Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
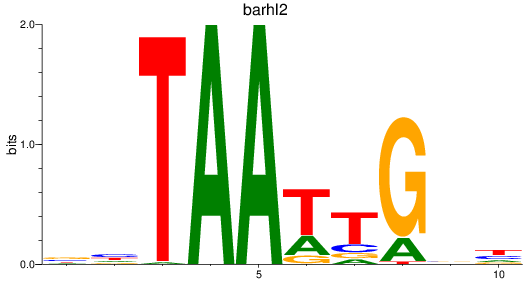
Results for barhl2
Z-value: 0.75
Transcription factors associated with barhl2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
barhl2
|
ENSDARG00000104361 | BarH-like homeobox 2 |
|
barhl2
|
ENSDARG00000114681 | BarH-like homeobox 2 |
|
barhl2
|
ENSDARG00000115723 | BarH-like homeobox 2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| barhl2 | dr11_v1_chr6_+_24817852_24817852 | -0.48 | 1.2e-06 | Click! |
Activity profile of barhl2 motif
Sorted Z-values of barhl2 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_+_16780643 | 4.76 |
ENSDART00000125647
ENSDART00000108611 ENSDART00000181245 ENSDART00000163194 |
tfa
|
transferrin-a |
| chr15_-_2640966 | 4.69 |
ENSDART00000063320
|
cldne
|
claudin e |
| chr15_-_12011390 | 4.61 |
ENSDART00000187403
|
si:dkey-202l22.6
|
si:dkey-202l22.6 |
| chr14_+_21107032 | 4.48 |
ENSDART00000138319
ENSDART00000139103 ENSDART00000184735 |
aldob
|
aldolase b, fructose-bisphosphate |
| chr7_+_22657566 | 4.32 |
ENSDART00000141048
|
ponzr5
|
plac8 onzin related protein 5 |
| chr18_+_9171778 | 4.24 |
ENSDART00000101192
|
sema3d
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3D |
| chr22_-_10586191 | 4.14 |
ENSDART00000148418
|
si:dkey-42i9.16
|
si:dkey-42i9.16 |
| chr1_-_52437056 | 4.14 |
ENSDART00000138337
|
si:ch211-217k17.12
|
si:ch211-217k17.12 |
| chr5_-_30615901 | 3.90 |
ENSDART00000147769
|
si:ch211-117m20.5
|
si:ch211-117m20.5 |
| chr8_-_38201415 | 3.89 |
ENSDART00000155189
|
pdlim2
|
PDZ and LIM domain 2 (mystique) |
| chr7_+_13988075 | 3.81 |
ENSDART00000186812
|
furina
|
furin (paired basic amino acid cleaving enzyme) a |
| chr24_-_25166720 | 3.72 |
ENSDART00000141601
|
phldb2b
|
pleckstrin homology-like domain, family B, member 2b |
| chr6_-_33075576 | 3.72 |
ENSDART00000154017
|
si:dkey-170g13.2
|
si:dkey-170g13.2 |
| chr6_-_8362419 | 3.66 |
ENSDART00000142752
ENSDART00000135810 |
acp5a
|
acid phosphatase 5a, tartrate resistant |
| chr15_+_34699585 | 3.55 |
ENSDART00000017569
|
tnfb
|
tumor necrosis factor b (TNF superfamily, member 2) |
| chr4_-_17669881 | 3.54 |
ENSDART00000066997
|
dram1
|
DNA-damage regulated autophagy modulator 1 |
| chr19_+_5640504 | 3.50 |
ENSDART00000179987
|
ft2
|
alpha(1,3)fucosyltransferase gene 2 |
| chr8_+_23738122 | 3.48 |
ENSDART00000062983
|
rpl10a
|
ribosomal protein L10a |
| chr22_-_24297510 | 3.37 |
ENSDART00000163297
|
si:ch211-117l17.6
|
si:ch211-117l17.6 |
| chr5_+_27897504 | 3.37 |
ENSDART00000130936
|
adam28
|
ADAM metallopeptidase domain 28 |
| chr25_-_21816269 | 3.35 |
ENSDART00000152014
|
zgc:158222
|
zgc:158222 |
| chr14_+_14568437 | 3.22 |
ENSDART00000164749
|
pcdh20
|
protocadherin 20 |
| chr17_+_16046314 | 3.22 |
ENSDART00000154554
ENSDART00000154338 ENSDART00000155336 |
si:ch73-204p21.2
|
si:ch73-204p21.2 |
| chr20_+_26880668 | 3.19 |
ENSDART00000077769
|
serpinb1
|
serpin peptidase inhibitor, clade B (ovalbumin), member 1 |
| chr16_-_9694822 | 3.18 |
ENSDART00000168748
|
COLEC10
|
si:ch211-93i7.4 |
| chr23_-_19715557 | 3.05 |
ENSDART00000143764
|
rpl10
|
ribosomal protein L10 |
| chr17_+_16046132 | 3.05 |
ENSDART00000155005
|
si:ch73-204p21.2
|
si:ch73-204p21.2 |
| chr17_-_30635298 | 2.95 |
ENSDART00000155478
|
sh3yl1
|
SH3 and SYLF domain containing 1 |
| chr5_+_6670945 | 2.94 |
ENSDART00000185686
|
pxna
|
paxillin a |
| chr11_+_6136220 | 2.93 |
ENSDART00000082223
|
tax1bp3
|
Tax1 (human T-cell leukemia virus type I) binding protein 3 |
| chr9_-_24031461 | 2.93 |
ENSDART00000021218
|
rpe
|
ribulose-5-phosphate-3-epimerase |
| chr13_-_50139916 | 2.90 |
ENSDART00000099475
|
nid1a
|
nidogen 1a |
| chr13_+_829585 | 2.86 |
ENSDART00000029051
|
gsta.2
|
glutathione S-transferase, alpha tandem duplicate 2 |
| chr11_-_7261717 | 2.85 |
ENSDART00000128959
|
zgc:113223
|
zgc:113223 |
| chr9_-_6991650 | 2.84 |
ENSDART00000081718
|
slc9a2
|
solute carrier family 9, subfamily A (NHE2, cation proton antiporter 2), member 2 |
| chr13_-_20381485 | 2.84 |
ENSDART00000131351
|
si:ch211-270n8.1
|
si:ch211-270n8.1 |
| chr1_-_19845378 | 2.76 |
ENSDART00000139314
ENSDART00000132958 ENSDART00000147502 |
grhprb
|
glyoxylate reductase/hydroxypyruvate reductase b |
| chr11_+_24703108 | 2.72 |
ENSDART00000159173
|
gpr25
|
G protein-coupled receptor 25 |
| chr22_-_10541372 | 2.70 |
ENSDART00000179708
|
si:dkey-42i9.4
|
si:dkey-42i9.4 |
| chr7_+_17443567 | 2.70 |
ENSDART00000060383
|
nitr2b
|
novel immune-type receptor 2b |
| chr18_+_3037998 | 2.70 |
ENSDART00000185844
ENSDART00000162657 |
rps3
|
ribosomal protein S3 |
| chr25_-_16724913 | 2.69 |
ENSDART00000157075
|
galnt8a.2
|
polypeptide N-acetylgalactosaminyltransferase 8a, tandem duplicate 2 |
| chr5_+_29794058 | 2.61 |
ENSDART00000045410
|
thy1
|
Thy-1 cell surface antigen |
| chr21_-_19919020 | 2.61 |
ENSDART00000147396
|
ppp1r3b
|
protein phosphatase 1, regulatory subunit 3B |
| chr8_-_24252933 | 2.58 |
ENSDART00000057624
|
zgc:110353
|
zgc:110353 |
| chr18_+_48423973 | 2.57 |
ENSDART00000184233
ENSDART00000147074 |
fli1a
|
Fli-1 proto-oncogene, ETS transcription factor a |
| chr22_+_10066846 | 2.57 |
ENSDART00000105916
ENSDART00000105914 ENSDART00000132480 |
si:ch211-222k6.3
|
si:ch211-222k6.3 |
| chr21_+_25236297 | 2.49 |
ENSDART00000112783
|
tmem45b
|
transmembrane protein 45B |
| chr3_-_29928695 | 2.46 |
ENSDART00000151275
ENSDART00000151083 |
grna
|
granulin a |
| chr11_-_30634286 | 2.46 |
ENSDART00000191019
|
zgc:153665
|
zgc:153665 |
| chr4_-_25257451 | 2.46 |
ENSDART00000142819
|
tmem110l
|
transmembrane protein 110, like |
| chr10_-_17988779 | 2.45 |
ENSDART00000132206
ENSDART00000144841 |
si:dkey-242g16.2
|
si:dkey-242g16.2 |
| chr16_+_7697878 | 2.45 |
ENSDART00000104176
ENSDART00000172977 |
ccr11.1
|
chemokine (C-C motif) receptor 11.1 |
| chr16_-_23797570 | 2.43 |
ENSDART00000077834
|
rps27.2
|
ribosomal protein S27, isoform 2 |
| chr18_-_18942098 | 2.41 |
ENSDART00000100458
|
si:dkey-73n10.1
|
si:dkey-73n10.1 |
| chr20_+_38257766 | 2.41 |
ENSDART00000147485
ENSDART00000149160 |
ccl38.6
|
chemokine (C-C motif) ligand 38, duplicate 6 |
| chr10_+_29840725 | 2.40 |
ENSDART00000127268
|
BX120005.1
|
|
| chr5_-_69620722 | 2.37 |
ENSDART00000097248
|
aldh2.2
|
aldehyde dehydrogenase 2 family (mitochondrial), tandem duplicate 2 |
| chr25_+_32473277 | 2.36 |
ENSDART00000146451
|
sqor
|
sulfide quinone oxidoreductase |
| chr18_-_34549721 | 2.35 |
ENSDART00000137101
ENSDART00000021880 |
ssr3
|
signal sequence receptor, gamma |
| chr3_-_32541033 | 2.34 |
ENSDART00000151476
ENSDART00000055324 |
rcn3
|
reticulocalbin 3, EF-hand calcium binding domain |
| chr12_-_11349899 | 2.30 |
ENSDART00000079645
|
zgc:174164
|
zgc:174164 |
