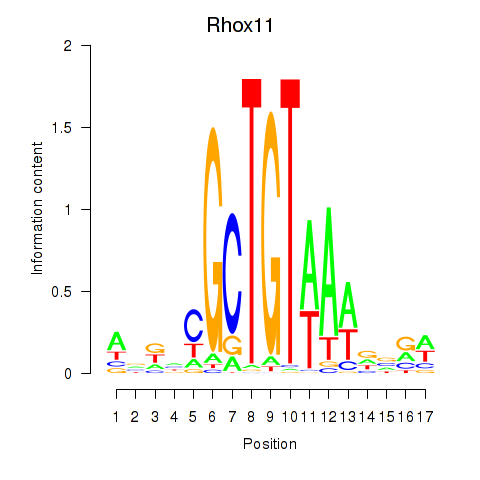
Project
GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
Results for Rhox11
Z-value: 0.47
Transcription factors associated with Rhox11
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Rhox11
|
ENSMUSG00000051038.11 | reproductive homeobox 11 |
Activity profile of Rhox11 motif
Sorted Z-values of Rhox11 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr17_+_41121979 | 2.91 |
ENSMUST00000024721.8
ENSMUST00000233740.2 |
Rhag
|
Rhesus blood group-associated A glycoprotein |
| chr9_+_110867807 | 2.85 |
ENSMUST00000197575.2
|
Ltf
|
lactotransferrin |
| chr11_-_101973288 | 1.41 |
ENSMUST00000100398.5
|
Mpp2
|
membrane protein, palmitoylated 2 (MAGUK p55 subfamily member 2) |
| chr11_+_70529944 | 1.26 |
ENSMUST00000055184.7
ENSMUST00000108551.3 |
Gp1ba
|
glycoprotein 1b, alpha polypeptide |
| chr18_-_60635059 | 1.18 |
ENSMUST00000042710.8
|
Smim3
|
small integral membrane protein 3 |
| chr18_-_42084249 | 1.10 |
ENSMUST00000070949.6
ENSMUST00000235606.2 |
Prelid2
|
PRELI domain containing 2 |
| chr19_-_39451509 | 1.04 |
ENSMUST00000035488.3
|
Cyp2c38
|
cytochrome P450, family 2, subfamily c, polypeptide 38 |
| chr5_+_121358254 | 0.99 |
ENSMUST00000042614.13
|
Hectd4
|
HECT domain E3 ubiquitin protein ligase 4 |
| chr8_-_106553822 | 0.98 |
ENSMUST00000239468.2
ENSMUST00000041400.6 |
Ranbp10
|
RAN binding protein 10 |
| chr9_-_107167046 | 0.89 |
ENSMUST00000035194.8
|
Mapkapk3
|
mitogen-activated protein kinase-activated protein kinase 3 |
| chr12_+_76884182 | 0.85 |
ENSMUST00000041008.10
|
Fntb
|
farnesyltransferase, CAAX box, beta |
| chr3_+_127584449 | 0.85 |
ENSMUST00000171621.3
|
Tifa
|
TRAF-interacting protein with forkhead-associated domain |
| chr1_+_153628598 | 0.83 |
ENSMUST00000182538.3
|
Rnasel
|
ribonuclease L (2', 5'-oligoisoadenylate synthetase-dependent) |
| chr4_+_126450728 | 0.76 |
ENSMUST00000048391.15
|
Clspn
|
claspin |
| chrX_-_101200670 | 0.69 |
ENSMUST00000056904.3
|
Ercc6l
|
excision repair cross-complementing rodent repair deficiency complementation group 6 like |
| chr1_-_169796709 | 0.67 |
ENSMUST00000027989.13
ENSMUST00000111353.4 |
Hsd17b7
|
hydroxysteroid (17-beta) dehydrogenase 7 |
| chr7_-_30364394 | 0.66 |
ENSMUST00000019697.9
|
Haus5
|
HAUS augmin-like complex, subunit 5 |
| chr3_-_122828592 | 0.65 |
ENSMUST00000029761.14
|
Myoz2
|
myozenin 2 |
| chr5_+_122239030 | 0.64 |
ENSMUST00000139213.8
ENSMUST00000111751.8 ENSMUST00000155612.8 |
Myl2
|
myosin, light polypeptide 2, regulatory, cardiac, slow |
| chr10_+_58230203 | 0.63 |
ENSMUST00000105468.2
|
Lims1
|
LIM and senescent cell antigen-like domains 1 |
| chr9_+_98178646 | 0.63 |
ENSMUST00000112938.8
ENSMUST00000112937.3 |
Nmnat3
|
nicotinamide nucleotide adenylyltransferase 3 |
| chr10_+_58230183 | 0.62 |
ENSMUST00000020077.11
|
Lims1
|
LIM and senescent cell antigen-like domains 1 |
| chr9_-_107166543 | 0.62 |
ENSMUST00000192054.2
|
Mapkapk3
|
mitogen-activated protein kinase-activated protein kinase 3 |
| chr11_-_97673203 | 0.60 |
ENSMUST00000128801.2
ENSMUST00000103146.5 |
Rpl23
|
ribosomal protein L23 |
