Project
GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
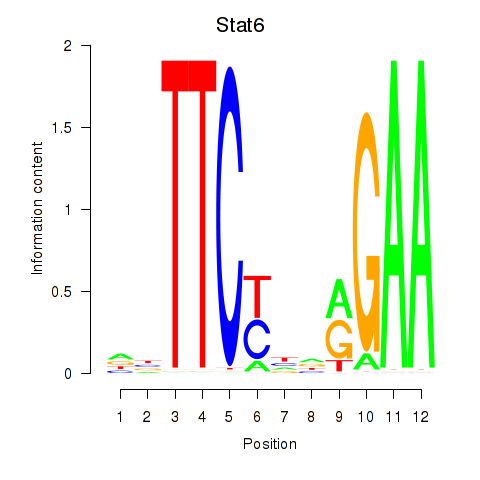
Results for Stat6
Z-value: 0.86
Transcription factors associated with Stat6
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Stat6
|
ENSMUSG00000002147.19 | signal transducer and activator of transcription 6 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Stat6 | mm39_v1_chr10_+_127478844_127478914 | -0.02 | 8.9e-01 | Click! |
Activity profile of Stat6 motif
Sorted Z-values of Stat6 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr11_+_87684299 | 8.31 |
ENSMUST00000020779.11
|
Mpo
|
myeloperoxidase |
| chr9_-_70328816 | 8.06 |
ENSMUST00000034742.8
|
Ccnb2
|
cyclin B2 |
| chr11_+_87684548 | 6.59 |
ENSMUST00000143021.9
|
Mpo
|
myeloperoxidase |
| chr4_+_120523758 | 5.51 |
ENSMUST00000094814.6
|
Cited4
|
Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 4 |
| chr15_-_82128888 | 4.10 |
ENSMUST00000089155.6
ENSMUST00000089157.11 |
Cenpm
|
centromere protein M |
| chr15_-_82128538 | 3.81 |
ENSMUST00000229747.2
ENSMUST00000230408.2 |
Cenpm
|
centromere protein M |
| chr14_-_56499690 | 3.80 |
ENSMUST00000015581.6
|
Gzmb
|
granzyme B |
| chr17_+_48607405 | 3.65 |
ENSMUST00000170941.3
|
Treml2
|
triggering receptor expressed on myeloid cells-like 2 |
| chr17_+_48606948 | 3.49 |
ENSMUST00000233092.2
|
Treml2
|
triggering receptor expressed on myeloid cells-like 2 |
| chr7_+_126690525 | 3.30 |
ENSMUST00000056288.7
ENSMUST00000206102.2 |
AI467606
|
expressed sequence AI467606 |
| chr18_+_50186349 | 3.22 |
ENSMUST00000148159.3
|
Tnfaip8
|
tumor necrosis factor, alpha-induced protein 8 |
| chr1_+_134110142 | 3.18 |
ENSMUST00000082060.10
ENSMUST00000153856.8 ENSMUST00000133701.8 ENSMUST00000132873.8 |
Chil1
|
chitinase-like 1 |
| chrX_-_47551990 | 3.17 |
ENSMUST00000033429.9
ENSMUST00000140486.2 |
Elf4
|
E74-like factor 4 (ets domain transcription factor) |
| chr5_+_114924106 | 2.94 |
ENSMUST00000137519.2
|
Ankrd13a
|
ankyrin repeat domain 13a |
| chr2_+_24235300 | 2.90 |
ENSMUST00000114485.9
ENSMUST00000114482.3 |
Il1rn
|
interleukin 1 receptor antagonist |
| chr7_-_126399574 | 2.74 |
ENSMUST00000106348.8
|
Aldoa
|
aldolase A, fructose-bisphosphate |
| chr1_+_134109888 | 2.74 |
ENSMUST00000156873.8
|
Chil1
|
chitinase-like 1 |
| chr2_-_113678999 | 2.72 |
ENSMUST00000102545.8
ENSMUST00000110948.8 |
Arhgap11a
|
Rho GTPase activating protein 11A |
| chr14_+_26722319 | 2.39 |
ENSMUST00000035433.10
|
Hesx1
|
homeobox gene expressed in ES cells |
| chr1_+_171594690 | 2.33 |
ENSMUST00000015460.5
|
Slamf1
|
signaling lymphocytic activation molecule family member 1 |
| chr3_+_96939732 | 2.12 |
ENSMUST00000132256.8
ENSMUST00000072600.7 |
Gja5
|
gap junction protein, alpha 5 |
| chr7_-_126399208 | 2.09 |
ENSMUST00000133514.8
ENSMUST00000151137.8 |
Aldoa
|
aldolase A, fructose-bisphosphate |
| chr1_-_23948764 | 2.09 |
ENSMUST00000129254.8
|
Smap1
|
small ArfGAP 1 |
| chr7_-_126399778 | 2.02 |
ENSMUST00000141355.4
|
Aldoa
|
aldolase A, fructose-bisphosphate |
| chr17_-_90217868 | 1.99 |
ENSMUST00000086423.6
|
Gm10184
|
predicted pseudogene 10184 |
| chr10_+_97315465 | 1.97 |
ENSMUST00000105287.11
|
Dcn
|
decorin |
| chr11_-_83421333 | 1.88 |
ENSMUST00000035938.3
|
Ccl5
|
chemokine (C-C motif) ligand 5 |
| chr19_+_10819896 | 1.88 |
ENSMUST00000025646.3
|
Slc15a3
|
solute carrier family 15, member 3 |
| chr6_+_8520006 | 1.85 |
ENSMUST00000162567.8
ENSMUST00000161217.8 |
Glcci1
|
glucocorticoid induced transcript 1 |
| chr5_-_148329615 | 1.83 |
ENSMUST00000138257.8
|
Slc7a1
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 1 |
| chr11_+_78079243 | 1.82 |
ENSMUST00000002128.14
ENSMUST00000150941.8 |
Rab34
|
RAB34, member RAS oncogene family |
| chr17_+_29309942 | 1.72 |
ENSMUST00000119901.9
|
Cdkn1a
|
cyclin-dependent kinase inhibitor 1A (P21) |
| chr11_+_75422516 | 1.71 |
ENSMUST00000149727.8
ENSMUST00000108433.8 ENSMUST00000042561.14 ENSMUST00000143035.8 |
Slc43a2
|
solute carrier family 43, member 2 |
| chr1_-_181670599 | 1.69 |
ENSMUST00000193030.6
|
Lbr
|
lamin B receptor |
| chr2_+_120331693 | 1.59 |
ENSMUST00000141181.9
|
Capn3
|
calpain 3 |
| chr16_-_22084700 | 1.51 |
ENSMUST00000161286.8
|
Tra2b
|
transformer 2 beta |
| chr19_-_5974825 | 1.48 |
ENSMUST00000055458.6
|
Cdc42ep2
|
CDC42 effector protein (Rho GTPase binding) 2 |
| chr11_-_78074377 | 1.48 |
ENSMUST00000102483.5
|
Rpl23a
|
ribosomal protein L23A |
| chr11_-_55310724 | 1.45 |
ENSMUST00000108858.8
ENSMUST00000141530.2 |
Sparc
|
secreted acidic cysteine rich glycoprotein |
| chr3_+_90383425 | 1.45 |
ENSMUST00000001042.10
|
Ilf2
|
interleukin enhancer binding factor 2 |
| chr3_-_96201248 | 1.44 |
ENSMUST00000029748.8
|
Fcgr1
|
Fc receptor, IgG, high affinity I |
| chr1_+_130659700 | 1.43 |
ENSMUST00000039323.8
|
AA986860
|
expressed sequence AA986860 |
| chr10_-_128462616 | 1.41 |
ENSMUST00000026420.7
|
Rps26
|
ribosomal protein S26 |
| chr16_+_97157934 | 1.36 |
ENSMUST00000047275.8
|
Bace2
|
beta-site APP-cleaving enzyme 2 |
| chr12_-_15866763 | 1.35 |
ENSMUST00000020922.8
ENSMUST00000221215.2 ENSMUST00000221518.2 |
Trib2
|
tribbles pseudokinase 2 |
| chr8_+_106331866 | 1.34 |
ENSMUST00000043531.10
|
Ripor1
|
RHO family interacting cell polarization regulator 1 |
| chr16_-_23709564 | 1.34 |
ENSMUST00000004480.5
|
Sst
|
somatostatin |
| chr2_-_148285450 | 1.33 |
ENSMUST00000099269.4
|
Cd93
|
CD93 antigen |
| chr6_+_57679621 | 1.30 |
ENSMUST00000050077.15
|
Lancl2
|
LanC (bacterial lantibiotic synthetase component C)-like 2 |
| chr2_+_29236815 | 1.27 |
ENSMUST00000028139.11
ENSMUST00000113830.11 |
Med27
|
mediator complex subunit 27 |
| chr4_-_116982804 | 1.26 |
ENSMUST00000183310.2
|
Btbd19
|
BTB (POZ) domain containing 19 |
| chr2_+_120331784 | 1.25 |
ENSMUST00000151342.3
|
Capn3
|
calpain 3 |
| chr2_-_113678945 | 1.25 |
ENSMUST00000110949.9
|
Arhgap11a
|
Rho GTPase activating protein 11A |
| chr4_+_116565784 | 1.24 |
ENSMUST00000138305.8
ENSMUST00000125671.8 ENSMUST00000130828.8 |
Ccdc163
|
coiled-coil domain containing 163 |
| chrX_+_162694397 | 1.24 |
ENSMUST00000140845.2
|
Ap1s2
|
adaptor-related protein complex 1, sigma 2 subunit |
| chr11_+_75422925 | 1.20 |
ENSMUST00000169547.9
|
Slc43a2
|
solute carrier family 43, member 2 |
| chr2_-_14060774 | 1.16 |
ENSMUST00000114753.8
ENSMUST00000091429.12 |
Hacd1
|
3-hydroxyacyl-CoA dehydratase 1 |
| chr15_+_6451721 | 1.13 |
ENSMUST00000163082.2
|
Dab2
|
disabled 2, mitogen-responsive phosphoprotein |
| chr4_+_116565706 | 1.13 |
ENSMUST00000030452.13
ENSMUST00000106462.9 |
Ccdc163
|
coiled-coil domain containing 163 |
| chr6_+_57679455 | 1.13 |
ENSMUST00000072954.8
|
Lancl2
|
LanC (bacterial lantibiotic synthetase component C)-like 2 |
| chr2_-_14060840 | 1.11 |
ENSMUST00000074854.9
|
Hacd1
|
3-hydroxyacyl-CoA dehydratase 1 |
| chr7_+_28050077 | 1.09 |
ENSMUST00000082134.6
|
Rps16
|
ribosomal protein S16 |
| chr18_+_42669322 | 1.06 |
ENSMUST00000236418.2
|
Tcerg1
|
transcription elongation regulator 1 (CA150) |
| chr11_+_68393845 | 1.04 |
ENSMUST00000102613.8
ENSMUST00000060441.7 |
Pik3r6
|
phosphoinositide-3-kinase regulatory subunit 5 |
| chr2_+_31578537 | 1.03 |
ENSMUST00000075759.13
|
Abl1
|
c-abl oncogene 1, non-receptor tyrosine kinase |
| chr6_-_87510200 | 1.03 |
ENSMUST00000113637.9
ENSMUST00000071024.7 |
Arhgap25
|
Rho GTPase activating protein 25 |
| chr1_+_171668173 | 1.02 |
ENSMUST00000136479.8
|
Cd84
|
CD84 antigen |
| chr9_+_86625694 | 1.02 |
ENSMUST00000179574.2
ENSMUST00000036426.13 |
Prss35
|
protease, serine 35 |
| chr5_+_31212165 | 1.00 |
ENSMUST00000202795.4
ENSMUST00000201182.4 ENSMUST00000200953.4 |
Cad
|
carbamoyl-phosphate synthetase 2, aspartate transcarbamylase, and dihydroorotase |
| chrX_+_133657312 | 0.99 |
ENSMUST00000081834.10
ENSMUST00000086880.11 ENSMUST00000086884.5 |
Armcx3
|
armadillo repeat containing, X-linked 3 |
| chr11_+_101136821 | 0.98 |
ENSMUST00000129680.8
|
Ramp2
|
receptor (calcitonin) activity modifying protein 2 |
| chr4_+_130202388 | 0.98 |
ENSMUST00000070532.8
|
Fabp3
|
fatty acid binding protein 3, muscle and heart |
| chr13_-_41513215 | 0.97 |
ENSMUST00000224803.2
|
Nedd9
|
neural precursor cell expressed, developmentally down-regulated gene 9 |
| chr19_+_12647803 | 0.97 |
ENSMUST00000207341.3
ENSMUST00000208494.3 ENSMUST00000208657.3 |
Olfr1442
|
olfactory receptor 1442 |
| chr13_+_43938251 | 0.95 |
ENSMUST00000015540.4
|
Cd83
|
CD83 antigen |
| chr14_-_56472102 | 0.94 |
ENSMUST00000015585.4
|
Gzmc
|
granzyme C |
| chr5_+_31212110 | 0.93 |
ENSMUST00000013773.12
ENSMUST00000201838.4 |
Cad
|
carbamoyl-phosphate synthetase 2, aspartate transcarbamylase, and dihydroorotase |
| chr5_-_140368482 | 0.92 |
ENSMUST00000196566.5
|
Snx8
|
sorting nexin 8 |
| chr4_+_116565819 | 0.90 |
ENSMUST00000106463.8
|
Ccdc163
|
coiled-coil domain containing 163 |
| chr10_+_80905869 | 0.88 |
ENSMUST00000005057.7
|
Thop1
|
thimet oligopeptidase 1 |
| chr10_-_79940168 | 0.86 |
ENSMUST00000219260.2
|
Sbno2
|
strawberry notch 2 |
| chr14_-_22039543 | 0.85 |
ENSMUST00000043409.9
|
Zfp503
|
zinc finger protein 503 |
| chr1_-_44157916 | 0.84 |
ENSMUST00000027213.14
ENSMUST00000065767.9 |
Poglut2
|
protein O-glucosyltransferase 2 |
| chr17_-_7639378 | 0.82 |
ENSMUST00000231397.2
|
Gm49630
|
predicted gene, 49630 |
| chr16_-_56533179 | 0.81 |
ENSMUST00000136394.8
|
Tfg
|
Trk-fused gene |
| chr8_-_85414528 | 0.81 |
ENSMUST00000001975.6
|
Nacc1
|
nucleus accumbens associated 1, BEN and BTB (POZ) domain containing |
| chr2_-_73284262 | 0.80 |
ENSMUST00000102679.8
|
Wipf1
|
WAS/WASL interacting protein family, member 1 |
| chr3_+_153549846 | 0.79 |
ENSMUST00000044089.4
|
Asb17
|
ankyrin repeat and SOCS box-containing 17 |
| chr3_+_14706781 | 0.71 |
ENSMUST00000029071.9
|
Car13
|
carbonic anhydrase 13 |
| chr15_-_81810349 | 0.71 |
ENSMUST00000023113.7
|
Polr3h
|
polymerase (RNA) III (DNA directed) polypeptide H |
| chr6_-_101354858 | 0.70 |
ENSMUST00000075994.11
|
Pdzrn3
|
PDZ domain containing RING finger 3 |
| chr12_-_57244096 | 0.70 |
ENSMUST00000044634.12
|
Slc25a21
|
solute carrier family 25 (mitochondrial oxodicarboxylate carrier), member 21 |
| chr6_-_141719536 | 0.68 |
ENSMUST00000148411.2
|
Gm5724
|
predicted gene 5724 |
| chr7_+_44465353 | 0.68 |
ENSMUST00000208626.2
ENSMUST00000057195.17 |
Nup62
|
nucleoporin 62 |
| chr15_-_93417380 | 0.68 |
ENSMUST00000109255.3
|
Prickle1
|
prickle planar cell polarity protein 1 |
| chr5_-_5315968 | 0.67 |
ENSMUST00000115451.8
ENSMUST00000115452.8 ENSMUST00000131392.8 |
Cdk14
|
cyclin-dependent kinase 14 |
| chr9_-_49710190 | 0.67 |
ENSMUST00000114476.8
ENSMUST00000193547.6 |
Ncam1
|
neural cell adhesion molecule 1 |
| chr19_+_29229147 | 0.67 |
ENSMUST00000025705.7
ENSMUST00000065796.10 ENSMUST00000236990.2 |
Jak2
|
Janus kinase 2 |
| chr9_+_53757448 | 0.66 |
ENSMUST00000048485.7
|
Sln
|
sarcolipin |
| chr11_+_96242422 | 0.64 |
ENSMUST00000100523.7
|
Hoxb2
|
homeobox B2 |
| chr14_-_104760051 | 0.63 |
ENSMUST00000022716.4
ENSMUST00000228448.2 ENSMUST00000227640.2 |
Obi1
|
ORC ubiquitin ligase 1 |
| chr7_+_44465806 | 0.62 |
ENSMUST00000207103.2
ENSMUST00000118125.9 |
Nup62
Il4i1
|
nucleoporin 62 interleukin 4 induced 1 |
| chr18_-_38417390 | 0.62 |
ENSMUST00000025311.8
|
Pcdh12
|
protocadherin 12 |
| chr19_-_8690330 | 0.57 |
ENSMUST00000206598.2
|
Slc3a2
|
solute carrier family 3 (activators of dibasic and neutral amino acid transport), member 2 |
| chr7_+_80764564 | 0.57 |
ENSMUST00000119083.2
|
Slc28a1
|
solute carrier family 28 (sodium-coupled nucleoside transporter), member 1 |
| chr14_-_55950939 | 0.56 |
ENSMUST00000168729.8
ENSMUST00000228123.2 ENSMUST00000178034.9 |
Tgm1
|
transglutaminase 1, K polypeptide |
| chr1_+_88154727 | 0.55 |
ENSMUST00000061013.13
ENSMUST00000113130.8 |
Mroh2a
|
maestro heat-like repeat family member 2A |
| chr5_+_117501557 | 0.55 |
ENSMUST00000111959.2
|
Wsb2
|
WD repeat and SOCS box-containing 2 |
| chr18_-_15536747 | 0.55 |
ENSMUST00000079081.8
|
Aqp4
|
aquaporin 4 |
| chr4_-_149747644 | 0.54 |
ENSMUST00000105689.8
|
Pik3cd
|
phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit delta |
| chr7_+_80764547 | 0.54 |
ENSMUST00000026820.11
|
Slc28a1
|
solute carrier family 28 (sodium-coupled nucleoside transporter), member 1 |
| chr18_-_36859732 | 0.53 |
ENSMUST00000061829.8
|
Cd14
|
CD14 antigen |
| chr6_+_48695535 | 0.53 |
ENSMUST00000127537.6
ENSMUST00000052503.8 |
Gm28053
Gimap7
|
predicted gene, 28053 GTPase, IMAP family member 7 |
| chr8_-_106427696 | 0.51 |
ENSMUST00000042608.8
|
Acd
|
adrenocortical dysplasia |
| chr7_+_44465714 | 0.47 |
ENSMUST00000208172.2
|
Nup62
|
nucleoporin 62 |
| chr18_+_32970278 | 0.46 |
ENSMUST00000053663.11
|
Wdr36
|
WD repeat domain 36 |
| chr3_-_8988854 | 0.46 |
ENSMUST00000042148.6
|
Mrps28
|
mitochondrial ribosomal protein S28 |
| chr9_-_49710058 | 0.46 |
ENSMUST00000192584.2
ENSMUST00000166811.9 |
Ncam1
|
neural cell adhesion molecule 1 |
| chr7_+_88079534 | 0.43 |
ENSMUST00000208478.2
|
Rab38
|
RAB38, member RAS oncogene family |
| chr5_+_21577640 | 0.42 |
ENSMUST00000035799.6
|
Fgl2
|
fibrinogen-like protein 2 |
| chr18_-_38417444 | 0.41 |
ENSMUST00000194012.2
|
Pcdh12
|
protocadherin 12 |
| chr1_+_120048890 | 0.41 |
ENSMUST00000027637.13
ENSMUST00000112644.9 ENSMUST00000056038.15 |
3110009E18Rik
|
RIKEN cDNA 3110009E18 gene |
| chr18_+_32970363 | 0.39 |
ENSMUST00000166214.9
|
Wdr36
|
WD repeat domain 36 |
| chr11_+_76297969 | 0.38 |
ENSMUST00000021203.7
ENSMUST00000152183.2 |
Timm22
|
translocase of inner mitochondrial membrane 22 |
| chr9_+_74959259 | 0.38 |
ENSMUST00000170310.2
ENSMUST00000166549.2 |
Arpp19
|
cAMP-regulated phosphoprotein 19 |
| chr5_+_8710059 | 0.36 |
ENSMUST00000047753.5
|
Abcb1a
|
ATP-binding cassette, sub-family B (MDR/TAP), member 1A |
| chr8_-_85414220 | 0.34 |
ENSMUST00000238449.2
ENSMUST00000238687.2 |
Nacc1
|
nucleus accumbens associated 1, BEN and BTB (POZ) domain containing |
| chr3_-_116601451 | 0.31 |
ENSMUST00000159670.3
|
Agl
|
amylo-1,6-glucosidase, 4-alpha-glucanotransferase |
| chr3_+_67799510 | 0.30 |
ENSMUST00000063263.5
ENSMUST00000182006.4 |
Iqcj
Iqschfp
|
IQ motif containing J Iqcj and Schip1 fusion protein |
| chr6_+_127430668 | 0.30 |
ENSMUST00000039680.7
|
Parp11
|
poly (ADP-ribose) polymerase family, member 11 |
| chr14_+_3576275 | 0.30 |
ENSMUST00000151926.8
|
Ube2e2
|
ubiquitin-conjugating enzyme E2E 2 |
| chr11_+_75422953 | 0.30 |
ENSMUST00000127226.3
|
Slc43a2
|
solute carrier family 43, member 2 |
| chr19_+_3818112 | 0.29 |
ENSMUST00000005518.16
ENSMUST00000237440.2 ENSMUST00000152935.8 ENSMUST00000176262.8 ENSMUST00000176407.8 ENSMUST00000176926.8 ENSMUST00000176512.8 |
Kmt5b
|
lysine methyltransferase 5B |
| chr7_+_127160751 | 0.28 |
ENSMUST00000190278.3
|
Tmem265
|
transmembrane protein 265 |
| chr9_+_38288382 | 0.28 |
ENSMUST00000214865.2
|
Olfr251
|
olfactory receptor 251 |
| chr6_+_149226891 | 0.26 |
ENSMUST00000189837.2
|
Resf1
|
retroelement silencing factor 1 |
| chr5_-_6926523 | 0.26 |
ENSMUST00000164784.2
|
Zfp804b
|
zinc finger protein 804B |
| chr9_+_100956734 | 0.25 |
ENSMUST00000085177.5
|
Msl2
|
MSL complex subunit 2 |
| chr10_-_100425067 | 0.23 |
ENSMUST00000218821.2
ENSMUST00000054471.10 |
4930430F08Rik
|
RIKEN cDNA 4930430F08 gene |
| chr2_+_118730823 | 0.22 |
ENSMUST00000151162.2
|
Bahd1
|
bromo adjacent homology domain containing 1 |
| chr1_-_160134873 | 0.21 |
ENSMUST00000193185.6
|
Rabgap1l
|
RAB GTPase activating protein 1-like |
| chr18_+_24338729 | 0.19 |
ENSMUST00000170243.8
|
Galnt1
|
polypeptide N-acetylgalactosaminyltransferase 1 |
| chr14_+_76082533 | 0.18 |
ENSMUST00000110894.9
|
Tpt1
|
tumor protein, translationally-controlled 1 |
| chr5_+_90708962 | 0.18 |
ENSMUST00000094615.8
ENSMUST00000200765.2 |
Albfm1
|
albumin superfamily member 1 |
| chr4_+_116565898 | 0.18 |
ENSMUST00000135499.8
|
Ccdc163
|
coiled-coil domain containing 163 |
| chr10_-_51507527 | 0.18 |
ENSMUST00000219286.2
ENSMUST00000020062.4 ENSMUST00000218684.2 |
Gprc6a
|
G protein-coupled receptor, family C, group 6, member A |
| chr5_-_125201872 | 0.18 |
ENSMUST00000055256.14
|
Ncor2
|
nuclear receptor co-repressor 2 |
| chr6_+_38358404 | 0.18 |
ENSMUST00000162554.8
ENSMUST00000161751.8 |
Ttc26
|
tetratricopeptide repeat domain 26 |
| chr4_-_43454600 | 0.17 |
ENSMUST00000098105.4
ENSMUST00000098104.10 ENSMUST00000030179.11 |
Cd72
|
CD72 antigen |
| chr13_-_77283534 | 0.17 |
ENSMUST00000159462.3
ENSMUST00000151524.9 |
Slf1
|
SMC5-SMC6 complex localization factor 1 |
| chr14_+_61844899 | 0.16 |
ENSMUST00000225582.2
ENSMUST00000051184.10 |
Kcnrg
|
potassium channel regulator |
| chr2_-_87466089 | 0.15 |
ENSMUST00000090711.3
|
Olfr1132
|
olfactory receptor 1132 |
| chr11_+_108477903 | 0.15 |
ENSMUST00000146912.9
|
Cep112
|
centrosomal protein 112 |
| chr11_+_83743746 | 0.15 |
ENSMUST00000108113.3
|
Hnf1b
|
HNF1 homeobox B |
| chr18_+_37827413 | 0.15 |
ENSMUST00000193414.2
|
Pcdhga5
|
protocadherin gamma subfamily A, 5 |
| chr15_+_51741138 | 0.14 |
ENSMUST00000136129.2
|
Utp23
|
UTP23 small subunit processome component |
| chr7_-_44465998 | 0.13 |
ENSMUST00000209072.2
ENSMUST00000047356.11 |
Atf5
|
activating transcription factor 5 |
| chr6_+_34840057 | 0.12 |
ENSMUST00000074949.4
|
Tmem140
|
transmembrane protein 140 |
| chr4_+_32982981 | 0.10 |
ENSMUST00000098190.10
ENSMUST00000029946.14 |
Rragd
|
Ras-related GTP binding D |
| chr10_+_126914755 | 0.09 |
ENSMUST00000039259.7
ENSMUST00000217941.2 |
Agap2
|
ArfGAP with GTPase domain, ankyrin repeat and PH domain 2 |
| chr5_-_88823049 | 0.08 |
ENSMUST00000133532.8
ENSMUST00000150438.2 |
Grsf1
|
G-rich RNA sequence binding factor 1 |
| chr7_+_140462343 | 0.07 |
ENSMUST00000163610.9
ENSMUST00000164681.8 |
Psmd13
|
proteasome (prosome, macropain) 26S subunit, non-ATPase, 13 |
| chr7_+_100435548 | 0.07 |
ENSMUST00000216021.2
|
Fam168a
|
family with sequence similarity 168, member A |
| chrX_-_104919201 | 0.05 |
ENSMUST00000198209.2
|
Atrx
|
ATRX, chromatin remodeler |
| chr6_-_78445846 | 0.05 |
ENSMUST00000032089.3
|
Reg3g
|
regenerating islet-derived 3 gamma |
| chr9_+_43978290 | 0.05 |
ENSMUST00000034508.14
|
Usp2
|
ubiquitin specific peptidase 2 |
| chr7_-_103535459 | 0.04 |
ENSMUST00000216303.2
|
Olfr66
|
olfactory receptor 66 |
| chr18_+_37488174 | 0.04 |
ENSMUST00000192867.2
ENSMUST00000051163.3 |
Pcdhb8
|
protocadherin beta 8 |
| chr7_-_65020655 | 0.03 |
ENSMUST00000032729.8
|
Tjp1
|
tight junction protein 1 |
| chr19_-_11852453 | 0.03 |
ENSMUST00000213954.2
ENSMUST00000217617.2 |
Olfr1419
|
olfactory receptor 1419 |
| chr1_+_37068387 | 0.02 |
ENSMUST00000067178.14
ENSMUST00000238500.2 |
Vwa3b
|
von Willebrand factor A domain containing 3B |
| chr1_+_179788675 | 0.02 |
ENSMUST00000076687.12
ENSMUST00000097450.10 ENSMUST00000212756.2 |
Cdc42bpa
|
CDC42 binding protein kinase alpha |
| chr17_+_37689924 | 0.01 |
ENSMUST00000215518.2
|
Olfr105-ps
|
olfactory receptor 105, pseudogene |
| chr7_+_66489465 | 0.00 |
ENSMUST00000098382.10
|
Adamts17
|
a disintegrin-like and metallopeptidase (reprolysin type) with thrombospondin type 1 motif, 17 |
Network of associatons between targets according to the STRING database.
First level regulatory network of Stat6
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.0 | 14.9 | GO:0002148 | hypochlorous acid metabolic process(GO:0002148) hypochlorous acid biosynthetic process(GO:0002149) |
| 0.9 | 4.7 | GO:0008626 | granzyme-mediated apoptotic signaling pathway(GO:0008626) |
| 0.8 | 2.3 | GO:0002277 | myeloid dendritic cell activation involved in immune response(GO:0002277) |
| 0.7 | 3.7 | GO:0070315 | G1 to G0 transition involved in cell differentiation(GO:0070315) |
| 0.7 | 2.1 | GO:0003294 | atrial ventricular junction remodeling(GO:0003294) regulation of cell communication by chemical coupling(GO:0010645) positive regulation of cell communication by chemical coupling(GO:0010652) |
| 0.6 | 2.4 | GO:1901421 | positive regulation of response to alcohol(GO:1901421) |
| 0.6 | 2.9 | GO:2000660 | negative regulation of interleukin-1-mediated signaling pathway(GO:2000660) |
| 0.5 | 1.4 | GO:0001788 | antibody-dependent cellular cytotoxicity(GO:0001788) |
| 0.5 | 1.9 | GO:2000503 | positive regulation of natural killer cell chemotaxis(GO:2000503) |
| 0.4 | 1.3 | GO:0042091 | interleukin-10 biosynthetic process(GO:0042091) regulation of interleukin-10 biosynthetic process(GO:0045074) |
| 0.4 | 6.9 | GO:0030388 | fructose 1,6-bisphosphate metabolic process(GO:0030388) |
| 0.4 | 1.8 | GO:1905171 | protein localization to phagocytic vesicle(GO:1905161) regulation of protein localization to phagocytic vesicle(GO:1905169) positive regulation of protein localization to phagocytic vesicle(GO:1905171) |
| 0.3 | 1.0 | GO:1990051 | negative regulation of phospholipase C activity(GO:1900275) activation of protein kinase C activity(GO:1990051) |
| 0.3 | 1.3 | GO:2001107 | negative regulation of Rho guanyl-nucleotide exchange factor activity(GO:2001107) |
| 0.3 | 1.8 | GO:1903436 | regulation of cytokinetic process(GO:0032954) regulation of mitotic cytokinetic process(GO:1903436) positive regulation of mitotic cytokinetic process(GO:1903438) positive regulation of mitotic cytokinesis(GO:1903490) |
| 0.3 | 1.1 | GO:0001928 | regulation of exocyst assembly(GO:0001928) |
| 0.3 | 1.1 | GO:0015851 | nucleobase transport(GO:0015851) pyrimidine nucleobase transport(GO:0015855) |
| 0.3 | 1.9 | GO:0006207 | 'de novo' pyrimidine nucleobase biosynthetic process(GO:0006207) pyrimidine nucleobase biosynthetic process(GO:0019856) |
| 0.3 | 0.5 | GO:0071725 | response to triacyl bacterial lipopeptide(GO:0071725) cellular response to triacyl bacterial lipopeptide(GO:0071727) |
| 0.3 | 3.2 | GO:0001866 | NK T cell proliferation(GO:0001866) |
| 0.3 | 5.9 | GO:0007250 | activation of NF-kappaB-inducing kinase activity(GO:0007250) |
| 0.2 | 2.0 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 0.2 | 0.7 | GO:0032685 | negative regulation of granulocyte macrophage colony-stimulating factor production(GO:0032685) |
| 0.2 | 1.0 | GO:2001245 | regulation of phosphatidylcholine biosynthetic process(GO:2001245) |
| 0.2 | 0.7 | GO:0051892 | negative regulation of cardioblast differentiation(GO:0051892) |
| 0.2 | 0.7 | GO:0035722 | interleukin-12-mediated signaling pathway(GO:0035722) cellular response to interleukin-12(GO:0071349) |
| 0.2 | 2.4 | GO:0030916 | otic vesicle formation(GO:0030916) |
| 0.2 | 0.9 | GO:0071348 | cellular response to interleukin-11(GO:0071348) |
| 0.2 | 0.6 | GO:0021570 | rhombomere 4 development(GO:0021570) |
| 0.2 | 1.1 | GO:0035026 | leading edge cell differentiation(GO:0035026) |
| 0.2 | 0.5 | GO:1904742 | regulation of telomeric DNA binding(GO:1904742) |
| 0.2 | 1.4 | GO:0042985 | negative regulation of amyloid precursor protein biosynthetic process(GO:0042985) |
| 0.1 | 1.0 | GO:0097647 | calcitonin family receptor signaling pathway(GO:0097646) amylin receptor signaling pathway(GO:0097647) |
| 0.1 | 0.7 | GO:0006384 | transcription initiation from RNA polymerase III promoter(GO:0006384) |
| 0.1 | 0.6 | GO:0060356 | leucine import(GO:0060356) |
| 0.1 | 0.5 | GO:0070295 | renal water absorption(GO:0070295) |
| 0.1 | 1.8 | GO:0015809 | arginine transport(GO:0015809) |
| 0.1 | 1.7 | GO:0071493 | cellular response to UV-B(GO:0071493) |
| 0.1 | 1.9 | GO:0006857 | oligopeptide transport(GO:0006857) |
| 0.1 | 1.0 | GO:0032713 | negative regulation of interleukin-4 production(GO:0032713) |
| 0.1 | 6.9 | GO:0043029 | T cell homeostasis(GO:0043029) |
| 0.1 | 1.5 | GO:0031274 | positive regulation of pseudopodium assembly(GO:0031274) |
| 0.1 | 2.3 | GO:0042761 | fatty acid elongation(GO:0030497) very long-chain fatty acid biosynthetic process(GO:0042761) |
| 0.1 | 0.4 | GO:2001025 | positive regulation of response to drug(GO:2001025) |
| 0.1 | 0.7 | GO:1901894 | regulation of calcium-transporting ATPase activity(GO:1901894) |
| 0.1 | 0.4 | GO:0045039 | protein import into mitochondrial inner membrane(GO:0045039) |
| 0.1 | 1.2 | GO:1990403 | embryonic brain development(GO:1990403) |
| 0.1 | 0.4 | GO:0090383 | platelet dense granule organization(GO:0060155) phagosome acidification(GO:0090383) melanosome assembly(GO:1903232) |
| 0.1 | 0.5 | GO:0033031 | positive regulation of neutrophil apoptotic process(GO:0033031) |
| 0.1 | 2.1 | GO:2000369 | regulation of clathrin-mediated endocytosis(GO:2000369) |
| 0.1 | 0.9 | GO:0034498 | early endosome to Golgi transport(GO:0034498) |
| 0.1 | 0.6 | GO:0010838 | positive regulation of keratinocyte proliferation(GO:0010838) |
| 0.0 | 0.2 | GO:2000384 | regulation of ectoderm development(GO:2000383) negative regulation of ectoderm development(GO:2000384) |
| 0.0 | 3.2 | GO:0015807 | L-amino acid transport(GO:0015807) |
| 0.0 | 0.2 | GO:0072365 | regulation of cellular ketone metabolic process by negative regulation of transcription from RNA polymerase II promoter(GO:0072365) |
| 0.0 | 0.2 | GO:0008594 | photoreceptor cell morphogenesis(GO:0008594) |
| 0.0 | 5.5 | GO:0043627 | response to estrogen(GO:0043627) |
| 0.0 | 1.5 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 0.0 | 0.3 | GO:0032020 | ISG15-protein conjugation(GO:0032020) |
| 0.0 | 0.1 | GO:0061228 | regulation of pronephros size(GO:0035565) mesonephros morphogenesis(GO:0061206) mesonephric nephron development(GO:0061215) mesonephric nephron morphogenesis(GO:0061228) mesenchymal stem cell maintenance involved in mesonephric nephron morphogenesis(GO:0061235) regulation of mesenchymal cell apoptotic process involved in mesonephric nephron morphogenesis(GO:0061295) negative regulation of mesenchymal cell apoptotic process involved in mesonephric nephron morphogenesis(GO:0061296) mesenchymal cell apoptotic process involved in mesonephric nephron morphogenesis(GO:1901146) |
| 0.0 | 1.5 | GO:0001937 | negative regulation of endothelial cell proliferation(GO:0001937) |
| 0.0 | 1.0 | GO:0016339 | calcium-dependent cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0016339) |
| 0.0 | 0.2 | GO:0018242 | protein O-linked glycosylation via serine(GO:0018242) |
| 0.0 | 0.1 | GO:0000480 | endonucleolytic cleavage in 5'-ETS of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000480) |
| 0.0 | 1.1 | GO:0000462 | maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000462) |
| 0.0 | 0.2 | GO:1990166 | positive regulation of maintenance of mitotic sister chromatid cohesion(GO:0034184) protein localization to site of double-strand break(GO:1990166) |
| 0.0 | 1.4 | GO:0033119 | negative regulation of RNA splicing(GO:0033119) |
| 0.0 | 2.5 | GO:0050830 | defense response to Gram-positive bacterium(GO:0050830) |
| 0.0 | 0.3 | GO:0034773 | histone H4-K20 trimethylation(GO:0034773) |
| 0.0 | 1.0 | GO:0043551 | regulation of phosphatidylinositol 3-kinase activity(GO:0043551) |
| 0.0 | 1.2 | GO:0036465 | synaptic vesicle recycling(GO:0036465) |
| 0.0 | 0.8 | GO:0046827 | positive regulation of protein export from nucleus(GO:0046827) |
| 0.0 | 0.1 | GO:1905051 | regulation of base-excision repair(GO:1905051) positive regulation of base-excision repair(GO:1905053) |
| 0.0 | 0.7 | GO:0043252 | sodium-independent organic anion transport(GO:0043252) |
| 0.0 | 0.7 | GO:0006730 | one-carbon metabolic process(GO:0006730) |
| 0.0 | 0.3 | GO:0044247 | glycogen catabolic process(GO:0005980) glucan catabolic process(GO:0009251) cellular polysaccharide catabolic process(GO:0044247) |
| 0.0 | 0.2 | GO:0031507 | heterochromatin assembly(GO:0031507) |
| 0.0 | 0.1 | GO:0043201 | response to leucine(GO:0043201) cellular response to leucine(GO:0071233) |
| 0.0 | 0.9 | GO:0001895 | retina homeostasis(GO:0001895) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 3.8 | GO:0044194 | cytolytic granule(GO:0044194) |
| 0.9 | 14.9 | GO:0005766 | primary lysosome(GO:0005766) azurophil granule(GO:0042582) |
| 0.6 | 1.7 | GO:0070557 | PCNA-p21 complex(GO:0070557) |
| 0.4 | 1.5 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.3 | 6.9 | GO:0035686 | sperm fibrous sheath(GO:0035686) |
| 0.2 | 1.1 | GO:0070022 | transforming growth factor beta receptor homodimeric complex(GO:0070022) |
| 0.2 | 1.0 | GO:0005944 | phosphatidylinositol 3-kinase complex, class IB(GO:0005944) |
| 0.2 | 1.0 | GO:1903440 | calcitonin family receptor complex(GO:1903439) amylin receptor complex(GO:1903440) |
| 0.2 | 1.8 | GO:0044613 | nuclear pore central transport channel(GO:0044613) |
| 0.1 | 0.7 | GO:0000308 | cytoplasmic cyclin-dependent protein kinase holoenzyme complex(GO:0000308) |
| 0.1 | 0.4 | GO:0042721 | mitochondrial inner membrane protein insertion complex(GO:0042721) |
| 0.1 | 0.9 | GO:0034388 | Pwp2p-containing subcomplex of 90S preribosome(GO:0034388) |
| 0.1 | 2.0 | GO:0098644 | complex of collagen trimers(GO:0098644) |
| 0.1 | 1.5 | GO:0031932 | TORC2 complex(GO:0031932) |
| 0.1 | 2.1 | GO:0005922 | connexon complex(GO:0005922) |
| 0.1 | 1.7 | GO:0005639 | integral component of nuclear inner membrane(GO:0005639) nuclear lamina(GO:0005652) intrinsic component of nuclear inner membrane(GO:0031229) |
| 0.1 | 0.5 | GO:0046696 | lipopolysaccharide receptor complex(GO:0046696) |
| 0.1 | 4.6 | GO:0045335 | phagocytic vesicle(GO:0045335) |
| 0.1 | 0.5 | GO:0070187 | telosome(GO:0070187) |
| 0.0 | 1.3 | GO:0005637 | nuclear inner membrane(GO:0005637) |
| 0.0 | 7.9 | GO:0000776 | kinetochore(GO:0000776) |
| 0.0 | 0.3 | GO:0072487 | MSL complex(GO:0072487) |
| 0.0 | 2.5 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 1.0 | GO:0031307 | integral component of mitochondrial outer membrane(GO:0031307) |
| 0.0 | 0.7 | GO:0005666 | DNA-directed RNA polymerase III complex(GO:0005666) |
| 0.0 | 3.8 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.0 | 1.3 | GO:0016592 | mediator complex(GO:0016592) |
| 0.0 | 0.3 | GO:0000780 | condensed nuclear chromosome, centromeric region(GO:0000780) |
| 0.0 | 0.9 | GO:0030904 | retromer complex(GO:0030904) |
| 0.0 | 3.1 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 0.7 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.0 | 3.1 | GO:0030315 | T-tubule(GO:0030315) |
| 0.0 | 0.8 | GO:0070971 | endoplasmic reticulum exit site(GO:0070971) |
| 0.0 | 0.2 | GO:0005677 | chromatin silencing complex(GO:0005677) |
| 0.0 | 0.5 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.0 | 0.2 | GO:0045298 | tubulin complex(GO:0045298) |
| 0.0 | 0.1 | GO:1990131 | Gtr1-Gtr2 GTPase complex(GO:1990131) |
| 0.0 | 1.3 | GO:0016528 | sarcoplasm(GO:0016528) |
| 0.0 | 0.5 | GO:0000314 | organellar small ribosomal subunit(GO:0000314) mitochondrial small ribosomal subunit(GO:0005763) |
| 0.0 | 0.2 | GO:0042405 | nuclear inclusion body(GO:0042405) |
| 0.0 | 1.2 | GO:0005905 | clathrin-coated pit(GO:0005905) |
| 0.0 | 0.9 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 2.9 | GO:0005152 | interleukin-1 receptor antagonist activity(GO:0005152) |
| 0.8 | 2.3 | GO:0080023 | 3R-hydroxyacyl-CoA dehydratase activity(GO:0080023) |
| 0.7 | 6.9 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 0.6 | 1.9 | GO:0004088 | carbamoyl-phosphate synthase (ammonia) activity(GO:0004087) carbamoyl-phosphate synthase (glutamine-hydrolyzing) activity(GO:0004088) |
| 0.6 | 1.9 | GO:0030298 | receptor signaling protein tyrosine kinase activator activity(GO:0030298) CCR1 chemokine receptor binding(GO:0031726) |
| 0.6 | 1.7 | GO:0019912 | cyclin-dependent protein kinase activating kinase activity(GO:0019912) |
| 0.5 | 2.1 | GO:0086077 | gap junction channel activity involved in AV node cell-bundle of His cell electrical coupling(GO:0086077) |
| 0.5 | 5.9 | GO:0008061 | chitin binding(GO:0008061) |
| 0.4 | 1.1 | GO:0015389 | pyrimidine- and adenine-specific:sodium symporter activity(GO:0015389) |
| 0.2 | 1.8 | GO:0051425 | PTB domain binding(GO:0051425) |
| 0.2 | 1.5 | GO:0070180 | large ribosomal subunit rRNA binding(GO:0070180) |
| 0.2 | 14.9 | GO:0004601 | peroxidase activity(GO:0004601) |
| 0.2 | 1.4 | GO:0019770 | IgG receptor activity(GO:0019770) |
| 0.2 | 1.8 | GO:0015181 | arginine transmembrane transporter activity(GO:0015181) |
| 0.2 | 2.8 | GO:0031432 | titin binding(GO:0031432) |
| 0.2 | 1.4 | GO:0001515 | opioid peptide activity(GO:0001515) |
| 0.2 | 1.3 | GO:0001849 | complement component C1q binding(GO:0001849) |
| 0.2 | 1.0 | GO:0097643 | amylin receptor activity(GO:0097643) |
| 0.1 | 1.6 | GO:0070087 | chromo shadow domain binding(GO:0070087) |
| 0.1 | 0.4 | GO:0036461 | BLOC-2 complex binding(GO:0036461) |
| 0.1 | 0.7 | GO:0005143 | interleukin-12 receptor binding(GO:0005143) |
| 0.1 | 0.4 | GO:0030943 | mitochondrion targeting sequence binding(GO:0030943) |
| 0.1 | 1.6 | GO:0052813 | phosphatidylinositol-4,5-bisphosphate 3-kinase activity(GO:0046934) phosphatidylinositol bisphosphate kinase activity(GO:0052813) |
| 0.1 | 2.4 | GO:0010314 | phosphatidylinositol-5-phosphate binding(GO:0010314) |
| 0.1 | 0.4 | GO:0090555 | phosphatidylethanolamine-translocating ATPase activity(GO:0090555) |
| 0.1 | 1.0 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 0.1 | 0.3 | GO:0004134 | glycogen debranching enzyme activity(GO:0004133) 4-alpha-glucanotransferase activity(GO:0004134) amylo-alpha-1,6-glucosidase activity(GO:0004135) |
| 0.1 | 0.5 | GO:0071723 | lipopeptide binding(GO:0071723) |
| 0.1 | 5.5 | GO:0001105 | RNA polymerase II transcription coactivator activity(GO:0001105) |
| 0.1 | 1.0 | GO:0070097 | delta-catenin binding(GO:0070097) |
| 0.1 | 1.1 | GO:0035615 | clathrin adaptor activity(GO:0035615) endocytic adaptor activity(GO:0098748) |
| 0.1 | 1.8 | GO:0017160 | Ral GTPase binding(GO:0017160) |
| 0.1 | 3.8 | GO:0015175 | neutral amino acid transmembrane transporter activity(GO:0015175) |
| 0.1 | 3.2 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.1 | 0.6 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 0.1 | 0.5 | GO:0015288 | porin activity(GO:0015288) |
| 0.1 | 1.0 | GO:0055106 | ligase regulator activity(GO:0055103) ubiquitin-protein transferase regulator activity(GO:0055106) |
| 0.1 | 0.3 | GO:0042296 | ISG15 transferase activity(GO:0042296) |
| 0.0 | 1.1 | GO:0030275 | LRR domain binding(GO:0030275) |
| 0.0 | 1.4 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) |
| 0.0 | 3.4 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.0 | 0.3 | GO:0042799 | histone methyltransferase activity (H4-K20 specific)(GO:0042799) |
| 0.0 | 0.7 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.0 | 1.5 | GO:0070717 | poly-purine tract binding(GO:0070717) |
| 0.0 | 0.7 | GO:0001056 | RNA polymerase III activity(GO:0001056) |
| 0.0 | 0.1 | GO:0070181 | small ribosomal subunit rRNA binding(GO:0070181) |
| 0.0 | 1.1 | GO:0001106 | RNA polymerase II transcription corepressor activity(GO:0001106) |
| 0.0 | 1.3 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.0 | 2.1 | GO:0030276 | clathrin binding(GO:0030276) |
| 0.0 | 0.5 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.0 | 0.7 | GO:0015347 | sodium-independent organic anion transmembrane transporter activity(GO:0015347) |
| 0.0 | 0.7 | GO:0005310 | dicarboxylic acid transmembrane transporter activity(GO:0005310) |
| 0.0 | 1.4 | GO:0003725 | double-stranded RNA binding(GO:0003725) |
| 0.0 | 2.5 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 0.7 | GO:0004693 | cyclin-dependent protein serine/threonine kinase activity(GO:0004693) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 15.0 | PID IL23 PATHWAY | IL23-mediated signaling events |
| 0.2 | 1.7 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.1 | 8.1 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.1 | 3.8 | PID IL12 2PATHWAY | IL12-mediated signaling events |
| 0.1 | 6.9 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.1 | 2.9 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.1 | 2.0 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.0 | 1.9 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 0.0 | 1.0 | PID ATM PATHWAY | ATM pathway |
| 0.0 | 1.0 | PID IL8 CXCR1 PATHWAY | IL8- and CXCR1-mediated signaling events |
| 0.0 | 0.5 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 0.5 | SA TRKA RECEPTOR | The TrkA receptor binds nerve growth factor to activate MAP kinase pathways and promote cell growth. |
| 0.0 | 1.9 | ST FAS SIGNALING PATHWAY | Fas Signaling Pathway |
| 0.0 | 0.8 | PID FCER1 PATHWAY | Fc-epsilon receptor I signaling in mast cells |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 8.1 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.1 | 2.0 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.1 | 6.9 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.1 | 2.1 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.1 | 1.2 | REACTOME NEF MEDIATED DOWNREGULATION OF MHC CLASS I COMPLEX CELL SURFACE EXPRESSION | Genes involved in Nef mediated downregulation of MHC class I complex cell surface expression |
| 0.1 | 7.1 | REACTOME AMINO ACID AND OLIGOPEPTIDE SLC TRANSPORTERS | Genes involved in Amino acid and oligopeptide SLC transporters |
| 0.1 | 0.5 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.1 | 7.9 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.1 | 3.4 | REACTOME INTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Intrinsic Pathway for Apoptosis |
| 0.1 | 1.1 | REACTOME GAP JUNCTION DEGRADATION | Genes involved in Gap junction degradation |
| 0.1 | 1.8 | REACTOME NEP NS2 INTERACTS WITH THE CELLULAR EXPORT MACHINERY | Genes involved in NEP/NS2 Interacts with the Cellular Export Machinery |
| 0.1 | 0.5 | REACTOME IKK COMPLEX RECRUITMENT MEDIATED BY RIP1 | Genes involved in IKK complex recruitment mediated by RIP1 |
| 0.1 | 1.7 | REACTOME AKT PHOSPHORYLATES TARGETS IN THE CYTOSOL | Genes involved in AKT phosphorylates targets in the cytosol |
| 0.1 | 1.7 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.1 | 1.9 | REACTOME PYRIMIDINE METABOLISM | Genes involved in Pyrimidine metabolism |
| 0.0 | 2.9 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.0 | 0.5 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.0 | 0.7 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 0.0 | 2.5 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.0 | 1.8 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.0 | 0.7 | REACTOME RNA POL III CHAIN ELONGATION | Genes involved in RNA Polymerase III Chain Elongation |
| 0.0 | 1.6 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.0 | 1.0 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 0.0 | 1.5 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.0 | 1.1 | REACTOME SIGNAL TRANSDUCTION BY L1 | Genes involved in Signal transduction by L1 |
| 0.0 | 0.8 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.0 | 0.3 | REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.0 | 3.0 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 1.5 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |