Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
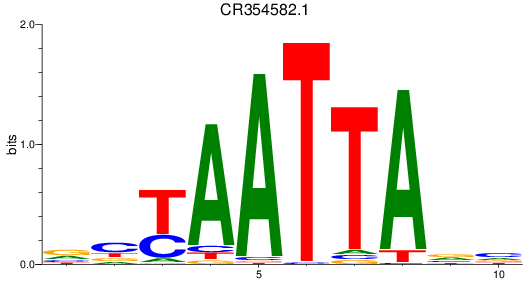
Results for CR354582.1
Z-value: 0.72
Transcription factors associated with CR354582.1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
CR354582.1
|
ENSDARG00000110598 | ENSDARG00000110598 |
Activity profile of CR354582.1 motif
Sorted Z-values of CR354582.1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr21_+_25777425 | 11.61 |
ENSDART00000021620
|
cldnd
|
claudin d |
| chr12_-_35830625 | 11.43 |
ENSDART00000180028
|
CU459056.1
|
|
| chr9_-_35633827 | 11.19 |
ENSDART00000077745
|
zp2l1
|
zona pellucida glycoprotein 2, like 1 |
| chr2_-_15324837 | 10.91 |
ENSDART00000015655
|
tecrl2b
|
trans-2,3-enoyl-CoA reductase-like 2b |
| chr20_-_23426339 | 10.58 |
ENSDART00000004625
|
zar1
|
zygote arrest 1 |
| chr23_+_20110086 | 8.52 |
ENSDART00000054664
|
tnnc1b
|
troponin C type 1b (slow) |
| chr21_-_5205617 | 8.45 |
ENSDART00000145554
ENSDART00000045284 |
rpl37
|
ribosomal protein L37 |
| chr3_+_18398876 | 7.90 |
ENSDART00000141100
ENSDART00000138107 |
rps2
|
ribosomal protein S2 |
| chr10_-_21362320 | 7.76 |
ENSDART00000189789
|
avd
|
avidin |
| chr10_-_21362071 | 7.52 |
ENSDART00000125167
|
avd
|
avidin |
| chr15_-_23376541 | 7.21 |
ENSDART00000078570
|
c1qtnf5
|
C1q and TNF related 5 |
| chr20_-_40755614 | 7.02 |
ENSDART00000061247
|
cx32.3
|
connexin 32.3 |
| chr24_+_38306010 | 6.74 |
ENSDART00000143184
|
mybpc2b
|
myosin binding protein C, fast type b |
| chr6_-_7720332 | 6.45 |
ENSDART00000135945
|
rpsa
|
ribosomal protein SA |
| chr12_+_22580579 | 6.31 |
ENSDART00000171725
ENSDART00000192290 |
capgb
|
capping protein (actin filament), gelsolin-like b |
| chr24_-_37640705 | 6.06 |
ENSDART00000066583
|
zgc:112496
|
zgc:112496 |
| chr16_-_42056137 | 5.81 |
ENSDART00000102798
|
zp3d.2
|
zona pellucida glycoprotein 3d tandem duplicate 2 |
| chr6_+_41191482 | 5.35 |
ENSDART00000000877
|
opn1mw3
|
opsin 1 (cone pigments), medium-wave-sensitive, 3 |
| chr10_-_34002185 | 5.34 |
ENSDART00000046599
|
zar1l
|
zygote arrest 1-like |
| chr2_+_6253246 | 5.31 |
ENSDART00000058256
ENSDART00000076700 |
zp3b
|
zona pellucida glycoprotein 3b |
| chr11_+_18183220 | 5.30 |
ENSDART00000113468
|
LO018315.10
|
|
| chr19_-_13808630 | 5.09 |
ENSDART00000166895
ENSDART00000187670 |
ctgfb
|
connective tissue growth factor b |
| chr8_-_23780334 | 5.01 |
ENSDART00000145179
ENSDART00000145894 |
zgc:195245
|
zgc:195245 |
| chr6_+_49926115 | 4.88 |
ENSDART00000018523
|
ahcy
|
adenosylhomocysteinase |
| chr23_+_44374041 | 4.86 |
ENSDART00000136056
|
ephb4b
|
eph receptor B4b |
| chr10_-_2971407 | 4.60 |
ENSDART00000132526
|
marveld2a
|
MARVEL domain containing 2a |
| chr18_+_9171778 | 4.45 |
ENSDART00000101192
|
sema3d
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3D |
| chr24_+_19415124 | 4.42 |
ENSDART00000186931
|
sulf1
|
sulfatase 1 |
| chr21_-_35419486 | 4.27 |
ENSDART00000138529
|
si:dkeyp-23e4.3
|
si:dkeyp-23e4.3 |
| chr15_-_47848544 | 4.23 |
ENSDART00000098711
|
eif3k
|
eukaryotic translation initiation factor 3, subunit K |
| chr10_+_21867307 | 4.14 |
ENSDART00000126629
|
cbln17
|
cerebellin 17 |
| chr21_+_11244068 | 4.13 |
ENSDART00000163432
|
arid6
|
AT-rich interaction domain 6 |
| chr17_+_16046132 | 4.12 |
ENSDART00000155005
|
si:ch73-204p21.2
|
si:ch73-204p21.2 |
| chr22_-_10121880 | 4.09 |
ENSDART00000002348
|
rdh5
|
retinol dehydrogenase 5 (11-cis/9-cis) |
| chr11_-_6048490 | 4.03 |
ENSDART00000066164
|
plvapb
|
plasmalemma vesicle associated protein b |
| chr18_+_48423973 | 4.03 |
ENSDART00000184233
ENSDART00000147074 |
fli1a
|
Fli-1 proto-oncogene, ETS transcription factor a |
| chr8_-_24252933 | 4.03 |
ENSDART00000057624
|
zgc:110353
|
zgc:110353 |
| chr17_+_16046314 | 3.95 |
ENSDART00000154554
ENSDART00000154338 ENSDART00000155336 |
si:ch73-204p21.2
|
si:ch73-204p21.2 |
| chr13_-_31017960 | 3.93 |
ENSDART00000145287
|
wdfy4
|
WDFY family member 4 |
| chr12_-_48477031 | 3.91 |
ENSDART00000105176
|
ndufb8
|
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 8 |
| chr10_+_6884627 | 3.89 |
ENSDART00000125262
ENSDART00000121729 ENSDART00000105384 |
ddx4
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 4 |
| chr8_+_11425048 | 3.88 |
ENSDART00000018739
|
tjp2b
|
tight junction protein 2b (zona occludens 2) |
| chr5_-_63302944 | 3.80 |
ENSDART00000047110
|
gsnb
|
gelsolin b |
| chr9_+_50001746 | 3.68 |
ENSDART00000058892
|
slc38a11
|
solute carrier family 38, member 11 |
| chr19_+_31585917 | 3.67 |
ENSDART00000132182
|
gmnn
|
geminin, DNA replication inhibitor |
| chr11_+_24703108 | 3.57 |
ENSDART00000159173
|
gpr25
|
G protein-coupled receptor 25 |
| chr22_-_10586191 | 3.56 |
ENSDART00000148418
|
si:dkey-42i9.16
|
si:dkey-42i9.16 |
| chr4_-_9891874 | 3.48 |
ENSDART00000067193
|
adm2a
|
adrenomedullin 2a |
| chr17_+_8799451 | 3.40 |
ENSDART00000189814
ENSDART00000191577 |
tonsl
|
tonsoku-like, DNA repair protein |
| chr12_+_20352400 | 3.39 |
ENSDART00000066383
|
hbae5
|
hemoglobin, alpha embryonic 5 |
| chr24_+_1023839 | 3.39 |
ENSDART00000082526
|
zgc:111976
|
zgc:111976 |
| chr12_-_14143344 | 3.34 |
ENSDART00000152742
|
buc2l
|
bucky ball 2-like |
| chr19_-_657439 | 3.27 |
ENSDART00000167100
|
slc6a18
|
solute carrier family 6 (neutral amino acid transporter), member 18 |
| chr10_-_35257458 | 3.18 |
ENSDART00000143890
ENSDART00000139107 ENSDART00000082445 |
prr11
|
proline rich 11 |
| chr3_+_17537352 | 3.15 |
ENSDART00000104549
|
hcrt
|
hypocretin (orexin) neuropeptide precursor |
| chr24_-_25144441 | 3.14 |
ENSDART00000152104
|
phldb2b
|
pleckstrin homology-like domain, family B, member 2b |
| chr11_-_30634286 | 3.10 |
ENSDART00000191019
|
zgc:153665
|
zgc:153665 |
| chr24_-_34680956 | 3.08 |
ENSDART00000171009
|
ctnna1
|
catenin (cadherin-associated protein), alpha 1 |
| chr13_+_35637048 | 3.05 |
ENSDART00000085037
|
thbs2a
|
thrombospondin 2a |
| chr23_-_24394719 | 3.04 |
ENSDART00000044918
|
epha2b
|
eph receptor A2 b |
| chr8_+_45334255 | 3.03 |
ENSDART00000126848
ENSDART00000134161 ENSDART00000142322 ENSDART00000145011 ENSDART00000183560 |
pabpc1l
|
poly(A) binding protein, cytoplasmic 1-like |
| chr22_+_20427170 | 3.00 |
ENSDART00000136744
|
foxq2
|
forkhead box Q2 |
| chr16_-_22930925 | 2.98 |
ENSDART00000133819
|
si:dkey-246i14.3
|
si:dkey-246i14.3 |
| chr24_+_37640626 | 2.98 |
ENSDART00000008047
|
wdr24
|
WD repeat domain 24 |
| chr1_-_18811517 | 2.97 |
ENSDART00000142026
|
si:dkey-167i21.2
|
si:dkey-167i21.2 |
| chr11_-_1550709 | 2.97 |
ENSDART00000110097
|
si:ch73-303b9.1
|
si:ch73-303b9.1 |
| chr9_-_19161982 | 2.94 |
ENSDART00000081878
|
pou1f1
|
POU class 1 homeobox 1 |
| chr10_+_39212898 | 2.94 |
ENSDART00000159501
|
stt3a
|
STT3A, subunit of the oligosaccharyltransferase complex (catalytic) |
| chr3_+_17933553 | 2.93 |
ENSDART00000167731
ENSDART00000165644 |
cnp
|
2',3'-cyclic nucleotide 3' phosphodiesterase |
| chr18_+_20560442 | 2.91 |
ENSDART00000151990
|
wee2
|
WEE1 homolog 2 (S. pombe) |
| chr4_+_11464255 | 2.83 |
ENSDART00000008584
|
gdi2
|
GDP dissociation inhibitor 2 |
| chr23_+_2728095 | 2.81 |
ENSDART00000066086
|
zgc:114123
|
zgc:114123 |
| chr14_+_45471642 | 2.80 |
ENSDART00000126979
ENSDART00000172952 ENSDART00000173284 |
ubxn1
|
UBX domain protein 1 |
| chr14_+_33329761 | 2.78 |
ENSDART00000161138
|
sowahd
|
sosondowah ankyrin repeat domain family d |
| chr8_-_14126646 | 2.70 |
ENSDART00000027225
|
bgna
|
biglycan a |
| chr18_-_40708537 | 2.69 |
ENSDART00000077577
|
si:ch211-132b12.8
|
si:ch211-132b12.8 |
| chr12_+_48803098 | 2.62 |
ENSDART00000074768
|
ppifb
|
peptidylprolyl isomerase Fb |
| chr19_+_43297546 | 2.62 |
ENSDART00000168002
|
laptm5
|
lysosomal protein transmembrane 5 |
| chr22_+_19407531 | 2.56 |
ENSDART00000141060
|
si:dkey-78l4.2
|
si:dkey-78l4.2 |
| chr9_-_50001606 | 2.56 |
ENSDART00000161648
ENSDART00000168514 |
scn1a
|
sodium channel, voltage-gated, type I, alpha |
| chr6_+_102506 | 2.56 |
ENSDART00000172678
|
ldlrb
|
low density lipoprotein receptor b |
| chr3_+_13624815 | 2.55 |
ENSDART00000161451
|
pglyrp6
|
peptidoglycan recognition protein 6 |
| chr9_-_22057658 | 2.53 |
ENSDART00000101944
|
crygmxl1
|
crystallin, gamma MX, like 1 |
| chr25_-_13703826 | 2.51 |
ENSDART00000163398
|
pla2g15
|
phospholipase A2, group XV |
| chr1_-_55248496 | 2.47 |
ENSDART00000098615
|
nanos3
|
nanos homolog 3 |
| chr8_+_20776654 | 2.41 |
ENSDART00000135850
|
nfic
|
nuclear factor I/C |
| chr5_-_9625459 | 2.40 |
ENSDART00000143347
|
sh2b3
|
SH2B adaptor protein 3 |
| chr4_-_4387012 | 2.38 |
ENSDART00000191836
|
CU468826.3
|
Danio rerio U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit-related protein 1-like (LOC100331497), mRNA. |
| chr25_-_6049339 | 2.36 |
ENSDART00000075184
|
snx1a
|
sorting nexin 1a |
| chr22_-_13350240 | 2.35 |
ENSDART00000154095
ENSDART00000155118 |
si:ch211-227m13.1
|
si:ch211-227m13.1 |
| chr24_-_2450597 | 2.33 |
ENSDART00000188080
ENSDART00000093331 |
rreb1a
|
ras responsive element binding protein 1a |
| chr10_-_13343831 | 2.31 |
ENSDART00000135941
|
il11ra
|
interleukin 11 receptor, alpha |
| chr10_+_6884123 | 2.27 |
ENSDART00000149095
ENSDART00000148772 ENSDART00000149334 |
ddx4
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 4 |
| chr3_-_23643751 | 2.26 |
ENSDART00000078425
ENSDART00000140264 |
eve1
|
even-skipped-like1 |
| chr17_-_10122204 | 2.21 |
ENSDART00000160751
|
BX088587.1
|
|
| chr19_+_390298 | 2.21 |
ENSDART00000136361
|
sema6d
|
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6D |
| chr11_-_45138857 | 2.21 |
ENSDART00000166501
|
cant1b
|
calcium activated nucleotidase 1b |
| chr7_+_34592526 | 2.19 |
ENSDART00000173959
|
fhod1
|
formin homology 2 domain containing 1 |
| chr24_+_6107901 | 2.16 |
ENSDART00000156419
|
si:ch211-37e10.2
|
si:ch211-37e10.2 |
| chr15_+_31344472 | 2.16 |
ENSDART00000146695
ENSDART00000159182 ENSDART00000060125 |
or107-1
|
odorant receptor, family D, subfamily 107, member 1 |
| chr14_+_33329420 | 2.16 |
ENSDART00000171090
ENSDART00000164062 |
sowahd
|
sosondowah ankyrin repeat domain family d |
| chr10_+_36178713 | 2.16 |
ENSDART00000140816
|
or108-3
|
odorant receptor, family D, subfamily 108, member 3 |
| chr6_+_50381665 | 2.14 |
ENSDART00000141128
|
cyc1
|
cytochrome c-1 |
| chr5_+_6954162 | 2.09 |
ENSDART00000086666
|
stpg2
|
sperm-tail PG-rich repeat containing 2 |
| chr10_-_11385155 | 2.09 |
ENSDART00000064214
|
plac8.1
|
placenta-specific 8, tandem duplicate 1 |
| chr24_+_13316737 | 2.07 |
ENSDART00000191658
|
SBSPON
|
somatomedin B and thrombospondin type 1 domain containing |
| chr7_+_2236317 | 2.05 |
ENSDART00000075859
|
zgc:172065
|
zgc:172065 |
| chr2_+_8779164 | 2.04 |
ENSDART00000134308
|
zzz3
|
zinc finger, ZZ-type containing 3 |
| chr2_+_39021282 | 2.04 |
ENSDART00000056577
|
RBP1 (1 of many)
|
si:ch211-119o8.7 |
| chr25_-_10503043 | 2.01 |
ENSDART00000155404
|
cox8b
|
cytochrome c oxidase subunit 8b |
| chr25_+_35891342 | 1.95 |
ENSDART00000147093
|
lsm14aa
|
LSM14A mRNA processing body assembly factor a |
| chr23_-_17003533 | 1.95 |
ENSDART00000080545
|
dnmt3bb.2
|
DNA (cytosine-5-)-methyltransferase 3 beta, duplicate b.2 |
| chr8_-_12867128 | 1.92 |
ENSDART00000142201
|
slc2a6
|
solute carrier family 2 (facilitated glucose transporter), member 6 |
| chr16_+_33902006 | 1.88 |
ENSDART00000161807
ENSDART00000159474 |
gnl2
|
guanine nucleotide binding protein-like 2 (nucleolar) |
| chr5_-_15851953 | 1.88 |
ENSDART00000173101
|
si:dkey-1k23.3
|
si:dkey-1k23.3 |
| chr11_+_31864921 | 1.88 |
ENSDART00000180252
|
diaph3
|
diaphanous-related formin 3 |
| chr10_+_25726694 | 1.86 |
ENSDART00000140308
|
ugt5d1
|
UDP glucuronosyltransferase 5 family, polypeptide D1 |
| chr20_+_37294112 | 1.85 |
ENSDART00000076293
ENSDART00000140450 |
cx23
|
connexin 23 |
| chr6_+_52873822 | 1.85 |
ENSDART00000103138
|
or137-3
|
odorant receptor, family H, subfamily 137, member 3 |
| chr16_-_16761164 | 1.79 |
ENSDART00000135872
|
si:dkey-27n14.1
|
si:dkey-27n14.1 |
| chr7_-_51368681 | 1.78 |
ENSDART00000146385
|
arhgap36
|
Rho GTPase activating protein 36 |
| chr9_+_22364997 | 1.77 |
ENSDART00000188054
ENSDART00000046116 |
crygs3
|
crystallin, gamma S3 |
| chr10_+_35257651 | 1.76 |
ENSDART00000028940
|
styxl1
|
serine/threonine/tyrosine interacting-like 1 |
| chr12_+_16168342 | 1.73 |
ENSDART00000079326
ENSDART00000170024 |
lrp2b
|
low density lipoprotein receptor-related protein 2b |
| chr14_+_40874608 | 1.73 |
ENSDART00000168448
|
si:ch211-106m9.1
|
si:ch211-106m9.1 |
| chr22_+_28337429 | 1.71 |
ENSDART00000166177
|
impg2b
|
interphotoreceptor matrix proteoglycan 2b |
| chr7_-_23768234 | 1.70 |
ENSDART00000173981
|
si:ch211-200p22.4
|
si:ch211-200p22.4 |
| chr19_-_5265155 | 1.70 |
ENSDART00000145003
|
prf1.3
|
perforin 1.3 |
| chr10_+_39212601 | 1.67 |
ENSDART00000168778
|
stt3a
|
STT3A, subunit of the oligosaccharyltransferase complex (catalytic) |
| chr20_-_37813863 | 1.66 |
ENSDART00000147529
|
batf3
|
basic leucine zipper transcription factor, ATF-like 3 |
| chr12_+_10631266 | 1.64 |
ENSDART00000161455
|
csf3a
|
colony stimulating factor 3 (granulocyte) a |
| chr13_+_38817871 | 1.63 |
ENSDART00000187708
|
col19a1
|
collagen, type XIX, alpha 1 |
| chr18_+_2228737 | 1.61 |
ENSDART00000165301
|
rab27a
|
RAB27A, member RAS oncogene family |
| chr6_+_28208973 | 1.61 |
ENSDART00000171216
ENSDART00000171377 ENSDART00000167389 ENSDART00000166988 |
LSM2 (1 of many)
|
si:ch73-14h10.2 |
| chr13_+_38814521 | 1.61 |
ENSDART00000110976
|
col19a1
|
collagen, type XIX, alpha 1 |
| chr4_+_13586689 | 1.58 |
ENSDART00000067161
ENSDART00000138201 |
tnpo3
|
transportin 3 |
| chr21_-_22827548 | 1.56 |
ENSDART00000079161
|
angptl5
|
angiopoietin-like 5 |
| chr13_+_35339182 | 1.56 |
ENSDART00000019323
|
jag1b
|
jagged 1b |
| chr12_-_33357655 | 1.54 |
ENSDART00000066233
ENSDART00000148165 |
slc16a3
|
solute carrier family 16 (monocarboxylate transporter), member 3 |
| chr10_-_40421050 | 1.50 |
ENSDART00000141834
ENSDART00000164847 |
taar20w
|
trace amine associated receptor 20w |
| chr9_-_3934963 | 1.49 |
ENSDART00000062336
|
ubr3
|
ubiquitin protein ligase E3 component n-recognin 3 |
| chr18_-_15551360 | 1.48 |
ENSDART00000159915
ENSDART00000172690 |
ppfibp1b
|
PTPRF interacting protein, binding protein 1b (liprin beta 1) |
| chr10_+_40660772 | 1.46 |
ENSDART00000148007
|
taar19l
|
trace amine associated receptor 19l |
| chr14_+_34490445 | 1.46 |
ENSDART00000132193
ENSDART00000148044 |
wnt8a
|
wingless-type MMTV integration site family, member 8a |
| chr15_-_2493771 | 1.45 |
ENSDART00000184906
|
neu4
|
sialidase 4 |
| chr13_+_255067 | 1.44 |
ENSDART00000102505
|
foxg1d
|
forkhead box G1d |
| chr24_-_16899406 | 1.44 |
ENSDART00000148753
|
mtrr
|
5-methyltetrahydrofolate-homocysteine methyltransferase reductase |
| chr1_-_29139141 | 1.44 |
ENSDART00000075546
ENSDART00000133246 |
hsf2bp
|
heat shock transcription factor 2 binding protein |
| chr7_-_7764287 | 1.44 |
ENSDART00000173021
ENSDART00000113131 |
intu
|
inturned planar cell polarity protein |
| chr19_+_10339538 | 1.43 |
ENSDART00000151808
ENSDART00000151235 |
rcvrn3
|
recoverin 3 |
| chr2_+_20793982 | 1.43 |
ENSDART00000014785
|
prg4a
|
proteoglycan 4a |
| chr21_-_26490186 | 1.40 |
ENSDART00000009889
|
zgc:110540
|
zgc:110540 |
| chr2_-_23778180 | 1.39 |
ENSDART00000136782
|
si:dkey-24c2.7
|
si:dkey-24c2.7 |
| chr3_-_50443607 | 1.38 |
ENSDART00000074036
|
rcvrna
|
recoverin a |
| chr13_+_27232848 | 1.38 |
ENSDART00000138043
|
rin2
|
Ras and Rab interactor 2 |
| chr4_-_20292821 | 1.35 |
ENSDART00000136069
ENSDART00000192504 |
cacna2d4a
|
calcium channel, voltage-dependent, alpha 2/delta subunit 4a |
| chr4_-_56954002 | 1.31 |
ENSDART00000160934
|
si:dkey-269o24.1
|
si:dkey-269o24.1 |
| chr1_-_17715493 | 1.31 |
ENSDART00000133027
|
si:dkey-256e7.8
|
si:dkey-256e7.8 |
| chr8_-_15129573 | 1.30 |
ENSDART00000142358
|
bcar3
|
BCAR3, NSP family adaptor protein |
| chr15_+_47903864 | 1.25 |
ENSDART00000063835
|
otx5
|
orthodenticle homolog 5 |
| chr18_-_25646286 | 1.24 |
ENSDART00000099511
ENSDART00000186890 |
si:ch211-13k12.2
|
si:ch211-13k12.2 |
| chr4_-_9852318 | 1.22 |
ENSDART00000080702
|
glt8d2
|
glycosyltransferase 8 domain containing 2 |
| chr3_+_17933132 | 1.21 |
ENSDART00000104299
ENSDART00000162144 ENSDART00000162242 ENSDART00000166289 ENSDART00000171101 ENSDART00000164853 |
cnp
|
2',3'-cyclic nucleotide 3' phosphodiesterase |
| chr12_-_19007834 | 1.21 |
ENSDART00000153248
|
chadlb
|
chondroadherin-like b |
| chr20_-_9095105 | 1.20 |
ENSDART00000140792
|
oma1
|
OMA1 zinc metallopeptidase |
| chr4_+_306036 | 1.20 |
ENSDART00000103659
|
msgn1
|
mesogenin 1 |
| chr3_+_46635527 | 1.20 |
ENSDART00000153971
|
si:dkey-248g21.1
|
si:dkey-248g21.1 |
| chr10_+_42423318 | 1.19 |
ENSDART00000134282
|
npy8ar
|
neuropeptide Y receptor Y8a |
| chr9_+_41156818 | 1.18 |
ENSDART00000105764
ENSDART00000147052 |
stat4
|
signal transducer and activator of transcription 4 |
| chr17_-_23616626 | 1.17 |
ENSDART00000104730
|
ifit14
|
interferon-induced protein with tetratricopeptide repeats 14 |
| chr16_-_17347727 | 1.16 |
ENSDART00000144392
|
zyx
|
zyxin |
| chr11_-_3865472 | 1.16 |
ENSDART00000161426
|
gata2a
|
GATA binding protein 2a |
| chr21_+_15592426 | 1.15 |
ENSDART00000138207
|
smarcb1b
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily b, member 1b |
| chr11_-_44801968 | 1.15 |
ENSDART00000161846
|
map1lc3c
|
microtubule-associated protein 1 light chain 3 gamma |
| chr18_+_1837668 | 1.14 |
ENSDART00000164210
|
CABZ01079192.1
|
|
| chr8_+_25034544 | 1.12 |
ENSDART00000123300
|
ngrn
|
neugrin, neurite outgrowth associated |
| chr23_-_19230627 | 1.11 |
ENSDART00000007122
|
guca1b
|
guanylate cyclase activator 1B |
| chr6_+_3280939 | 1.11 |
ENSDART00000151359
|
kdm4aa
|
lysine (K)-specific demethylase 4A, genome duplicate a |
| chr17_+_37227936 | 1.08 |
ENSDART00000076009
|
hadhab
|
hydroxyacyl-CoA dehydrogenase trifunctional multienzyme complex subunit alpha b |
| chr18_+_2554666 | 1.08 |
ENSDART00000167218
|
p2ry2.1
|
purinergic receptor P2Y2, tandem duplicate 1 |
| chr22_+_28337204 | 1.07 |
ENSDART00000163352
|
impg2b
|
interphotoreceptor matrix proteoglycan 2b |
| chr21_-_17296789 | 1.06 |
ENSDART00000192180
|
gfi1b
|
growth factor independent 1B transcription repressor |
| chr11_-_15296805 | 1.05 |
ENSDART00000124968
|
rpn2
|
ribophorin II |
| chr7_+_48667081 | 1.03 |
ENSDART00000083473
|
trpm5
|
transient receptor potential cation channel, subfamily M, member 5 |
| chr22_-_5518117 | 1.01 |
ENSDART00000164613
|
CABZ01064972.1
|
|
| chr10_+_2799285 | 0.99 |
ENSDART00000030709
|
pnx
|
posterior neuron-specific homeobox |
| chr15_+_34988148 | 0.99 |
ENSDART00000076269
|
ccdc105
|
coiled-coil domain containing 105 |
| chr18_-_25568994 | 0.99 |
ENSDART00000133029
|
si:ch211-13k12.2
|
si:ch211-13k12.2 |
| chr24_-_35282568 | 0.98 |
ENSDART00000167406
ENSDART00000088609 |
sntg1
|
syntrophin, gamma 1 |
| chr4_-_71912457 | 0.96 |
ENSDART00000179685
|
si:dkey-92c21.1
|
si:dkey-92c21.1 |
| chr5_-_25733745 | 0.94 |
ENSDART00000051566
|
zgc:101016
|
zgc:101016 |
| chr1_-_5455498 | 0.94 |
ENSDART00000040368
ENSDART00000114035 |
mnx2b
|
motor neuron and pancreas homeobox 2b |
| chr15_+_1796313 | 0.92 |
ENSDART00000126253
|
fam124b
|
family with sequence similarity 124B |
| chr10_-_26512993 | 0.89 |
ENSDART00000188549
ENSDART00000193316 |
si:dkey-5g14.1
|
si:dkey-5g14.1 |
| chr16_+_42471455 | 0.88 |
ENSDART00000166640
|
si:ch211-215k15.5
|
si:ch211-215k15.5 |
| chr15_+_36309070 | 0.88 |
ENSDART00000157034
|
gmnc
|
geminin coiled-coil domain containing |
| chr6_+_50381347 | 0.85 |
ENSDART00000055504
|
cyc1
|
cytochrome c-1 |
| chr7_+_19495379 | 0.85 |
ENSDART00000180514
|
si:ch211-212k18.8
|
si:ch211-212k18.8 |
| chr10_-_32494499 | 0.85 |
ENSDART00000129395
|
uvrag
|
UV radiation resistance associated gene |
| chr10_+_17714866 | 0.85 |
ENSDART00000039969
|
slc20a1b
|
solute carrier family 20 (phosphate transporter), member 1b |
Network of associatons between targets according to the STRING database.
First level regulatory network of CR354582.1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.3 | 11.6 | GO:0016338 | calcium-independent cell-cell adhesion via plasma membrane cell-adhesion molecules(GO:0016338) |
| 1.1 | 3.4 | GO:0097067 | cellular response to thyroid hormone stimulus(GO:0097067) |
| 1.0 | 2.9 | GO:0060631 | regulation of meiosis I(GO:0060631) |
| 0.9 | 4.4 | GO:0007508 | larval development(GO:0002164) larval heart development(GO:0007508) |
| 0.9 | 2.6 | GO:0098581 | detection of biotic stimulus(GO:0009595) detection of bacterium(GO:0016045) detection of other organism(GO:0098543) detection of external biotic stimulus(GO:0098581) |
| 0.8 | 4.9 | GO:0033353 | S-adenosylmethionine cycle(GO:0033353) |
| 0.8 | 3.1 | GO:0042755 | eating behavior(GO:0042755) |
| 0.7 | 4.1 | GO:0042362 | fat-soluble vitamin biosynthetic process(GO:0042362) |
| 0.7 | 11.1 | GO:0035803 | binding of sperm to zona pellucida(GO:0007339) egg coat formation(GO:0035803) regulation of acrosome reaction(GO:0060046) positive regulation of acrosome reaction(GO:2000344) |
| 0.6 | 2.5 | GO:0030718 | germ-line stem cell population maintenance(GO:0030718) |
| 0.6 | 4.4 | GO:0021910 | smoothened signaling pathway involved in ventral spinal cord patterning(GO:0021910) regulation of myotome development(GO:2000290) |
| 0.5 | 1.5 | GO:0039015 | spinal cord anterior/posterior patterning(GO:0021512) cell proliferation involved in pronephros development(GO:0039015) cell proliferation involved in kidney development(GO:0072111) |
| 0.5 | 1.9 | GO:0000448 | cleavage in ITS2 between 5.8S rRNA and LSU-rRNA of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000448) |
| 0.5 | 8.5 | GO:0097034 | mitochondrial respiratory chain complex IV assembly(GO:0033617) mitochondrial respiratory chain complex IV biogenesis(GO:0097034) |
| 0.4 | 1.8 | GO:2001244 | positive regulation of intrinsic apoptotic signaling pathway(GO:2001244) |
| 0.4 | 1.2 | GO:0048340 | paraxial mesoderm morphogenesis(GO:0048340) paraxial mesoderm formation(GO:0048341) |
| 0.4 | 2.4 | GO:0034498 | early endosome to Golgi transport(GO:0034498) |
| 0.4 | 1.9 | GO:0032776 | DNA methylation on cytosine(GO:0032776) |
| 0.4 | 1.2 | GO:0070589 | cell wall biogenesis(GO:0042546) cell wall macromolecule biosynthetic process(GO:0044038) cellular component macromolecule biosynthetic process(GO:0070589) |
| 0.4 | 1.2 | GO:1902893 | pri-miRNA transcription from RNA polymerase II promoter(GO:0061614) regulation of pri-miRNA transcription from RNA polymerase II promoter(GO:1902893) positive regulation of pri-miRNA transcription from RNA polymerase II promoter(GO:1902895) |
| 0.4 | 6.2 | GO:0051445 | regulation of meiotic cell cycle(GO:0051445) |
| 0.4 | 2.8 | GO:0040038 | meiotic cytokinesis(GO:0033206) polar body extrusion after meiotic divisions(GO:0040038) |
| 0.3 | 11.5 | GO:0042761 | very long-chain fatty acid biosynthetic process(GO:0042761) |
| 0.3 | 2.6 | GO:0034063 | stress granule assembly(GO:0034063) |
| 0.3 | 3.0 | GO:0006122 | mitochondrial electron transport, ubiquinol to cytochrome c(GO:0006122) |
| 0.3 | 1.5 | GO:0071596 | ubiquitin-dependent protein catabolic process via the N-end rule pathway(GO:0071596) |
| 0.3 | 8.7 | GO:0051014 | actin filament severing(GO:0051014) |
| 0.3 | 6.4 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 0.3 | 1.1 | GO:0070257 | regulation of mucus secretion(GO:0070255) positive regulation of mucus secretion(GO:0070257) |
| 0.3 | 4.6 | GO:0018279 | protein N-linked glycosylation via asparagine(GO:0018279) post-translational protein modification(GO:0043687) |
| 0.3 | 2.8 | GO:0036368 | cone photoresponse recovery(GO:0036368) |
| 0.2 | 2.6 | GO:0070301 | cellular response to hydrogen peroxide(GO:0070301) |
| 0.2 | 4.0 | GO:0043114 | regulation of vascular permeability(GO:0043114) |
| 0.2 | 3.9 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
| 0.2 | 2.3 | GO:0042661 | regulation of mesodermal cell fate specification(GO:0042661) |
| 0.2 | 4.5 | GO:0008156 | negative regulation of DNA replication(GO:0008156) |
| 0.2 | 3.4 | GO:0031297 | replication fork processing(GO:0031297) |
| 0.2 | 3.9 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.2 | 8.5 | GO:0003009 | skeletal muscle contraction(GO:0003009) |
| 0.2 | 2.8 | GO:0031397 | negative regulation of protein ubiquitination(GO:0031397) |
| 0.2 | 4.2 | GO:0006446 | regulation of translational initiation(GO:0006446) |
| 0.2 | 3.4 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.2 | 4.1 | GO:0009214 | cyclic nucleotide catabolic process(GO:0009214) |
| 0.1 | 1.2 | GO:1990845 | adaptive thermogenesis(GO:1990845) |
| 0.1 | 2.2 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.1 | 1.6 | GO:0060325 | face morphogenesis(GO:0060325) |
| 0.1 | 1.6 | GO:0071425 | hematopoietic stem cell proliferation(GO:0071425) |
| 0.1 | 1.4 | GO:0009313 | oligosaccharide catabolic process(GO:0009313) |
| 0.1 | 1.4 | GO:0009086 | methionine biosynthetic process(GO:0009086) |
| 0.1 | 11.3 | GO:0017148 | negative regulation of translation(GO:0017148) |
| 0.1 | 3.0 | GO:0032008 | positive regulation of TOR signaling(GO:0032008) |
| 0.1 | 0.3 | GO:0060005 | reflex(GO:0060004) vestibular reflex(GO:0060005) |
| 0.1 | 1.4 | GO:0035493 | SNARE complex assembly(GO:0035493) |
| 0.1 | 1.0 | GO:0050909 | sensory perception of taste(GO:0050909) |
| 0.1 | 1.3 | GO:0046549 | retinal cone cell development(GO:0046549) |
| 0.1 | 0.4 | GO:0030205 | dermatan sulfate metabolic process(GO:0030205) |
| 0.1 | 4.0 | GO:0018298 | protein-chromophore linkage(GO:0018298) |
| 0.1 | 0.9 | GO:0021520 | spinal cord motor neuron cell fate specification(GO:0021520) |
| 0.1 | 3.1 | GO:0016525 | negative regulation of angiogenesis(GO:0016525) |
| 0.1 | 1.6 | GO:0045921 | positive regulation of exocytosis(GO:0045921) |
| 0.1 | 2.0 | GO:0043967 | histone H4 acetylation(GO:0043967) |
| 0.1 | 3.2 | GO:0007050 | cell cycle arrest(GO:0007050) |
| 0.1 | 3.7 | GO:0003333 | amino acid transmembrane transport(GO:0003333) |
| 0.1 | 0.7 | GO:1900052 | regulation of isoprenoid metabolic process(GO:0019747) regulation of vitamin metabolic process(GO:0030656) regulation of retinoic acid biosynthetic process(GO:1900052) |
| 0.1 | 1.4 | GO:0060173 | limb development(GO:0060173) |
| 0.1 | 1.3 | GO:0033138 | positive regulation of peptidyl-serine phosphorylation(GO:0033138) |
| 0.1 | 1.7 | GO:0019835 | cytolysis(GO:0019835) |
| 0.1 | 1.7 | GO:0048268 | clathrin coat assembly(GO:0048268) |
| 0.1 | 4.8 | GO:0007605 | sensory perception of sound(GO:0007605) |
| 0.0 | 2.0 | GO:0006119 | oxidative phosphorylation(GO:0006119) |
| 0.0 | 4.3 | GO:0035023 | regulation of Rho protein signal transduction(GO:0035023) |
| 0.0 | 2.2 | GO:0006368 | transcription elongation from RNA polymerase II promoter(GO:0006368) |
| 0.0 | 0.5 | GO:0051382 | kinetochore assembly(GO:0051382) |
| 0.0 | 3.1 | GO:0001894 | tissue homeostasis(GO:0001894) |
| 0.0 | 4.3 | GO:0002088 | lens development in camera-type eye(GO:0002088) |
| 0.0 | 4.6 | GO:0001756 | somitogenesis(GO:0001756) |
| 0.0 | 1.4 | GO:0006635 | fatty acid beta-oxidation(GO:0006635) |
| 0.0 | 2.2 | GO:0050919 | negative chemotaxis(GO:0050919) |
| 0.0 | 1.6 | GO:0030217 | T cell differentiation(GO:0030217) |
| 0.0 | 12.5 | GO:0006412 | translation(GO:0006412) |
| 0.0 | 0.2 | GO:0006370 | 7-methylguanosine mRNA capping(GO:0006370) |
| 0.0 | 2.9 | GO:0001708 | cell fate specification(GO:0001708) |
| 0.0 | 0.7 | GO:0051784 | negative regulation of mitotic nuclear division(GO:0045839) negative regulation of nuclear division(GO:0051784) |
| 0.0 | 1.2 | GO:0097696 | JAK-STAT cascade(GO:0007259) STAT cascade(GO:0097696) |
| 0.0 | 0.2 | GO:0035871 | protein K11-linked deubiquitination(GO:0035871) |
| 0.0 | 3.1 | GO:0070507 | regulation of microtubule cytoskeleton organization(GO:0070507) |
| 0.0 | 1.5 | GO:0015718 | monocarboxylic acid transport(GO:0015718) |
| 0.0 | 1.5 | GO:0007528 | neuromuscular junction development(GO:0007528) |
| 0.0 | 2.5 | GO:0006338 | chromatin remodeling(GO:0006338) |
| 0.0 | 0.6 | GO:0010499 | proteasomal ubiquitin-independent protein catabolic process(GO:0010499) |
| 0.0 | 0.4 | GO:0035435 | phosphate ion transmembrane transport(GO:0035435) |
| 0.0 | 2.4 | GO:0008284 | positive regulation of cell proliferation(GO:0008284) |
| 0.0 | 2.4 | GO:0006260 | DNA replication(GO:0006260) |
| 0.0 | 1.1 | GO:0006487 | protein N-linked glycosylation(GO:0006487) |
| 0.0 | 1.2 | GO:0071560 | transforming growth factor beta receptor signaling pathway(GO:0007179) response to transforming growth factor beta(GO:0071559) cellular response to transforming growth factor beta stimulus(GO:0071560) |
| 0.0 | 0.2 | GO:0090309 | methylation-dependent chromatin silencing(GO:0006346) positive regulation of chromatin silencing(GO:0031937) regulation of methylation-dependent chromatin silencing(GO:0090308) positive regulation of methylation-dependent chromatin silencing(GO:0090309) |
| 0.0 | 0.2 | GO:0006995 | cellular response to nitrogen starvation(GO:0006995) cellular response to nitrogen levels(GO:0043562) |
| 0.0 | 3.6 | GO:0043062 | extracellular matrix organization(GO:0030198) extracellular structure organization(GO:0043062) |
| 0.0 | 3.0 | GO:0007169 | transmembrane receptor protein tyrosine kinase signaling pathway(GO:0007169) |
| 0.0 | 2.8 | GO:0060537 | muscle tissue development(GO:0060537) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 4.6 | GO:0061689 | tricellular tight junction(GO:0061689) |
| 0.6 | 2.8 | GO:0033165 | interphotoreceptor matrix(GO:0033165) |
| 0.5 | 1.5 | GO:0097189 | apoptotic body(GO:0097189) |
| 0.5 | 3.4 | GO:0005662 | DNA replication factor A complex(GO:0005662) |
| 0.3 | 3.0 | GO:0005750 | mitochondrial respiratory chain complex III(GO:0005750) respiratory chain complex III(GO:0045275) |
| 0.3 | 14.3 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.3 | 3.4 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 0.3 | 8.6 | GO:0043186 | P granule(GO:0043186) |
| 0.2 | 2.0 | GO:0005671 | Ada2/Gcn5/Ada3 transcription activator complex(GO:0005671) |
| 0.2 | 2.4 | GO:0030904 | retromer complex(GO:0030904) |
| 0.2 | 8.5 | GO:0005861 | troponin complex(GO:0005861) |
| 0.2 | 4.2 | GO:0016282 | eukaryotic 43S preinitiation complex(GO:0016282) |
| 0.2 | 3.1 | GO:0030057 | desmosome(GO:0030057) |
| 0.2 | 8.5 | GO:0031305 | integral component of mitochondrial inner membrane(GO:0031305) |
| 0.2 | 1.7 | GO:0098894 | presynaptic endocytic zone(GO:0098833) presynaptic endocytic zone membrane(GO:0098835) extrinsic component of presynaptic membrane(GO:0098888) extrinsic component of presynaptic endocytic zone(GO:0098894) |
| 0.1 | 1.2 | GO:0035060 | brahma complex(GO:0035060) |
| 0.1 | 3.1 | GO:0045180 | basal cortex(GO:0045180) |
| 0.1 | 2.0 | GO:1990124 | messenger ribonucleoprotein complex(GO:1990124) |
| 0.1 | 5.4 | GO:0001750 | photoreceptor outer segment(GO:0001750) |
| 0.1 | 2.0 | GO:0070069 | respiratory chain complex IV(GO:0045277) cytochrome complex(GO:0070069) |
| 0.1 | 3.6 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.1 | 1.1 | GO:0016602 | CCAAT-binding factor complex(GO:0016602) |
| 0.1 | 3.9 | GO:0005747 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.1 | 10.8 | GO:0070160 | bicellular tight junction(GO:0005923) occluding junction(GO:0070160) |
| 0.1 | 6.3 | GO:0001726 | ruffle(GO:0001726) |
| 0.1 | 1.1 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.1 | 3.4 | GO:0016592 | mediator complex(GO:0016592) |
| 0.1 | 0.4 | GO:0038039 | G-protein coupled receptor dimeric complex(GO:0038037) G-protein coupled receptor heterodimeric complex(GO:0038039) G-protein coupled receptor complex(GO:0097648) |
| 0.1 | 0.6 | GO:0019773 | proteasome core complex, alpha-subunit complex(GO:0019773) |
| 0.0 | 4.4 | GO:0005795 | Golgi stack(GO:0005795) |
| 0.0 | 1.6 | GO:0048770 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.0 | 15.9 | GO:0031012 | extracellular matrix(GO:0031012) |
| 0.0 | 0.2 | GO:0070743 | interleukin-12 complex(GO:0043514) interleukin-23 complex(GO:0070743) |
| 0.0 | 2.2 | GO:0008023 | transcription elongation factor complex(GO:0008023) |
| 0.0 | 4.2 | GO:0098852 | lysosomal membrane(GO:0005765) lytic vacuole membrane(GO:0098852) |
| 0.0 | 1.2 | GO:0001725 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.0 | 0.2 | GO:0002141 | stereocilia coupling link(GO:0002139) stereocilia ankle link(GO:0002141) stereocilia ankle link complex(GO:0002142) stereocilium tip(GO:0032426) |
| 0.0 | 0.4 | GO:0016010 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.0 | 2.8 | GO:0000323 | lytic vacuole(GO:0000323) |
| 0.0 | 0.2 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.0 | 1.7 | GO:0016324 | apical plasma membrane(GO:0016324) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.1 | 15.3 | GO:0009374 | biotin binding(GO:0009374) |
| 1.6 | 4.9 | GO:0016802 | adenosylhomocysteinase activity(GO:0004013) trialkylsulfonium hydrolase activity(GO:0016802) |
| 1.4 | 4.1 | GO:0004113 | 2',3'-cyclic-nucleotide 3'-phosphodiesterase activity(GO:0004113) |
| 1.2 | 4.6 | GO:0004579 | oligosaccharyl transferase activity(GO:0004576) dolichyl-diphosphooligosaccharide-protein glycotransferase activity(GO:0004579) |
| 0.9 | 2.8 | GO:0005093 | Rab GDP-dissociation inhibitor activity(GO:0005093) |
| 0.9 | 2.6 | GO:0016019 | peptidoglycan receptor activity(GO:0016019) |
| 0.7 | 3.6 | GO:0060182 | apelin receptor activity(GO:0060182) |
| 0.7 | 3.4 | GO:0042809 | vitamin D receptor binding(GO:0042809) |
| 0.7 | 11.1 | GO:0032190 | acrosin binding(GO:0032190) |
| 0.4 | 2.4 | GO:0005068 | transmembrane receptor protein tyrosine kinase adaptor activity(GO:0005068) |
| 0.4 | 1.4 | GO:0030586 | [methionine synthase] reductase activity(GO:0030586) |
| 0.4 | 1.1 | GO:0016509 | long-chain-3-hydroxyacyl-CoA dehydrogenase activity(GO:0016509) |
| 0.4 | 7.9 | GO:0005003 | ephrin receptor activity(GO:0005003) |
| 0.4 | 2.5 | GO:0055056 | D-glucose transmembrane transporter activity(GO:0055056) |
| 0.3 | 8.5 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.3 | 4.4 | GO:0038191 | neuropilin binding(GO:0038191) |
| 0.3 | 3.4 | GO:0031720 | haptoglobin binding(GO:0031720) |
| 0.3 | 6.4 | GO:0043236 | laminin binding(GO:0043236) |
| 0.3 | 4.4 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 0.2 | 0.7 | GO:0001588 | dopamine neurotransmitter receptor activity, coupled via Gs(GO:0001588) |
| 0.2 | 3.0 | GO:0008266 | poly(U) RNA binding(GO:0008266) |
| 0.2 | 1.4 | GO:0019136 | deoxynucleoside kinase activity(GO:0019136) |
| 0.2 | 3.9 | GO:0050136 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.2 | 8.5 | GO:0019843 | rRNA binding(GO:0019843) |
| 0.2 | 2.6 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.2 | 11.5 | GO:0016627 | oxidoreductase activity, acting on the CH-CH group of donors(GO:0016627) |
| 0.1 | 1.9 | GO:0003886 | DNA (cytosine-5-)-methyltransferase activity(GO:0003886) |
| 0.1 | 0.4 | GO:0070735 | protein-glycine ligase activity(GO:0070735) tubulin-glycine ligase activity(GO:0070738) |
| 0.1 | 1.1 | GO:0008048 | calcium sensitive guanylate cyclase activator activity(GO:0008048) |
| 0.1 | 1.4 | GO:0052794 | exo-alpha-(2->3)-sialidase activity(GO:0052794) exo-alpha-(2->6)-sialidase activity(GO:0052795) exo-alpha-(2->8)-sialidase activity(GO:0052796) |
| 0.1 | 1.7 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.1 | 1.5 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.1 | 5.3 | GO:0008020 | G-protein coupled photoreceptor activity(GO:0008020) |
| 0.1 | 2.5 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.1 | 3.8 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.1 | 5.0 | GO:0009055 | electron carrier activity(GO:0009055) |
| 0.1 | 2.2 | GO:0017110 | nucleoside-diphosphatase activity(GO:0017110) |
| 0.1 | 0.4 | GO:0047757 | chondroitin-glucuronate 5-epimerase activity(GO:0047757) |
| 0.1 | 1.1 | GO:0051864 | histone demethylase activity (H3-K36 specific)(GO:0051864) |
| 0.1 | 2.8 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.1 | 5.9 | GO:0003724 | RNA helicase activity(GO:0003724) |
| 0.1 | 1.1 | GO:0045028 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.1 | 1.2 | GO:0004983 | neuropeptide Y receptor activity(GO:0004983) |
| 0.1 | 21.2 | GO:0044822 | mRNA binding(GO:0003729) poly(A) RNA binding(GO:0044822) |
| 0.1 | 4.2 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.1 | 3.1 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.1 | 1.6 | GO:0005112 | Notch binding(GO:0005112) |
| 0.1 | 4.3 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.1 | 0.5 | GO:0043515 | kinetochore binding(GO:0043515) |
| 0.1 | 0.9 | GO:0005315 | inorganic phosphate transmembrane transporter activity(GO:0005315) |
| 0.1 | 0.4 | GO:0004965 | G-protein coupled GABA receptor activity(GO:0004965) |
| 0.1 | 2.2 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.1 | 1.4 | GO:0001871 | pattern binding(GO:0001871) polysaccharide binding(GO:0030247) |
| 0.1 | 8.0 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 1.8 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.0 | 3.0 | GO:0001594 | trace-amine receptor activity(GO:0001594) |
| 0.0 | 0.2 | GO:0019972 | interleukin-12 binding(GO:0019972) interleukin-12 alpha subunit binding(GO:0042164) |
| 0.0 | 0.3 | GO:0016416 | carnitine O-palmitoyltransferase activity(GO:0004095) O-palmitoyltransferase activity(GO:0016416) |
| 0.0 | 2.9 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 1.0 | GO:0005227 | calcium activated cation channel activity(GO:0005227) |
| 0.0 | 1.9 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.0 | 3.7 | GO:0015171 | amino acid transmembrane transporter activity(GO:0015171) |
| 0.0 | 3.1 | GO:0008201 | heparin binding(GO:0008201) |
| 0.0 | 0.1 | GO:0001596 | angiotensin type I receptor activity(GO:0001596) angiotensin type II receptor activity(GO:0004945) |
| 0.0 | 1.2 | GO:0016755 | transferase activity, transferring amino-acyl groups(GO:0016755) |
| 0.0 | 3.2 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 1.7 | GO:0042562 | hormone binding(GO:0042562) |
| 0.0 | 2.3 | GO:0019955 | cytokine binding(GO:0019955) |
| 0.0 | 6.5 | GO:0005198 | structural molecule activity(GO:0005198) |
| 0.0 | 7.1 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.0 | 5.8 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.0 | 0.7 | GO:0031593 | polyubiquitin binding(GO:0031593) |
| 0.0 | 2.3 | GO:0001228 | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001228) |
| 0.0 | 2.4 | GO:0035091 | phosphatidylinositol binding(GO:0035091) |
| 0.0 | 2.6 | GO:0050839 | cell adhesion molecule binding(GO:0050839) |
| 0.0 | 3.5 | GO:0003779 | actin binding(GO:0003779) |
| 0.0 | 1.5 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 1.6 | GO:0005125 | cytokine activity(GO:0005125) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 5.2 | PID CONE PATHWAY | Visual signal transduction: Cones |
| 0.2 | 4.2 | PID ECADHERIN KERATINOCYTE PATHWAY | E-cadherin signaling in keratinocytes |
| 0.2 | 6.4 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.1 | 1.7 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.1 | 8.5 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.1 | 1.2 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.1 | 3.4 | ST TUMOR NECROSIS FACTOR PATHWAY | Tumor Necrosis Factor Pathway. |
| 0.1 | 3.2 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.1 | 2.4 | PID EPO PATHWAY | EPO signaling pathway |
| 0.1 | 1.4 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.1 | 2.0 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 1.3 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.0 | 1.4 | PID AJDISS 2PATHWAY | Posttranslational regulation of adherens junction stability and dissassembly |
| 0.0 | 2.9 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
| 0.0 | 1.9 | PID CDC42 PATHWAY | CDC42 signaling events |
| 0.0 | 4.8 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 1.5 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.0 | 2.1 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 1.6 | NABA MATRISOME ASSOCIATED | Ensemble of genes encoding ECM-associated proteins including ECM-affilaited proteins, ECM regulators and secreted factors |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 18.6 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.4 | 2.5 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.2 | 3.3 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.2 | 3.7 | REACTOME ASSOCIATION OF LICENSING FACTORS WITH THE PRE REPLICATIVE COMPLEX | Genes involved in Association of licensing factors with the pre-replicative complex |
| 0.2 | 4.9 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.1 | 3.1 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.1 | 2.4 | REACTOME REGULATION OF KIT SIGNALING | Genes involved in Regulation of KIT signaling |
| 0.1 | 2.2 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.1 | 6.9 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.1 | 8.5 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.1 | 1.5 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.1 | 1.6 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.1 | 3.2 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 4.6 | REACTOME ASPARAGINE N LINKED GLYCOSYLATION | Genes involved in Asparagine N-linked glycosylation |
| 0.0 | 1.4 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.0 | 0.6 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |
| 0.0 | 0.3 | REACTOME GRB2 SOS PROVIDES LINKAGE TO MAPK SIGNALING FOR INTERGRINS | Genes involved in GRB2:SOS provides linkage to MAPK signaling for Intergrins |
| 0.0 | 0.4 | REACTOME CHONDROITIN SULFATE BIOSYNTHESIS | Genes involved in Chondroitin sulfate biosynthesis |
| 0.0 | 0.5 | REACTOME DEPOSITION OF NEW CENPA CONTAINING NUCLEOSOMES AT THE CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.0 | 2.2 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 0.3 | REACTOME ACTIVATED AMPK STIMULATES FATTY ACID OXIDATION IN MUSCLE | Genes involved in Activated AMPK stimulates fatty-acid oxidation in muscle |
| 0.0 | 0.4 | REACTOME CHONDROITIN SULFATE DERMATAN SULFATE METABOLISM | Genes involved in Chondroitin sulfate/dermatan sulfate metabolism |
| 0.0 | 0.6 | REACTOME CROSS PRESENTATION OF SOLUBLE EXOGENOUS ANTIGENS ENDOSOMES | Genes involved in Cross-presentation of soluble exogenous antigens (endosomes) |