| chr12_+_104067346 | 0.60 |
ENSMUST00000021495.4
|
Serpina5
|
serine (or cysteine) peptidase inhibitor, clade A, member 5 |
| chr9_+_98178608 | 0.58 |
ENSMUST00000112935.8
|
Nmnat3
|
nicotinamide nucleotide adenylyltransferase 3 |
| chr11_-_102771806 | 0.57 |
ENSMUST00000107060.8
|
Eftud2
|
elongation factor Tu GTP binding domain containing 2 |
| chr7_-_142253247 | 0.56 |
ENSMUST00000105934.8
|
Ins2
|
insulin II |
| chr14_-_21898992 | 0.54 |
ENSMUST00000124549.9
|
Comtd1
|
catechol-O-methyltransferase domain containing 1 |
| chr8_-_84184978 | 0.53 |
ENSMUST00000081506.11
|
Scoc
|
short coiled-coil protein |
| chr6_+_78382131 | 0.51 |
ENSMUST00000023906.4
|
Reg2
|
regenerating islet-derived 2 |
| chr5_+_43976218 | 0.50 |
ENSMUST00000101237.8
|
Bst1
|
bone marrow stromal cell antigen 1 |
| chr12_+_102521225 | 0.48 |
ENSMUST00000021610.7
|
Chga
|
chromogranin A |
| chrX_+_104123341 | 0.48 |
ENSMUST00000033577.11
|
Pbdc1
|
polysaccharide biosynthesis domain containing 1 |
| chr11_-_102771751 | 0.47 |
ENSMUST00000021306.14
|
Eftud2
|
elongation factor Tu GTP binding domain containing 2 |
| chr11_+_23234644 | 0.46 |
ENSMUST00000150750.3
|
Xpo1
|
exportin 1 |
| chr8_+_105558204 | 0.45 |
ENSMUST00000059449.7
|
Ces2b
|
carboxyesterase 2B |
| chr13_-_95661726 | 0.44 |
ENSMUST00000022185.10
|
F2rl1
|
coagulation factor II (thrombin) receptor-like 1 |
| chr9_-_78396407 | 0.42 |
ENSMUST00000154207.8
|
Eef1a1
|
eukaryotic translation elongation factor 1 alpha 1 |
| chr11_-_97591150 | 0.42 |
ENSMUST00000018681.14
|
Pcgf2
|
polycomb group ring finger 2 |
| chr8_+_14938022 | 0.40 |
ENSMUST00000123990.2
ENSMUST00000027554.8 |
Cln8
|
CLN8 transmembrane ER and ERGIC protein |
| chr10_-_59787646 | 0.37 |
ENSMUST00000020308.5
|
Ddit4
|
DNA-damage-inducible transcript 4 |
| chr9_+_50679528 | 0.33 |
ENSMUST00000042391.13
|
Fdxacb1
|
ferredoxin-fold anticodon binding domain containing 1 |
| chr19_-_8796288 | 0.33 |
ENSMUST00000153281.2
|
Ttc9c
|
tetratricopeptide repeat domain 9C |
| chr4_+_138926577 | 0.33 |
ENSMUST00000145368.8
|
Capzb
|
capping protein (actin filament) muscle Z-line, beta |
| chr12_-_111452242 | 0.32 |
ENSMUST00000010673.7
|
Gm266
|
predicted gene 266 |
| chr14_-_47059694 | 0.29 |
ENSMUST00000111817.8
ENSMUST00000079314.12 |
Gmfb
|
glia maturation factor, beta |
| chr11_-_69786324 | 0.29 |
ENSMUST00000001631.7
|
Acap1
|
ArfGAP with coiled-coil, ankyrin repeat and PH domains 1 |
| chr10_-_77845571 | 0.29 |
ENSMUST00000020522.9
|
Pfkl
|
phosphofructokinase, liver, B-type |
| chr7_-_119078472 | 0.27 |
ENSMUST00000209095.2
ENSMUST00000033263.6 ENSMUST00000207261.2 |
Umod
|
uromodulin |
| chr1_+_40844739 | 0.26 |
ENSMUST00000114765.4
|
Tmem182
|
transmembrane protein 182 |
| chr3_+_87950408 | 0.26 |
ENSMUST00000029707.13
ENSMUST00000166021.8 ENSMUST00000194258.6 ENSMUST00000193398.2 |
Gpatch4
|
G patch domain containing 4 |
| chr12_-_99849660 | 0.25 |
ENSMUST00000221929.2
ENSMUST00000046485.5 |
Efcab11
|
EF-hand calcium binding domain 11 |
| chr15_-_38079089 | 0.24 |
ENSMUST00000110336.4
|
Ubr5
|
ubiquitin protein ligase E3 component n-recognin 5 |
| chr14_-_20546848 | 0.22 |
ENSMUST00000022353.5
|
Mss51
|
MSS51 mitochondrial translational activator |
| chr11_+_98632631 | 0.22 |
ENSMUST00000064187.12
|
Thra
|
thyroid hormone receptor alpha |
| chr13_-_23746543 | 0.22 |
ENSMUST00000105107.2
|
H3c6
|
H3 clustered histone 6 |
| chr16_+_20470402 | 0.21 |
ENSMUST00000007212.9
ENSMUST00000232629.2 |
Psmd2
|
proteasome (prosome, macropain) 26S subunit, non-ATPase, 2 |
| chr7_+_126184108 | 0.20 |
ENSMUST00000039522.8
|
Apobr
|
apolipoprotein B receptor |
| chr4_+_126450762 | 0.19 |
ENSMUST00000147675.2
|
Clspn
|
claspin |
| chr7_-_119078330 | 0.18 |
ENSMUST00000207460.2
|
Umod
|
uromodulin |
| chr13_+_95461703 | 0.18 |
ENSMUST00000045909.8
|
Zbed3
|
zinc finger, BED type containing 3 |
| chr1_+_118409769 | 0.18 |
ENSMUST00000191823.6
|
Clasp1
|
CLIP associating protein 1 |
| chr8_-_81741495 | 0.18 |
ENSMUST00000042724.8
|
Usp38
|
ubiquitin specific peptidase 38 |
| chrX_+_104123367 | 0.17 |
ENSMUST00000119477.2
|
Pbdc1
|
polysaccharide biosynthesis domain containing 1 |
| chr15_+_91722524 | 0.17 |
ENSMUST00000109276.8
ENSMUST00000088555.10 ENSMUST00000100293.9 ENSMUST00000126508.8 ENSMUST00000239545.1 |
Smgc
ENSMUSG00000118670.1
|
submandibular gland protein C mucin 19 |
| chr3_-_92393193 | 0.17 |
ENSMUST00000054599.8
|
Sprr1a
|
small proline-rich protein 1A |
| chr17_-_46558894 | 0.15 |
ENSMUST00000142706.9
ENSMUST00000173349.8 ENSMUST00000087026.13 |
Polr1c
|
polymerase (RNA) I polypeptide C |
| chr10_-_127147609 | 0.15 |
ENSMUST00000037290.12
ENSMUST00000171564.8 |
Mars1
|
methionine-tRNA synthetase 1 |
| chr10_+_102376109 | 0.14 |
ENSMUST00000055355.6
|
Rassf9
|
Ras association (RalGDS/AF-6) domain family (N-terminal) member 9 |
| chr5_-_139115914 | 0.14 |
ENSMUST00000129851.8
|
Prkar1b
|
protein kinase, cAMP dependent regulatory, type I beta |
| chr5_+_93241385 | 0.14 |
ENSMUST00000201421.4
ENSMUST00000202415.4 ENSMUST00000202217.3 |
Septin11
|
septin 11 |
| chr4_+_86493905 | 0.14 |
ENSMUST00000091064.8
|
Rraga
|
Ras-related GTP binding A |
| chr1_+_180769890 | 0.14 |
ENSMUST00000161847.8
|
Tmem63a
|
transmembrane protein 63a |
| chr9_-_20432562 | 0.14 |
ENSMUST00000215908.2
ENSMUST00000068296.8 ENSMUST00000174462.8 ENSMUST00000213418.2 |
Zfp266
|
zinc finger protein 266 |
| chr12_-_114398864 | 0.12 |
ENSMUST00000103489.2
|
Ighv6-6
|
immunoglobulin heavy variable 6-6 |
| chr18_+_63110924 | 0.12 |
ENSMUST00000150267.2
ENSMUST00000236925.2 |
Napg
|
N-ethylmaleimide sensitive fusion protein attachment protein gamma |
| chr6_+_139564196 | 0.12 |
ENSMUST00000188066.2
ENSMUST00000190962.7 |
Pik3c2g
|
phosphatidylinositol-4-phosphate 3-kinase catalytic subunit type 2 gamma |
| chr6_-_129428869 | 0.12 |
ENSMUST00000203162.3
|
Clec1a
|
C-type lectin domain family 1, member a |
| chr7_-_44773750 | 0.12 |
ENSMUST00000211725.2
ENSMUST00000003521.10 |
Rps11
|
ribosomal protein S11 |
| chr2_-_5867734 | 0.12 |
ENSMUST00000071016.3
|
Gm13199
|
predicted gene 13199 |
| chr6_-_57827328 | 0.11 |
ENSMUST00000203310.3
ENSMUST00000203488.3 |
Vmn1r21
|
vomeronasal 1 receptor 21 |
| chr5_+_93241287 | 0.11 |
ENSMUST00000074733.11
ENSMUST00000201700.4 ENSMUST00000202196.4 ENSMUST00000202308.4 |
Septin11
|
septin 11 |
| chr19_+_9824919 | 0.11 |
ENSMUST00000179814.3
|
Scgb2a2
|
secretoglobin, family 2A, member 2 |
| chr1_+_132433968 | 0.10 |
ENSMUST00000058167.3
|
Tmem81
|
transmembrane protein 81 |
| chr12_-_30423356 | 0.09 |
ENSMUST00000021004.14
|
Sntg2
|
syntrophin, gamma 2 |
| chr13_-_3661064 | 0.07 |
ENSMUST00000096069.5
|
Tasor2
|
transcription activation suppressor family member 2 |
| chr3_-_144680801 | 0.07 |
ENSMUST00000029923.10
ENSMUST00000238960.2 |
Clca4a
|
chloride channel accessory 4A |
| chr9_+_20555629 | 0.06 |
ENSMUST00000161887.8
|
Ubl5
|
ubiquitin-like 5 |
| chr3_+_64856617 | 0.06 |
ENSMUST00000238901.2
|
Kcnab1
|
potassium voltage-gated channel, shaker-related subfamily, beta member 1 |
| chr11_+_29323618 | 0.06 |
ENSMUST00000040182.13
ENSMUST00000109477.2 |
Ccdc88a
|
coiled coil domain containing 88A |
| chr7_+_46445512 | 0.05 |
ENSMUST00000006774.11
ENSMUST00000165031.8 |
Gtf2h1
|
general transcription factor II H, polypeptide 1 |
| chr14_-_30890544 | 0.05 |
ENSMUST00000036618.14
|
Stab1
|
stabilin 1 |
| chr1_+_34275665 | 0.04 |
ENSMUST00000194192.3
|
Dst
|
dystonin |
| chr6_-_129637519 | 0.04 |
ENSMUST00000119533.2
ENSMUST00000145984.8 ENSMUST00000118401.8 ENSMUST00000112057.9 ENSMUST00000071920.11 |
Klrc2
|
killer cell lectin-like receptor subfamily C, member 2 |
| chr1_+_109911467 | 0.03 |
ENSMUST00000172005.8
|
Cdh7
|
cadherin 7, type 2 |
| chr19_-_10952028 | 0.03 |
ENSMUST00000072748.13
|
Ms4a10
|
membrane-spanning 4-domains, subfamily A, member 10 |
| chr16_-_88609108 | 0.03 |
ENSMUST00000232664.2
|
Gm20741
|
predicted gene, 20741 |
| chr13_+_49806772 | 0.03 |
ENSMUST00000223264.2
ENSMUST00000221142.2 ENSMUST00000222333.2 ENSMUST00000021824.8 |
Nol8
|
nucleolar protein 8 |
| chr9_-_20556031 | 0.03 |
ENSMUST00000148631.8
ENSMUST00000131128.2 ENSMUST00000151861.9 ENSMUST00000131343.8 ENSMUST00000086458.10 |
Fbxl12
|
F-box and leucine-rich repeat protein 12 |
| chr4_+_155048571 | 0.02 |
ENSMUST00000030931.11
ENSMUST00000070953.11 |
Pank4
|
pantothenate kinase 4 |
| chr9_-_14651985 | 0.01 |
ENSMUST00000076946.4
ENSMUST00000115644.10 |
Piwil4
|
piwi-like RNA-mediated gene silencing 4 |
| chr6_+_41204430 | 0.01 |
ENSMUST00000193064.2
ENSMUST00000103280.3 |
Trbv26
|
T cell receptor beta, variable 26 |
| chr14_-_51311892 | 0.01 |
ENSMUST00000216202.2
|
Olfr750
|
olfactory receptor 750 |
| chr10_+_41179966 | 0.00 |
ENSMUST00000173494.4
|
Ak9
|
adenylate kinase 9 |
| chr4_-_52919172 | 0.00 |
ENSMUST00000107667.3
ENSMUST00000213989.2 |
Olfr272
|
olfactory receptor 272 |
| chr18_-_70377653 | 0.00 |
ENSMUST00000025390.4
|
Dynap
|
dynactin associated protein |
| chr3_-_105908789 | 0.00 |
ENSMUST00000066319.8
|
Pifo
|
primary cilia formation |
Network of associatons between targets according to the STRING database.
First level regulatory network of Rhox11
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.9 | GO:0072488 | ammonium transmembrane transport(GO:0072488) |
| 0.7 | 2.9 | GO:1900191 | biofilm formation(GO:0042710) single-species biofilm formation(GO:0044010) single-species biofilm formation in or on host organism(GO:0044407) membrane disruption in other organism(GO:0051673) regulation of single-species biofilm formation(GO:1900190) negative regulation of single-species biofilm formation(GO:1900191) regulation of single-species biofilm formation in or on host organism(GO:1900228) negative regulation of single-species biofilm formation in or on host organism(GO:1900229) |
| 0.2 | 0.6 | GO:0042694 | muscle cell fate specification(GO:0042694) |
| 0.2 | 1.2 | GO:0034627 | nicotinamide nucleotide biosynthetic process from aspartate(GO:0019355) 'de novo' NAD biosynthetic process(GO:0034627) 'de novo' NAD biosynthetic process from aspartate(GO:0034628) |
| 0.2 | 0.6 | GO:0033861 | negative regulation of NAD(P)H oxidase activity(GO:0033861) neuron projection maintenance(GO:1990535) |
| 0.2 | 0.9 | GO:0018343 | protein farnesylation(GO:0018343) |
| 0.2 | 0.6 | GO:0072717 | cellular response to actinomycin D(GO:0072717) |
| 0.1 | 0.4 | GO:1900135 | positive regulation of eosinophil degranulation(GO:0043311) regulation of neutrophil mediated cytotoxicity(GO:0070948) regulation of neutrophil mediated killing of symbiont cell(GO:0070949) positive regulation of renin secretion into blood stream(GO:1900135) positive regulation of eosinophil activation(GO:1902568) |
| 0.1 | 1.5 | GO:0044351 | macropinocytosis(GO:0044351) |
| 0.1 | 0.5 | GO:2000705 | positive regulation of relaxation of muscle(GO:1901079) regulation of dense core granule biogenesis(GO:2000705) |
| 0.1 | 1.3 | GO:2000346 | negative regulation of hepatocyte proliferation(GO:2000346) |
| 0.1 | 0.4 | GO:0072021 | ascending thin limb development(GO:0072021) thick ascending limb development(GO:0072023) metanephric ascending thin limb development(GO:0072218) metanephric thick ascending limb development(GO:0072233) |
| 0.1 | 0.9 | GO:0033314 | mitotic DNA replication checkpoint(GO:0033314) |
| 0.1 | 0.4 | GO:0007571 | age-dependent response to oxidative stress(GO:0001306) age-dependent general metabolic decline(GO:0007571) amino acid neurotransmitter reuptake(GO:0051933) glutamate reuptake(GO:0051935) |
| 0.1 | 0.3 | GO:0006432 | phenylalanyl-tRNA aminoacylation(GO:0006432) |
| 0.1 | 0.2 | GO:1905077 | negative regulation of interleukin-17 secretion(GO:1905077) |
| 0.1 | 0.2 | GO:0017055 | negative regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0017055) |
| 0.0 | 1.3 | GO:0042730 | fibrinolysis(GO:0042730) |
| 0.0 | 0.5 | GO:0000056 | ribosomal small subunit export from nucleus(GO:0000056) |
| 0.0 | 0.5 | GO:0061635 | regulation of protein complex stability(GO:0061635) |
| 0.0 | 0.8 | GO:2001275 | positive regulation of glucose import in response to insulin stimulus(GO:2001275) |
| 0.0 | 0.3 | GO:0034316 | negative regulation of Arp2/3 complex-mediated actin nucleation(GO:0034316) |
| 0.0 | 1.0 | GO:0019373 | epoxygenase P450 pathway(GO:0019373) |
| 0.0 | 0.3 | GO:0006002 | fructose 6-phosphate metabolic process(GO:0006002) |
| 0.0 | 0.3 | GO:0051490 | negative regulation of filopodium assembly(GO:0051490) |
| 0.0 | 0.4 | GO:0051195 | negative regulation of glycolytic process(GO:0045820) negative regulation of cofactor metabolic process(GO:0051195) negative regulation of coenzyme metabolic process(GO:0051198) |
| 0.0 | 0.5 | GO:0001967 | suckling behavior(GO:0001967) |
| 0.0 | 0.2 | GO:1903690 | negative regulation of wound healing, spreading of epidermal cells(GO:1903690) |
| 0.0 | 0.1 | GO:0045919 | positive regulation of cytolysis(GO:0045919) |
| 0.0 | 1.1 | GO:0015914 | phospholipid transport(GO:0015914) |
| 0.0 | 1.4 | GO:0060291 | long-term synaptic potentiation(GO:0060291) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 2.9 | GO:0044279 | other organism cell membrane(GO:0044218) other organism membrane(GO:0044279) |
| 0.3 | 0.9 | GO:0005965 | protein farnesyltransferase complex(GO:0005965) |
| 0.1 | 0.3 | GO:0009328 | phenylalanine-tRNA ligase complex(GO:0009328) |
| 0.1 | 0.5 | GO:0042583 | chromaffin granule(GO:0042583) |
| 0.1 | 0.7 | GO:0070652 | HAUS complex(GO:0070652) |
| 0.1 | 0.3 | GO:0005945 | 6-phosphofructokinase complex(GO:0005945) |
| 0.1 | 0.6 | GO:0031094 | platelet dense tubular network(GO:0031094) |
| 0.0 | 1.3 | GO:0031362 | anchored component of external side of plasma membrane(GO:0031362) |
| 0.0 | 0.5 | GO:0005642 | annulate lamellae(GO:0005642) |
| 0.0 | 0.4 | GO:0001739 | sex chromatin(GO:0001739) |
| 0.0 | 0.6 | GO:0097512 | cardiac myofibril(GO:0097512) |
| 0.0 | 1.0 | GO:0046540 | U4/U6 x U5 tri-snRNP complex(GO:0046540) |
| 0.0 | 0.5 | GO:0001931 | uropod(GO:0001931) cell trailing edge(GO:0031254) |
| 0.0 | 1.4 | GO:0032590 | dendrite membrane(GO:0032590) |
| 0.0 | 0.1 | GO:1990131 | Iml1 complex(GO:1990130) Gtr1-Gtr2 GTPase complex(GO:1990131) |
| 0.0 | 0.2 | GO:0005828 | kinetochore microtubule(GO:0005828) |
| 0.0 | 0.4 | GO:0031143 | pseudopodium(GO:0031143) |
| 0.0 | 0.3 | GO:0071203 | F-actin capping protein complex(GO:0008290) WASH complex(GO:0071203) |
| 0.0 | 0.6 | GO:0005732 | small nucleolar ribonucleoprotein complex(GO:0005732) |
| 0.0 | 0.2 | GO:0008540 | proteasome regulatory particle, base subcomplex(GO:0008540) cytosolic proteasome complex(GO:0031597) |
| 0.0 | 0.2 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 0.0 | 1.0 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.9 | GO:0004660 | protein farnesyltransferase activity(GO:0004660) |
| 0.2 | 1.2 | GO:0004515 | nicotinamide-nucleotide adenylyltransferase activity(GO:0000309) nicotinate-nucleotide adenylyltransferase activity(GO:0004515) |
| 0.2 | 0.6 | GO:0032190 | acrosin binding(GO:0032190) |
| 0.2 | 1.1 | GO:1990050 | phosphatidic acid transporter activity(GO:1990050) |
| 0.2 | 1.0 | GO:0033695 | oxidoreductase activity, acting on CH or CH2 groups, quinone or similar compound as acceptor(GO:0033695) caffeine oxidase activity(GO:0034875) |
| 0.2 | 0.9 | GO:0010997 | anaphase-promoting complex binding(GO:0010997) |
| 0.1 | 1.0 | GO:0005087 | Ran guanyl-nucleotide exchange factor activity(GO:0005087) |
| 0.1 | 2.9 | GO:0022840 | leak channel activity(GO:0022840) narrow pore channel activity(GO:0022842) |
| 0.1 | 0.6 | GO:0051373 | FATZ binding(GO:0051373) |
| 0.1 | 0.5 | GO:0003953 | NAD+ nucleosidase activity(GO:0003953) |
| 0.1 | 0.7 | GO:0000253 | 3-keto sterol reductase activity(GO:0000253) |
| 0.1 | 0.4 | GO:0015057 | thrombin receptor activity(GO:0015057) |
| 0.1 | 1.5 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 0.1 | 2.9 | GO:0001530 | lipopolysaccharide binding(GO:0001530) |
| 0.1 | 0.6 | GO:0070180 | large ribosomal subunit rRNA binding(GO:0070180) |
| 0.1 | 0.3 | GO:0003872 | 6-phosphofructokinase activity(GO:0003872) |
| 0.1 | 0.3 | GO:0004826 | phenylalanine-tRNA ligase activity(GO:0004826) |
| 0.1 | 0.2 | GO:0002153 | steroid receptor RNA activator RNA binding(GO:0002153) |
| 0.0 | 0.4 | GO:0097027 | ubiquitin-protein transferase activator activity(GO:0097027) |
| 0.0 | 0.4 | GO:0019864 | IgG binding(GO:0019864) |
| 0.0 | 0.5 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.0 | 0.2 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.0 | 0.2 | GO:0043515 | kinetochore binding(GO:0043515) |
| 0.0 | 0.1 | GO:0005483 | soluble NSF attachment protein activity(GO:0005483) |
| 0.0 | 0.3 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 0.0 | 0.2 | GO:0030229 | very-low-density lipoprotein particle receptor activity(GO:0030229) |
| 0.0 | 0.5 | GO:0008171 | O-methyltransferase activity(GO:0008171) |
| 0.0 | 0.4 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.0 | 0.6 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.0 | 0.2 | GO:0034450 | ubiquitin-ubiquitin ligase activity(GO:0034450) |
| 0.0 | 0.9 | GO:0019843 | rRNA binding(GO:0019843) |
| 0.0 | 0.6 | GO:0005159 | insulin-like growth factor receptor binding(GO:0005159) |
| 0.0 | 0.2 | GO:0001054 | RNA polymerase I activity(GO:0001054) |
| 0.0 | 1.4 | GO:0004860 | protein kinase inhibitor activity(GO:0004860) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.3 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.0 | 1.5 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.0 | 0.6 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
| 0.0 | 1.5 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.0 | 0.5 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
| 0.0 | 1.3 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.0 | 0.4 | ST GA12 PATHWAY | G alpha 12 Pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.9 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.1 | 1.3 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.1 | 2.9 | REACTOME AMINE COMPOUND SLC TRANSPORTERS | Genes involved in Amine compound SLC transporters |
| 0.1 | 1.5 | REACTOME P38MAPK EVENTS | Genes involved in p38MAPK events |
| 0.0 | 1.3 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.0 | 0.5 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.0 | 0.7 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 1.0 | REACTOME MRNA SPLICING MINOR PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |
| 0.0 | 1.2 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.0 | 0.8 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.0 | 0.2 | REACTOME RNA POL III CHAIN ELONGATION | Genes involved in RNA Polymerase III Chain Elongation |