
|
nurim (nuclear envelope membrane protein) |
| chr5_-_134485081 | 0.30 |
ENSMUST00000111244.5
|
Gtf2ird1
|
general transcription factor II I repeat domain-containing 1 |
| chr3_-_95789505 | 0.28 |
ENSMUST00000159863.2
ENSMUST00000159739.8 ENSMUST00000036418.10 ENSMUST00000161866.8 |
Ciart
|
circadian associated repressor of transcription |
| chr3_-_95789338 | 0.27 |
ENSMUST00000161994.2
|
Ciart
|
circadian associated repressor of transcription |
| chr11_+_61395964 | 0.27 |
ENSMUST00000102657.10
|
B9d1
|
B9 protein domain 1 |
| chr7_-_45343785 | 0.17 |
ENSMUST00000040636.9
|
Sec1
|
secretory blood group 1 |
| chr7_-_101570382 | 0.15 |
ENSMUST00000098236.9
|
Lrrc51
|
leucine rich repeat containing 51 |
| chr3_-_57599956 | 0.13 |
ENSMUST00000238789.2
ENSMUST00000197088.5 ENSMUST00000099091.4 |
Ankub1
|
ankrin repeat and ubiquitin domain containing 1 |
| chr5_+_104318542 | 0.12 |
ENSMUST00000112771.2
|
Dspp
|
dentin sialophosphoprotein |
| chr10_+_90665270 | 0.11 |
ENSMUST00000182202.8
ENSMUST00000182966.8 |
Anks1b
|
ankyrin repeat and sterile alpha motif domain containing 1B |
| chr7_-_101570393 | 0.10 |
ENSMUST00000106965.8
ENSMUST00000106968.8 ENSMUST00000106967.8 |
Lrrc51
|
leucine rich repeat containing 51 |
| chr10_-_128579879 | 0.08 |
ENSMUST00000026414.9
|
Dgka
|
diacylglycerol kinase, alpha |
| chr7_+_100186259 | 0.07 |
ENSMUST00000054310.4
|
Coa4
|
cytochrome c oxidase assembly factor 4 |
| chr7_+_46490899 | 0.07 |
ENSMUST00000147535.8
|
Ldha
|
lactate dehydrogenase A |
| chr1_+_194302123 | 0.07 |
ENSMUST00000027952.12
|
Plxna2
|
plexin A2 |
| chr17_+_36172235 | 0.05 |
ENSMUST00000172931.2
|
Nrm
|
nurim (nuclear envelope membrane protein) |
| chr10_+_90665399 | 0.04 |
ENSMUST00000179694.9
|
Anks1b
|
ankyrin repeat and sterile alpha motif domain containing 1B |
| chr10_-_7539333 | 0.04 |
ENSMUST00000162606.8
|
Pcmt1
|
protein-L-isoaspartate (D-aspartate) O-methyltransferase 1 |
| chr8_-_105236506 | 0.03 |
ENSMUST00000064576.8
|
Terb1
|
telomere repeat binding bouquet formation protein 1 |
| chr14_+_111912529 | 0.02 |
ENSMUST00000042767.9
|
Slitrk5
|
SLIT and NTRK-like family, member 5 |
| chr1_+_133279647 | 0.02 |
ENSMUST00000112287.2
|
Ren1
|
renin 1 structural |
| chr4_+_137989783 | 0.01 |
ENSMUST00000105821.3
|
Kif17
|
kinesin family member 17 |
| chrX_+_111150171 | 0.00 |
ENSMUST00000164272.3
ENSMUST00000132037.2 |
4933403O08Rik
|
RIKEN cDNA 4933403O08 gene |
Network of associatons between targets according to the STRING database.
First level regulatory network of Rfx5
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.8 | 5.4 | GO:0046967 | cytosol to ER transport(GO:0046967) |
| 1.2 | 3.6 | GO:0002344 | peripheral B cell selection(GO:0002343) B cell affinity maturation(GO:0002344) |
| 1.2 | 3.5 | GO:0018879 | biphenyl metabolic process(GO:0018879) |
| 0.9 | 17.6 | GO:0019886 | antigen processing and presentation of exogenous peptide antigen via MHC class II(GO:0019886) |
| 0.9 | 1.7 | GO:0002481 | antigen processing and presentation of exogenous peptide antigen via MHC class Ib(GO:0002477) antigen processing and presentation of exogenous protein antigen via MHC class Ib, TAP-dependent(GO:0002481) |
| 0.5 | 1.1 | GO:0002581 | negative regulation of antigen processing and presentation of peptide or polysaccharide antigen via MHC class II(GO:0002581) |
| 0.5 | 8.1 | GO:0006707 | cholesterol catabolic process(GO:0006707) sterol catabolic process(GO:0016127) |
| 0.4 | 1.3 | GO:0032416 | negative regulation of sodium:proton antiporter activity(GO:0032416) |
| 0.4 | 1.4 | GO:0002485 | antigen processing and presentation of endogenous peptide antigen via MHC class I via ER pathway(GO:0002484) antigen processing and presentation of endogenous peptide antigen via MHC class I via ER pathway, TAP-dependent(GO:0002485) |
| 0.2 | 0.7 | GO:2000850 | negative regulation of corticosteroid hormone secretion(GO:2000847) negative regulation of glucocorticoid secretion(GO:2000850) |
| 0.2 | 4.9 | GO:0001539 | cilium or flagellum-dependent cell motility(GO:0001539) cilium-dependent cell motility(GO:0060285) |
| 0.2 | 0.6 | GO:0051182 | coenzyme transport(GO:0051182) |
| 0.1 | 10.0 | GO:0019882 | antigen processing and presentation(GO:0019882) |
| 0.1 | 0.3 | GO:0014886 | transition between slow and fast fiber(GO:0014886) |
| 0.0 | 0.4 | GO:0006013 | mannose metabolic process(GO:0006013) |
| 0.0 | 0.1 | GO:0071460 | cellular response to cell-matrix adhesion(GO:0071460) |
| 0.0 | 0.6 | GO:0034498 | early endosome to Golgi transport(GO:0034498) |
| 0.0 | 0.1 | GO:0021934 | hindbrain tangential cell migration(GO:0021934) |
| 0.0 | 0.6 | GO:0045475 | locomotor rhythm(GO:0045475) |
| 0.0 | 0.6 | GO:0001921 | positive regulation of receptor recycling(GO:0001921) |
| 0.0 | 0.1 | GO:1900383 | regulation of synaptic plasticity by receptor localization to synapse(GO:1900383) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.9 | 22.3 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 1.3 | 7.9 | GO:1990111 | spermatoproteasome complex(GO:1990111) |
| 0.7 | 5.4 | GO:0042825 | TAP complex(GO:0042825) |
| 0.4 | 5.8 | GO:0042611 | MHC protein complex(GO:0042611) |
| 0.2 | 3.5 | GO:0034663 | endoplasmic reticulum chaperone complex(GO:0034663) |
| 0.2 | 6.9 | GO:0042588 | zymogen granule(GO:0042588) |
| 0.1 | 5.2 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 0.1 | 2.4 | GO:0031528 | microvillus membrane(GO:0031528) |
| 0.0 | 0.4 | GO:0005652 | nuclear lamina(GO:0005652) |
| 0.0 | 0.3 | GO:0036038 | MKS complex(GO:0036038) |
| 0.0 | 0.6 | GO:0030904 | retromer complex(GO:0030904) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.3 | 6.9 | GO:0050253 | sterol esterase activity(GO:0004771) retinyl-palmitate esterase activity(GO:0050253) |
| 1.8 | 5.4 | GO:0015433 | peptide antigen-transporting ATPase activity(GO:0015433) tapasin binding(GO:0046980) |
| 0.7 | 4.2 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 0.4 | 3.6 | GO:0050649 | testosterone 6-beta-hydroxylase activity(GO:0050649) |
| 0.3 | 1.3 | GO:0045159 | myosin II binding(GO:0045159) |
| 0.3 | 7.9 | GO:0004298 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 0.3 | 1.2 | GO:0070506 | high-density lipoprotein particle receptor activity(GO:0070506) |
| 0.3 | 4.1 | GO:0046977 | TAP binding(GO:0046977) |
| 0.3 | 6.4 | GO:0023026 | MHC class II protein complex binding(GO:0023026) |
| 0.2 | 3.6 | GO:1990405 | protein antigen binding(GO:1990405) |
| 0.1 | 3.5 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.1 | 0.6 | GO:0051185 | coenzyme transporter activity(GO:0051185) |
| 0.1 | 0.2 | GO:0008107 | galactoside 2-alpha-L-fucosyltransferase activity(GO:0008107) alpha-(1,2)-fucosyltransferase activity(GO:0031127) |
| 0.1 | 0.7 | GO:0009881 | photoreceptor activity(GO:0009881) |
| 0.0 | 1.0 | GO:0003841 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) |
| 0.0 | 0.3 | GO:0008158 | hedgehog receptor activity(GO:0008158) |
| 0.0 | 0.7 | GO:0042605 | peptide antigen binding(GO:0042605) |
| 0.0 | 0.6 | GO:0031702 | type 1 angiotensin receptor binding(GO:0031702) |
| 0.0 | 0.4 | GO:0004559 | alpha-mannosidase activity(GO:0004559) |
| 0.0 | 1.0 | GO:0030170 | pyridoxal phosphate binding(GO:0030170) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.7 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.0 | 5.4 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
| 0.0 | 1.3 | PID TXA2PATHWAY | Thromboxane A2 receptor signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 3.5 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
| 0.3 | 5.9 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.1 | 7.8 | REACTOME CROSS PRESENTATION OF SOLUBLE EXOGENOUS ANTIGENS ENDOSOMES | Genes involved in Cross-presentation of soluble exogenous antigens (endosomes) |
| 0.1 | 1.2 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.0 | 1.0 | REACTOME SYNTHESIS OF PA | Genes involved in Synthesis of PA |
| 0.0 | 1.7 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.0 | 0.7 | REACTOME BMAL1 CLOCK NPAS2 ACTIVATES CIRCADIAN EXPRESSION | Genes involved in BMAL1:CLOCK/NPAS2 Activates Circadian Expression |