Project
GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
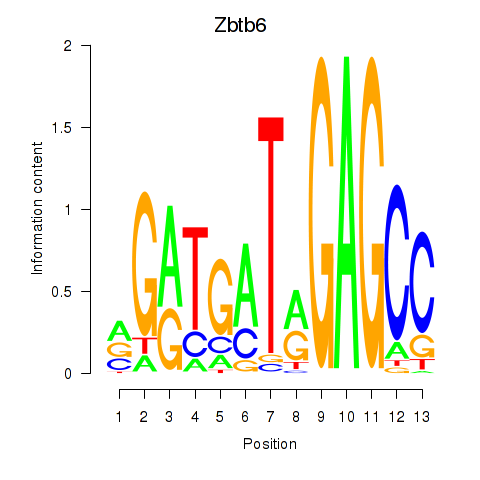
Results for Zbtb6
Z-value: 0.73
Transcription factors associated with Zbtb6
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Zbtb6
|
ENSMUSG00000066798.4 | zinc finger and BTB domain containing 6 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Zbtb6 | mm39_v1_chr2_-_37333109_37333210 | -0.12 | 4.7e-01 | Click! |
Activity profile of Zbtb6 motif
Sorted Z-values of Zbtb6 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_-_87724533 | 4.96 |
ENSMUST00000126353.8
ENSMUST00000149357.8 |
Mllt3
|
myeloid/lymphoid or mixed-lineage leukemia; translocated to, 3 |
| chr17_+_37180437 | 4.21 |
ENSMUST00000060524.11
|
Trim10
|
tripartite motif-containing 10 |
| chr6_-_60806810 | 3.61 |
ENSMUST00000163779.8
|
Snca
|
synuclein, alpha |
| chr4_-_87724512 | 3.60 |
ENSMUST00000148059.2
|
Mllt3
|
myeloid/lymphoid or mixed-lineage leukemia; translocated to, 3 |
| chr11_-_102787950 | 3.54 |
ENSMUST00000067444.10
|
Gfap
|
glial fibrillary acidic protein |
| chr6_+_86005729 | 3.32 |
ENSMUST00000203366.3
|
Add2
|
adducin 2 (beta) |
| chr6_+_86005663 | 2.99 |
ENSMUST00000204059.3
|
Add2
|
adducin 2 (beta) |
| chr8_-_86091970 | 2.93 |
ENSMUST00000121972.8
|
Mylk3
|
myosin light chain kinase 3 |
| chr8_-_86091946 | 2.92 |
ENSMUST00000034133.14
|
Mylk3
|
myosin light chain kinase 3 |
| chr3_-_100396635 | 2.71 |
ENSMUST00000061455.9
|
Tent5c
|
terminal nucleotidyltransferase 5C |
| chr4_+_117692583 | 2.20 |
ENSMUST00000169885.8
|
Slc6a9
|
solute carrier family 6 (neurotransmitter transporter, glycine), member 9 |
| chr9_-_21671571 | 2.17 |
ENSMUST00000217382.2
ENSMUST00000214149.2 ENSMUST00000098942.6 ENSMUST00000216057.2 |
Spc24
|
SPC24, NDC80 kinetochore complex component, homolog (S. cerevisiae) |
| chr9_+_107468146 | 2.14 |
ENSMUST00000195746.2
|
Ifrd2
|
interferon-related developmental regulator 2 |
| chr1_+_134110142 | 2.08 |
ENSMUST00000082060.10
ENSMUST00000153856.8 ENSMUST00000133701.8 ENSMUST00000132873.8 |
Chil1
|
chitinase-like 1 |
| chr11_-_102787972 | 2.06 |
ENSMUST00000077902.5
|
Gfap
|
glial fibrillary acidic protein |
| chr7_-_97827461 | 2.05 |
ENSMUST00000040971.14
|
Capn5
|
calpain 5 |
| chr8_+_94899292 | 2.01 |
ENSMUST00000034214.8
ENSMUST00000212806.2 |
Mt2
|
metallothionein 2 |
| chr10_-_40018243 | 1.83 |
ENSMUST00000092566.8
ENSMUST00000213488.2 |
Slc16a10
|
solute carrier family 16 (monocarboxylic acid transporters), member 10 |
| chr11_-_79971750 | 1.76 |
ENSMUST00000103233.10
ENSMUST00000061283.15 |
Crlf3
|
cytokine receptor-like factor 3 |
| chr15_+_78798116 | 1.70 |
ENSMUST00000089378.5
|
Pdxp
|
pyridoxal (pyridoxine, vitamin B6) phosphatase |
| chr17_-_46464441 | 1.69 |
ENSMUST00000171172.3
|
Mad2l1bp
|
MAD2L1 binding protein |
| chr4_-_156285247 | 1.67 |
ENSMUST00000085425.6
|
Isg15
|
ISG15 ubiquitin-like modifier |
| chr7_-_100504610 | 1.60 |
ENSMUST00000156855.8
|
Relt
|
RELT tumor necrosis factor receptor |
| chr9_+_54606832 | 1.55 |
ENSMUST00000070070.8
|
Dnaja4
|
DnaJ heat shock protein family (Hsp40) member A4 |
| chr9_+_54606144 | 1.48 |
ENSMUST00000120452.8
|
Dnaja4
|
DnaJ heat shock protein family (Hsp40) member A4 |
| chr7_-_44785603 | 1.44 |
ENSMUST00000209467.2
|
Gm45713
|
predicted gene 45713 |
| chr17_+_35069347 | 1.43 |
ENSMUST00000097343.11
ENSMUST00000173357.8 ENSMUST00000173065.8 ENSMUST00000165953.3 |
Nelfe
|
negative elongation factor complex member E, Rdbp |
| chr1_+_134109888 | 1.41 |
ENSMUST00000156873.8
|
Chil1
|
chitinase-like 1 |
| chr7_+_24246575 | 1.40 |
ENSMUST00000063249.9
|
Xrcc1
|
X-ray repair complementing defective repair in Chinese hamster cells 1 |
| chr4_+_129083553 | 1.38 |
ENSMUST00000106054.4
|
Yars
|
tyrosyl-tRNA synthetase |
| chr17_+_29171386 | 1.33 |
ENSMUST00000118762.9
ENSMUST00000057174.16 ENSMUST00000232874.2 ENSMUST00000232772.2 ENSMUST00000233334.2 ENSMUST00000150858.2 ENSMUST00000233064.2 |
Kctd20
|
potassium channel tetramerisation domain containing 20 |
| chr9_+_54606798 | 1.31 |
ENSMUST00000154690.8
|
Dnaja4
|
DnaJ heat shock protein family (Hsp40) member A4 |
| chr2_+_78699360 | 1.14 |
ENSMUST00000028398.14
|
Ube2e3
|
ubiquitin-conjugating enzyme E2E 3 |
| chr1_-_74040723 | 1.13 |
ENSMUST00000190389.7
|
Tns1
|
tensin 1 |
| chr15_+_98006346 | 1.06 |
ENSMUST00000051226.8
|
Pfkm
|
phosphofructokinase, muscle |
| chr6_+_113508636 | 1.04 |
ENSMUST00000036340.12
ENSMUST00000204827.3 |
Fancd2
|
Fanconi anemia, complementation group D2 |
| chr15_+_58004793 | 1.02 |
ENSMUST00000227142.2
ENSMUST00000226955.2 |
Wdyhv1
|
WDYHV motif containing 1 |
| chr11_-_75918551 | 1.02 |
ENSMUST00000021207.7
|
Rflnb
|
refilin B |
| chr15_-_79571977 | 1.00 |
ENSMUST00000023061.7
|
Josd1
|
Josephin domain containing 1 |
| chr9_-_22046970 | 0.93 |
ENSMUST00000165735.9
|
Acp5
|
acid phosphatase 5, tartrate resistant |
| chr2_-_126460575 | 0.93 |
ENSMUST00000028838.5
|
Hdc
|
histidine decarboxylase |
| chr15_+_58004753 | 0.91 |
ENSMUST00000067563.9
|
Wdyhv1
|
WDYHV motif containing 1 |
| chr3_-_95214443 | 0.90 |
ENSMUST00000015846.9
|
Anxa9
|
annexin A9 |
| chr17_+_29171655 | 0.89 |
ENSMUST00000117672.9
ENSMUST00000153831.9 |
Kctd20
|
potassium channel tetramerisation domain containing 20 |
| chr8_+_75820240 | 0.88 |
ENSMUST00000005548.8
|
Hmox1
|
heme oxygenase 1 |
| chr7_-_30738471 | 0.86 |
ENSMUST00000162250.8
|
Fxyd5
|
FXYD domain-containing ion transport regulator 5 |
| chr8_-_106434565 | 0.84 |
ENSMUST00000013299.11
|
Enkd1
|
enkurin domain containing 1 |
| chr7_+_28488380 | 0.84 |
ENSMUST00000209035.2
ENSMUST00000059857.8 |
Rinl
|
Ras and Rab interactor-like |
| chr19_+_4242064 | 0.81 |
ENSMUST00000046094.6
|
Ppp1ca
|
protein phosphatase 1 catalytic subunit alpha |
| chr7_-_44785480 | 0.76 |
ENSMUST00000211246.2
ENSMUST00000210197.2 |
Flt3l
|
FMS-like tyrosine kinase 3 ligand |
| chr14_-_79718890 | 0.65 |
ENSMUST00000022601.7
|
Wbp4
|
WW domain binding protein 4 |
| chr7_-_44785709 | 0.65 |
ENSMUST00000211429.2
|
Flt3l
|
FMS-like tyrosine kinase 3 ligand |
| chr13_+_55517545 | 0.63 |
ENSMUST00000063771.14
|
Rgs14
|
regulator of G-protein signaling 14 |
| chr9_-_21982637 | 0.62 |
ENSMUST00000179605.9
ENSMUST00000043922.7 |
Zfp653
|
zinc finger protein 653 |
| chr9_+_66257747 | 0.61 |
ENSMUST00000042824.13
|
Herc1
|
HECT and RLD domain containing E3 ubiquitin protein ligase family member 1 |
| chr16_+_18695787 | 0.59 |
ENSMUST00000120532.9
ENSMUST00000004222.14 |
Hira
|
histone cell cycle regulator |
| chr9_+_113760376 | 0.59 |
ENSMUST00000214095.2
ENSMUST00000116492.3 ENSMUST00000216558.2 |
Ubp1
|
upstream binding protein 1 |
| chr11_+_69857722 | 0.59 |
ENSMUST00000151515.2
|
Cldn7
|
claudin 7 |
| chr6_+_123239076 | 0.57 |
ENSMUST00000032240.4
|
Clec4d
|
C-type lectin domain family 4, member d |
| chr17_-_35069136 | 0.57 |
ENSMUST00000046022.16
|
Skiv2l
|
superkiller viralicidic activity 2-like (S. cerevisiae) |
| chr7_-_44785815 | 0.55 |
ENSMUST00000146760.7
|
Flt3l
|
FMS-like tyrosine kinase 3 ligand |
| chrX_-_52328963 | 0.54 |
ENSMUST00000074861.9
|
Plac1
|
placental specific protein 1 |
| chr11_+_3438274 | 0.52 |
ENSMUST00000064265.13
|
Pla2g3
|
phospholipase A2, group III |
| chr7_-_140859034 | 0.51 |
ENSMUST00000211667.2
ENSMUST00000167790.3 ENSMUST00000046156.13 |
Sct
|
secretin |
| chr16_+_38383154 | 0.50 |
ENSMUST00000171687.8
ENSMUST00000002924.15 |
Tmem39a
|
transmembrane protein 39a |
| chr8_-_95564881 | 0.48 |
ENSMUST00000034233.15
ENSMUST00000162538.9 |
Ciapin1
|
cytokine induced apoptosis inhibitor 1 |
| chr19_-_46561532 | 0.47 |
ENSMUST00000026009.10
ENSMUST00000236255.2 |
Arl3
|
ADP-ribosylation factor-like 3 |
| chr1_+_74627445 | 0.47 |
ENSMUST00000113733.10
ENSMUST00000027358.11 |
Bcs1l
|
BCS1-like (yeast) |
| chr16_+_17095416 | 0.47 |
ENSMUST00000232364.2
|
Tmem191c
|
transmembrane protein 191C |
| chr17_-_25652750 | 0.46 |
ENSMUST00000159610.8
ENSMUST00000159048.8 ENSMUST00000078496.12 |
Cacna1h
|
calcium channel, voltage-dependent, T type, alpha 1H subunit |
| chr14_-_55204023 | 0.46 |
ENSMUST00000124930.8
|
Myh6
|
myosin, heavy polypeptide 6, cardiac muscle, alpha |
| chr18_-_42395207 | 0.46 |
ENSMUST00000097590.5
|
Lars
|
leucyl-tRNA synthetase |
| chr17_+_36134450 | 0.45 |
ENSMUST00000172846.2
|
Flot1
|
flotillin 1 |
| chr15_-_101621332 | 0.45 |
ENSMUST00000023709.7
|
Krt5
|
keratin 5 |
| chr16_-_19334384 | 0.45 |
ENSMUST00000054606.2
|
Olfr167
|
olfactory receptor 167 |
| chr4_+_107659361 | 0.44 |
ENSMUST00000106731.4
|
Lrp8
|
low density lipoprotein receptor-related protein 8, apolipoprotein e receptor |
| chr2_+_127205117 | 0.44 |
ENSMUST00000104934.2
|
Adra2b
|
adrenergic receptor, alpha 2b |
| chr9_-_99599312 | 0.42 |
ENSMUST00000112882.9
ENSMUST00000131922.2 |
Cldn18
|
claudin 18 |
| chr12_-_69629758 | 0.42 |
ENSMUST00000058639.11
|
Vcpkmt
|
valosin containing protein lysine (K) methyltransferase |
| chr15_-_51728893 | 0.42 |
ENSMUST00000022925.10
|
Eif3h
|
eukaryotic translation initiation factor 3, subunit H |
| chr2_-_146927365 | 0.41 |
ENSMUST00000067020.3
|
Nkx2-4
|
NK2 homeobox 4 |
| chr16_-_56748424 | 0.40 |
ENSMUST00000210579.2
|
Lnp1
|
leukemia NUP98 fusion partner 1 |
| chr9_+_108660989 | 0.40 |
ENSMUST00000192307.6
ENSMUST00000193560.6 ENSMUST00000194875.6 |
Ip6k2
|
inositol hexaphosphate kinase 2 |
| chrX_-_165992311 | 0.39 |
ENSMUST00000112172.4
|
Tmsb4x
|
thymosin, beta 4, X chromosome |
| chr13_-_40887444 | 0.38 |
ENSMUST00000224665.2
|
Tfap2a
|
transcription factor AP-2, alpha |
| chrX_+_49930311 | 0.38 |
ENSMUST00000114887.9
|
Stk26
|
serine/threonine kinase 26 |
| chr4_+_63478478 | 0.37 |
ENSMUST00000080336.4
|
Tmem268
|
transmembrane protein 268 |
| chr10_-_82459024 | 0.36 |
ENSMUST00000183363.2
ENSMUST00000079648.12 ENSMUST00000185168.8 |
1190007I07Rik
|
RIKEN cDNA 1190007I07 gene |
| chr17_+_25946644 | 0.36 |
ENSMUST00000237183.2
ENSMUST00000237785.2 ENSMUST00000047273.3 |
Rpusd1
|
RNA pseudouridylate synthase domain containing 1 |
| chr8_+_106331866 | 0.35 |
ENSMUST00000043531.10
|
Ripor1
|
RHO family interacting cell polarization regulator 1 |
| chr10_-_82458760 | 0.35 |
ENSMUST00000176200.8
ENSMUST00000183416.2 |
1190007I07Rik
|
RIKEN cDNA 1190007I07 gene |
| chr9_+_22322802 | 0.34 |
ENSMUST00000058868.9
|
9530077C05Rik
|
RIKEN cDNA 9530077C05 gene |
| chr8_+_84293300 | 0.33 |
ENSMUST00000036996.6
|
Ndufb7
|
NADH:ubiquinone oxidoreductase subunit B7 |
| chr5_-_71253107 | 0.32 |
ENSMUST00000197284.5
|
Gabra2
|
gamma-aminobutyric acid (GABA) A receptor, subunit alpha 2 |
| chr5_-_107745404 | 0.32 |
ENSMUST00000124140.2
|
Glmn
|
glomulin, FKBP associated protein |
| chr16_+_17716480 | 0.32 |
ENSMUST00000055374.8
|
Tssk2
|
testis-specific serine kinase 2 |
| chr1_+_74627506 | 0.31 |
ENSMUST00000113732.2
|
Bcs1l
|
BCS1-like (yeast) |
| chr7_+_79939747 | 0.31 |
ENSMUST00000205864.2
|
Vps33b
|
vacuolar protein sorting 33B |
| chr14_+_55132061 | 0.31 |
ENSMUST00000172557.2
|
Pabpn1
|
poly(A) binding protein, nuclear 1 |
| chr17_-_56607250 | 0.31 |
ENSMUST00000233911.2
|
Arrdc5
|
arrestin domain containing 5 |
| chr17_+_35354655 | 0.30 |
ENSMUST00000174478.8
ENSMUST00000174281.9 ENSMUST00000173550.8 |
Bag6
|
BCL2-associated athanogene 6 |
| chr16_-_18066591 | 0.30 |
ENSMUST00000115645.10
|
Ranbp1
|
RAN binding protein 1 |
| chr15_+_102381929 | 0.30 |
ENSMUST00000229746.2
ENSMUST00000230211.2 ENSMUST00000230539.2 |
Pcbp2
|
poly(rC) binding protein 2 |
| chr17_+_24257118 | 0.29 |
ENSMUST00000059482.6
|
Prss27
|
protease, serine 27 |
| chr3_-_92594516 | 0.29 |
ENSMUST00000029524.4
|
Lce1d
|
late cornified envelope 1D |
| chr17_-_56607286 | 0.29 |
ENSMUST00000097303.3
|
Arrdc5
|
arrestin domain containing 5 |
| chr7_-_29894471 | 0.29 |
ENSMUST00000126116.3
|
Capns1
|
calpain, small subunit 1 |
| chr4_+_63478454 | 0.28 |
ENSMUST00000124332.8
ENSMUST00000150360.8 |
Tmem268
|
transmembrane protein 268 |
| chr2_+_177834868 | 0.28 |
ENSMUST00000103065.2
|
Phactr3
|
phosphatase and actin regulator 3 |
| chr10_-_116808514 | 0.28 |
ENSMUST00000092165.5
|
Gm10271
|
predicted gene 10271 |
| chr1_+_78488440 | 0.27 |
ENSMUST00000152111.2
|
Mogat1
|
monoacylglycerol O-acyltransferase 1 |
| chr7_-_21161866 | 0.27 |
ENSMUST00000177741.4
|
Gm6882
|
predicted gene 6882 |
| chr8_+_72837021 | 0.27 |
ENSMUST00000213940.2
|
Olfr373
|
olfactory receptor 373 |
| chr1_-_74627264 | 0.26 |
ENSMUST00000066986.13
|
Zfp142
|
zinc finger protein 142 |
| chr15_+_103411689 | 0.26 |
ENSMUST00000226493.2
|
Pde1b
|
phosphodiesterase 1B, Ca2+-calmodulin dependent |
| chr2_+_139335829 | 0.26 |
ENSMUST00000110083.8
ENSMUST00000047370.3 |
Sptlc3
|
serine palmitoyltransferase, long chain base subunit 3 |
| chr15_+_58004831 | 0.26 |
ENSMUST00000226889.2
|
Wdyhv1
|
WDYHV motif containing 1 |
| chr18_-_61147272 | 0.26 |
ENSMUST00000025520.10
|
Slc6a7
|
solute carrier family 6 (neurotransmitter transporter, L-proline), member 7 |
| chr10_-_128505096 | 0.26 |
ENSMUST00000238610.2
ENSMUST00000238712.2 |
Ikzf4
|
IKAROS family zinc finger 4 |
| chr15_+_102381705 | 0.26 |
ENSMUST00000229958.2
ENSMUST00000229184.2 ENSMUST00000230728.2 ENSMUST00000230114.2 |
Pcbp2
|
poly(rC) binding protein 2 |
| chr18_-_42395131 | 0.26 |
ENSMUST00000236102.2
|
Lars
|
leucyl-tRNA synthetase |
| chr17_-_46044768 | 0.26 |
ENSMUST00000178179.2
|
1600014C23Rik
|
RIKEN cDNA 1600014C23 gene |
| chr10_+_21469039 | 0.25 |
ENSMUST00000057341.6
|
1700020N01Rik
|
RIKEN cDNA 1700020N01 gene |
| chr8_+_11847411 | 0.25 |
ENSMUST00000239427.2
|
Arhgef7
|
Rho guanine nucleotide exchange factor (GEF7) |
| chr11_-_115158062 | 0.25 |
ENSMUST00000106554.2
|
Grin2c
|
glutamate receptor, ionotropic, NMDA2C (epsilon 3) |
| chr11_-_83320281 | 0.25 |
ENSMUST00000052521.9
|
Gas2l2
|
growth arrest-specific 2 like 2 |
| chr4_-_140781015 | 0.25 |
ENSMUST00000102491.10
|
Crocc
|
ciliary rootlet coiled-coil, rootletin |
| chr2_-_92201342 | 0.24 |
ENSMUST00000176810.8
ENSMUST00000090582.11 ENSMUST00000068586.13 |
Large2
|
LARGE xylosyl- and glucuronyltransferase 2 |
| chr2_+_84867783 | 0.24 |
ENSMUST00000168266.8
ENSMUST00000130729.3 |
Ssrp1
|
structure specific recognition protein 1 |
| chr4_-_140780972 | 0.24 |
ENSMUST00000040222.14
|
Crocc
|
ciliary rootlet coiled-coil, rootletin |
| chr14_+_55131568 | 0.24 |
ENSMUST00000116476.9
ENSMUST00000022808.14 ENSMUST00000150975.8 |
Pabpn1
|
poly(A) binding protein, nuclear 1 |
| chr7_+_140427729 | 0.23 |
ENSMUST00000106049.2
|
Odf3
|
outer dense fiber of sperm tails 3 |
| chr15_+_99024394 | 0.23 |
ENSMUST00000063517.6
|
Spats2
|
spermatogenesis associated, serine-rich 2 |
| chr11_-_102837514 | 0.23 |
ENSMUST00000057849.6
|
C1ql1
|
complement component 1, q subcomponent-like 1 |
| chr12_-_8589545 | 0.22 |
ENSMUST00000095863.10
ENSMUST00000165657.3 |
Slc7a15
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 15 |
| chr3_+_108000425 | 0.22 |
ENSMUST00000151326.8
|
Gnat2
|
guanine nucleotide binding protein, alpha transducing 2 |
| chr7_-_103734672 | 0.22 |
ENSMUST00000057104.7
|
Olfr645
|
olfactory receptor 645 |
| chr12_+_111373243 | 0.21 |
ENSMUST00000150384.3
|
Lbhd2
|
LBH domain containing 2 |
| chr13_-_40887244 | 0.21 |
ENSMUST00000110193.9
|
Tfap2a
|
transcription factor AP-2, alpha |
| chr5_+_73563418 | 0.20 |
ENSMUST00000031040.13
ENSMUST00000065543.8 |
Cwh43
|
cell wall biogenesis 43 C-terminal homolog |
| chr1_+_166829001 | 0.20 |
ENSMUST00000126198.3
|
Fam78b
|
family with sequence similarity 78, member B |
| chr1_+_92545510 | 0.20 |
ENSMUST00000213247.2
|
Olfr12
|
olfactory receptor 12 |
| chr13_-_22193900 | 0.20 |
ENSMUST00000006341.4
|
Prss16
|
protease, serine 16 (thymus) |
| chr14_-_55204383 | 0.20 |
ENSMUST00000111456.2
|
Myh6
|
myosin, heavy polypeptide 6, cardiac muscle, alpha |
| chr15_+_103181311 | 0.20 |
ENSMUST00000100162.5
ENSMUST00000230893.2 |
Copz1
|
coatomer protein complex, subunit zeta 1 |
| chr17_+_35643818 | 0.19 |
ENSMUST00000174699.8
|
H2-Q6
|
histocompatibility 2, Q region locus 6 |
| chr2_-_179976458 | 0.19 |
ENSMUST00000015771.3
|
Gata5
|
GATA binding protein 5 |
| chr4_+_124880223 | 0.19 |
ENSMUST00000030687.8
|
Rspo1
|
R-spondin 1 |
| chr18_+_37637317 | 0.19 |
ENSMUST00000052179.8
|
Pcdhb20
|
protocadherin beta 20 |
| chrX_+_135723531 | 0.19 |
ENSMUST00000113085.2
|
Plp1
|
proteolipid protein (myelin) 1 |
| chr10_+_36383008 | 0.19 |
ENSMUST00000168572.8
|
Hs3st5
|
heparan sulfate (glucosamine) 3-O-sulfotransferase 5 |
| chr15_-_48655329 | 0.19 |
ENSMUST00000160658.8
ENSMUST00000100670.10 ENSMUST00000162830.8 |
Csmd3
|
CUB and Sushi multiple domains 3 |
| chr9_+_58395850 | 0.19 |
ENSMUST00000085658.5
|
Insyn1
|
inhibitory synaptic factor 1 |
| chr7_+_140427711 | 0.19 |
ENSMUST00000026555.12
|
Odf3
|
outer dense fiber of sperm tails 3 |
| chr7_+_126832399 | 0.19 |
ENSMUST00000056232.7
|
Zfp553
|
zinc finger protein 553 |
| chr13_-_64236994 | 0.18 |
ENSMUST00000039832.7
|
Hsd17b3
|
hydroxysteroid (17-beta) dehydrogenase 3 |
| chr1_-_74627123 | 0.18 |
ENSMUST00000027315.14
ENSMUST00000113737.8 |
Zfp142
|
zinc finger protein 142 |
| chr17_-_36953554 | 0.18 |
ENSMUST00000087165.11
ENSMUST00000087167.5 |
H2-M9
|
histocompatibility 2, M region locus 9 |
| chr18_+_37580692 | 0.18 |
ENSMUST00000052387.5
|
Pcdhb14
|
protocadherin beta 14 |
| chr14_-_24054352 | 0.18 |
ENSMUST00000190339.2
|
Kcnma1
|
potassium large conductance calcium-activated channel, subfamily M, alpha member 1 |
| chr1_+_166828982 | 0.17 |
ENSMUST00000165874.8
ENSMUST00000190081.7 |
Fam78b
|
family with sequence similarity 78, member B |
| chr10_-_33920300 | 0.17 |
ENSMUST00000048052.7
|
Calhm4
|
calcium homeostasis modulator family member 4 |
| chr11_+_74661647 | 0.16 |
ENSMUST00000010698.13
ENSMUST00000141755.8 |
Mettl16
|
methyltransferase like 16 |
| chr6_-_139478919 | 0.16 |
ENSMUST00000170650.3
|
Rergl
|
RERG/RAS-like |
| chr13_-_64237036 | 0.16 |
ENSMUST00000222783.2
|
Hsd17b3
|
hydroxysteroid (17-beta) dehydrogenase 3 |
| chr9_+_110485561 | 0.16 |
ENSMUST00000019803.9
ENSMUST00000196347.5 |
Ccdc12
|
coiled-coil domain containing 12 |
| chr13_-_59879548 | 0.16 |
ENSMUST00000052978.6
|
Spata31d1d
|
spermatogenesis associated 31 subfamily D, member 1D |
| chr19_-_5507532 | 0.16 |
ENSMUST00000236881.2
|
Ccdc85b
|
coiled-coil domain containing 85B |
| chr18_+_37575553 | 0.16 |
ENSMUST00000056915.3
|
Pcdhb13
|
protocadherin beta 13 |
| chr3_+_54063459 | 0.16 |
ENSMUST00000029311.11
ENSMUST00000200048.5 |
Trpc4
|
transient receptor potential cation channel, subfamily C, member 4 |
| chrX_+_135723420 | 0.16 |
ENSMUST00000033800.13
|
Plp1
|
proteolipid protein (myelin) 1 |
| chr4_+_103000248 | 0.16 |
ENSMUST00000106855.2
|
Mier1
|
MEIR1 treanscription regulator |
| chr10_-_95509251 | 0.16 |
ENSMUST00000218771.2
ENSMUST00000099328.2 |
Anapc15-ps
|
anaphase promoting complex C subunit 15, pseudogene |
| chr3_+_92864693 | 0.15 |
ENSMUST00000059053.11
|
Lce3d
|
late cornified envelope 3D |
| chr7_+_108064533 | 0.15 |
ENSMUST00000217616.3
|
Olfr498
|
olfactory receptor 498 |
| chr16_+_38383182 | 0.15 |
ENSMUST00000163948.8
|
Tmem39a
|
transmembrane protein 39a |
| chr13_-_64237011 | 0.14 |
ENSMUST00000166224.8
ENSMUST00000222810.2 |
Hsd17b3
|
hydroxysteroid (17-beta) dehydrogenase 3 |
| chr18_-_15196612 | 0.14 |
ENSMUST00000168989.9
|
Kctd1
|
potassium channel tetramerisation domain containing 1 |
| chr18_+_36498826 | 0.14 |
ENSMUST00000144158.2
|
Cystm1
|
cysteine-rich transmembrane module containing 1 |
| chr11_-_72302520 | 0.14 |
ENSMUST00000108500.8
ENSMUST00000050226.7 |
Smtnl2
|
smoothelin-like 2 |
| chr1_+_75526225 | 0.14 |
ENSMUST00000154101.8
|
Slc4a3
|
solute carrier family 4 (anion exchanger), member 3 |
| chr2_-_85509111 | 0.14 |
ENSMUST00000099917.4
|
Olfr1006
|
olfactory receptor 1006 |
| chr8_-_61407760 | 0.14 |
ENSMUST00000110302.8
|
Clcn3
|
chloride channel, voltage-sensitive 3 |
| chr15_-_99355623 | 0.14 |
ENSMUST00000023747.14
|
Nckap5l
|
NCK-associated protein 5-like |
| chr6_-_67931692 | 0.13 |
ENSMUST00000103313.3
|
Igkv9-123
|
immunoglobulin kappa variable 9-123 |
| chr3_-_129548954 | 0.13 |
ENSMUST00000029653.7
|
Egf
|
epidermal growth factor |
| chr10_+_19588318 | 0.13 |
ENSMUST00000020185.5
|
Il20ra
|
interleukin 20 receptor, alpha |
| chrX_-_71318353 | 0.13 |
ENSMUST00000064780.4
|
Gabre
|
gamma-aminobutyric acid (GABA) A receptor, subunit epsilon |
| chr4_+_104224774 | 0.12 |
ENSMUST00000106830.9
|
Dab1
|
disabled 1 |
| chr7_+_3337591 | 0.12 |
ENSMUST00000203328.4
|
Myadm
|
myeloid-associated differentiation marker |
| chr19_-_45034156 | 0.12 |
ENSMUST00000169459.4
|
Pdzd7
|
PDZ domain containing 7 |
| chr2_+_30485048 | 0.12 |
ENSMUST00000102853.4
|
Cstad
|
CSA-conditional, T cell activation-dependent protein |
| chr15_-_101555815 | 0.12 |
ENSMUST00000229963.2
ENSMUST00000100184.4 |
Gm5478
|
predicted pseudogene 5478 |
| chr14_+_55132030 | 0.12 |
ENSMUST00000141446.8
ENSMUST00000139985.8 |
Pabpn1
|
poly(A) binding protein, nuclear 1 |
| chr11_-_51153870 | 0.12 |
ENSMUST00000054226.3
|
BC049762
|
cDNA sequence BC049762 |
| chrX_-_7185424 | 0.12 |
ENSMUST00000115746.8
|
Clcn5
|
chloride channel, voltage-sensitive 5 |
| chr14_+_56905698 | 0.11 |
ENSMUST00000116468.2
|
Mphosph8
|
M-phase phosphoprotein 8 |
| chr8_+_120339440 | 0.11 |
ENSMUST00000098361.4
|
Adad2
|
adenosine deaminase domain containing 2 |
| chr4_+_33078796 | 0.10 |
ENSMUST00000131920.8
|
Gabrr2
|
gamma-aminobutyric acid (GABA) C receptor, subunit rho 2 |
| chr7_+_126832215 | 0.10 |
ENSMUST00000106312.4
|
Zfp553
|
zinc finger protein 553 |
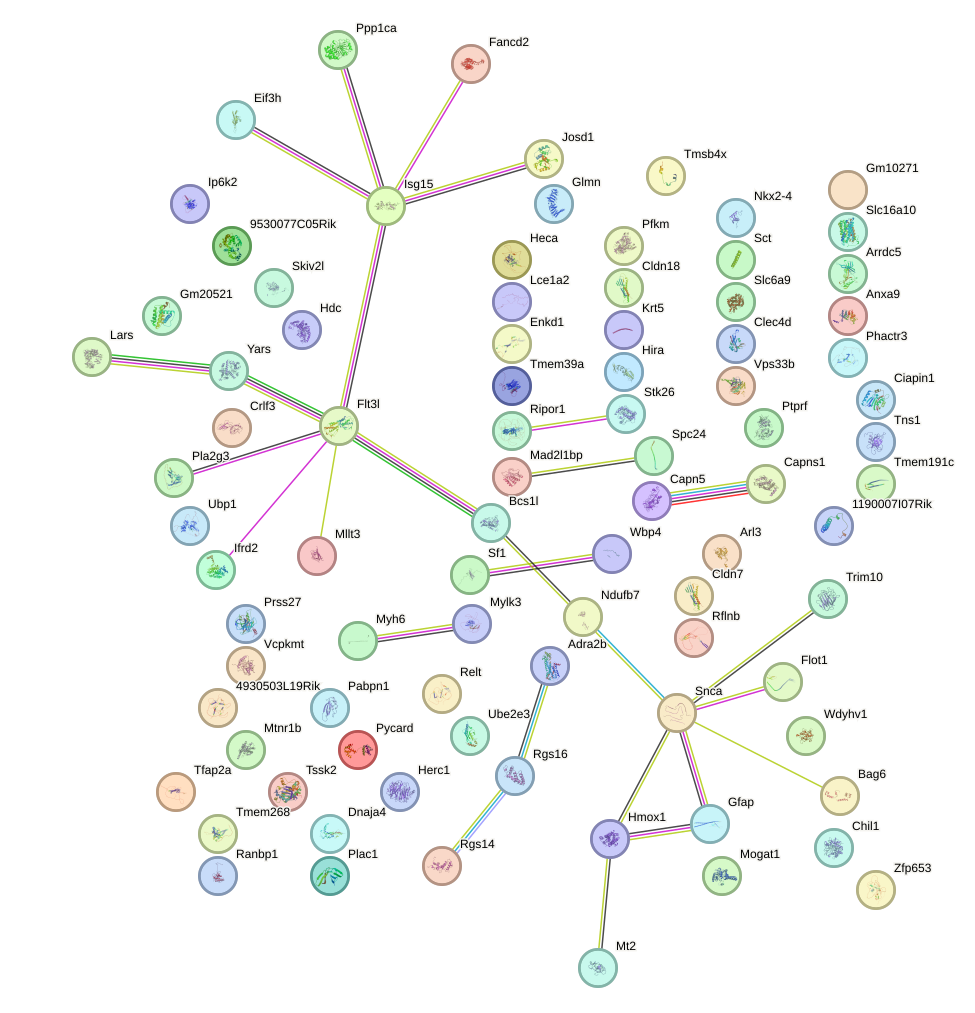
Network of associatons between targets according to the STRING database.
First level regulatory network of Zbtb6
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 3.6 | GO:2000295 | regulation of hydrogen peroxide catabolic process(GO:2000295) |
| 1.0 | 5.8 | GO:0002528 | regulation of vascular permeability involved in acute inflammatory response(GO:0002528) |
| 0.9 | 5.6 | GO:1904714 | regulation of chaperone-mediated autophagy(GO:1904714) |
| 0.7 | 2.2 | GO:0061536 | glycine secretion(GO:0061536) glycine secretion, neurotransmission(GO:0061537) |
| 0.6 | 1.7 | GO:0042822 | pyridoxal phosphate metabolic process(GO:0042822) |
| 0.5 | 2.0 | GO:0090290 | positive regulation of osteoclast proliferation(GO:0090290) |
| 0.4 | 8.6 | GO:0007379 | segment specification(GO:0007379) |
| 0.3 | 1.0 | GO:1900158 | negative regulation of bone mineralization involved in bone maturation(GO:1900158) |
| 0.3 | 0.9 | GO:0006788 | heme oxidation(GO:0006788) |
| 0.3 | 4.3 | GO:0090084 | negative regulation of inclusion body assembly(GO:0090084) |
| 0.3 | 2.0 | GO:0010273 | detoxification of copper ion(GO:0010273) stress response to copper ion(GO:1990169) |
| 0.2 | 0.7 | GO:0006429 | leucyl-tRNA aminoacylation(GO:0006429) |
| 0.2 | 1.4 | GO:0010836 | negative regulation of protein ADP-ribosylation(GO:0010836) |
| 0.2 | 1.1 | GO:0061622 | glycolytic process through glucose-1-phosphate(GO:0061622) |
| 0.2 | 1.7 | GO:0032020 | ISG15-protein conjugation(GO:0032020) |
| 0.2 | 0.9 | GO:0006548 | histamine metabolic process(GO:0001692) histidine catabolic process(GO:0006548) |
| 0.2 | 0.7 | GO:0007522 | visceral muscle development(GO:0007522) |
| 0.2 | 0.5 | GO:0001698 | gastrin-induced gastric acid secretion(GO:0001698) |
| 0.2 | 4.2 | GO:0046597 | negative regulation of viral entry into host cell(GO:0046597) |
| 0.2 | 0.7 | GO:1904247 | positive regulation of polynucleotide adenylyltransferase activity(GO:1904247) |
| 0.2 | 6.3 | GO:0051016 | barbed-end actin filament capping(GO:0051016) |
| 0.2 | 3.5 | GO:0007250 | activation of NF-kappaB-inducing kinase activity(GO:0007250) |
| 0.1 | 1.4 | GO:1900364 | negative regulation of mRNA polyadenylation(GO:1900364) |
| 0.1 | 0.6 | GO:0003409 | optic cup structural organization(GO:0003409) |
| 0.1 | 0.9 | GO:0032929 | negative regulation of superoxide anion generation(GO:0032929) |
| 0.1 | 1.0 | GO:2000348 | regulation of CD40 signaling pathway(GO:2000348) |
| 0.1 | 0.5 | GO:0032053 | ciliary basal body organization(GO:0032053) |
| 0.1 | 0.1 | GO:0021623 | oculomotor nerve morphogenesis(GO:0021622) oculomotor nerve formation(GO:0021623) |
| 0.1 | 0.5 | GO:0090214 | spongiotrophoblast layer developmental growth(GO:0090214) |
| 0.1 | 0.4 | GO:2001107 | negative regulation of Rho guanyl-nucleotide exchange factor activity(GO:2001107) |
| 0.1 | 1.7 | GO:0007096 | regulation of exit from mitosis(GO:0007096) |
| 0.1 | 0.3 | GO:0046604 | positive regulation of mitotic centrosome separation(GO:0046604) |
| 0.1 | 0.6 | GO:0038094 | Fc-gamma receptor signaling pathway(GO:0038094) |
| 0.1 | 0.3 | GO:0035524 | proline transmembrane transport(GO:0035524) |
| 0.1 | 0.4 | GO:2000049 | positive regulation of cell-cell adhesion mediated by cadherin(GO:2000049) |
| 0.1 | 0.4 | GO:0035624 | receptor transactivation(GO:0035624) epidermal growth factor-activated receptor transactivation by G-protein coupled receptor signaling pathway(GO:0035625) |
| 0.1 | 0.8 | GO:0005981 | regulation of glycogen catabolic process(GO:0005981) |
| 0.1 | 0.4 | GO:2001205 | negative regulation of osteoclast development(GO:2001205) |
| 0.1 | 0.6 | GO:0034427 | nuclear-transcribed mRNA catabolic process, exonucleolytic, 3'-5'(GO:0034427) |
| 0.1 | 0.7 | GO:0034551 | respiratory chain complex III assembly(GO:0017062) mitochondrial respiratory chain complex III assembly(GO:0034551) |
| 0.0 | 0.4 | GO:0038026 | reelin-mediated signaling pathway(GO:0038026) |
| 0.0 | 0.5 | GO:0061370 | testosterone biosynthetic process(GO:0061370) |
| 0.0 | 0.3 | GO:0006651 | diacylglycerol biosynthetic process(GO:0006651) |
| 0.0 | 0.1 | GO:0097401 | synaptic vesicle lumen acidification(GO:0097401) |
| 0.0 | 0.5 | GO:0034651 | cortisol biosynthetic process(GO:0034651) |
| 0.0 | 0.2 | GO:0099558 | maintenance of synapse structure(GO:0099558) |
| 0.0 | 0.6 | GO:0031914 | negative regulation of synaptic plasticity(GO:0031914) |
| 0.0 | 0.3 | GO:0070889 | platelet alpha granule organization(GO:0070889) |
| 0.0 | 0.5 | GO:0019372 | lipoxygenase pathway(GO:0019372) |
| 0.0 | 0.1 | GO:0097477 | hindbrain structural organization(GO:0021577) cerebellum structural organization(GO:0021589) lateral motor column neuron migration(GO:0097477) |
| 0.0 | 1.8 | GO:0071158 | positive regulation of cell cycle arrest(GO:0071158) |
| 0.0 | 0.3 | GO:0036506 | maintenance of unfolded protein(GO:0036506) maintenance of unfolded protein involved in ERAD pathway(GO:1904378) |
| 0.0 | 0.2 | GO:0033058 | directional locomotion(GO:0033058) |
| 0.0 | 0.4 | GO:0031118 | rRNA pseudouridine synthesis(GO:0031118) tRNA pseudouridine synthesis(GO:0031119) |
| 0.0 | 0.6 | GO:0021702 | cerebellar Purkinje cell differentiation(GO:0021702) |
| 0.0 | 0.7 | GO:0045292 | mRNA cis splicing, via spliceosome(GO:0045292) |
| 0.0 | 0.2 | GO:0006420 | arginyl-tRNA aminoacylation(GO:0006420) |
| 0.0 | 0.2 | GO:0015015 | heparan sulfate proteoglycan biosynthetic process, enzymatic modification(GO:0015015) |
| 0.0 | 1.1 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
| 0.0 | 1.4 | GO:0006418 | tRNA aminoacylation for protein translation(GO:0006418) |
| 0.0 | 0.2 | GO:0060082 | response to carbon monoxide(GO:0034465) eye blink reflex(GO:0060082) |
| 0.0 | 0.1 | GO:1990768 | regulation of gastric mucosal blood circulation(GO:1904344) positive regulation of gastric mucosal blood circulation(GO:1904346) gastric mucosal blood circulation(GO:1990768) |
| 0.0 | 0.6 | GO:0032463 | negative regulation of protein homooligomerization(GO:0032463) |
| 0.0 | 0.6 | GO:0006336 | DNA replication-independent nucleosome assembly(GO:0006336) |
| 0.0 | 0.1 | GO:0090370 | negative regulation of cholesterol efflux(GO:0090370) |
| 0.0 | 0.6 | GO:0075522 | IRES-dependent viral translational initiation(GO:0075522) |
| 0.0 | 0.3 | GO:0035269 | protein O-linked mannosylation(GO:0035269) |
| 0.0 | 0.4 | GO:0051152 | positive regulation of smooth muscle cell differentiation(GO:0051152) |
| 0.0 | 0.5 | GO:0031163 | iron-sulfur cluster assembly(GO:0016226) metallo-sulfur cluster assembly(GO:0031163) |
| 0.0 | 0.3 | GO:0045086 | positive regulation of interleukin-2 biosynthetic process(GO:0045086) |
| 0.0 | 0.3 | GO:0042759 | long-chain fatty acid biosynthetic process(GO:0042759) |
| 0.0 | 0.4 | GO:0032780 | negative regulation of ATPase activity(GO:0032780) |
| 0.0 | 0.3 | GO:2000394 | positive regulation of lamellipodium morphogenesis(GO:2000394) |
| 0.0 | 0.3 | GO:0046520 | sphingoid biosynthetic process(GO:0046520) |
| 0.0 | 0.1 | GO:0090038 | negative regulation of protein kinase C signaling(GO:0090038) |
| 0.0 | 0.2 | GO:0046549 | retinal cone cell development(GO:0046549) |
| 0.0 | 0.2 | GO:0036005 | response to macrophage colony-stimulating factor(GO:0036005) cellular response to macrophage colony-stimulating factor stimulus(GO:0036006) |
| 0.0 | 0.2 | GO:2000254 | regulation of male germ cell proliferation(GO:2000254) |
| 0.0 | 0.5 | GO:0007214 | gamma-aminobutyric acid signaling pathway(GO:0007214) |
| 0.0 | 0.4 | GO:0030033 | microvillus assembly(GO:0030033) |
| 0.0 | 0.1 | GO:0015824 | proline transport(GO:0015824) |
| 0.0 | 0.2 | GO:0031937 | positive regulation of chromatin silencing(GO:0031937) |
| 0.0 | 0.4 | GO:0046854 | phosphatidylinositol phosphorylation(GO:0046854) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 3.6 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.7 | 5.6 | GO:0098574 | cytoplasmic side of lysosomal membrane(GO:0098574) |
| 0.4 | 6.3 | GO:0008290 | F-actin capping protein complex(GO:0008290) |
| 0.4 | 2.2 | GO:0031262 | Ndc80 complex(GO:0031262) |
| 0.4 | 1.4 | GO:0032021 | NELF complex(GO:0032021) |
| 0.2 | 1.1 | GO:0005945 | 6-phosphofructokinase complex(GO:0005945) |
| 0.2 | 1.4 | GO:0070522 | ERCC4-ERCC1 complex(GO:0070522) |
| 0.2 | 0.6 | GO:0055087 | Ski complex(GO:0055087) |
| 0.1 | 8.6 | GO:0008023 | transcription elongation factor complex(GO:0008023) |
| 0.1 | 2.1 | GO:0017101 | aminoacyl-tRNA synthetase multienzyme complex(GO:0017101) |
| 0.1 | 0.8 | GO:0072357 | PTW/PP1 phosphatase complex(GO:0072357) |
| 0.1 | 1.7 | GO:0070938 | contractile ring(GO:0070938) |
| 0.1 | 2.0 | GO:0031233 | intrinsic component of external side of plasma membrane(GO:0031233) |
| 0.0 | 0.2 | GO:0044301 | climbing fiber(GO:0044301) |
| 0.0 | 0.4 | GO:0071541 | eukaryotic translation initiation factor 3 complex, eIF3m(GO:0071541) |
| 0.0 | 0.3 | GO:0072379 | BAT3 complex(GO:0071818) ER membrane insertion complex(GO:0072379) |
| 0.0 | 0.4 | GO:0001520 | outer dense fiber(GO:0001520) |
| 0.0 | 0.7 | GO:0042405 | nuclear inclusion body(GO:0042405) |
| 0.0 | 0.4 | GO:0001931 | uropod(GO:0001931) cell trailing edge(GO:0031254) |
| 0.0 | 0.3 | GO:0017059 | serine C-palmitoyltransferase complex(GO:0017059) endoplasmic reticulum palmitoyltransferase complex(GO:0031211) |
| 0.0 | 0.3 | GO:0031464 | Cul4A-RING E3 ubiquitin ligase complex(GO:0031464) |
| 0.0 | 0.3 | GO:0000322 | storage vacuole(GO:0000322) |
| 0.0 | 0.1 | GO:1990696 | USH2 complex(GO:1990696) |
| 0.0 | 0.3 | GO:0071439 | clathrin complex(GO:0071439) |
| 0.0 | 0.5 | GO:0005859 | muscle myosin complex(GO:0005859) |
| 0.0 | 0.5 | GO:0035253 | ciliary rootlet(GO:0035253) |
| 0.0 | 0.6 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.0 | 0.4 | GO:1902711 | GABA-A receptor complex(GO:1902711) |
| 0.0 | 0.4 | GO:0045095 | keratin filament(GO:0045095) |
| 0.0 | 1.3 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.0 | 0.4 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.0 | 0.2 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.2 | GO:0070773 | protein-N-terminal glutamine amidohydrolase activity(GO:0070773) |
| 0.6 | 5.8 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.4 | 1.7 | GO:0033883 | pyridoxal phosphatase activity(GO:0033883) |
| 0.4 | 1.8 | GO:0015173 | aromatic amino acid transmembrane transporter activity(GO:0015173) |
| 0.3 | 3.6 | GO:1903136 | cuprous ion binding(GO:1903136) |
| 0.3 | 0.9 | GO:0004392 | heme oxygenase (decyclizing) activity(GO:0004392) |
| 0.3 | 3.5 | GO:0008061 | chitin binding(GO:0008061) |
| 0.2 | 0.7 | GO:0004823 | glutamine-tRNA ligase activity(GO:0004819) leucine-tRNA ligase activity(GO:0004823) |
| 0.2 | 1.4 | GO:1990599 | 3' overhang single-stranded DNA endodeoxyribonuclease activity(GO:1990599) |
| 0.2 | 1.1 | GO:0003872 | 6-phosphofructokinase activity(GO:0003872) |
| 0.2 | 2.2 | GO:0015187 | glycine transmembrane transporter activity(GO:0015187) |
| 0.1 | 0.5 | GO:0008332 | low voltage-gated calcium channel activity(GO:0008332) |
| 0.1 | 0.3 | GO:0005171 | hepatocyte growth factor receptor binding(GO:0005171) |
| 0.1 | 0.4 | GO:0038025 | reelin receptor activity(GO:0038025) |
| 0.1 | 0.4 | GO:0004938 | alpha2-adrenergic receptor activity(GO:0004938) |
| 0.1 | 0.7 | GO:0030899 | calcium-dependent ATPase activity(GO:0030899) |
| 0.1 | 1.7 | GO:0031386 | protein tag(GO:0031386) |
| 0.1 | 2.3 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.1 | 6.4 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.1 | 0.5 | GO:0047045 | testosterone 17-beta-dehydrogenase (NADP+) activity(GO:0047045) |
| 0.1 | 0.4 | GO:0000832 | inositol hexakisphosphate kinase activity(GO:0000828) inositol hexakisphosphate 5-kinase activity(GO:0000832) inositol hexakisphosphate 1-kinase activity(GO:0052723) inositol hexakisphosphate 3-kinase activity(GO:0052724) |
| 0.1 | 0.2 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.1 | 0.9 | GO:0003993 | acid phosphatase activity(GO:0003993) |
| 0.1 | 0.3 | GO:0004144 | diacylglycerol O-acyltransferase activity(GO:0004144) |
| 0.1 | 0.3 | GO:0016454 | serine C-palmitoyltransferase activity(GO:0004758) C-palmitoyltransferase activity(GO:0016454) |
| 0.1 | 0.3 | GO:0015193 | L-proline transmembrane transporter activity(GO:0015193) |
| 0.0 | 0.1 | GO:0042015 | interleukin-20 binding(GO:0042015) |
| 0.0 | 5.4 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.0 | 0.6 | GO:1990829 | C-rich single-stranded DNA binding(GO:1990829) |
| 0.0 | 1.0 | GO:0031005 | filamin binding(GO:0031005) |
| 0.0 | 1.0 | GO:0015464 | acetylcholine receptor activity(GO:0015464) |
| 0.0 | 0.1 | GO:0072320 | volume-sensitive chloride channel activity(GO:0072320) |
| 0.0 | 4.3 | GO:0051087 | chaperone binding(GO:0051087) |
| 0.0 | 0.8 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 0.0 | 0.2 | GO:0004814 | arginine-tRNA ligase activity(GO:0004814) |
| 0.0 | 0.2 | GO:0008467 | [heparan sulfate]-glucosamine 3-sulfotransferase 1 activity(GO:0008467) |
| 0.0 | 1.0 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.0 | 0.3 | GO:0022851 | GABA-gated chloride ion channel activity(GO:0022851) |
| 0.0 | 1.4 | GO:0016876 | aminoacyl-tRNA ligase activity(GO:0004812) ligase activity, forming carbon-oxygen bonds(GO:0016875) ligase activity, forming aminoacyl-tRNA and related compounds(GO:0016876) |
| 0.0 | 0.1 | GO:0031768 | ghrelin receptor binding(GO:0031768) |
| 0.0 | 0.9 | GO:0005035 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.0 | 0.4 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 0.0 | 0.3 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.0 | 0.7 | GO:0008143 | poly(A) binding(GO:0008143) |
| 0.0 | 0.2 | GO:0060072 | large conductance calcium-activated potassium channel activity(GO:0060072) |
| 0.0 | 0.3 | GO:1990381 | ubiquitin-specific protease binding(GO:1990381) |
| 0.0 | 1.1 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.0 | 0.4 | GO:0009982 | pseudouridine synthase activity(GO:0009982) |
| 0.0 | 0.2 | GO:0004972 | NMDA glutamate receptor activity(GO:0004972) |
| 0.0 | 0.2 | GO:0004117 | calmodulin-dependent cyclic-nucleotide phosphodiesterase activity(GO:0004117) calcium- and calmodulin-regulated 3',5'-cyclic-GMP phosphodiesterase activity(GO:0048101) |
| 0.0 | 0.6 | GO:0030159 | receptor signaling complex scaffold activity(GO:0030159) |
| 0.0 | 0.9 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.0 | 0.7 | GO:0070064 | proline-rich region binding(GO:0070064) |
| 0.0 | 0.5 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 0.0 | 0.3 | GO:0002162 | dystroglycan binding(GO:0002162) |
| 0.0 | 0.3 | GO:0008137 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.0 | 0.2 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 0.0 | 1.2 | GO:0030971 | receptor tyrosine kinase binding(GO:0030971) |
| 0.0 | 0.9 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.0 | 0.2 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.0 | 0.2 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 3.6 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.0 | 2.2 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.0 | 1.4 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.0 | 1.0 | PID ATM PATHWAY | ATM pathway |
| 0.0 | 0.9 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.0 | 0.8 | PID ALK1 PATHWAY | ALK1 signaling events |
| 0.0 | 1.4 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.0 | 0.6 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 3.6 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.1 | 2.5 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.1 | 1.4 | REACTOME RESOLUTION OF AP SITES VIA THE SINGLE NUCLEOTIDE REPLACEMENT PATHWAY | Genes involved in Resolution of AP sites via the single-nucleotide replacement pathway |
| 0.1 | 1.0 | REACTOME REGULATION OF THE FANCONI ANEMIA PATHWAY | Genes involved in Regulation of the Fanconi anemia pathway |
| 0.1 | 5.6 | REACTOME NUCLEAR SIGNALING BY ERBB4 | Genes involved in Nuclear signaling by ERBB4 |
| 0.1 | 2.1 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 0.0 | 1.9 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |
| 0.0 | 0.9 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.0 | 1.8 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.0 | 0.5 | REACTOME ANDROGEN BIOSYNTHESIS | Genes involved in Androgen biosynthesis |
| 0.0 | 0.7 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.0 | 0.8 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.0 | 0.5 | REACTOME ACYL CHAIN REMODELLING OF PG | Genes involved in Acyl chain remodelling of PG |
| 0.0 | 0.4 | REACTOME PLATELET SENSITIZATION BY LDL | Genes involved in Platelet sensitization by LDL |
| 0.0 | 0.8 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.0 | 2.2 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.0 | 0.2 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.0 | 0.8 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.0 | 0.3 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.0 | 0.2 | REACTOME ROLE OF SECOND MESSENGERS IN NETRIN1 SIGNALING | Genes involved in Role of second messengers in netrin-1 signaling |
| 0.0 | 0.4 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.0 | 0.9 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.0 | 0.7 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |