Project
GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
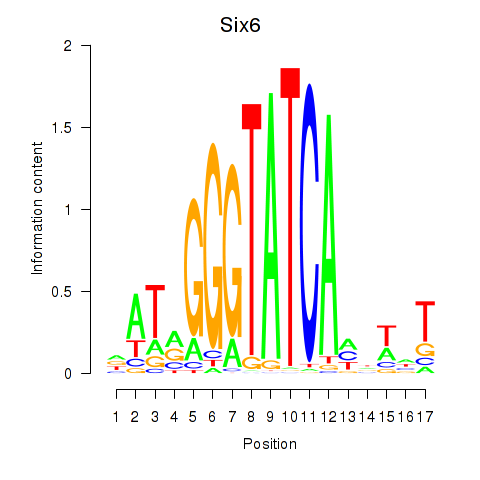
Results for Six6
Z-value: 1.49
Transcription factors associated with Six6
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Six6
|
ENSMUSG00000021099.7 | sine oculis-related homeobox 6 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Six6 | mm39_v1_chr12_+_72986665_72986674 | 0.10 | 5.6e-01 | Click! |
Activity profile of Six6 motif
Sorted Z-values of Six6 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr6_+_34389269 | 20.04 |
ENSMUST00000007449.9
|
Akr1b7
|
aldo-keto reductase family 1, member B7 |
| chr11_-_102255999 | 18.77 |
ENSMUST00000006749.10
|
Slc4a1
|
solute carrier family 4 (anion exchanger), member 1 |
| chr11_+_117688486 | 17.98 |
ENSMUST00000106331.2
|
6030468B19Rik
|
RIKEN cDNA 6030468B19 gene |
| chr17_+_48554786 | 15.96 |
ENSMUST00000048065.6
|
Trem3
|
triggering receptor expressed on myeloid cells 3 |
| chr17_+_41121979 | 12.82 |
ENSMUST00000024721.8
ENSMUST00000233740.2 |
Rhag
|
Rhesus blood group-associated A glycoprotein |
| chr15_+_79779218 | 9.13 |
ENSMUST00000023054.14
|
Apobec3
|
apolipoprotein B mRNA editing enzyme, catalytic polypeptide 3 |
| chr9_+_110856425 | 7.35 |
ENSMUST00000199313.2
|
Ltf
|
lactotransferrin |
| chr2_+_131333672 | 6.97 |
ENSMUST00000028806.12
|
Smox
|
spermine oxidase |
| chr6_+_41369290 | 6.96 |
ENSMUST00000049079.9
|
Gm5771
|
predicted gene 5771 |
| chr12_-_113386312 | 6.54 |
ENSMUST00000177715.8
ENSMUST00000103426.3 |
Ighm
|
immunoglobulin heavy constant mu |
| chr3_-_113166153 | 6.03 |
ENSMUST00000098673.5
|
Amy2a5
|
amylase 2a5 |
| chr17_+_43879496 | 5.91 |
ENSMUST00000169694.2
|
Pla2g7
|
phospholipase A2, group VII (platelet-activating factor acetylhydrolase, plasma) |
| chr8_+_95720864 | 5.88 |
ENSMUST00000212141.2
|
Adgrg1
|
adhesion G protein-coupled receptor G1 |
| chr19_+_4204605 | 5.85 |
ENSMUST00000061086.9
|
Ptprcap
|
protein tyrosine phosphatase, receptor type, C polypeptide-associated protein |
| chr14_-_70864666 | 5.19 |
ENSMUST00000022694.17
|
Dmtn
|
dematin actin binding protein |
| chr7_+_127661835 | 5.13 |
ENSMUST00000106242.10
ENSMUST00000120355.8 ENSMUST00000106240.9 ENSMUST00000098015.10 |
Itgam
Gm49368
|
integrin alpha M predicted gene, 49368 |
| chr7_-_97827461 | 5.00 |
ENSMUST00000040971.14
|
Capn5
|
calpain 5 |
| chr1_-_131066004 | 4.98 |
ENSMUST00000016670.9
|
Dyrk3
|
dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 3 |
| chr6_+_123099615 | 4.96 |
ENSMUST00000161636.8
ENSMUST00000161365.8 |
Clec4a2
|
C-type lectin domain family 4, member a2 |
| chr3_-_113263974 | 4.74 |
ENSMUST00000098667.5
|
Amy2a2
|
amylase 2a2 |
| chr3_-_37778470 | 4.67 |
ENSMUST00000108105.2
ENSMUST00000079755.5 ENSMUST00000099128.2 |
Gm5148
|
predicted gene 5148 |
| chr7_-_133304244 | 4.35 |
ENSMUST00000209636.2
ENSMUST00000153698.3 |
Uros
|
uroporphyrinogen III synthase |
| chr1_+_152683568 | 4.18 |
ENSMUST00000190323.7
|
Ncf2
|
neutrophil cytosolic factor 2 |
| chr7_-_44198157 | 4.17 |
ENSMUST00000145956.2
ENSMUST00000049343.15 |
Pold1
|
polymerase (DNA directed), delta 1, catalytic subunit |
| chr17_+_43879366 | 4.14 |
ENSMUST00000167418.8
|
Pla2g7
|
phospholipase A2, group VII (platelet-activating factor acetylhydrolase, plasma) |
| chr7_+_43086432 | 4.12 |
ENSMUST00000070518.4
|
Nkg7
|
natural killer cell group 7 sequence |
| chr17_+_35460722 | 3.87 |
ENSMUST00000068056.12
ENSMUST00000174757.8 ENSMUST00000173731.8 |
Ddx39b
|
DEAD box helicase 39b |
| chr1_-_128287347 | 3.76 |
ENSMUST00000190495.2
ENSMUST00000027601.11 |
Mcm6
|
minichromosome maintenance complex component 6 |
| chr7_+_130633776 | 3.66 |
ENSMUST00000084509.7
ENSMUST00000213064.3 ENSMUST00000208311.4 |
Dmbt1
|
deleted in malignant brain tumors 1 |
| chr13_+_100261358 | 3.59 |
ENSMUST00000022147.15
ENSMUST00000091321.6 ENSMUST00000143937.2 |
Smn1
|
survival motor neuron 1 |
| chr9_+_98305014 | 3.57 |
ENSMUST00000052068.11
|
Rbp1
|
retinol binding protein 1, cellular |
| chr7_-_100504610 | 3.53 |
ENSMUST00000156855.8
|
Relt
|
RELT tumor necrosis factor receptor |
| chr1_-_165535654 | 3.47 |
ENSMUST00000097474.9
|
Rcsd1
|
RCSD domain containing 1 |
| chr12_-_114252202 | 3.40 |
ENSMUST00000195124.6
ENSMUST00000103481.3 |
Ighv3-6
|
immunoglobulin heavy variable 3-6 |
| chr15_-_74635423 | 3.36 |
ENSMUST00000040404.8
|
Ly6d
|
lymphocyte antigen 6 complex, locus D |
| chr7_+_43086554 | 3.34 |
ENSMUST00000206741.2
|
Nkg7
|
natural killer cell group 7 sequence |
| chr12_-_113589576 | 3.24 |
ENSMUST00000103446.2
|
Ighv5-6
|
immunoglobulin heavy variable 5-6 |
| chr8_+_57908920 | 3.24 |
ENSMUST00000034023.4
|
Scrg1
|
scrapie responsive gene 1 |
| chr3_+_32760447 | 3.13 |
ENSMUST00000194781.6
|
Actl6a
|
actin-like 6A |
| chr6_-_120893771 | 3.10 |
ENSMUST00000004560.12
|
Bid
|
BH3 interacting domain death agonist |
| chr3_+_90173813 | 3.04 |
ENSMUST00000098914.10
|
Dennd4b
|
DENN/MADD domain containing 4B |
| chr6_+_123100382 | 3.04 |
ENSMUST00000032248.8
|
Clec4a2
|
C-type lectin domain family 4, member a2 |
| chr4_-_87724512 | 2.99 |
ENSMUST00000148059.2
|
Mllt3
|
myeloid/lymphoid or mixed-lineage leukemia; translocated to, 3 |
| chr11_+_115455260 | 2.99 |
ENSMUST00000021085.11
|
Nup85
|
nucleoporin 85 |
| chr9_+_120400510 | 2.99 |
ENSMUST00000165532.3
|
Rpl14
|
ribosomal protein L14 |
| chr6_-_69792108 | 2.94 |
ENSMUST00000103367.3
|
Igkv12-44
|
immunoglobulin kappa variable 12-44 |
| chr6_-_69741999 | 2.93 |
ENSMUST00000103365.3
|
Igkv12-46
|
immunoglobulin kappa variable 12-46 |
| chr6_+_123100272 | 2.91 |
ENSMUST00000041779.13
|
Clec4a2
|
C-type lectin domain family 4, member a2 |
| chr7_+_89814713 | 2.84 |
ENSMUST00000207084.2
|
Picalm
|
phosphatidylinositol binding clathrin assembly protein |
| chr13_-_97897139 | 2.83 |
ENSMUST00000074072.5
|
Rps18-ps6
|
ribosomal protein S18, pseudogene 6 |
| chr15_-_82128888 | 2.80 |
ENSMUST00000089155.6
ENSMUST00000089157.11 |
Cenpm
|
centromere protein M |
| chr10_+_128073900 | 2.77 |
ENSMUST00000105245.3
|
Timeless
|
timeless circadian clock 1 |
| chr4_+_119052548 | 2.69 |
ENSMUST00000106345.3
|
Svbp
|
small vasohibin binding protein |
| chr1_-_165462020 | 2.56 |
ENSMUST00000194437.6
ENSMUST00000068705.13 ENSMUST00000111435.9 ENSMUST00000193023.2 |
Mpzl1
|
myelin protein zero-like 1 |
| chr9_+_108820846 | 2.55 |
ENSMUST00000198140.5
ENSMUST00000051873.15 |
Pfkfb4
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 4 |
| chr2_+_127178072 | 2.51 |
ENSMUST00000028846.7
|
Dusp2
|
dual specificity phosphatase 2 |
| chr11_+_98303287 | 2.47 |
ENSMUST00000058295.6
|
Erbb2
|
erb-b2 receptor tyrosine kinase 2 |
| chr5_+_21748523 | 2.29 |
ENSMUST00000035651.6
|
Lrrc17
|
leucine rich repeat containing 17 |
| chrX_-_141749704 | 2.15 |
ENSMUST00000041317.3
|
Ammecr1
|
Alport syndrome, mental retardation, midface hypoplasia and elliptocytosis chromosomal region gene 1 |
| chr3_-_101195213 | 2.15 |
ENSMUST00000029456.5
|
Cd2
|
CD2 antigen |
| chr5_+_121342544 | 2.09 |
ENSMUST00000031617.13
|
Rpl6
|
ribosomal protein L6 |
| chr16_+_23109213 | 2.08 |
ENSMUST00000115335.2
|
St6gal1
|
beta galactoside alpha 2,6 sialyltransferase 1 |
| chr5_-_123126550 | 2.07 |
ENSMUST00000086200.11
ENSMUST00000156474.8 |
Kdm2b
|
lysine (K)-specific demethylase 2B |
| chr17_+_36176485 | 2.01 |
ENSMUST00000127442.8
ENSMUST00000144382.8 |
Ppp1r18
|
protein phosphatase 1, regulatory subunit 18 |
| chr5_+_24569802 | 1.97 |
ENSMUST00000115090.6
ENSMUST00000030834.7 |
Nos3
|
nitric oxide synthase 3, endothelial cell |
| chr2_+_30171055 | 1.86 |
ENSMUST00000143119.3
|
Gm28038
|
predicted gene, 28038 |
| chrX_-_73416869 | 1.85 |
ENSMUST00000073067.11
ENSMUST00000037967.6 |
Slc10a3
|
solute carrier family 10 (sodium/bile acid cotransporter family), member 3 |
| chrX_-_73416824 | 1.85 |
ENSMUST00000178691.2
ENSMUST00000114146.8 |
Ubl4a
Slc10a3
|
ubiquitin-like 4A solute carrier family 10 (sodium/bile acid cotransporter family), member 3 |
| chr2_-_114005752 | 1.71 |
ENSMUST00000102543.5
ENSMUST00000043160.13 |
Aqr
|
aquarius |
| chr7_+_80764547 | 1.59 |
ENSMUST00000026820.11
|
Slc28a1
|
solute carrier family 28 (sodium-coupled nucleoside transporter), member 1 |
| chr9_-_119170456 | 1.55 |
ENSMUST00000139870.2
|
Myd88
|
myeloid differentiation primary response gene 88 |
| chr6_-_119925387 | 1.54 |
ENSMUST00000162541.8
|
Wnk1
|
WNK lysine deficient protein kinase 1 |
| chr11_-_99121822 | 1.54 |
ENSMUST00000103133.4
|
Smarce1
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily e, member 1 |
| chr19_+_12647803 | 1.53 |
ENSMUST00000207341.3
ENSMUST00000208494.3 ENSMUST00000208657.3 |
Olfr1442
|
olfactory receptor 1442 |
| chr1_+_139382485 | 1.49 |
ENSMUST00000200083.5
ENSMUST00000053364.12 |
Aspm
|
abnormal spindle microtubule assembly |
| chr19_-_6002210 | 1.48 |
ENSMUST00000236013.2
|
Pola2
|
polymerase (DNA directed), alpha 2 |
| chr16_+_10652910 | 1.44 |
ENSMUST00000037913.9
|
Rmi2
|
RecQ mediated genome instability 2 |
| chr10_-_81463631 | 1.42 |
ENSMUST00000042923.9
|
Sirt6
|
sirtuin 6 |
| chr7_+_108265625 | 1.38 |
ENSMUST00000213979.3
ENSMUST00000216331.2 ENSMUST00000217170.2 |
Olfr510
|
olfactory receptor 510 |
| chr2_-_118859821 | 1.38 |
ENSMUST00000110833.2
ENSMUST00000036470.14 ENSMUST00000110834.8 |
Ccdc32
|
coiled-coil domain containing 32 |
| chr12_-_114646685 | 1.36 |
ENSMUST00000194350.6
ENSMUST00000103504.3 |
Ighv1-18
|
immunoglobulin heavy variable V1-18 |
| chr1_+_45350698 | 1.33 |
ENSMUST00000087883.13
|
Col3a1
|
collagen, type III, alpha 1 |
| chr15_+_101152078 | 1.32 |
ENSMUST00000228985.2
|
Nr4a1
|
nuclear receptor subfamily 4, group A, member 1 |
| chr2_-_180928867 | 1.32 |
ENSMUST00000130475.8
|
Gmeb2
|
glucocorticoid modulatory element binding protein 2 |
| chr5_+_139197689 | 1.31 |
ENSMUST00000148772.8
ENSMUST00000110882.8 |
Sun1
|
Sad1 and UNC84 domain containing 1 |
| chrX_+_168468186 | 1.30 |
ENSMUST00000112107.8
ENSMUST00000112104.8 |
Mid1
|
midline 1 |
| chr19_-_11313471 | 1.28 |
ENSMUST00000056035.9
ENSMUST00000067532.11 |
Ms4a7
|
membrane-spanning 4-domains, subfamily A, member 7 |
| chr15_+_82136598 | 1.21 |
ENSMUST00000136948.3
|
1500009C09Rik
|
RIKEN cDNA 1500009C09 gene |
| chrX_-_12539778 | 1.20 |
ENSMUST00000060108.7
|
1810030O07Rik
|
RIKEN cDNA 1810030O07 gene |
| chr19_-_4675352 | 1.17 |
ENSMUST00000224707.2
ENSMUST00000237059.2 |
Rce1
|
Ras converting CAAX endopeptidase 1 |
| chr2_-_13276074 | 1.16 |
ENSMUST00000137670.3
ENSMUST00000114791.9 |
Rsu1
|
Ras suppressor protein 1 |
| chr8_+_70275079 | 1.14 |
ENSMUST00000164890.8
ENSMUST00000034325.6 ENSMUST00000238452.2 |
Lpar2
|
lysophosphatidic acid receptor 2 |
| chr7_-_13856967 | 1.13 |
ENSMUST00000098809.4
|
Sult2a3
|
sulfotransferase family 2A, dehydroepiandrosterone (DHEA)-preferring, member 3 |
| chr18_+_37840092 | 1.12 |
ENSMUST00000195823.2
|
Pcdhga6
|
protocadherin gamma subfamily A, 6 |
| chr14_-_66071412 | 1.11 |
ENSMUST00000022613.10
|
Esco2
|
establishment of sister chromatid cohesion N-acetyltransferase 2 |
| chr12_-_103956176 | 1.10 |
ENSMUST00000151709.3
ENSMUST00000176246.3 ENSMUST00000074693.13 ENSMUST00000120251.9 |
Serpina11
|
serine (or cysteine) peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 11 |
| chr2_-_13276205 | 1.10 |
ENSMUST00000191959.6
ENSMUST00000028059.9 |
Rsu1
|
Ras suppressor protein 1 |
| chr7_+_110226857 | 1.09 |
ENSMUST00000033054.10
|
Adm
|
adrenomedullin |
| chr6_+_56979285 | 1.09 |
ENSMUST00000079669.7
|
Vmn1r6
|
vomeronasal 1 receptor 6 |
| chr3_+_87283687 | 1.08 |
ENSMUST00000163661.8
ENSMUST00000072480.9 |
Fcrl1
|
Fc receptor-like 1 |
| chr2_-_110781268 | 1.08 |
ENSMUST00000099623.10
|
Ano3
|
anoctamin 3 |
| chr6_+_48514518 | 1.05 |
ENSMUST00000040361.8
|
Atp6v0e2
|
ATPase, H+ transporting, lysosomal V0 subunit E2 |
| chr11_+_100210705 | 1.04 |
ENSMUST00000049385.14
|
Eif1
|
eukaryotic translation initiation factor 1 |
| chr19_-_4675300 | 1.02 |
ENSMUST00000225264.2
ENSMUST00000237022.2 ENSMUST00000224675.3 |
Rce1
|
Ras converting CAAX endopeptidase 1 |
| chr18_+_74349189 | 0.98 |
ENSMUST00000025444.8
|
Cxxc1
|
CXXC finger 1 (PHD domain) |
| chr8_+_70285282 | 0.97 |
ENSMUST00000131637.9
|
Pbx4
|
pre B cell leukemia homeobox 4 |
| chr7_-_101552989 | 0.96 |
ENSMUST00000106969.9
|
Tomt
|
transmembrane O-methyltransferase |
| chr6_+_48514578 | 0.93 |
ENSMUST00000203011.2
|
Atp6v0e2
|
ATPase, H+ transporting, lysosomal V0 subunit E2 |
| chr12_-_115611981 | 0.92 |
ENSMUST00000103540.3
ENSMUST00000199266.2 |
Ighv8-12
|
immunoglobulin heavy variable V8-12 |
| chr2_-_174314672 | 0.92 |
ENSMUST00000117442.8
ENSMUST00000141100.2 ENSMUST00000120822.2 |
Prelid3b
|
PRELI domain containing 3B |
| chr11_-_4799345 | 0.91 |
ENSMUST00000053079.13
ENSMUST00000109910.9 |
Nf2
|
neurofibromin 2 |
| chr8_+_85558151 | 0.91 |
ENSMUST00000036734.6
|
Gadd45gip1
|
growth arrest and DNA-damage-inducible, gamma interacting protein 1 |
| chr16_-_18052937 | 0.87 |
ENSMUST00000076957.7
|
Zdhhc8
|
zinc finger, DHHC domain containing 8 |
| chr6_-_68887957 | 0.85 |
ENSMUST00000200454.2
|
Igkv4-86
|
immunoglobulin kappa variable 4-86 |
| chr5_-_23821523 | 0.85 |
ENSMUST00000088392.9
|
Srpk2
|
serine/arginine-rich protein specific kinase 2 |
| chr4_+_11758147 | 0.84 |
ENSMUST00000029871.12
ENSMUST00000108303.2 |
Cdh17
|
cadherin 17 |
| chr5_+_140404997 | 0.83 |
ENSMUST00000100507.8
|
Eif3b
|
eukaryotic translation initiation factor 3, subunit B |
| chr2_-_174314741 | 0.82 |
ENSMUST00000016401.15
|
Prelid3b
|
PRELI domain containing 3B |
| chr19_-_4675631 | 0.80 |
ENSMUST00000225375.2
ENSMUST00000025823.6 |
Rce1
|
Ras converting CAAX endopeptidase 1 |
| chr8_+_31640332 | 0.79 |
ENSMUST00000209851.2
ENSMUST00000098842.3 ENSMUST00000210129.2 |
Tti2
|
TELO2 interacting protein 2 |
| chr2_-_88534814 | 0.79 |
ENSMUST00000216928.2
ENSMUST00000216977.2 |
Olfr1196
|
olfactory receptor 1196 |
| chr9_+_106099797 | 0.79 |
ENSMUST00000062241.11
|
Tlr9
|
toll-like receptor 9 |
| chr18_-_88945571 | 0.79 |
ENSMUST00000147313.2
|
Socs6
|
suppressor of cytokine signaling 6 |
| chr6_+_67816777 | 0.79 |
ENSMUST00000200578.5
ENSMUST00000103308.3 |
Igkv9-129
|
immunoglobulin kappa variable 9-129 |
| chr8_-_87611849 | 0.76 |
ENSMUST00000034074.8
|
N4bp1
|
NEDD4 binding protein 1 |
| chr2_-_103609703 | 0.76 |
ENSMUST00000143188.2
|
Caprin1
|
cell cycle associated protein 1 |
| chr12_-_114710326 | 0.74 |
ENSMUST00000103507.2
|
Ighv1-22
|
immunoglobulin heavy variable 1-22 |
| chr11_+_76836545 | 0.74 |
ENSMUST00000125145.8
|
Blmh
|
bleomycin hydrolase |
| chr2_-_89408791 | 0.72 |
ENSMUST00000217402.2
|
Olfr1245
|
olfactory receptor 1245 |
| chr2_+_31135813 | 0.70 |
ENSMUST00000000199.8
|
Ncs1
|
neuronal calcium sensor 1 |
| chr13_-_23806530 | 0.68 |
ENSMUST00000062045.4
|
H1f4
|
H1.4 linker histone, cluster member |
| chr13_+_94219934 | 0.68 |
ENSMUST00000156071.2
|
Lhfpl2
|
lipoma HMGIC fusion partner-like 2 |
| chr5_-_108943211 | 0.67 |
ENSMUST00000004943.2
|
Tmed11
|
transmembrane p24 trafficking protein 11 |
| chr2_+_117080212 | 0.67 |
ENSMUST00000028825.5
|
Fam98b
|
family with sequence similarity 98, member B |
| chr1_+_40305738 | 0.67 |
ENSMUST00000114795.3
|
Il1r1
|
interleukin 1 receptor, type I |
| chr6_-_68857658 | 0.63 |
ENSMUST00000198756.2
|
Gm42543
|
predicted gene 42543 |
| chr16_-_18695267 | 0.62 |
ENSMUST00000119273.2
|
Mrpl40
|
mitochondrial ribosomal protein L40 |
| chr7_-_119058489 | 0.61 |
ENSMUST00000207887.3
ENSMUST00000239424.2 ENSMUST00000033255.8 |
Gp2
|
glycoprotein 2 (zymogen granule membrane) |
| chr14_+_53607470 | 0.60 |
ENSMUST00000103652.5
|
Trav14n-3
|
T cell receptor alpha variable 14N-3 |
| chr8_+_22224506 | 0.59 |
ENSMUST00000080533.6
|
Defa24
|
defensin, alpha, 24 |
| chr19_-_32173824 | 0.58 |
ENSMUST00000151822.2
|
Sgms1
|
sphingomyelin synthase 1 |
| chrX_-_56438322 | 0.56 |
ENSMUST00000114730.8
|
Rbmx
|
RNA binding motif protein, X chromosome |
| chr8_+_70285133 | 0.55 |
ENSMUST00000081503.13
|
Pbx4
|
pre B cell leukemia homeobox 4 |
| chr10_-_7162196 | 0.54 |
ENSMUST00000015346.12
|
Cnksr3
|
Cnksr family member 3 |
| chr1_+_83022653 | 0.54 |
ENSMUST00000222567.2
|
Gm47969
|
predicted gene, 47969 |
| chr2_-_86061745 | 0.51 |
ENSMUST00000216056.2
|
Olfr1047
|
olfactory receptor 1047 |
| chr2_+_86128161 | 0.51 |
ENSMUST00000054746.5
|
Olfr1052
|
olfactory receptor 1052 |
| chr14_+_51181956 | 0.50 |
ENSMUST00000178092.2
ENSMUST00000227052.2 |
Pnp
Gm49342
|
purine-nucleoside phosphorylase predicted gene, 49342 |
| chr3_-_75177378 | 0.50 |
ENSMUST00000039047.5
|
Serpini2
|
serine (or cysteine) peptidase inhibitor, clade I, member 2 |
| chr9_+_32547529 | 0.50 |
ENSMUST00000238819.2
|
Ets1
|
E26 avian leukemia oncogene 1, 5' domain |
| chr7_+_37885660 | 0.48 |
ENSMUST00000179503.5
|
1600014C10Rik
|
RIKEN cDNA 1600014C10 gene |
| chr18_-_43820759 | 0.47 |
ENSMUST00000082254.8
|
Jakmip2
|
janus kinase and microtubule interacting protein 2 |
| chr6_-_57821483 | 0.47 |
ENSMUST00000226191.2
|
Vmn1r21
|
vomeronasal 1 receptor 21 |
| chr19_+_46329552 | 0.46 |
ENSMUST00000128041.8
|
Mfsd13a
|
major facilitator superfamily domain containing 13a |
| chr18_+_37794819 | 0.46 |
ENSMUST00000194888.2
ENSMUST00000194190.2 |
Pcdhga1
|
protocadherin gamma subfamily A, 1 |
| chr6_-_50359797 | 0.44 |
ENSMUST00000114468.9
|
Osbpl3
|
oxysterol binding protein-like 3 |
| chr15_+_12205095 | 0.43 |
ENSMUST00000038172.16
|
Mtmr12
|
myotubularin related protein 12 |
| chr7_-_79974166 | 0.43 |
ENSMUST00000047362.11
ENSMUST00000121882.8 |
Rccd1
|
RCC1 domain containing 1 |
| chr5_-_100521343 | 0.40 |
ENSMUST00000182433.8
|
Sec31a
|
Sec31 homolog A (S. cerevisiae) |
| chr6_+_57180275 | 0.40 |
ENSMUST00000226892.2
ENSMUST00000227421.2 |
Vmn1r13
|
vomeronasal 1 receptor 13 |
| chr11_+_78215026 | 0.40 |
ENSMUST00000102478.4
|
Aldoc
|
aldolase C, fructose-bisphosphate |
| chr9_-_38585897 | 0.39 |
ENSMUST00000215461.3
|
Olfr918
|
olfactory receptor 918 |
| chr19_-_11637880 | 0.38 |
ENSMUST00000135994.2
ENSMUST00000121793.2 |
Oosp2
|
oocyte secreted protein 2 |
| chrX_-_52672363 | 0.38 |
ENSMUST00000088778.5
|
Rtl8b
|
retrotransposon Gag like 8B |
| chr19_+_13595285 | 0.38 |
ENSMUST00000216688.3
|
Olfr1487
|
olfactory receptor 1487 |
| chr8_-_4325886 | 0.38 |
ENSMUST00000003029.14
|
Timm44
|
translocase of inner mitochondrial membrane 44 |
| chr6_-_120893725 | 0.36 |
ENSMUST00000145948.2
|
Bid
|
BH3 interacting domain death agonist |
| chr7_+_37883300 | 0.36 |
ENSMUST00000179992.10
|
1600014C10Rik
|
RIKEN cDNA 1600014C10 gene |
| chr7_+_23330147 | 0.35 |
ENSMUST00000227774.2
ENSMUST00000226771.2 ENSMUST00000228681.2 ENSMUST00000228559.2 ENSMUST00000228674.2 ENSMUST00000227866.2 ENSMUST00000227386.2 ENSMUST00000228484.2 ENSMUST00000226321.2 ENSMUST00000226128.2 ENSMUST00000226733.2 ENSMUST00000228228.2 |
Vmn1r171
|
vomeronasal 1 receptor 171 |
| chr6_+_57234937 | 0.34 |
ENSMUST00000228297.2
|
Vmn1r15
|
vomeronasal 1 receptor 15 |
| chr1_-_173703424 | 0.34 |
ENSMUST00000186442.7
|
Mndal
|
myeloid nuclear differentiation antigen like |
| chr11_+_60244132 | 0.34 |
ENSMUST00000070805.13
ENSMUST00000094140.9 ENSMUST00000108723.9 ENSMUST00000108722.5 |
Drc3
|
dynein regulatory complex subunit 3 |
| chr17_-_53846438 | 0.33 |
ENSMUST00000056198.4
|
Pp2d1
|
protein phosphatase 2C-like domain containing 1 |
| chr7_-_66915756 | 0.33 |
ENSMUST00000207715.2
|
Mef2a
|
myocyte enhancer factor 2A |
| chr9_+_38725910 | 0.29 |
ENSMUST00000213164.2
|
Olfr922
|
olfactory receptor 922 |
| chr3_+_87283767 | 0.27 |
ENSMUST00000194786.6
ENSMUST00000191666.2 |
Fcrl1
|
Fc receptor-like 1 |
| chr12_+_84034628 | 0.27 |
ENSMUST00000021649.8
|
Acot2
|
acyl-CoA thioesterase 2 |
| chr3_+_82915031 | 0.26 |
ENSMUST00000048486.13
ENSMUST00000194175.2 |
Fgg
|
fibrinogen gamma chain |
| chr3_-_92594516 | 0.26 |
ENSMUST00000029524.4
|
Lce1d
|
late cornified envelope 1D |
| chr3_-_92407800 | 0.26 |
ENSMUST00000062129.2
|
Sprr4
|
small proline-rich protein 4 |
| chr6_+_90246088 | 0.26 |
ENSMUST00000058039.3
|
Vmn1r54
|
vomeronasal 1 receptor 54 |
| chr2_+_106523532 | 0.24 |
ENSMUST00000111063.8
|
Mpped2
|
metallophosphoesterase domain containing 2 |
| chr12_-_103409912 | 0.23 |
ENSMUST00000055071.9
|
Ifi27l2a
|
interferon, alpha-inducible protein 27 like 2A |
| chr19_+_9824919 | 0.23 |
ENSMUST00000179814.3
|
Scgb2a2
|
secretoglobin, family 2A, member 2 |
| chr2_+_87185159 | 0.21 |
ENSMUST00000215163.3
|
Olfr1120
|
olfactory receptor 1120 |
| chr3_+_92158054 | 0.20 |
ENSMUST00000071805.4
|
Sprr2a2
|
small proline-rich protein 2A2 |
| chr6_-_81942906 | 0.20 |
ENSMUST00000032124.9
|
Mrpl19
|
mitochondrial ribosomal protein L19 |
| chr11_-_101877832 | 0.19 |
ENSMUST00000107173.9
ENSMUST00000107172.8 |
Dusp3
|
dual specificity phosphatase 3 (vaccinia virus phosphatase VH1-related) |
| chr11_+_49327451 | 0.18 |
ENSMUST00000215226.2
|
Olfr1388
|
olfactory receptor 1388 |
| chr6_-_28449250 | 0.18 |
ENSMUST00000164519.9
ENSMUST00000171089.9 ENSMUST00000031718.14 |
Pax4
|
paired box 4 |
| chr4_-_58499398 | 0.18 |
ENSMUST00000107570.2
|
Lpar1
|
lysophosphatidic acid receptor 1 |
| chr3_-_9069745 | 0.18 |
ENSMUST00000120143.8
|
Tpd52
|
tumor protein D52 |
| chr11_-_106163753 | 0.18 |
ENSMUST00000021052.16
|
Smarcd2
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 2 |
| chr2_+_87137925 | 0.17 |
ENSMUST00000216396.3
|
Olfr1118
|
olfactory receptor 1118 |
| chr11_+_85243970 | 0.16 |
ENSMUST00000108056.8
ENSMUST00000108061.8 ENSMUST00000108062.8 ENSMUST00000138423.8 ENSMUST00000092821.10 ENSMUST00000074875.11 |
Bcas3
|
breast carcinoma amplified sequence 3 |
| chr3_+_68479578 | 0.16 |
ENSMUST00000170788.9
|
Schip1
|
schwannomin interacting protein 1 |
| chr6_-_146855880 | 0.16 |
ENSMUST00000111622.2
ENSMUST00000036592.15 |
1700034J05Rik
|
RIKEN cDNA 1700034J05 gene |
| chr3_-_96634880 | 0.14 |
ENSMUST00000029741.9
|
Polr3c
|
polymerase (RNA) III (DNA directed) polypeptide C |
| chr7_-_102507962 | 0.13 |
ENSMUST00000213481.2
ENSMUST00000209952.2 |
Olfr566
|
olfactory receptor 566 |
| chr2_-_127324419 | 0.13 |
ENSMUST00000088538.6
|
Kcnip3
|
Kv channel interacting protein 3, calsenilin |
Network of associatons between targets according to the STRING database.
First level regulatory network of Six6
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.2 | 12.8 | GO:0072488 | ammonium transmembrane transport(GO:0072488) |
| 2.2 | 6.5 | GO:0002344 | peripheral B cell selection(GO:0002343) B cell affinity maturation(GO:0002344) |
| 2.0 | 10.0 | GO:0034441 | plasma lipoprotein particle oxidation(GO:0034441) |
| 2.0 | 16.0 | GO:0070945 | neutrophil mediated killing of gram-negative bacterium(GO:0070945) |
| 1.8 | 7.3 | GO:0042710 | biofilm formation(GO:0042710) single-species biofilm formation(GO:0044010) single-species biofilm formation in or on host organism(GO:0044407) membrane disruption in other organism(GO:0051673) regulation of single-species biofilm formation(GO:1900190) negative regulation of single-species biofilm formation(GO:1900191) regulation of single-species biofilm formation in or on host organism(GO:1900228) negative regulation of single-species biofilm formation in or on host organism(GO:1900229) |
| 1.4 | 4.3 | GO:0006780 | uroporphyrinogen III biosynthetic process(GO:0006780) |
| 1.4 | 4.2 | GO:0045004 | DNA replication proofreading(GO:0045004) |
| 1.2 | 3.6 | GO:0033189 | response to vitamin A(GO:0033189) |
| 1.0 | 5.0 | GO:0035617 | stress granule disassembly(GO:0035617) |
| 0.9 | 5.2 | GO:0035585 | calcium-mediated signaling using extracellular calcium source(GO:0035585) |
| 0.8 | 7.0 | GO:0046208 | spermine catabolic process(GO:0046208) |
| 0.7 | 3.0 | GO:0071586 | CAAX-box protein processing(GO:0071586) CAAX-box protein maturation(GO:0080120) |
| 0.7 | 4.2 | GO:0006742 | NADP catabolic process(GO:0006742) |
| 0.7 | 18.8 | GO:0015701 | bicarbonate transport(GO:0015701) |
| 0.7 | 2.0 | GO:0014805 | smooth muscle adaptation(GO:0014805) |
| 0.6 | 5.1 | GO:0014004 | microglia differentiation(GO:0014004) microglia development(GO:0014005) |
| 0.6 | 2.8 | GO:1904975 | response to bleomycin(GO:1904975) cellular response to bleomycin(GO:1904976) |
| 0.6 | 3.9 | GO:2000002 | negative regulation of DNA damage checkpoint(GO:2000002) |
| 0.5 | 2.1 | GO:0021993 | initiation of neural tube closure(GO:0021993) |
| 0.5 | 3.0 | GO:0032483 | regulation of Rab protein signal transduction(GO:0032483) |
| 0.5 | 2.5 | GO:0044849 | estrous cycle(GO:0044849) |
| 0.5 | 1.4 | GO:0003245 | cardiac muscle tissue growth involved in heart morphogenesis(GO:0003245) |
| 0.5 | 2.8 | GO:1902962 | regulation of metalloendopeptidase activity involved in amyloid precursor protein catabolic process(GO:1902962) negative regulation of metalloendopeptidase activity involved in amyloid precursor protein catabolic process(GO:1902963) |
| 0.4 | 9.1 | GO:0010528 | regulation of transposition(GO:0010528) negative regulation of transposition(GO:0010529) |
| 0.4 | 2.1 | GO:1990743 | protein sialylation(GO:1990743) |
| 0.4 | 1.6 | GO:0015851 | nucleobase transport(GO:0015851) pyrimidine nucleobase transport(GO:0015855) |
| 0.4 | 0.8 | GO:0032741 | positive regulation of interleukin-18 production(GO:0032741) |
| 0.4 | 1.5 | GO:0072566 | chemokine (C-X-C motif) ligand 1 production(GO:0072566) regulation of chemokine (C-X-C motif) ligand 1 production(GO:2000338) |
| 0.4 | 3.5 | GO:0071474 | cellular hyperosmotic response(GO:0071474) |
| 0.4 | 3.5 | GO:2000271 | positive regulation of fibroblast apoptotic process(GO:2000271) |
| 0.3 | 3.8 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 0.3 | 1.5 | GO:0023016 | signal transduction by trans-phosphorylation(GO:0023016) |
| 0.3 | 2.3 | GO:0048539 | bone marrow development(GO:0048539) |
| 0.3 | 0.8 | GO:0002314 | germinal center B cell differentiation(GO:0002314) |
| 0.3 | 1.3 | GO:0060414 | aorta smooth muscle tissue morphogenesis(GO:0060414) |
| 0.3 | 2.6 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 0.3 | 0.5 | GO:0046122 | purine deoxyribonucleoside metabolic process(GO:0046122) |
| 0.2 | 1.5 | GO:0051661 | maintenance of centrosome location(GO:0051661) |
| 0.2 | 5.9 | GO:0021796 | cerebral cortex regionalization(GO:0021796) |
| 0.2 | 1.1 | GO:0034421 | post-translational protein acetylation(GO:0034421) |
| 0.2 | 2.1 | GO:1902715 | positive regulation of interferon-gamma secretion(GO:1902715) |
| 0.2 | 1.8 | GO:0071816 | tail-anchored membrane protein insertion into ER membrane(GO:0071816) |
| 0.2 | 0.6 | GO:0002386 | immune response in mucosal-associated lymphoid tissue(GO:0002386) |
| 0.2 | 1.3 | GO:0071376 | response to corticotropin-releasing hormone(GO:0043435) cellular response to corticotropin-releasing hormone stimulus(GO:0071376) |
| 0.2 | 1.3 | GO:0030473 | nucleokinesis involved in cell motility in cerebral cortex radial glia guided migration(GO:0021817) nuclear migration along microtubule(GO:0030473) nuclear matrix organization(GO:0043578) nuclear matrix anchoring at nuclear membrane(GO:0090292) |
| 0.2 | 0.7 | GO:0043418 | homocysteine catabolic process(GO:0043418) |
| 0.2 | 0.8 | GO:0035063 | nuclear speck organization(GO:0035063) |
| 0.2 | 0.7 | GO:0010286 | heat acclimation(GO:0010286) |
| 0.1 | 3.0 | GO:2000096 | positive regulation of Wnt signaling pathway, planar cell polarity pathway(GO:2000096) |
| 0.1 | 3.7 | GO:0001833 | inner cell mass cell proliferation(GO:0001833) |
| 0.1 | 1.1 | GO:0097647 | calcitonin family receptor signaling pathway(GO:0097646) amylin receptor signaling pathway(GO:0097647) |
| 0.1 | 1.1 | GO:0061590 | calcium activated phospholipid scrambling(GO:0061588) calcium activated phosphatidylcholine scrambling(GO:0061590) calcium activated galactosylceramide scrambling(GO:0061591) |
| 0.1 | 1.0 | GO:0042424 | catechol-containing compound catabolic process(GO:0019614) catecholamine catabolic process(GO:0042424) |
| 0.1 | 0.8 | GO:0075525 | viral translational termination-reinitiation(GO:0075525) |
| 0.1 | 3.1 | GO:0043968 | histone H2A acetylation(GO:0043968) |
| 0.1 | 0.3 | GO:0055005 | ventricular cardiac myofibril assembly(GO:0055005) |
| 0.1 | 3.4 | GO:0035634 | response to stilbenoid(GO:0035634) |
| 0.1 | 1.0 | GO:0007549 | dosage compensation(GO:0007549) dosage compensation by inactivation of X chromosome(GO:0009048) |
| 0.1 | 1.3 | GO:0035372 | protein localization to microtubule(GO:0035372) |
| 0.1 | 1.7 | GO:0006337 | nucleosome disassembly(GO:0006337) |
| 0.1 | 9.6 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.1 | 0.9 | GO:0035330 | regulation of hippo signaling(GO:0035330) |
| 0.1 | 0.9 | GO:1903862 | positive regulation of oxidative phosphorylation(GO:1903862) |
| 0.1 | 0.6 | GO:0006686 | sphingomyelin biosynthetic process(GO:0006686) |
| 0.1 | 2.1 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 0.1 | 4.8 | GO:0006405 | RNA export from nucleus(GO:0006405) |
| 0.1 | 0.5 | GO:1904996 | positive regulation of leukocyte adhesion to vascular endothelial cell(GO:1904996) |
| 0.1 | 3.6 | GO:0033120 | positive regulation of RNA splicing(GO:0033120) |
| 0.1 | 2.0 | GO:0015991 | ATP hydrolysis coupled proton transport(GO:0015991) |
| 0.0 | 2.5 | GO:0001706 | endoderm formation(GO:0001706) |
| 0.0 | 0.2 | GO:1904565 | response to 1-oleoyl-sn-glycerol 3-phosphate(GO:1904565) cellular response to 1-oleoyl-sn-glycerol 3-phosphate(GO:1904566) |
| 0.0 | 6.5 | GO:0007586 | digestion(GO:0007586) |
| 0.0 | 1.5 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.0 | 2.1 | GO:0042273 | ribosomal large subunit biogenesis(GO:0042273) |
| 0.0 | 2.7 | GO:0010596 | negative regulation of endothelial cell migration(GO:0010596) |
| 0.0 | 1.0 | GO:0035025 | positive regulation of Rho protein signal transduction(GO:0035025) |
| 0.0 | 0.3 | GO:0042760 | very long-chain fatty acid catabolic process(GO:0042760) |
| 0.0 | 1.7 | GO:0015914 | phospholipid transport(GO:0015914) |
| 0.0 | 0.4 | GO:0030388 | fructose 1,6-bisphosphate metabolic process(GO:0030388) |
| 0.0 | 0.4 | GO:0090110 | cargo loading into COPII-coated vesicle(GO:0090110) |
| 0.0 | 0.3 | GO:0072378 | blood coagulation, fibrin clot formation(GO:0072378) |
| 0.0 | 0.8 | GO:0051560 | mitochondrial calcium ion homeostasis(GO:0051560) |
| 0.0 | 0.4 | GO:0030150 | protein import into mitochondrial matrix(GO:0030150) |
| 0.0 | 0.5 | GO:2000651 | positive regulation of sodium ion transmembrane transporter activity(GO:2000651) |
| 0.0 | 0.9 | GO:1903146 | regulation of mitophagy(GO:1903146) |
| 0.0 | 0.4 | GO:0019731 | antibacterial humoral response(GO:0019731) |
| 0.0 | 1.0 | GO:0051568 | histone H3-K4 methylation(GO:0051568) |
| 0.0 | 1.9 | GO:0032526 | response to retinoic acid(GO:0032526) |
| 0.0 | 0.8 | GO:0061003 | positive regulation of dendritic spine morphogenesis(GO:0061003) |
| 0.0 | 0.6 | GO:0006376 | mRNA splice site selection(GO:0006376) |
| 0.0 | 1.5 | GO:0019236 | response to pheromone(GO:0019236) |
| 0.0 | 0.8 | GO:0046426 | negative regulation of JAK-STAT cascade(GO:0046426) negative regulation of STAT cascade(GO:1904893) |
| 0.0 | 5.0 | GO:0002250 | adaptive immune response(GO:0002250) |
| 0.0 | 0.2 | GO:0000188 | inactivation of MAPK activity(GO:0000188) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 7.3 | GO:0044218 | other organism cell membrane(GO:0044218) other organism membrane(GO:0044279) |
| 0.8 | 4.2 | GO:0043625 | delta DNA polymerase complex(GO:0043625) |
| 0.7 | 5.2 | GO:0031095 | platelet dense tubular network membrane(GO:0031095) |
| 0.7 | 6.6 | GO:0019815 | B cell receptor complex(GO:0019815) |
| 0.5 | 5.9 | GO:0097451 | glial limiting end-foot(GO:0097451) |
| 0.4 | 3.6 | GO:0097504 | Gemini of coiled bodies(GO:0097504) |
| 0.4 | 2.8 | GO:0070381 | endosome to plasma membrane transport vesicle(GO:0070381) |
| 0.4 | 10.0 | GO:0034362 | low-density lipoprotein particle(GO:0034362) |
| 0.3 | 3.9 | GO:0005687 | U4 snRNP(GO:0005687) |
| 0.3 | 3.7 | GO:0030670 | phagocytic vesicle membrane(GO:0030670) |
| 0.3 | 4.2 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 0.3 | 1.9 | GO:0044611 | nuclear pore inner ring(GO:0044611) |
| 0.2 | 3.8 | GO:0042555 | MCM complex(GO:0042555) |
| 0.2 | 3.0 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 0.2 | 1.5 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 0.2 | 1.8 | GO:0072379 | BAT3 complex(GO:0071818) ER membrane insertion complex(GO:0072379) |
| 0.2 | 0.8 | GO:0036019 | endolysosome(GO:0036019) |
| 0.2 | 2.7 | GO:0000138 | Golgi trans cisterna(GO:0000138) |
| 0.2 | 4.7 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.2 | 2.5 | GO:0043219 | lateral loop(GO:0043219) |
| 0.1 | 1.5 | GO:0072687 | meiotic spindle(GO:0072687) |
| 0.1 | 18.8 | GO:0014704 | intercalated disc(GO:0014704) |
| 0.1 | 2.0 | GO:0033179 | proton-transporting V-type ATPase, V0 domain(GO:0033179) |
| 0.1 | 1.3 | GO:0034992 | microtubule organizing center attachment site(GO:0034992) LINC complex(GO:0034993) |
| 0.1 | 5.1 | GO:0008305 | integrin complex(GO:0008305) |
| 0.1 | 1.3 | GO:0098643 | fibrillar collagen trimer(GO:0005583) banded collagen fibril(GO:0098643) |
| 0.1 | 0.6 | GO:0044530 | supraspliceosomal complex(GO:0044530) |
| 0.1 | 9.5 | GO:0019814 | immunoglobulin complex(GO:0019814) immunoglobulin complex, circulating(GO:0042571) |
| 0.1 | 0.8 | GO:0071541 | eukaryotic translation initiation factor 3 complex, eIF3m(GO:0071541) |
| 0.1 | 0.7 | GO:0072669 | tRNA-splicing ligase complex(GO:0072669) |
| 0.1 | 2.6 | GO:0001741 | XY body(GO:0001741) |
| 0.1 | 5.7 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.1 | 1.0 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.1 | 4.2 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.0 | 3.0 | GO:0008023 | transcription elongation factor complex(GO:0008023) |
| 0.0 | 0.2 | GO:0016514 | SWI/SNF complex(GO:0016514) |
| 0.0 | 0.7 | GO:0031045 | dense core granule(GO:0031045) |
| 0.0 | 0.3 | GO:0031933 | telomeric heterochromatin(GO:0031933) |
| 0.0 | 10.7 | GO:0031965 | nuclear membrane(GO:0031965) |
| 0.0 | 2.1 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.0 | 0.3 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.0 | 2.1 | GO:0031519 | PcG protein complex(GO:0031519) |
| 0.0 | 4.0 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 0.4 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.0 | 1.5 | GO:0032592 | integral component of mitochondrial membrane(GO:0032592) |
| 0.0 | 2.0 | GO:0005901 | caveola(GO:0005901) |
| 0.0 | 9.8 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.0 | 0.1 | GO:0000221 | vacuolar proton-transporting V-type ATPase, V1 domain(GO:0000221) |
| 0.0 | 1.5 | GO:0000313 | organellar ribosome(GO:0000313) mitochondrial ribosome(GO:0005761) |
| 0.0 | 1.7 | GO:0071013 | catalytic step 2 spliceosome(GO:0071013) |
| 0.0 | 0.2 | GO:0035327 | transcriptionally active chromatin(GO:0035327) |
| 0.0 | 3.0 | GO:0044306 | neuron projection terminus(GO:0044306) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.3 | 7.0 | GO:0052895 | norspermine:oxygen oxidoreductase activity(GO:0052894) N1-acetylspermine:oxygen oxidoreductase (N1-acetylspermidine-forming) activity(GO:0052895) |
| 2.1 | 19.0 | GO:0004032 | alditol:NADP+ 1-oxidoreductase activity(GO:0004032) |
| 1.8 | 16.0 | GO:0070891 | lipoteichoic acid binding(GO:0070891) |
| 1.5 | 10.8 | GO:0004556 | alpha-amylase activity(GO:0004556) |
| 1.5 | 9.1 | GO:0047844 | deoxycytidine deaminase activity(GO:0047844) |
| 1.0 | 10.0 | GO:0047499 | calcium-independent phospholipase A2 activity(GO:0047499) |
| 0.9 | 18.8 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.8 | 2.5 | GO:0001042 | RNA polymerase I core binding(GO:0001042) |
| 0.8 | 3.9 | GO:0030621 | U4 snRNA binding(GO:0030621) |
| 0.7 | 3.0 | GO:0031727 | CCR2 chemokine receptor binding(GO:0031727) |
| 0.7 | 5.1 | GO:0001851 | complement component C3b binding(GO:0001851) |
| 0.7 | 4.3 | GO:0004852 | uroporphyrinogen-III synthase activity(GO:0004852) |
| 0.6 | 12.8 | GO:0022840 | leak channel activity(GO:0022840) narrow pore channel activity(GO:0022842) |
| 0.5 | 1.6 | GO:0015389 | pyrimidine- and adenine-specific:sodium symporter activity(GO:0015389) |
| 0.5 | 2.1 | GO:1990932 | 5.8S rRNA binding(GO:1990932) |
| 0.5 | 2.0 | GO:0034617 | tetrahydrobiopterin binding(GO:0034617) |
| 0.5 | 1.4 | GO:0046969 | histone deacetylase activity (H3-K9 specific)(GO:0032129) NAD-dependent histone deacetylase activity (H3-K9 specific)(GO:0046969) |
| 0.5 | 3.7 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.4 | 1.5 | GO:0070976 | TIR domain binding(GO:0070976) |
| 0.3 | 2.1 | GO:0003835 | beta-galactoside alpha-2,6-sialyltransferase activity(GO:0003835) |
| 0.3 | 7.3 | GO:0001530 | lipopolysaccharide binding(GO:0001530) |
| 0.3 | 4.2 | GO:0016175 | superoxide-generating NADPH oxidase activity(GO:0016175) superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 0.3 | 1.0 | GO:0016206 | catechol O-methyltransferase activity(GO:0016206) |
| 0.3 | 1.8 | GO:0045322 | unmethylated CpG binding(GO:0045322) |
| 0.3 | 1.7 | GO:1990050 | phosphatidic acid transporter activity(GO:1990050) |
| 0.3 | 2.6 | GO:0003873 | 6-phosphofructo-2-kinase activity(GO:0003873) |
| 0.2 | 4.2 | GO:0008296 | 3'-5'-exodeoxyribonuclease activity(GO:0008296) |
| 0.2 | 0.8 | GO:0035673 | oligopeptide transmembrane transporter activity(GO:0035673) |
| 0.2 | 3.6 | GO:0016918 | retinal binding(GO:0016918) retinol binding(GO:0019841) |
| 0.2 | 2.8 | GO:0032050 | clathrin heavy chain binding(GO:0032050) |
| 0.2 | 0.5 | GO:0004731 | purine-nucleoside phosphorylase activity(GO:0004731) |
| 0.2 | 0.7 | GO:0004909 | interleukin-1, Type I, activating receptor activity(GO:0004909) |
| 0.2 | 1.1 | GO:0004027 | alcohol sulfotransferase activity(GO:0004027) |
| 0.2 | 2.5 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.2 | 4.9 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.1 | 16.2 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.1 | 2.1 | GO:0051864 | histone demethylase activity (H3-K36 specific)(GO:0051864) |
| 0.1 | 4.8 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 0.1 | 1.5 | GO:0019869 | chloride channel inhibitor activity(GO:0019869) |
| 0.1 | 1.1 | GO:0004468 | lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0004468) |
| 0.1 | 0.6 | GO:0033188 | sphingomyelin synthase activity(GO:0033188) ceramide cholinephosphotransferase activity(GO:0047493) |
| 0.1 | 0.2 | GO:0033549 | MAP kinase phosphatase activity(GO:0033549) |
| 0.1 | 3.2 | GO:0005031 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.1 | 3.8 | GO:0004003 | ATP-dependent DNA helicase activity(GO:0004003) |
| 0.1 | 3.6 | GO:0017134 | fibroblast growth factor binding(GO:0017134) |
| 0.1 | 1.3 | GO:0048407 | platelet-derived growth factor binding(GO:0048407) |
| 0.1 | 5.9 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.1 | 1.3 | GO:0043495 | protein anchor(GO:0043495) |
| 0.1 | 3.7 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.1 | 5.0 | GO:0004712 | protein serine/threonine/tyrosine kinase activity(GO:0004712) |
| 0.1 | 1.9 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.1 | 7.3 | GO:0043621 | protein self-association(GO:0043621) |
| 0.1 | 2.0 | GO:0046961 | proton-transporting ATPase activity, rotational mechanism(GO:0046961) |
| 0.1 | 3.0 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.1 | 0.2 | GO:0010698 | acetyltransferase activator activity(GO:0010698) |
| 0.1 | 3.7 | GO:0004197 | cysteine-type endopeptidase activity(GO:0004197) |
| 0.0 | 0.4 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 0.0 | 1.3 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.0 | 1.1 | GO:0017128 | phospholipid scramblase activity(GO:0017128) |
| 0.0 | 5.3 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 1.9 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.0 | 0.7 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.0 | 0.2 | GO:0035727 | lysophosphatidic acid binding(GO:0035727) |
| 0.0 | 0.8 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.0 | 3.2 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 1.3 | GO:0005550 | pheromone binding(GO:0005550) |
| 0.0 | 0.8 | GO:0035035 | histone acetyltransferase binding(GO:0035035) |
| 0.0 | 0.1 | GO:0019958 | C-X-C chemokine binding(GO:0019958) |
| 0.0 | 0.3 | GO:0016290 | palmitoyl-CoA hydrolase activity(GO:0016290) |
| 0.0 | 1.2 | GO:0003823 | antigen binding(GO:0003823) |
| 0.0 | 0.2 | GO:0001206 | transcriptional repressor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001206) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 9.8 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.2 | 5.4 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.2 | 3.5 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 0.1 | 2.8 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.1 | 0.9 | PID ERBB2 ERBB3 PATHWAY | ErbB2/ErbB3 signaling events |
| 0.1 | 1.5 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.1 | 2.5 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.1 | 3.1 | PID MYC PATHWAY | C-MYC pathway |
| 0.1 | 3.6 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
| 0.1 | 2.0 | PID VEGFR1 PATHWAY | VEGFR1 specific signals |
| 0.0 | 2.2 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.0 | 4.2 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.0 | 1.3 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.0 | 1.3 | PID SYNDECAN 1 PATHWAY | Syndecan-1-mediated signaling events |
| 0.0 | 0.6 | PID IL2 PI3K PATHWAY | IL2 signaling events mediated by PI3K |
| 0.0 | 1.6 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.0 | 1.4 | PID HDAC CLASSI PATHWAY | Signaling events mediated by HDAC Class I |
| 0.0 | 2.8 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 10.0 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.4 | 7.0 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.3 | 12.8 | REACTOME AMINE COMPOUND SLC TRANSPORTERS | Genes involved in Amine compound SLC transporters |
| 0.3 | 4.2 | REACTOME REMOVAL OF THE FLAP INTERMEDIATE FROM THE C STRAND | Genes involved in Removal of the Flap Intermediate from the C-strand |
| 0.3 | 13.6 | REACTOME LATENT INFECTION OF HOMO SAPIENS WITH MYCOBACTERIUM TUBERCULOSIS | Genes involved in Latent infection of Homo sapiens with Mycobacterium tuberculosis |
| 0.2 | 3.8 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 0.2 | 1.5 | REACTOME PROCESSIVE SYNTHESIS ON THE LAGGING STRAND | Genes involved in Processive synthesis on the lagging strand |
| 0.2 | 4.3 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.2 | 2.3 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION IN TLR7 8 OR 9 SIGNALING | Genes involved in TRAF6 mediated IRF7 activation in TLR7/8 or 9 signaling |
| 0.1 | 18.8 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.1 | 3.5 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.1 | 6.6 | REACTOME METABOLISM OF NON CODING RNA | Genes involved in Metabolism of non-coding RNA |
| 0.1 | 2.1 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.1 | 2.3 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.1 | 2.5 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.1 | 3.0 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.1 | 7.3 | REACTOME CELL SURFACE INTERACTIONS AT THE VASCULAR WALL | Genes involved in Cell surface interactions at the vascular wall |
| 0.0 | 0.8 | REACTOME REGULATION OF KIT SIGNALING | Genes involved in Regulation of KIT signaling |
| 0.0 | 0.8 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.0 | 5.1 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.0 | 3.0 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.0 | 1.3 | REACTOME PIP3 ACTIVATES AKT SIGNALING | Genes involved in PIP3 activates AKT signaling |
| 0.0 | 0.5 | REACTOME PURINE CATABOLISM | Genes involved in Purine catabolism |
| 0.0 | 1.4 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.0 | 0.3 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.0 | 1.3 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.0 | 1.8 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.0 | 0.2 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.0 | 0.7 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.0 | 0.6 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 0.3 | REACTOME ERK MAPK TARGETS | Genes involved in ERK/MAPK targets |