| chr3_-_32596394 | 2.28 |
ENSDART00000103239
|
tspan4b
|
tetraspanin 4b |
| chr6_-_7720332 | 2.24 |
ENSDART00000135945
|
rpsa
|
ribosomal protein SA |
| chr15_+_24795473 | 2.23 |
ENSDART00000139689
ENSDART00000141033 ENSDART00000100746 ENSDART00000135677 |
gosr1
|
golgi SNAP receptor complex member 1 |
| chr15_+_22394074 | 2.23 |
ENSDART00000109931
|
oafa
|
OAF homolog a (Drosophila) |
| chr7_+_17816006 | 2.17 |
ENSDART00000080834
|
eml3
|
echinoderm microtubule associated protein like 3 |
| chr3_+_22381955 | 2.16 |
ENSDART00000185315
|
arhgap27l
|
Rho GTPase activating protein 27, like |
| chr6_+_39098397 | 2.16 |
ENSDART00000003716
ENSDART00000188655 |
prss60.2
|
protease, serine, 60.2 |
| chr22_-_10121880 | 2.15 |
ENSDART00000002348
|
rdh5
|
retinol dehydrogenase 5 (11-cis/9-cis) |
| chr5_-_34993242 | 2.13 |
ENSDART00000134516
ENSDART00000051295 |
btf3
|
basic transcription factor 3 |
| chr7_-_26270014 | 2.10 |
ENSDART00000079347
|
serpine1
|
serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 1 |
| chr4_-_77218637 | 2.10 |
ENSDART00000174325
|
psmb10
|
proteasome subunit beta 10 |
| chr6_+_41191482 | 2.08 |
ENSDART00000000877
|
opn1mw3
|
opsin 1 (cone pigments), medium-wave-sensitive, 3 |
| chr22_+_3156386 | 2.07 |
ENSDART00000161212
|
rpl36
|
ribosomal protein L36 |
| chr24_-_34680956 | 2.03 |
ENSDART00000171009
|
ctnna1
|
catenin (cadherin-associated protein), alpha 1 |
| chr7_-_51368681 | 2.03 |
ENSDART00000146385
|
arhgap36
|
Rho GTPase activating protein 36 |
| chr21_-_8085635 | 2.02 |
ENSDART00000082790
|
si:dkey-163m14.2
|
si:dkey-163m14.2 |
| chr16_-_26140768 | 2.02 |
ENSDART00000143960
|
cd79a
|
CD79a molecule, immunoglobulin-associated alpha |
| chr16_+_21790870 | 2.01 |
ENSDART00000155039
|
trim108
|
tripartite motif containing 108 |
| chr23_+_9220436 | 2.01 |
ENSDART00000033663
ENSDART00000139870 |
rps21
|
ribosomal protein S21 |
| chr16_+_23398369 | 2.01 |
ENSDART00000037694
|
s100a10b
|
S100 calcium binding protein A10b |
| chr19_+_43297546 | 2.00 |
ENSDART00000168002
|
laptm5
|
lysosomal protein transmembrane 5 |
| chr20_-_40755614 | 1.99 |
ENSDART00000061247
|
cx32.3
|
connexin 32.3 |
| chr18_-_25568994 | 1.98 |
ENSDART00000133029
|
si:ch211-13k12.2
|
si:ch211-13k12.2 |
| chr2_-_29994726 | 1.97 |
ENSDART00000163350
|
cnpy1
|
canopy1 |
| chr11_+_13166750 | 1.97 |
ENSDART00000169961
|
mob3c
|
MOB kinase activator 3C |
| chr10_-_22127942 | 1.96 |
ENSDART00000133374
|
ponzr2
|
plac8 onzin related protein 2 |
| chr13_-_15702672 | 1.96 |
ENSDART00000144445
ENSDART00000168950 |
ckba
|
creatine kinase, brain a |
| chr21_-_11654422 | 1.94 |
ENSDART00000081614
ENSDART00000132699 |
cast
|
calpastatin |
| chr13_-_40401870 | 1.94 |
ENSDART00000128951
|
nkx3.3
|
NK3 homeobox 3 |
| chr16_+_50972803 | 1.94 |
ENSDART00000178189
|
si:dkeyp-97a10.2
|
si:dkeyp-97a10.2 |
| chr20_-_48700370 | 1.92 |
ENSDART00000185195
ENSDART00000184898 |
pax1b
|
paired box 1b |
| chr6_+_42475730 | 1.89 |
ENSDART00000150226
|
mst1ra
|
macrophage stimulating 1 receptor a |
| chr5_-_67762434 | 1.88 |
ENSDART00000167301
|
b4galt4
|
UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 4 |
| chr17_+_25849332 | 1.88 |
ENSDART00000191994
|
acss1
|
acyl-CoA synthetase short chain family member 1 |
| chr17_+_43867889 | 1.88 |
ENSDART00000132673
ENSDART00000167214 |
zgc:66313
|
zgc:66313 |
| chr16_-_7443388 | 1.87 |
ENSDART00000017445
|
prdm1a
|
PR domain containing 1a, with ZNF domain |
| chr1_-_513762 | 1.86 |
ENSDART00000148162
ENSDART00000144606 |
trmt10c
|
tRNA methyltransferase 10C, mitochondrial RNase P subunit |
| chr10_-_13343831 | 1.82 |
ENSDART00000135941
|
il11ra
|
interleukin 11 receptor, alpha |
| chr8_+_13389115 | 1.79 |
ENSDART00000184428
ENSDART00000154266 ENSDART00000049469 |
jak3
|
Janus kinase 3 (a protein tyrosine kinase, leukocyte) |
| chr10_+_17714866 | 1.79 |
ENSDART00000039969
|
slc20a1b
|
solute carrier family 20 (phosphate transporter), member 1b |
| chr18_-_48547564 | 1.78 |
ENSDART00000138607
|
kcnj1a.1
|
potassium inwardly-rectifying channel, subfamily J, member 1a, tandem duplicate 1 |
| chr11_-_40457325 | 1.76 |
ENSDART00000128442
|
tnfrsf1b
|
tumor necrosis factor receptor superfamily, member 1B |
| chr11_-_37691449 | 1.76 |
ENSDART00000185340
|
zgc:158258
|
zgc:158258 |
| chr17_+_6563307 | 1.75 |
ENSDART00000156454
|
adgrf3a
|
adhesion G protein-coupled receptor F3a |
| chr19_+_12915498 | 1.74 |
ENSDART00000132892
|
cthrc1a
|
collagen triple helix repeat containing 1a |
| chr23_+_19813677 | 1.74 |
ENSDART00000139192
ENSDART00000142308 |
emd
|
emerin (Emery-Dreifuss muscular dystrophy) |
| chr8_-_10949847 | 1.73 |
ENSDART00000123209
|
pqlc2
|
PQ loop repeat containing 2 |
| chr22_-_17595310 | 1.72 |
ENSDART00000099056
|
gpx4a
|
glutathione peroxidase 4a |
| chr13_+_835390 | 1.71 |
ENSDART00000171329
|
gsta.1
|
glutathione S-transferase, alpha tandem duplicate 1 |
| chr10_-_15854743 | 1.69 |
ENSDART00000092343
|
tjp2a
|
tight junction protein 2a (zona occludens 2) |
| chr12_+_20352400 | 1.69 |
ENSDART00000066383
|
hbae5
|
hemoglobin, alpha embryonic 5 |
| chr5_-_72390259 | 1.66 |
ENSDART00000172302
|
wbp1
|
WW domain binding protein 1 |
| chr25_-_18435481 | 1.66 |
ENSDART00000004771
|
poc1b
|
POC1 centriolar protein B |
| chr16_-_48430272 | 1.64 |
ENSDART00000005927
|
rad21a
|
RAD21 cohesin complex component a |
| chr3_+_39663987 | 1.64 |
ENSDART00000184614
ENSDART00000184573 ENSDART00000183127 |
epn2
|
epsin 2 |
| chr22_+_31039091 | 1.63 |
ENSDART00000060005
|
rpl32
|
ribosomal protein L32 |
| chr19_+_31904836 | 1.63 |
ENSDART00000162297
ENSDART00000088340 ENSDART00000151280 ENSDART00000151218 |
tpd52
|
tumor protein D52 |
| chr20_-_49889111 | 1.62 |
ENSDART00000058858
|
kif13bb
|
kinesin family member 13Bb |
| chr25_+_32473433 | 1.61 |
ENSDART00000152326
|
sqor
|
sulfide quinone oxidoreductase |
| chr20_-_23426339 | 1.60 |
ENSDART00000004625
|
zar1
|
zygote arrest 1 |
| chr25_+_3327071 | 1.59 |
ENSDART00000136131
ENSDART00000133243 |
ldhbb
|
lactate dehydrogenase Bb |
| chr8_-_46486009 | 1.59 |
ENSDART00000140431
|
sult1st9
|
sulfotransferase family 1, cytosolic sulfotransferase 9 |
| chr23_+_2666944 | 1.59 |
ENSDART00000192861
|
CABZ01057928.1
|
|
| chr15_+_7054754 | 1.58 |
ENSDART00000149800
|
foxl2a
|
forkhead box L2a |
| chr5_+_61998641 | 1.57 |
ENSDART00000137110
|
selenow1
|
selenoprotein W, 1 |
| chr1_-_10821584 | 1.57 |
ENSDART00000167452
ENSDART00000162137 |
crfb15
|
cytokine receptor family member B15 |
| chr17_+_4398102 | 1.56 |
ENSDART00000152387
|
si:zfos-364h11.1
|
si:zfos-364h11.1 |
| chr2_+_50999477 | 1.55 |
ENSDART00000190111
|
eef1da
|
eukaryotic translation elongation factor 1 delta a (guanine nucleotide exchange protein) |
| chr16_+_42471455 | 1.55 |
ENSDART00000166640
|
si:ch211-215k15.5
|
si:ch211-215k15.5 |
| chr5_+_9246458 | 1.54 |
ENSDART00000081772
|
susd1
|
sushi domain containing 1 |
| chr2_-_37140423 | 1.53 |
ENSDART00000144220
|
tspan37
|
tetraspanin 37 |
| chr6_+_28877306 | 1.52 |
ENSDART00000065137
ENSDART00000123189 ENSDART00000065135 ENSDART00000181512 ENSDART00000130799 |
tp63
|
tumor protein p63 |
| chr4_+_73606482 | 1.52 |
ENSDART00000150765
|
si:ch211-165i18.2
|
si:ch211-165i18.2 |
| chr10_-_7785930 | 1.51 |
ENSDART00000043961
ENSDART00000111058 |
mpx
|
myeloid-specific peroxidase |
| chr14_+_30773764 | 1.49 |
ENSDART00000186961
|
atl3
|
atlastin 3 |
| chr5_+_9428876 | 1.49 |
ENSDART00000081791
|
ugt2a7
|
UDP glucuronosyltransferase 2 family, polypeptide A7 |
| chr3_-_26244256 | 1.48 |
ENSDART00000103741
|
ppp4ca
|
protein phosphatase 4, catalytic subunit a |
| chr7_-_51321126 | 1.47 |
ENSDART00000067647
|
rasl11a
|
RAS-like, family 11, member A |
| chr23_-_24394719 | 1.47 |
ENSDART00000044918
|
epha2b
|
eph receptor A2 b |
| chr13_-_11984867 | 1.46 |
ENSDART00000157538
|
npm3
|
nucleophosmin/nucleoplasmin, 3 |
| chr14_-_3381303 | 1.45 |
ENSDART00000171601
|
im:7150988
|
im:7150988 |
| chr7_+_57108823 | 1.45 |
ENSDART00000184943
ENSDART00000055956 |
enosf1
|
enolase superfamily member 1 |
| chr11_+_25539698 | 1.45 |
ENSDART00000035602
|
cxxc1b
|
CXXC finger protein 1b |
| chr10_+_8690936 | 1.45 |
ENSDART00000144218
|
si:dkey-27b3.4
|
si:dkey-27b3.4 |
| chr20_-_9580446 | 1.43 |
ENSDART00000014168
|
zfp36l1b
|
zinc finger protein 36, C3H type-like 1b |
| chr22_+_883678 | 1.43 |
ENSDART00000140588
|
stk38b
|
serine/threonine kinase 38b |
| chr11_+_30295582 | 1.43 |
ENSDART00000122424
|
ugt1b7
|
UDP glucuronosyltransferase 1 family, polypeptide B7 |
| chr3_+_27798094 | 1.43 |
ENSDART00000075100
ENSDART00000151437 |
carhsp1
|
calcium regulated heat stable protein 1 |
| chr6_+_40922572 | 1.41 |
ENSDART00000133599
ENSDART00000002728 ENSDART00000145153 |
eif4enif1
|
eukaryotic translation initiation factor 4E nuclear import factor 1 |
| chr22_-_28668442 | 1.41 |
ENSDART00000182377
|
col8a1b
|
collagen, type VIII, alpha 1b |
| chr23_-_20126257 | 1.40 |
ENSDART00000005021
|
tktb
|
transketolase b |
| chr19_-_29832876 | 1.40 |
ENSDART00000005119
|
eif3i
|
eukaryotic translation initiation factor 3, subunit I |
| chr17_+_50701748 | 1.40 |
ENSDART00000191938
ENSDART00000183220 ENSDART00000049464 |
fermt2
|
fermitin family member 2 |
| chr3_+_43373867 | 1.40 |
ENSDART00000159455
ENSDART00000172425 |
zfand2a
|
zinc finger, AN1-type domain 2A |
| chr19_+_7045033 | 1.40 |
ENSDART00000146579
|
mhc1uka
|
major histocompatibility complex class I UKA |
| chr25_+_35553542 | 1.38 |
ENSDART00000113723
|
spi1a
|
Spi-1 proto-oncogene a |
| chr23_-_20363261 | 1.37 |
ENSDART00000054651
|
si:rp71-17i16.5
|
si:rp71-17i16.5 |
| chr2_+_20793982 | 1.37 |
ENSDART00000014785
|
prg4a
|
proteoglycan 4a |
| chr11_-_21030070 | 1.36 |
ENSDART00000186322
|
fmoda
|
fibromodulin a |
| chr22_+_31025096 | 1.35 |
ENSDART00000185953
|
zmp:0000000735
|
zmp:0000000735 |
| chr4_-_14192254 | 1.35 |
ENSDART00000143804
|
pus7l
|
pseudouridylate synthase 7-like |
| chr14_-_32884138 | 1.33 |
ENSDART00000105726
|
slc25a5
|
solute carrier family 25 (mitochondrial carrier; adenine nucleotide translocator), member 5 |
| chr7_+_2797795 | 1.32 |
ENSDART00000161208
|
si:ch211-217i17.1
|
si:ch211-217i17.1 |
| chr19_+_11217279 | 1.31 |
ENSDART00000181859
|
si:ch73-109i22.2
|
si:ch73-109i22.2 |
| chr3_-_19367081 | 1.31 |
ENSDART00000191369
|
s1pr5a
|
sphingosine-1-phosphate receptor 5a |
| chr5_+_33519943 | 1.30 |
ENSDART00000131316
|
ms4a17c.1
|
membrane-spanning 4-domains, subfamily A, member 17C.1 |
| chr7_-_20158380 | 1.29 |
ENSDART00000142891
|
acap1
|
ArfGAP with coiled-coil, ankyrin repeat and PH domains 1 |
| chr7_-_54679595 | 1.29 |
ENSDART00000165320
|
ccnd1
|
cyclin D1 |
| chr23_+_17387325 | 1.29 |
ENSDART00000083947
|
ptk6b
|
PTK6 protein tyrosine kinase 6b |
| chr23_+_24973773 | 1.28 |
ENSDART00000047020
|
casp9
|
caspase 9, apoptosis-related cysteine peptidase |
| chr3_+_6443992 | 1.28 |
ENSDART00000169325
ENSDART00000162255 |
nup85
|
nucleoporin 85 |
| chr3_-_42125655 | 1.27 |
ENSDART00000040753
|
nudt1
|
nudix (nucleoside diphosphate linked moiety X)-type motif 1 |
| chr15_-_1746963 | 1.27 |
ENSDART00000156225
|
stx1a
|
syntaxin 1A (brain) |
| chr11_-_40519886 | 1.27 |
ENSDART00000172819
|
miip
|
migration and invasion inhibitory protein |
| chr10_+_15024772 | 1.26 |
ENSDART00000135667
|
si:dkey-88l16.5
|
si:dkey-88l16.5 |
| chr4_-_4259079 | 1.25 |
ENSDART00000135352
ENSDART00000026559 |
cd9b
|
CD9 molecule b |
| chr24_+_36414829 | 1.23 |
ENSDART00000062699
|
si:ch211-253b1.3
|
si:ch211-253b1.3 |
| chr2_-_55779927 | 1.22 |
ENSDART00000168579
|
CABZ01066725.1
|
|
| chr9_+_14010823 | 1.22 |
ENSDART00000143837
|
si:ch211-67e16.3
|
si:ch211-67e16.3 |
| chr21_+_43702016 | 1.21 |
ENSDART00000017176
|
dkc1
|
dyskeratosis congenita 1, dyskerin |
| chr14_+_33329761 | 1.21 |
ENSDART00000161138
|
sowahd
|
sosondowah ankyrin repeat domain family d |
| chr15_-_28802140 | 1.21 |
ENSDART00000156120
ENSDART00000156708 ENSDART00000122536 ENSDART00000155008 ENSDART00000153825 |
cd3eap
|
CD3e molecule, epsilon associated protein |
| chr1_+_40802454 | 1.21 |
ENSDART00000193568
|
cpz
|
carboxypeptidase Z |
| chr2_-_51634431 | 1.21 |
ENSDART00000165568
|
pigr
|
polymeric immunoglobulin receptor |
| chr16_+_33902006 | 1.20 |
ENSDART00000161807
ENSDART00000159474 |
gnl2
|
guanine nucleotide binding protein-like 2 (nucleolar) |
| chr17_+_44697604 | 1.19 |
ENSDART00000156625
|
pgfb
|
placental growth factor b |
| chr23_+_38171186 | 1.18 |
ENSDART00000148188
|
zgc:112994
|
zgc:112994 |
| chr6_-_43817761 | 1.17 |
ENSDART00000183945
|
foxp1b
|
forkhead box P1b |
| chr19_-_11336782 | 1.17 |
ENSDART00000131014
|
sept7a
|
septin 7a |
| chr3_+_26244353 | 1.17 |
ENSDART00000103733
|
atad5a
|
ATPase family, AAA domain containing 5a |
| chr7_-_64971839 | 1.17 |
ENSDART00000164682
|
sinhcafl
|
SIN3-HDAC complex associated factor, like |
| chr11_-_22303678 | 1.16 |
ENSDART00000159527
ENSDART00000159681 |
tfeb
|
transcription factor EB |
| chr11_+_37251825 | 1.15 |
ENSDART00000169804
|
il17rc
|
interleukin 17 receptor C |
| chr11_+_24716837 | 1.15 |
ENSDART00000145217
|
zgc:153953
|
zgc:153953 |
| chr21_+_7298687 | 1.15 |
ENSDART00000187746
|
CU929160.1
|
|
| chr20_+_2039518 | 1.14 |
ENSDART00000043157
|
CABZ01088134.1
|
|
| chr17_+_24821627 | 1.13 |
ENSDART00000112389
|
wdr43
|
WD repeat domain 43 |
| chr14_+_26247319 | 1.13 |
ENSDART00000192793
|
CCDC69
|
coiled-coil domain containing 69 |
Network of associatons between targets according to the STRING database.
First level regulatory network of barhl2
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.3 | 4.0 | GO:0070221 | sulfide oxidation(GO:0019418) sulfide oxidation, using sulfide:quinone oxidoreductase(GO:0070221) |
| 1.2 | 3.6 | GO:0019323 | pentose catabolic process(GO:0019323) |
| 1.0 | 3.1 | GO:1990403 | embryonic brain development(GO:1990403) |
| 1.0 | 2.9 | GO:2000008 | regulation of protein localization to cell surface(GO:2000008) negative regulation of protein localization to cell surface(GO:2000009) |
| 0.9 | 2.6 | GO:0005981 | regulation of glycogen catabolic process(GO:0005981) |
| 0.8 | 3.9 | GO:0007508 | larval development(GO:0002164) larval heart development(GO:0007508) |
| 0.6 | 1.8 | GO:0001779 | natural killer cell differentiation(GO:0001779) |
| 0.6 | 2.2 | GO:0010755 | regulation of plasminogen activation(GO:0010755) |
| 0.5 | 1.6 | GO:0048496 | maintenance of organ identity(GO:0048496) |
| 0.4 | 2.2 | GO:0048209 | regulation of vesicle targeting, to, from or within Golgi(GO:0048209) |
| 0.4 | 0.9 | GO:0032755 | positive regulation of interleukin-6 production(GO:0032755) |
| 0.4 | 4.8 | GO:0019731 | antibacterial humoral response(GO:0019731) |
| 0.4 | 1.2 | GO:0031120 | snRNA pseudouridine synthesis(GO:0031120) |
| 0.4 | 1.9 | GO:0090646 | mitochondrial tRNA processing(GO:0090646) |
| 0.4 | 2.2 | GO:0042362 | fat-soluble vitamin biosynthetic process(GO:0042362) |
| 0.4 | 1.8 | GO:0032776 | DNA methylation on cytosine(GO:0032776) |
| 0.3 | 2.0 | GO:0000447 | endonucleolytic cleavage in ITS1 to separate SSU-rRNA from 5.8S rRNA and LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000447) |
| 0.3 | 3.7 | GO:0045453 | bone resorption(GO:0045453) |
| 0.3 | 1.3 | GO:1990544 | intracellular nucleoside transport(GO:0015859) purine nucleoside transmembrane transport(GO:0015860) purine nucleotide transport(GO:0015865) ATP transport(GO:0015867) purine ribonucleotide transport(GO:0015868) adenine nucleotide transport(GO:0051503) mitochondrial ATP transmembrane transport(GO:1990544) |
| 0.3 | 1.6 | GO:0003173 | ventriculo bulbo valve development(GO:0003173) |
| 0.3 | 1.0 | GO:0034473 | U1 snRNA 3'-end processing(GO:0034473) U5 snRNA 3'-end processing(GO:0034476) nuclear polyadenylation-dependent mRNA catabolic process(GO:0071042) polyadenylation-dependent mRNA catabolic process(GO:0071047) |
| 0.3 | 2.8 | GO:0006971 | hypotonic response(GO:0006971) hypotonic salinity response(GO:0042539) |
| 0.3 | 1.9 | GO:0006083 | acetate metabolic process(GO:0006083) acetyl-CoA biosynthetic process from acetate(GO:0019427) |
| 0.3 | 2.5 | GO:0071340 | skeletal muscle acetylcholine-gated channel clustering(GO:0071340) |
| 0.3 | 1.2 | GO:0000448 | cleavage in ITS2 between 5.8S rRNA and LSU-rRNA of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000448) |
| 0.3 | 0.9 | GO:0036336 | dendritic cell chemotaxis(GO:0002407) dendritic cell migration(GO:0036336) |
| 0.3 | 0.9 | GO:0006041 | glucosamine metabolic process(GO:0006041) glucosamine catabolic process(GO:0006043) |
| 0.3 | 1.5 | GO:0016338 | calcium-independent cell-cell adhesion via plasma membrane cell-adhesion molecules(GO:0016338) |
| 0.3 | 2.0 | GO:0050820 | positive regulation of blood coagulation(GO:0030194) positive regulation of coagulation(GO:0050820) positive regulation of hemostasis(GO:1900048) |
| 0.3 | 1.9 | GO:0097341 | inhibition of cysteine-type endopeptidase activity(GO:0097340) zymogen inhibition(GO:0097341) |
| 0.3 | 1.1 | GO:0016267 | O-glycan processing, core 1(GO:0016267) |
| 0.3 | 3.8 | GO:0060971 | embryonic heart tube left/right pattern formation(GO:0060971) |
| 0.3 | 1.6 | GO:0010269 | response to selenium ion(GO:0010269) |
| 0.3 | 1.0 | GO:0070098 | chemokine-mediated signaling pathway(GO:0070098) |
| 0.3 | 0.8 | GO:0006145 | purine nucleobase catabolic process(GO:0006145) |
| 0.3 | 1.3 | GO:0031571 | mitotic G1 DNA damage checkpoint(GO:0031571) |
| 0.3 | 1.5 | GO:0060965 | negative regulation of posttranscriptional gene silencing(GO:0060149) negative regulation of gene silencing by miRNA(GO:0060965) negative regulation of gene silencing by RNA(GO:0060967) |
| 0.2 | 4.7 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 0.2 | 3.5 | GO:0036065 | fucosylation(GO:0036065) |
| 0.2 | 0.7 | GO:1900364 | negative regulation of mRNA 3'-end processing(GO:0031441) negative regulation of mRNA polyadenylation(GO:1900364) |
| 0.2 | 1.5 | GO:0045943 | positive regulation of transcription from RNA polymerase I promoter(GO:0045943) |
| 0.2 | 0.6 | GO:1903392 | epicardial cell to mesenchymal cell transition(GO:0003347) negative regulation of adherens junction organization(GO:1903392) |
| 0.2 | 2.0 | GO:0001709 | cell fate determination(GO:0001709) |
| 0.2 | 2.9 | GO:0061055 | myotome development(GO:0061055) |
| 0.2 | 0.8 | GO:2001295 | malonyl-CoA biosynthetic process(GO:2001295) |
| 0.2 | 1.5 | GO:0048730 | epidermis morphogenesis(GO:0048730) |
| 0.2 | 4.5 | GO:0030388 | fructose 1,6-bisphosphate metabolic process(GO:0030388) |
| 0.2 | 3.5 | GO:0042541 | hemoglobin biosynthetic process(GO:0042541) |
| 0.2 | 3.6 | GO:0030183 | B cell differentiation(GO:0030183) |
| 0.2 | 1.1 | GO:0019563 | glycerol catabolic process(GO:0019563) |
| 0.2 | 0.5 | GO:0042998 | regulation of Golgi to plasma membrane protein transport(GO:0042996) positive regulation of Golgi to plasma membrane protein transport(GO:0042998) regulation of establishment of protein localization to plasma membrane(GO:0090003) positive regulation of establishment of protein localization to plasma membrane(GO:0090004) |
| 0.2 | 1.0 | GO:0035332 | positive regulation of hippo signaling(GO:0035332) |
| 0.2 | 0.5 | GO:0045218 | zonula adherens maintenance(GO:0045218) |
| 0.2 | 1.0 | GO:0045852 | pH elevation(GO:0045852) intracellular pH elevation(GO:0051454) |
| 0.2 | 1.6 | GO:0072383 | plus-end-directed vesicle transport along microtubule(GO:0072383) plus-end-directed organelle transport along microtubule(GO:0072386) |
| 0.2 | 4.0 | GO:0042743 | hydrogen peroxide metabolic process(GO:0042743) |
| 0.2 | 4.2 | GO:0001732 | formation of cytoplasmic translation initiation complex(GO:0001732) |
| 0.2 | 0.6 | GO:0042780 | tRNA 3'-end processing(GO:0042780) |
| 0.2 | 2.0 | GO:0006599 | phosphagen metabolic process(GO:0006599) phosphocreatine metabolic process(GO:0006603) phosphagen biosynthetic process(GO:0042396) phosphocreatine biosynthetic process(GO:0046314) |
| 0.1 | 0.9 | GO:1902765 | L-arginine import(GO:0043091) amino acid import across plasma membrane(GO:0089718) arginine import(GO:0090467) L-arginine import across plasma membrane(GO:0097638) L-arginine transport(GO:1902023) L-arginine import into cell(GO:1902765) amino acid import into cell(GO:1902837) L-arginine transmembrane transport(GO:1903400) arginine transmembrane transport(GO:1903826) |
| 0.1 | 0.4 | GO:0042546 | cell wall biogenesis(GO:0042546) cell wall macromolecule biosynthetic process(GO:0044038) cellular component macromolecule biosynthetic process(GO:0070589) |
| 0.1 | 0.7 | GO:0046485 | ether lipid metabolic process(GO:0046485) |
| 0.1 | 2.3 | GO:0006614 | SRP-dependent cotranslational protein targeting to membrane(GO:0006614) |
| 0.1 | 1.9 | GO:0015721 | bile acid and bile salt transport(GO:0015721) |
| 0.1 | 0.4 | GO:0050666 | regulation of sulfur amino acid metabolic process(GO:0031335) regulation of homocysteine metabolic process(GO:0050666) |
| 0.1 | 0.9 | GO:0006283 | transcription-coupled nucleotide-excision repair(GO:0006283) |
| 0.1 | 0.8 | GO:0038065 | collagen-activated tyrosine kinase receptor signaling pathway(GO:0038063) collagen-activated signaling pathway(GO:0038065) |
| 0.1 | 2.4 | GO:0005979 | regulation of glycogen biosynthetic process(GO:0005979) regulation of glucan biosynthetic process(GO:0010962) |
| 0.1 | 2.7 | GO:2001235 | positive regulation of apoptotic signaling pathway(GO:2001235) |
| 0.1 | 0.5 | GO:1990511 | piRNA biosynthetic process(GO:1990511) |
| 0.1 | 2.8 | GO:0060287 | epithelial cilium movement involved in determination of left/right asymmetry(GO:0060287) |
| 0.1 | 0.7 | GO:0097241 | hematopoietic stem cell migration to bone marrow(GO:0097241) |
| 0.1 | 1.1 | GO:0046627 | negative regulation of insulin receptor signaling pathway(GO:0046627) negative regulation of cellular response to insulin stimulus(GO:1900077) |
| 0.1 | 4.6 | GO:0006749 | glutathione metabolic process(GO:0006749) |
| 0.1 | 0.8 | GO:1903076 | regulation of protein localization to plasma membrane(GO:1903076) regulation of protein localization to cell periphery(GO:1904375) |
| 0.1 | 1.0 | GO:0072498 | embryonic skeletal joint development(GO:0072498) |
| 0.1 | 1.9 | GO:0051897 | positive regulation of protein kinase B signaling(GO:0051897) |
| 0.1 | 1.8 | GO:1901534 | positive regulation of hematopoietic progenitor cell differentiation(GO:1901534) |
| 0.1 | 1.2 | GO:0097531 | mast cell chemotaxis(GO:0002551) regulation of mast cell chemotaxis(GO:0060753) positive regulation of mast cell chemotaxis(GO:0060754) mast cell migration(GO:0097531) |
| 0.1 | 0.3 | GO:0051973 | positive regulation of telomerase activity(GO:0051973) |
| 0.1 | 0.8 | GO:0033687 | osteoblast proliferation(GO:0033687) |
| 0.1 | 0.5 | GO:1990564 | protein polyufmylation(GO:1990564) protein K69-linked ufmylation(GO:1990592) |
| 0.1 | 1.2 | GO:0070212 | protein poly-ADP-ribosylation(GO:0070212) |
| 0.1 | 4.7 | GO:0070830 | apical junction assembly(GO:0043297) bicellular tight junction assembly(GO:0070830) |
| 0.1 | 1.0 | GO:0015937 | coenzyme A biosynthetic process(GO:0015937) |
| 0.1 | 0.9 | GO:0048635 | negative regulation of muscle organ development(GO:0048635) |
| 0.1 | 0.7 | GO:0048478 | replication fork protection(GO:0048478) |
| 0.1 | 0.8 | GO:0000722 | telomere maintenance via recombination(GO:0000722) |
| 0.1 | 1.0 | GO:0018120 | peptidyl-arginine ADP-ribosylation(GO:0018120) |
| 0.1 | 0.5 | GO:0015936 | coenzyme A metabolic process(GO:0015936) |
| 0.1 | 1.3 | GO:0048669 | collateral sprouting in absence of injury(GO:0048669) regulation of collateral sprouting in absence of injury(GO:0048696) |
| 0.1 | 0.4 | GO:1900026 | positive regulation of substrate adhesion-dependent cell spreading(GO:1900026) |
| 0.1 | 1.4 | GO:0036092 | phosphatidylinositol-3-phosphate biosynthetic process(GO:0036092) |
| 0.1 | 1.0 | GO:0042664 | locus ceruleus development(GO:0021703) negative regulation of endodermal cell fate specification(GO:0042664) |
| 0.1 | 1.6 | GO:0002761 | regulation of myeloid leukocyte differentiation(GO:0002761) |
| 0.1 | 0.7 | GO:0009146 | dGTP catabolic process(GO:0006203) purine nucleoside triphosphate catabolic process(GO:0009146) purine deoxyribonucleoside triphosphate catabolic process(GO:0009217) dGTP metabolic process(GO:0046070) |
| 0.1 | 0.4 | GO:0035988 | chondrocyte proliferation(GO:0035988) |
| 0.1 | 0.3 | GO:0030718 | germ-line stem cell population maintenance(GO:0030718) |
| 0.1 | 1.1 | GO:0030311 | poly-N-acetyllactosamine metabolic process(GO:0030309) poly-N-acetyllactosamine biosynthetic process(GO:0030311) |
| 0.1 | 0.2 | GO:0000212 | meiotic spindle organization(GO:0000212) |
| 0.1 | 1.8 | GO:0035435 | phosphate ion transmembrane transport(GO:0035435) |
| 0.1 | 0.4 | GO:0000395 | mRNA 5'-splice site recognition(GO:0000395) |
| 0.1 | 0.5 | GO:2000178 | negative regulation of neural precursor cell proliferation(GO:2000178) |
| 0.1 | 1.7 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.1 | 3.2 | GO:0006368 | transcription elongation from RNA polymerase II promoter(GO:0006368) |
| 0.1 | 0.3 | GO:0097242 | regulation of nitric-oxide synthase activity(GO:0050999) positive regulation of nitric-oxide synthase activity(GO:0051000) beta-amyloid clearance(GO:0097242) regulation of beta-amyloid clearance(GO:1900221) |
| 0.1 | 0.5 | GO:0006556 | S-adenosylmethionine biosynthetic process(GO:0006556) |
| 0.1 | 2.2 | GO:0010499 | proteasomal ubiquitin-independent protein catabolic process(GO:0010499) |
| 0.1 | 0.4 | GO:1904668 | positive regulation of ubiquitin protein ligase activity(GO:1904668) |
| 0.1 | 0.4 | GO:0061484 | hematopoietic stem cell homeostasis(GO:0061484) |
| 0.1 | 0.8 | GO:0042026 | protein refolding(GO:0042026) |
| 0.1 | 1.1 | GO:0030150 | protein import into mitochondrial matrix(GO:0030150) |
| 0.1 | 0.3 | GO:0000012 | single strand break repair(GO:0000012) |
| 0.1 | 0.2 | GO:0002283 | neutrophil activation involved in immune response(GO:0002283) |
| 0.1 | 1.3 | GO:0001522 | pseudouridine synthesis(GO:0001522) |
| 0.1 | 0.4 | GO:0006054 | N-acetylneuraminate metabolic process(GO:0006054) |
| 0.1 | 0.8 | GO:0060236 | regulation of mitotic spindle organization(GO:0060236) |
| 0.1 | 0.4 | GO:0060282 | positive regulation of oocyte development(GO:0060282) |
| 0.1 | 0.4 | GO:0001887 | selenium compound metabolic process(GO:0001887) |
| 0.1 | 1.5 | GO:0048264 | determination of ventral identity(GO:0048264) |
| 0.1 | 1.6 | GO:0051923 | sulfation(GO:0051923) |
| 0.1 | 0.2 | GO:0072679 | chemokine production(GO:0032602) negative T cell selection(GO:0043383) thymocyte migration(GO:0072679) |
| 0.1 | 1.3 | GO:0010972 | negative regulation of G2/M transition of mitotic cell cycle(GO:0010972) |
| 0.1 | 0.2 | GO:0061400 | positive regulation of transcription from RNA polymerase II promoter in response to calcium ion(GO:0061400) |
| 0.1 | 0.2 | GO:1990918 | double-strand break repair involved in meiotic recombination(GO:1990918) |
| 0.1 | 0.5 | GO:0045947 | negative regulation of translational initiation(GO:0045947) |
| 0.1 | 1.0 | GO:0043687 | post-translational protein modification(GO:0043687) |
| 0.1 | 0.2 | GO:0036306 | embryonic heart tube elongation(GO:0036306) |
| 0.1 | 0.2 | GO:0018343 | protein farnesylation(GO:0018343) |
| 0.1 | 1.0 | GO:0051445 | regulation of meiotic cell cycle(GO:0051445) |
| 0.1 | 0.8 | GO:2000134 | negative regulation of cell cycle G1/S phase transition(GO:1902807) negative regulation of G1/S transition of mitotic cell cycle(GO:2000134) |
| 0.1 | 0.4 | GO:0001958 | endochondral ossification(GO:0001958) replacement ossification(GO:0036075) |
| 0.0 | 4.8 | GO:0007229 | integrin-mediated signaling pathway(GO:0007229) |
| 0.0 | 0.6 | GO:0045740 | positive regulation of DNA replication(GO:0045740) |
| 0.0 | 0.1 | GO:0002926 | tRNA wobble base 5-methoxycarbonylmethyl-2-thiouridine biosynthesis.(GO:0002926) |
| 0.0 | 0.3 | GO:0061635 | regulation of protein complex stability(GO:0061635) |
| 0.0 | 0.8 | GO:0018216 | peptidyl-arginine methylation(GO:0018216) |
| 0.0 | 0.6 | GO:0030214 | hyaluronan catabolic process(GO:0030214) |
| 0.0 | 1.0 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 0.0 | 0.1 | GO:0046098 | guanine metabolic process(GO:0046098) |
| 0.0 | 2.1 | GO:0006959 | humoral immune response(GO:0006959) |
| 0.0 | 1.8 | GO:0060028 | convergent extension involved in axis elongation(GO:0060028) |
| 0.0 | 1.3 | GO:0071427 | mRNA export from nucleus(GO:0006406) mRNA-containing ribonucleoprotein complex export from nucleus(GO:0071427) |
| 0.0 | 0.4 | GO:0097374 | sensory neuron axon guidance(GO:0097374) |
| 0.0 | 0.9 | GO:0051209 | release of sequestered calcium ion into cytosol(GO:0051209) |
| 0.0 | 3.2 | GO:0010951 | negative regulation of endopeptidase activity(GO:0010951) |
| 0.0 | 2.0 | GO:0018298 | protein-chromophore linkage(GO:0018298) |
| 0.0 | 0.6 | GO:0016926 | protein desumoylation(GO:0016926) |
| 0.0 | 2.0 | GO:0040036 | regulation of fibroblast growth factor receptor signaling pathway(GO:0040036) |
| 0.0 | 2.1 | GO:0002181 | cytoplasmic translation(GO:0002181) |
| 0.0 | 0.3 | GO:0046247 | carotene metabolic process(GO:0016119) carotene catabolic process(GO:0016121) terpene metabolic process(GO:0042214) terpene catabolic process(GO:0046247) |
| 0.0 | 1.3 | GO:0031629 | synaptic vesicle fusion to presynaptic active zone membrane(GO:0031629) vesicle fusion to plasma membrane(GO:0099500) |
| 0.0 | 0.5 | GO:0035493 | SNARE complex assembly(GO:0035493) |
| 0.0 | 0.4 | GO:0006488 | dolichol-linked oligosaccharide biosynthetic process(GO:0006488) |
| 0.0 | 3.9 | GO:0007160 | cell-matrix adhesion(GO:0007160) |
| 0.0 | 0.1 | GO:1900864 | mitochondrial tRNA modification(GO:0070900) mitochondrial RNA modification(GO:1900864) |
| 0.0 | 0.6 | GO:0002574 | thrombocyte differentiation(GO:0002574) |
| 0.0 | 0.2 | GO:1902369 | negative regulation of RNA catabolic process(GO:1902369) |
| 0.0 | 0.5 | GO:0007004 | RNA-dependent DNA biosynthetic process(GO:0006278) telomere maintenance via telomerase(GO:0007004) |
| 0.0 | 0.2 | GO:0015846 | polyamine transport(GO:0015846) polyamine transmembrane transport(GO:1902047) regulation of polyamine transmembrane transport(GO:1902267) positive regulation of polyamine transmembrane transport(GO:1902269) |
| 0.0 | 1.0 | GO:0030574 | collagen catabolic process(GO:0030574) multicellular organism catabolic process(GO:0044243) |
| 0.0 | 0.8 | GO:0051568 | histone H3-K4 methylation(GO:0051568) |
| 0.0 | 0.1 | GO:0070863 | positive regulation of protein exit from endoplasmic reticulum(GO:0070863) |
| 0.0 | 0.4 | GO:0042493 | response to drug(GO:0042493) |
| 0.0 | 0.4 | GO:2000601 | positive regulation of actin nucleation(GO:0051127) positive regulation of Arp2/3 complex-mediated actin nucleation(GO:2000601) |
| 0.0 | 4.3 | GO:0060326 | cell chemotaxis(GO:0060326) |
| 0.0 | 0.3 | GO:0042542 | response to hydrogen peroxide(GO:0042542) |
| 0.0 | 1.9 | GO:0033339 | pectoral fin development(GO:0033339) |
| 0.0 | 1.4 | GO:0043488 | regulation of mRNA stability(GO:0043488) |
| 0.0 | 1.9 | GO:0009617 | response to bacterium(GO:0009617) |
| 0.0 | 0.3 | GO:0036368 | cone photoresponse recovery(GO:0036368) |
| 0.0 | 0.7 | GO:0048268 | clathrin coat assembly(GO:0048268) |
| 0.0 | 2.7 | GO:0006979 | response to oxidative stress(GO:0006979) |
| 0.0 | 3.8 | GO:0051604 | protein maturation(GO:0051604) |
| 0.0 | 0.3 | GO:0007076 | mitotic chromosome condensation(GO:0007076) |
| 0.0 | 3.4 | GO:0042742 | defense response to bacterium(GO:0042742) |
| 0.0 | 3.4 | GO:0019221 | cytokine-mediated signaling pathway(GO:0019221) |
| 0.0 | 0.7 | GO:1901215 | negative regulation of neuron death(GO:1901215) |
| 0.0 | 0.8 | GO:0006289 | nucleotide-excision repair(GO:0006289) |
| 0.0 | 1.0 | GO:0007029 | endoplasmic reticulum organization(GO:0007029) |
| 0.0 | 1.4 | GO:0001894 | tissue homeostasis(GO:0001894) |
| 0.0 | 0.4 | GO:0031468 | nuclear envelope reassembly(GO:0031468) |
| 0.0 | 1.8 | GO:1990573 | potassium ion import across plasma membrane(GO:1990573) |
| 0.0 | 1.9 | GO:0071466 | cellular response to xenobiotic stimulus(GO:0071466) |
| 0.0 | 0.5 | GO:0048665 | neuron fate specification(GO:0048665) |
| 0.0 | 1.0 | GO:0003333 | amino acid transmembrane transport(GO:0003333) |
| 0.0 | 1.4 | GO:0006413 | translational initiation(GO:0006413) |
| 0.0 | 0.7 | GO:0003094 | glomerular filtration(GO:0003094) |
| 0.0 | 5.0 | GO:0043413 | protein glycosylation(GO:0006486) macromolecule glycosylation(GO:0043413) |
| 0.0 | 0.5 | GO:0097529 | myeloid leukocyte migration(GO:0097529) neutrophil migration(GO:1990266) |
| 0.0 | 1.7 | GO:0008285 | negative regulation of cell proliferation(GO:0008285) |
| 0.0 | 0.6 | GO:0031398 | positive regulation of protein ubiquitination(GO:0031398) |
| 0.0 | 0.1 | GO:0043504 | mitochondrial DNA repair(GO:0043504) |
| 0.0 | 0.2 | GO:0046959 | nonassociative learning(GO:0046958) habituation(GO:0046959) |
| 0.0 | 0.7 | GO:0031532 | actin cytoskeleton reorganization(GO:0031532) |
| 0.0 | 0.4 | GO:0006378 | mRNA polyadenylation(GO:0006378) RNA polyadenylation(GO:0043631) |
| 0.0 | 1.3 | GO:0048916 | posterior lateral line development(GO:0048916) |
| 0.0 | 0.5 | GO:0006376 | mRNA splice site selection(GO:0006376) |
| 0.0 | 0.4 | GO:0006506 | GPI anchor biosynthetic process(GO:0006506) |
| 0.0 | 0.1 | GO:0009957 | epidermal cell fate specification(GO:0009957) lateral line ganglion development(GO:0048890) |
| 0.0 | 2.0 | GO:0001889 | liver development(GO:0001889) |
| 0.0 | 0.1 | GO:1901073 | chitin biosynthetic process(GO:0006031) glucosamine-containing compound biosynthetic process(GO:1901073) |
| 0.0 | 0.2 | GO:0008089 | anterograde axonal transport(GO:0008089) |
| 0.0 | 0.8 | GO:0043044 | ATP-dependent chromatin remodeling(GO:0043044) |
| 0.0 | 1.1 | GO:0060218 | hematopoietic stem cell differentiation(GO:0060218) |
| 0.0 | 0.2 | GO:0006405 | RNA export from nucleus(GO:0006405) |
| 0.0 | 1.1 | GO:0035023 | regulation of Rho protein signal transduction(GO:0035023) |
| 0.0 | 0.1 | GO:0051382 | kinetochore assembly(GO:0051382) |
| 0.0 | 0.8 | GO:0007528 | neuromuscular junction development(GO:0007528) |
| 0.0 | 0.1 | GO:0018027 | peptidyl-lysine dimethylation(GO:0018027) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 2.0 | GO:0019815 | B cell receptor complex(GO:0019815) |
| 0.4 | 2.2 | GO:0005797 | Golgi medial cisterna(GO:0005797) |
| 0.4 | 4.1 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 0.3 | 1.4 | GO:0071541 | eukaryotic translation initiation factor 3 complex, eIF3m(GO:0071541) |
| 0.3 | 1.2 | GO:0072588 | box H/ACA snoRNP complex(GO:0031429) box H/ACA RNP complex(GO:0072588) |
| 0.3 | 5.1 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.3 | 0.8 | GO:0033065 | Rad51C-XRCC3 complex(GO:0033065) |
| 0.2 | 1.6 | GO:0030892 | nuclear cohesin complex(GO:0000798) mitotic cohesin complex(GO:0030892) nuclear mitotic cohesin complex(GO:0034990) nuclear meiotic cohesin complex(GO:0034991) |
| 0.2 | 1.4 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.2 | 1.8 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.2 | 1.6 | GO:0005853 | eukaryotic translation elongation factor 1 complex(GO:0005853) |
| 0.2 | 1.1 | GO:0033596 | TSC1-TSC2 complex(GO:0033596) |
| 0.2 | 0.8 | GO:0005880 | nuclear microtubule(GO:0005880) |
| 0.2 | 2.3 | GO:0005944 | phosphatidylinositol 3-kinase complex, class IB(GO:0005944) phosphatidylinositol 3-kinase complex, class I(GO:0097651) |
| 0.2 | 10.2 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.2 | 7.4 | GO:0022626 | cytosolic ribosome(GO:0022626) |
| 0.2 | 2.8 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.2 | 3.7 | GO:0045180 | basal cortex(GO:0045180) |
| 0.1 | 1.7 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 0.1 | 1.8 | GO:0030128 | clathrin coat of endocytic vesicle(GO:0030128) clathrin-coated endocytic vesicle membrane(GO:0030669) |
| 0.1 | 0.8 | GO:0098645 | collagen type IV trimer(GO:0005587) network-forming collagen trimer(GO:0098642) collagen network(GO:0098645) basement membrane collagen trimer(GO:0098651) |
| 0.1 | 0.8 | GO:0005782 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.1 | 0.5 | GO:0005913 | cell-cell adherens junction(GO:0005913) zonula adherens(GO:0005915) |
| 0.1 | 0.9 | GO:0005790 | smooth endoplasmic reticulum(GO:0005790) |
| 0.1 | 1.3 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 0.1 | 2.0 | GO:0030057 | desmosome(GO:0030057) |
| 0.1 | 0.4 | GO:0070209 | ASTRA complex(GO:0070209) |
| 0.1 | 1.1 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.1 | 0.7 | GO:0005662 | DNA replication factor A complex(GO:0005662) |
| 0.1 | 1.0 | GO:0000177 | cytoplasmic exosome (RNase complex)(GO:0000177) |
| 0.1 | 0.5 | GO:0070390 | transcription export complex 2(GO:0070390) |
| 0.1 | 5.4 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.1 | 3.9 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.1 | 2.2 | GO:0005839 | proteasome core complex(GO:0005839) |
| 0.1 | 1.6 | GO:0045178 | basal part of cell(GO:0045178) |
| 0.1 | 3.0 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.1 | 0.3 | GO:0034363 | intermediate-density lipoprotein particle(GO:0034363) |
| 0.1 | 0.2 | GO:0005953 | CAAX-protein geranylgeranyltransferase complex(GO:0005953) |
| 0.1 | 2.1 | GO:0043186 | P granule(GO:0043186) |
| 0.1 | 0.8 | GO:0005956 | protein kinase CK2 complex(GO:0005956) |
| 0.1 | 1.0 | GO:0031011 | Ino80 complex(GO:0031011) DNA helicase complex(GO:0033202) |
| 0.1 | 0.7 | GO:0098888 | presynaptic endocytic zone(GO:0098833) presynaptic endocytic zone membrane(GO:0098835) extrinsic component of presynaptic membrane(GO:0098888) extrinsic component of presynaptic endocytic zone(GO:0098894) |
| 0.1 | 7.4 | GO:0070160 | bicellular tight junction(GO:0005923) occluding junction(GO:0070160) |
| 0.1 | 0.9 | GO:0071014 | post-mRNA release spliceosomal complex(GO:0071014) |
| 0.1 | 2.2 | GO:0017053 | transcriptional repressor complex(GO:0017053) |
| 0.1 | 1.3 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
| 0.1 | 2.4 | GO:0001750 | photoreceptor outer segment(GO:0001750) |
| 0.1 | 0.7 | GO:0005922 | connexon complex(GO:0005922) |
| 0.0 | 0.2 | GO:0005850 | eukaryotic translation initiation factor 2 complex(GO:0005850) |
| 0.0 | 1.4 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.0 | 4.3 | GO:0005802 | trans-Golgi network(GO:0005802) |
| 0.0 | 0.5 | GO:0045095 | keratin filament(GO:0045095) |
| 0.0 | 0.6 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.0 | 1.0 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.0 | 2.4 | GO:0008023 | transcription elongation factor complex(GO:0008023) |
| 0.0 | 2.3 | GO:0072686 | mitotic spindle(GO:0072686) |
| 0.0 | 0.6 | GO:0043189 | NuA4 histone acetyltransferase complex(GO:0035267) H4/H2A histone acetyltransferase complex(GO:0043189) H4 histone acetyltransferase complex(GO:1902562) |
| 0.0 | 2.9 | GO:0055037 | recycling endosome(GO:0055037) |
| 0.0 | 1.2 | GO:0005940 | septin ring(GO:0005940) septin complex(GO:0031105) septin cytoskeleton(GO:0032156) |
| 0.0 | 0.7 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.0 | 2.2 | GO:0005814 | centriole(GO:0005814) |
| 0.0 | 1.0 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.0 | 7.6 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.0 | 0.4 | GO:0071339 | MLL1/2 complex(GO:0044665) MLL1 complex(GO:0071339) |
| 0.0 | 0.4 | GO:0000815 | ESCRT III complex(GO:0000815) |
| 0.0 | 0.6 | GO:0033116 | endoplasmic reticulum-Golgi intermediate compartment membrane(GO:0033116) |
| 0.0 | 0.2 | GO:0000930 | gamma-tubulin complex(GO:0000930) |
| 0.0 | 0.1 | GO:0000506 | glycosylphosphatidylinositol-N-acetylglucosaminyltransferase (GPI-GnT) complex(GO:0000506) |
| 0.0 | 2.5 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.0 | 0.1 | GO:0035517 | PR-DUB complex(GO:0035517) |
| 0.0 | 1.5 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.0 | 0.4 | GO:0005686 | U2 snRNP(GO:0005686) |
| 0.0 | 0.1 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.0 | 0.4 | GO:0071144 | SMAD2-SMAD3 protein complex(GO:0071144) |
| 0.0 | 0.2 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.0 | 1.5 | GO:0005604 | basement membrane(GO:0005604) |
| 0.0 | 2.3 | GO:0009898 | cytoplasmic side of plasma membrane(GO:0009898) |
| 0.0 | 3.7 | GO:0005815 | microtubule organizing center(GO:0005815) |
| 0.0 | 0.8 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.0 | 0.6 | GO:0016592 | mediator complex(GO:0016592) |
| 0.0 | 3.0 | GO:0000323 | lytic vacuole(GO:0000323) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.3 | 4.0 | GO:0070224 | sulfide:quinone oxidoreductase activity(GO:0070224) |
| 0.9 | 2.8 | GO:0016618 | hydroxypyruvate reductase activity(GO:0016618) glyoxylate reductase (NADP) activity(GO:0030267) |
| 0.6 | 4.5 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 0.5 | 2.7 | GO:0060182 | apelin receptor activity(GO:0060182) |
| 0.5 | 2.6 | GO:0071074 | eukaryotic initiation factor eIF2 binding(GO:0071074) |
| 0.5 | 4.0 | GO:0003993 | acid phosphatase activity(GO:0003993) |
| 0.5 | 1.9 | GO:0052905 | tRNA (guanine(9)-N(1))-methyltransferase activity(GO:0052905) |
| 0.4 | 2.6 | GO:0004459 | L-lactate dehydrogenase activity(GO:0004459) |
| 0.3 | 1.7 | GO:0047066 | phospholipid-hydroperoxide glutathione peroxidase activity(GO:0047066) |
| 0.3 | 5.1 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.3 | 1.3 | GO:0005471 | ATP:ADP antiporter activity(GO:0005471) |
| 0.3 | 1.9 | GO:0047429 | nucleoside-triphosphate diphosphatase activity(GO:0047429) |
| 0.3 | 1.9 | GO:0003987 | acetate-CoA ligase activity(GO:0003987) |
| 0.3 | 0.9 | GO:0004342 | glucosamine-6-phosphate deaminase activity(GO:0004342) |
| 0.3 | 3.5 | GO:0046920 | alpha-(1->3)-fucosyltransferase activity(GO:0046920) |
| 0.3 | 1.9 | GO:0010859 | calcium-dependent cysteine-type endopeptidase inhibitor activity(GO:0010859) |
| 0.3 | 1.1 | GO:0016263 | glycoprotein-N-acetylgalactosamine 3-beta-galactosyltransferase activity(GO:0016263) beta-1,3-galactosyltransferase activity(GO:0048531) |
| 0.3 | 0.8 | GO:0032574 | 5'-3' RNA helicase activity(GO:0032574) |
| 0.3 | 2.8 | GO:0005451 | monovalent cation:proton antiporter activity(GO:0005451) sodium:proton antiporter activity(GO:0015385) potassium:proton antiporter activity(GO:0015386) |
| 0.3 | 2.0 | GO:0015057 | thrombin receptor activity(GO:0015057) |
| 0.2 | 1.0 | GO:0004576 | oligosaccharyl transferase activity(GO:0004576) dolichyl-diphosphooligosaccharide-protein glycotransferase activity(GO:0004579) |
| 0.2 | 1.4 | GO:0016744 | transferase activity, transferring aldehyde or ketonic groups(GO:0016744) |
| 0.2 | 0.7 | GO:0036310 | annealing helicase activity(GO:0036310) |
| 0.2 | 0.9 | GO:0097108 | hedgehog family protein binding(GO:0097108) |
| 0.2 | 3.3 | GO:0038191 | neuropilin binding(GO:0038191) |
| 0.2 | 1.7 | GO:0072518 | Rho-dependent protein serine/threonine kinase activity(GO:0072518) |
| 0.2 | 3.9 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) alpha-actinin binding(GO:0051393) |
| 0.2 | 0.8 | GO:0003989 | acetyl-CoA carboxylase activity(GO:0003989) |
| 0.2 | 3.6 | GO:0016857 | racemase and epimerase activity, acting on carbohydrates and derivatives(GO:0016857) |
| 0.2 | 0.5 | GO:0071568 | UFM1 transferase activity(GO:0071568) |
| 0.2 | 1.6 | GO:0004062 | aryl sulfotransferase activity(GO:0004062) |
| 0.2 | 0.9 | GO:0048763 | ryanodine-sensitive calcium-release channel activity(GO:0005219) calcium-induced calcium release activity(GO:0048763) |
| 0.2 | 2.4 | GO:0004029 | aldehyde dehydrogenase (NAD) activity(GO:0004029) |
| 0.2 | 0.5 | GO:0070097 | delta-catenin binding(GO:0070097) |
| 0.2 | 1.5 | GO:0008494 | translation activator activity(GO:0008494) |
| 0.2 | 0.5 | GO:0001734 | mRNA (N6-adenosine)-methyltransferase activity(GO:0001734) |
| 0.2 | 1.3 | GO:0038036 | sphingosine-1-phosphate receptor activity(GO:0038036) |
| 0.2 | 1.9 | GO:0015125 | bile acid transmembrane transporter activity(GO:0015125) |
| 0.2 | 0.8 | GO:0008821 | crossover junction endodeoxyribonuclease activity(GO:0008821) |
| 0.2 | 2.0 | GO:0016775 | creatine kinase activity(GO:0004111) phosphotransferase activity, nitrogenous group as acceptor(GO:0016775) |
| 0.1 | 0.1 | GO:0009019 | tRNA (guanine-N1-)-methyltransferase activity(GO:0009019) |
| 0.1 | 1.7 | GO:0031720 | haptoglobin binding(GO:0031720) |
| 0.1 | 0.5 | GO:0071253 | connexin binding(GO:0071253) |
| 0.1 | 2.6 | GO:0009982 | pseudouridine synthase activity(GO:0009982) |
| 0.1 | 4.6 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.1 | 0.8 | GO:0019107 | myristoyltransferase activity(GO:0019107) |
| 0.1 | 0.5 | GO:0005173 | stem cell factor receptor binding(GO:0005173) |
| 0.1 | 0.9 | GO:0015189 | L-lysine transmembrane transporter activity(GO:0015189) |
| 0.1 | 1.8 | GO:0015272 | ATP-activated inward rectifier potassium channel activity(GO:0015272) |
| 0.1 | 1.8 | GO:0005315 | inorganic phosphate transmembrane transporter activity(GO:0005315) |
| 0.1 | 1.9 | GO:0033613 | activating transcription factor binding(GO:0033613) |
| 0.1 | 0.6 | GO:1990757 | anaphase-promoting complex binding(GO:0010997) ubiquitin ligase activator activity(GO:1990757) |
| 0.1 | 0.6 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.1 | 1.4 | GO:0001871 | pattern binding(GO:0001871) polysaccharide binding(GO:0030247) |
| 0.1 | 2.1 | GO:0070003 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 0.1 | 0.8 | GO:0008379 | thioredoxin peroxidase activity(GO:0008379) |
| 0.1 | 1.2 | GO:0005172 | vascular endothelial growth factor receptor binding(GO:0005172) |
| 0.1 | 0.4 | GO:0004652 | polynucleotide adenylyltransferase activity(GO:0004652) |
| 0.1 | 1.2 | GO:1990404 | protein ADP-ribosylase activity(GO:1990404) |
| 0.1 | 3.6 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.1 | 3.6 | GO:0045182 | translation regulator activity(GO:0045182) |
| 0.1 | 4.0 | GO:0019957 | C-C chemokine binding(GO:0019957) |
| 0.1 | 2.2 | GO:0043236 | laminin binding(GO:0043236) |
| 0.1 | 1.0 | GO:0070700 | BMP receptor binding(GO:0070700) |
| 0.1 | 1.2 | GO:0030368 | interleukin-17 receptor activity(GO:0030368) |
| 0.1 | 1.0 | GO:0003956 | NAD(P)+-protein-arginine ADP-ribosyltransferase activity(GO:0003956) |
| 0.1 | 1.3 | GO:0097199 | cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:0097199) |
| 0.1 | 0.4 | GO:0004122 | cystathionine beta-synthase activity(GO:0004122) |
| 0.1 | 1.9 | GO:0008378 | galactosyltransferase activity(GO:0008378) |
| 0.1 | 0.7 | GO:0016714 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced pteridine as one donor, and incorporation of one atom of oxygen(GO:0016714) |
| 0.1 | 1.1 | GO:0008532 | N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase activity(GO:0008532) |
| 0.1 | 2.1 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.1 | 1.4 | GO:0035005 | 1-phosphatidylinositol-4-phosphate 3-kinase activity(GO:0035005) |
| 0.1 | 4.0 | GO:0005178 | integrin binding(GO:0005178) |
| 0.1 | 1.0 | GO:0061575 | cyclin-dependent protein serine/threonine kinase activator activity(GO:0061575) |
| 0.1 | 2.0 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.1 | 4.1 | GO:0016209 | antioxidant activity(GO:0016209) |
| 0.1 | 0.4 | GO:0008430 | selenium binding(GO:0008430) |
| 0.1 | 0.2 | GO:0004662 | CAAX-protein geranylgeranyltransferase activity(GO:0004662) |
| 0.1 | 13.3 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.1 | 1.4 | GO:0008143 | poly(A) binding(GO:0008143) |
| 0.1 | 1.0 | GO:0019843 | rRNA binding(GO:0019843) |
| 0.1 | 0.4 | GO:0015616 | DNA translocase activity(GO:0015616) |
| 0.1 | 0.5 | GO:0004366 | glycerol-3-phosphate O-acyltransferase activity(GO:0004366) |
| 0.1 | 1.0 | GO:0031726 | CCR1 chemokine receptor binding(GO:0031726) CCR4 chemokine receptor binding(GO:0031729) |
| 0.1 | 0.8 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.1 | 1.5 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.1 | 0.9 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.1 | 0.8 | GO:0032041 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.1 | 1.0 | GO:0019531 | oxalate transmembrane transporter activity(GO:0019531) |
| 0.1 | 0.7 | GO:0008327 | methyl-CpG binding(GO:0008327) |
| 0.1 | 0.7 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.1 | 2.9 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.1 | 1.0 | GO:0016307 | phosphatidylinositol phosphate kinase activity(GO:0016307) |
| 0.0 | 0.3 | GO:0016316 | phosphatidylinositol-3,4-bisphosphate 4-phosphatase activity(GO:0016316) |
| 0.0 | 0.7 | GO:0005504 | fatty acid binding(GO:0005504) |
| 0.0 | 0.7 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.0 | 0.7 | GO:0005243 | gap junction channel activity(GO:0005243) |
| 0.0 | 3.4 | GO:0004896 | cytokine receptor activity(GO:0004896) |
| 0.0 | 0.8 | GO:0016273 | arginine N-methyltransferase activity(GO:0016273) protein-arginine N-methyltransferase activity(GO:0016274) |
| 0.0 | 0.6 | GO:0004415 | hyalurononglucosaminidase activity(GO:0004415) |
| 0.0 | 0.7 | GO:0015106 | bicarbonate transmembrane transporter activity(GO:0015106) |
| 0.0 | 0.7 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.0 | 0.1 | GO:0004422 | hypoxanthine phosphoribosyltransferase activity(GO:0004422) |
| 0.0 | 2.1 | GO:0008020 | G-protein coupled photoreceptor activity(GO:0008020) |
| 0.0 | 0.2 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.0 | 2.8 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.0 | 0.6 | GO:0016929 | SUMO-specific protease activity(GO:0016929) |
| 0.0 | 0.2 | GO:0061656 | SUMO conjugating enzyme activity(GO:0061656) |
| 0.0 | 0.4 | GO:0070567 | cytidylyltransferase activity(GO:0070567) |
| 0.0 | 2.0 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.0 | 0.3 | GO:0003834 | beta-carotene 15,15'-monooxygenase activity(GO:0003834) carotenoid dioxygenase activity(GO:0010436) |
| 0.0 | 0.5 | GO:0070628 | proteasome binding(GO:0070628) |
| 0.0 | 0.2 | GO:0047192 | 1-alkylglycerophosphocholine O-acetyltransferase activity(GO:0047192) |
| 0.0 | 3.0 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 3.0 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.0 | 0.2 | GO:0042978 | ornithine decarboxylase activator activity(GO:0042978) |
| 0.0 | 0.5 | GO:0008320 | protein transmembrane transporter activity(GO:0008320) |
| 0.0 | 3.9 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 0.3 | GO:0001222 | transcription corepressor binding(GO:0001222) |
| 0.0 | 4.8 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 2.0 | GO:0001594 | trace-amine receptor activity(GO:0001594) |
| 0.0 | 0.2 | GO:0008048 | calcium sensitive guanylate cyclase activator activity(GO:0008048) |
| 0.0 | 0.9 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.0 | 0.1 | GO:0015217 | ADP transmembrane transporter activity(GO:0015217) coenzyme A transmembrane transporter activity(GO:0015228) adenosine 3',5'-bisphosphate transmembrane transporter activity(GO:0071077) AMP transmembrane transporter activity(GO:0080122) |
| 0.0 | 0.3 | GO:0003886 | DNA (cytosine-5-)-methyltransferase activity(GO:0003886) |
| 0.0 | 0.1 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.0 | 1.3 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.0 | 0.7 | GO:0016881 | acid-amino acid ligase activity(GO:0016881) |
| 0.0 | 1.5 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.0 | 1.5 | GO:0016836 | hydro-lyase activity(GO:0016836) |
| 0.0 | 2.0 | GO:0001227 | transcriptional repressor activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001227) |
| 0.0 | 0.1 | GO:0070698 | type I activin receptor binding(GO:0070698) |
| 0.0 | 0.1 | GO:0004311 | farnesyltranstransferase activity(GO:0004311) |
| 0.0 | 0.3 | GO:0015145 | glucose transmembrane transporter activity(GO:0005355) monosaccharide transmembrane transporter activity(GO:0015145) hexose transmembrane transporter activity(GO:0015149) |
| 0.0 | 1.0 | GO:0004866 | endopeptidase inhibitor activity(GO:0004866) |
| 0.0 | 0.1 | GO:0003910 | DNA ligase activity(GO:0003909) DNA ligase (ATP) activity(GO:0003910) |
| 0.0 | 0.1 | GO:0042903 | tubulin deacetylase activity(GO:0042903) |
| 0.0 | 4.4 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 1.8 | GO:0030276 | clathrin binding(GO:0030276) |
| 0.0 | 0.4 | GO:0070411 | I-SMAD binding(GO:0070411) |
| 0.0 | 0.1 | GO:0017176 | phosphatidylinositol N-acetylglucosaminyltransferase activity(GO:0017176) |
| 0.0 | 0.2 | GO:0004724 | magnesium-dependent protein serine/threonine phosphatase activity(GO:0004724) |
| 0.0 | 0.3 | GO:0003684 | damaged DNA binding(GO:0003684) |
| 0.0 | 0.4 | GO:0017147 | Wnt-protein binding(GO:0017147) |
| 0.0 | 1.4 | GO:0004714 | transmembrane receptor protein tyrosine kinase activity(GO:0004714) |
| 0.0 | 0.1 | GO:0004051 | arachidonate 5-lipoxygenase activity(GO:0004051) |
| 0.0 | 0.8 | GO:0008094 | DNA-dependent ATPase activity(GO:0008094) |
| 0.0 | 0.4 | GO:0042162 | telomeric DNA binding(GO:0042162) |
| 0.0 | 0.5 | GO:0019239 | deaminase activity(GO:0019239) |
| 0.0 | 0.3 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.0 | 0.4 | GO:0016755 | transferase activity, transferring amino-acyl groups(GO:0016755) |
| 0.0 | 1.3 | GO:0004519 | endonuclease activity(GO:0004519) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.8 | ST JAK STAT PATHWAY | Jak-STAT Pathway |
| 0.2 | 1.8 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 0.2 | 2.3 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 0.2 | 1.3 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 0.2 | 2.6 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.2 | 3.4 | PID INTEGRIN A4B1 PATHWAY | Alpha4 beta1 integrin signaling events |
| 0.1 | 2.1 | PID CONE PATHWAY | Visual signal transduction: Cones |
| 0.1 | 2.0 | PID NECTIN PATHWAY | Nectin adhesion pathway |
| 0.1 | 1.3 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.1 | 4.5 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 1.4 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.1 | 2.0 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.1 | 2.2 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.1 | 1.3 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.1 | 1.0 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.1 | 0.9 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.1 | 0.8 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
| 0.1 | 0.9 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.1 | 0.8 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.0 | 0.8 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.0 | 3.2 | PID P73PATHWAY | p73 transcription factor network |
| 0.0 | 0.9 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 0.8 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.0 | 1.2 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.0 | 0.6 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.0 | 1.0 | PID IL12 2PATHWAY | IL12-mediated signaling events |
| 0.0 | 2.3 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 1.1 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.0 | 0.8 | PID ATR PATHWAY | ATR signaling pathway |
| 0.0 | 0.8 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.0 | 1.3 | PID BCR 5PATHWAY | BCR signaling pathway |
| 0.0 | 0.7 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.0 | 0.3 | PID RB 1PATHWAY | Regulation of retinoblastoma protein |
| 0.0 | 0.7 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.0 | 2.5 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 0.4 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.0 | 0.5 | PID FGF PATHWAY | FGF signaling pathway |
| 0.0 | 0.3 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.0 | 0.3 | PID PLK1 PATHWAY | PLK1 signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.3 | REACTOME ACETYLCHOLINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Acetylcholine Neurotransmitter Release Cycle |
| 0.4 | 1.9 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.2 | 4.5 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.2 | 0.8 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.2 | 8.6 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.2 | 1.8 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.2 | 0.8 | REACTOME PURINE CATABOLISM | Genes involved in Purine catabolism |
| 0.1 | 0.9 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 0.1 | 10.2 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.1 | 1.8 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.1 | 1.0 | REACTOME TRANSPORT OF ORGANIC ANIONS | Genes involved in Transport of organic anions |
| 0.1 | 3.2 | REACTOME SMAD2 SMAD3 SMAD4 HETEROTRIMER REGULATES TRANSCRIPTION | Genes involved in SMAD2/SMAD3:SMAD4 heterotrimer regulates transcription |
| 0.1 | 1.6 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.1 | 2.7 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.1 | 4.5 | REACTOME TRANSLATION | Genes involved in Translation |
| 0.1 | 1.0 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 0.1 | 0.8 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.1 | 0.8 | REACTOME INHIBITION OF REPLICATION INITIATION OF DAMAGED DNA BY RB1 E2F1 | Genes involved in Inhibition of replication initiation of damaged DNA by RB1/E2F1 |
| 0.1 | 2.3 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.1 | 0.8 | REACTOME FATTY ACYL COA BIOSYNTHESIS | Genes involved in Fatty Acyl-CoA Biosynthesis |
| 0.1 | 3.8 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.1 | 1.9 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.1 | 2.8 | REACTOME INTERACTIONS OF VPR WITH HOST CELLULAR PROTEINS | Genes involved in Interactions of Vpr with host cellular proteins |
| 0.1 | 0.4 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.1 | 2.0 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 0.1 | 0.5 | REACTOME FGFR4 LIGAND BINDING AND ACTIVATION | Genes involved in FGFR4 ligand binding and activation |
| 0.1 | 1.3 | REACTOME PRE NOTCH TRANSCRIPTION AND TRANSLATION | Genes involved in Pre-NOTCH Transcription and Translation |
| 0.1 | 2.2 | REACTOME CROSS PRESENTATION OF SOLUBLE EXOGENOUS ANTIGENS ENDOSOMES | Genes involved in Cross-presentation of soluble exogenous antigens (endosomes) |
| 0.1 | 1.3 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.1 | 1.0 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.1 | 0.6 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.1 | 2.0 | REACTOME NUCLEOTIDE EXCISION REPAIR | Genes involved in Nucleotide Excision Repair |
| 0.1 | 0.9 | REACTOME GPVI MEDIATED ACTIVATION CASCADE | Genes involved in GPVI-mediated activation cascade |
| 0.0 | 0.6 | REACTOME RNA POL I PROMOTER OPENING | Genes involved in RNA Polymerase I Promoter Opening |
| 0.0 | 1.2 | REACTOME EXTENSION OF TELOMERES | Genes involved in Extension of Telomeres |
| 0.0 | 0.9 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.0 | 0.7 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.0 | 1.6 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.0 | 0.9 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.0 | 0.2 | REACTOME REGULATION OF THE FANCONI ANEMIA PATHWAY | Genes involved in Regulation of the Fanconi anemia pathway |
| 0.0 | 0.1 | REACTOME N GLYCAN ANTENNAE ELONGATION IN THE MEDIAL TRANS GOLGI | Genes involved in N-glycan antennae elongation in the medial/trans-Golgi |
| 0.0 | 0.2 | REACTOME RECRUITMENT OF NUMA TO MITOTIC CENTROSOMES | Genes involved in Recruitment of NuMA to mitotic centrosomes |
| 0.0 | 0.3 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.0 | 0.5 | REACTOME SYNTHESIS OF PA | Genes involved in Synthesis of PA |
| 0.0 | 1.3 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 2.5 | REACTOME METABOLISM OF CARBOHYDRATES | Genes involved in Metabolism of carbohydrates |
| 0.0 | 0.3 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.0 | 0.8 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.0 | 0.2 | REACTOME ACYL CHAIN REMODELLING OF PC | Genes involved in Acyl chain remodelling of PC |
| 0.0 | 0.7 | REACTOME CELL SURFACE INTERACTIONS AT THE VASCULAR WALL | Genes involved in Cell surface interactions at the vascular wall |
| 0.0 | 0.2 | REACTOME RETROGRADE NEUROTROPHIN SIGNALLING | Genes involved in Retrograde neurotrophin signalling |
| 0.0 | 0.8 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 0.7 | REACTOME ASPARAGINE N LINKED GLYCOSYLATION | Genes involved in Asparagine N-linked glycosylation |
| 0.0 | 0.1 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.0 | 0.6 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.0 | 1.2 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 0.6 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.0 | 0.5 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |