Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
Results for creb3l1
Z-value: 0.90
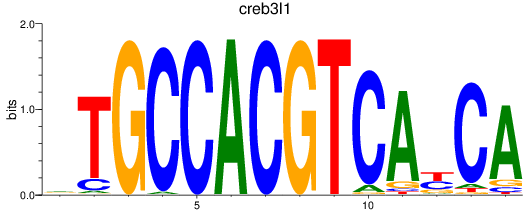
Transcription factors associated with creb3l1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
creb3l1
|
ENSDARG00000015793 | cAMP responsive element binding protein 3-like 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| creb3l1 | dr11_v1_chr7_+_38897836_38897836 | -0.15 | 1.5e-01 | Click! |
Activity profile of creb3l1 motif
Sorted Z-values of creb3l1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr22_-_2886937 | 17.27 |
ENSDART00000063533
|
aqp12
|
aquaporin 12 |
| chr1_-_7930679 | 8.10 |
ENSDART00000146090
|
si:dkey-79f11.10
|
si:dkey-79f11.10 |
| chr18_-_34549721 | 8.09 |
ENSDART00000137101
ENSDART00000021880 |
ssr3
|
signal sequence receptor, gamma |
| chr21_+_10702031 | 8.05 |
ENSDART00000102304
|
lman1
|
lectin, mannose-binding, 1 |
| chr21_+_10701834 | 7.66 |
ENSDART00000192473
|
lman1
|
lectin, mannose-binding, 1 |
| chr16_+_19637384 | 7.28 |
ENSDART00000184773
ENSDART00000191895 ENSDART00000182020 ENSDART00000135359 |
macc1
|
metastasis associated in colon cancer 1 |
| chr18_-_7031409 | 5.81 |
ENSDART00000148485
ENSDART00000005405 |
calub
|
calumenin b |
| chr23_-_31266586 | 5.49 |
ENSDART00000139746
|
si:dkey-261l7.2
|
si:dkey-261l7.2 |
| chr6_+_13506841 | 5.44 |
ENSDART00000032331
|
gmppab
|
GDP-mannose pyrophosphorylase Ab |
| chr14_-_45464112 | 5.43 |
ENSDART00000173209
|
ehd1a
|
EH-domain containing 1a |
| chr4_+_25651720 | 5.19 |
ENSDART00000100693
ENSDART00000100717 |
acot16
|
acyl-CoA thioesterase 16 |
| chr4_-_26095755 | 4.86 |
ENSDART00000100611
ENSDART00000191266 |
si:ch211-244b2.3
|
si:ch211-244b2.3 |
| chr6_-_50685862 | 4.82 |
ENSDART00000134146
|
mtss1
|
metastasis suppressor 1 |
| chr12_-_17479078 | 4.65 |
ENSDART00000079115
|
papss2b
|
3'-phosphoadenosine 5'-phosphosulfate synthase 2b |
| chr21_-_43636595 | 4.48 |
ENSDART00000151115
ENSDART00000151486 ENSDART00000151778 |
si:ch1073-263o8.2
|
si:ch1073-263o8.2 |
| chr1_+_54124209 | 4.43 |
ENSDART00000187730
|
LO017722.1
|
|
| chr9_-_29497916 | 4.23 |
ENSDART00000060246
|
dnajc3a
|
DnaJ (Hsp40) homolog, subfamily C, member 3a |
| chr25_+_37443194 | 3.96 |
ENSDART00000163178
ENSDART00000190262 |
slc10a3
|
solute carrier family 10, member 3 |
| chr25_-_3034668 | 3.90 |
ENSDART00000053405
|
scamp2
|
secretory carrier membrane protein 2 |
| chr18_-_7481036 | 3.88 |
ENSDART00000101292
|
si:dkey-238c7.16
|
si:dkey-238c7.16 |
| chr5_-_48680580 | 3.86 |
ENSDART00000031194
|
lysmd3
|
LysM, putative peptidoglycan-binding, domain containing 3 |
| chr4_+_13568469 | 3.75 |
ENSDART00000171235
ENSDART00000136152 |
calua
|
calumenin a |
| chr4_-_17812131 | 3.66 |
ENSDART00000025731
|
spic
|
Spi-C transcription factor (Spi-1/PU.1 related) |
| chr5_+_32924669 | 3.66 |
ENSDART00000085219
|
lmo4a
|
LIM domain only 4a |
| chr4_+_11464255 | 3.58 |
ENSDART00000008584
|
gdi2
|
GDP dissociation inhibitor 2 |
| chr3_+_32862730 | 3.25 |
ENSDART00000144939
ENSDART00000125126 ENSDART00000144531 ENSDART00000141717 ENSDART00000137599 |
zgc:162613
|
zgc:162613 |
| chr3_-_26183699 | 3.14 |
ENSDART00000147517
ENSDART00000140731 |
si:ch211-11k18.4
|
si:ch211-11k18.4 |
| chr1_+_277731 | 3.11 |
ENSDART00000133431
|
cenpe
|
centromere protein E |
| chr4_+_68562464 | 3.10 |
ENSDART00000192954
|
BX548011.4
|
|
| chr23_-_25798099 | 2.95 |
ENSDART00000041833
|
fitm2
|
fat storage-inducing transmembrane protein 2 |
| chr1_-_18592068 | 2.70 |
ENSDART00000082063
|
fam114a1
|
family with sequence similarity 114, member A1 |
| chr20_-_22464250 | 2.70 |
ENSDART00000165904
|
pdgfra
|
platelet-derived growth factor receptor, alpha polypeptide |
| chr18_+_2153530 | 2.70 |
ENSDART00000158255
|
pygo1
|
pygopus family PHD finger 1 |
| chr7_-_30177691 | 2.68 |
ENSDART00000046689
|
tmed3
|
transmembrane p24 trafficking protein 3 |
| chr22_+_30446987 | 2.67 |
ENSDART00000146471
|
si:dkey-169i5.4
|
si:dkey-169i5.4 |
| chr5_-_39474235 | 2.67 |
ENSDART00000171557
|
antxr2a
|
anthrax toxin receptor 2a |
| chr15_-_18115540 | 2.63 |
ENSDART00000131639
ENSDART00000047902 |
arcn1b
|
archain 1b |
| chr10_-_3427589 | 2.57 |
ENSDART00000133452
ENSDART00000037183 |
tmed2
|
transmembrane p24 trafficking protein 2 |
| chr22_+_31059919 | 2.54 |
ENSDART00000077063
|
sec13
|
SEC13 homolog, nuclear pore and COPII coat complex component |
| chr7_+_18100996 | 2.42 |
ENSDART00000055810
|
rab1ba
|
zRAB1B, member RAS oncogene family a |
| chr10_+_31646020 | 2.34 |
ENSDART00000115251
|
esama
|
endothelial cell adhesion molecule a |
| chr23_-_306796 | 2.30 |
ENSDART00000143125
|
anks1aa
|
ankyrin repeat and sterile alpha motif domain containing 1Aa |
| chr19_+_33139164 | 2.30 |
ENSDART00000043039
|
fam84b
|
family with sequence similarity 84, member B |
| chr7_-_45990681 | 2.25 |
ENSDART00000165441
|
si:ch211-260e23.7
|
si:ch211-260e23.7 |
| chr17_-_28797395 | 2.25 |
ENSDART00000134735
|
scfd1
|
sec1 family domain containing 1 |
| chr13_-_9875538 | 2.20 |
ENSDART00000041609
|
tm9sf3
|
transmembrane 9 superfamily member 3 |
| chr5_-_30516646 | 2.18 |
ENSDART00000014666
|
arcn1a
|
archain 1a |
| chr8_+_47219107 | 2.15 |
ENSDART00000146018
ENSDART00000075068 |
mthfr
|
methylenetetrahydrofolate reductase (NAD(P)H) |
| chr22_-_21897203 | 2.10 |
ENSDART00000158501
ENSDART00000105566 ENSDART00000136795 |
gna11a
|
guanine nucleotide binding protein (G protein), alpha 11a (Gq class) |
| chr3_-_180860 | 1.99 |
ENSDART00000059956
ENSDART00000192506 |
kdelr3
|
KDEL (Lys-Asp-Glu-Leu) endoplasmic reticulum protein retention receptor 3 |
| chr5_-_28968964 | 1.93 |
ENSDART00000184936
ENSDART00000016628 |
fam129bb
|
family with sequence similarity 129, member Bb |
| chr17_+_10090638 | 1.85 |
ENSDART00000169522
ENSDART00000160156 |
sec23a
|
Sec23 homolog A, COPII coat complex component |
| chr3_-_26184018 | 1.84 |
ENSDART00000191604
|
si:ch211-11k18.4
|
si:ch211-11k18.4 |
| chr11_-_2456656 | 1.84 |
ENSDART00000171219
ENSDART00000167938 |
slc2a10
|
solute carrier family 2 (facilitated glucose transporter), member 10 |
| chr17_-_11417551 | 1.82 |
ENSDART00000128291
|
arid4a
|
AT rich interactive domain 4A (RBP1-like) |
| chr5_+_58996765 | 1.80 |
ENSDART00000062175
|
derl2
|
derlin 2 |
| chr12_-_35582683 | 1.72 |
ENSDART00000167933
|
sec24c
|
SEC24 homolog C, COPII coat complex component |
| chr3_+_26123736 | 1.70 |
ENSDART00000010477
|
hsp70.3
|
heat shock cognate 70-kd protein, tandem duplicate 3 |
| chr12_+_30789611 | 1.68 |
ENSDART00000181501
|
aldh18a1
|
aldehyde dehydrogenase 18 family, member A1 |
| chr18_-_18587745 | 1.51 |
ENSDART00000191973
|
sf3b3
|
splicing factor 3b, subunit 3 |
| chr12_-_35582521 | 1.51 |
ENSDART00000162175
ENSDART00000168958 |
sec24c
|
SEC24 homolog C, COPII coat complex component |
| chr20_+_22799857 | 1.50 |
ENSDART00000058527
|
scfd2
|
sec1 family domain containing 2 |
| chr4_+_58576146 | 1.42 |
ENSDART00000164911
|
si:ch211-212k5.4
|
si:ch211-212k5.4 |
| chr24_+_854877 | 1.37 |
ENSDART00000188408
|
piezo2a.2
|
piezo-type mechanosensitive ion channel component 2a, tandem duplicate 2 |
| chr25_-_37331513 | 1.34 |
ENSDART00000111862
|
ldlrad3
|
low density lipoprotein receptor class A domain containing 3 |
| chr7_-_45990386 | 1.32 |
ENSDART00000186008
|
si:ch211-260e23.7
|
si:ch211-260e23.7 |
| chr22_-_10482253 | 1.24 |
ENSDART00000143164
|
aspn
|
asporin (LRR class 1) |
| chr15_-_41245962 | 1.17 |
ENSDART00000155359
|
smco4
|
single-pass membrane protein with coiled-coil domains 4 |
| chr22_-_1168291 | 1.11 |
ENSDART00000167724
|
si:ch1073-181h11.4
|
si:ch1073-181h11.4 |
| chr14_-_26411918 | 1.11 |
ENSDART00000020582
|
tmed9
|
transmembrane p24 trafficking protein 9 |
| chr4_-_5795309 | 1.08 |
ENSDART00000039987
|
pgm3
|
phosphoglucomutase 3 |
| chr20_+_43083745 | 1.00 |
ENSDART00000139014
ENSDART00000153438 |
moxd1l
|
monooxygenase, DBH-like 1, like |
| chr14_+_8594367 | 0.87 |
ENSDART00000037749
|
stx5a
|
syntaxin 5A |
| chr21_+_11105007 | 0.86 |
ENSDART00000128859
ENSDART00000193105 |
prlra
|
prolactin receptor a |
| chr18_+_7553950 | 0.80 |
ENSDART00000193420
ENSDART00000062143 |
zgc:77650
|
zgc:77650 |
| chr6_+_3864040 | 0.60 |
ENSDART00000013743
|
gorasp2
|
golgi reassembly stacking protein 2 |
| chr20_+_22799641 | 0.53 |
ENSDART00000131132
|
scfd2
|
sec1 family domain containing 2 |
| chr4_-_2637689 | 0.43 |
ENSDART00000192550
ENSDART00000021953 ENSDART00000150344 |
cog5
|
component of oligomeric golgi complex 5 |
| chr14_-_45595711 | 0.42 |
ENSDART00000074038
|
scyl1
|
SCY1-like, kinase-like 1 |
| chr4_+_2637947 | 0.41 |
ENSDART00000130623
|
dus4l
|
dihydrouridine synthase 4-like (S. cerevisiae) |
| chr21_+_5932140 | 0.41 |
ENSDART00000193767
|
rexo4
|
REX4 homolog, 3'-5' exonuclease |
| chr18_-_21097663 | 0.38 |
ENSDART00000060196
|
anpepa
|
alanyl (membrane) aminopeptidase a |
| chr15_+_23784842 | 0.30 |
ENSDART00000192889
ENSDART00000138375 |
ift20
|
intraflagellar transport 20 homolog (Chlamydomonas) |
| chr20_+_34511678 | 0.27 |
ENSDART00000061588
|
prrx1b
|
paired related homeobox 1b |
| chr18_-_3542435 | 0.17 |
ENSDART00000184017
|
dcun1d5
|
DCN1, defective in cullin neddylation 1, domain containing 5 (S. cerevisiae) |
| chr1_-_49649122 | 0.16 |
ENSDART00000134399
|
slkb
|
STE20-like kinase b |
| chr17_+_30448452 | 0.12 |
ENSDART00000153939
|
lpin1
|
lipin 1 |
| chr9_+_8898835 | 0.10 |
ENSDART00000147820
|
naxd
|
NAD(P)HX dehydratase |
| chr22_-_14346575 | 0.07 |
ENSDART00000136326
|
si:dkeyp-122a9.1
|
si:dkeyp-122a9.1 |
| chr16_+_24684107 | 0.02 |
ENSDART00000183920
|
ywhabl
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, beta polypeptide like |
Network of associatons between targets according to the STRING database.
First level regulatory network of creb3l1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 4.7 | GO:0050428 | sulfate assimilation(GO:0000103) purine ribonucleoside bisphosphate metabolic process(GO:0034035) purine ribonucleoside bisphosphate biosynthetic process(GO:0034036) 3'-phosphoadenosine 5'-phosphosulfate metabolic process(GO:0050427) 3'-phosphoadenosine 5'-phosphosulfate biosynthetic process(GO:0050428) |
| 1.2 | 4.8 | GO:0051645 | Golgi localization(GO:0051645) |
| 0.9 | 5.4 | GO:0009298 | GDP-mannose biosynthetic process(GO:0009298) |
| 0.8 | 2.5 | GO:0070278 | extracellular matrix constituent secretion(GO:0070278) |
| 0.8 | 2.3 | GO:1901187 | regulation of ephrin receptor signaling pathway(GO:1901187) |
| 0.7 | 4.2 | GO:0034975 | protein folding in endoplasmic reticulum(GO:0034975) |
| 0.6 | 33.0 | GO:0071391 | cellular response to estrogen stimulus(GO:0071391) |
| 0.5 | 2.7 | GO:0003261 | cardiac muscle progenitor cell migration to the midline involved in heart field formation(GO:0003261) |
| 0.5 | 8.1 | GO:0006614 | SRP-dependent cotranslational protein targeting to membrane(GO:0006614) |
| 0.4 | 3.0 | GO:0034389 | lipid particle organization(GO:0034389) |
| 0.4 | 1.1 | GO:0035964 | COPI-coated vesicle budding(GO:0035964) Golgi transport vesicle coating(GO:0048200) COPI coating of Golgi vesicle(GO:0048205) |
| 0.4 | 5.1 | GO:0090110 | cargo loading into COPII-coated vesicle(GO:0090110) |
| 0.3 | 1.0 | GO:0006589 | octopamine biosynthetic process(GO:0006589) dopamine catabolic process(GO:0042420) norepinephrine biosynthetic process(GO:0042421) octopamine metabolic process(GO:0046333) |
| 0.3 | 2.7 | GO:1905066 | regulation of canonical Wnt signaling pathway involved in heart development(GO:1905066) |
| 0.3 | 2.7 | GO:1901998 | toxin transport(GO:1901998) |
| 0.3 | 2.0 | GO:0006621 | protein retention in ER lumen(GO:0006621) |
| 0.3 | 4.0 | GO:0015721 | bile acid and bile salt transport(GO:0015721) |
| 0.2 | 0.9 | GO:0038161 | prolactin signaling pathway(GO:0038161) |
| 0.2 | 1.7 | GO:0055129 | proline biosynthetic process(GO:0006561) L-proline biosynthetic process(GO:0055129) |
| 0.2 | 1.4 | GO:0071260 | cellular response to mechanical stimulus(GO:0071260) |
| 0.1 | 2.3 | GO:0070527 | platelet aggregation(GO:0070527) |
| 0.1 | 2.2 | GO:0035999 | methionine biosynthetic process(GO:0009086) tetrahydrofolate interconversion(GO:0035999) |
| 0.1 | 5.4 | GO:0032456 | endocytic recycling(GO:0032456) |
| 0.1 | 1.7 | GO:0042026 | protein refolding(GO:0042026) |
| 0.1 | 9.7 | GO:0007030 | Golgi organization(GO:0007030) |
| 0.1 | 4.3 | GO:0006904 | vesicle docking involved in exocytosis(GO:0006904) |
| 0.1 | 2.1 | GO:0060158 | phospholipase C-activating dopamine receptor signaling pathway(GO:0060158) |
| 0.1 | 1.1 | GO:0006048 | UDP-N-acetylglucosamine biosynthetic process(GO:0006048) |
| 0.1 | 0.4 | GO:0000738 | DNA catabolic process, exonucleolytic(GO:0000738) |
| 0.1 | 0.3 | GO:0035790 | regulation of platelet-derived growth factor receptor signaling pathway(GO:0010640) platelet-derived growth factor receptor-alpha signaling pathway(GO:0035790) regulation of platelet-derived growth factor receptor-alpha signaling pathway(GO:2000583) |
| 0.0 | 5.2 | GO:0006637 | acyl-CoA metabolic process(GO:0006637) thioester metabolic process(GO:0035383) |
| 0.0 | 2.7 | GO:0050852 | T cell receptor signaling pathway(GO:0050852) |
| 0.0 | 1.8 | GO:0030968 | endoplasmic reticulum unfolded protein response(GO:0030968) |
| 0.0 | 3.7 | GO:0031076 | embryonic camera-type eye development(GO:0031076) |
| 0.0 | 1.8 | GO:0008643 | carbohydrate transport(GO:0008643) |
| 0.0 | 7.3 | GO:0048701 | embryonic cranial skeleton morphogenesis(GO:0048701) |
| 0.0 | 2.4 | GO:0000045 | autophagosome assembly(GO:0000045) autophagosome organization(GO:1905037) |
| 0.0 | 0.9 | GO:0048278 | vesicle docking(GO:0048278) |
| 0.0 | 0.4 | GO:0006891 | intra-Golgi vesicle-mediated transport(GO:0006891) |
| 0.0 | 4.8 | GO:0007009 | plasma membrane organization(GO:0007009) |
| 0.0 | 4.5 | GO:0016197 | endosomal transport(GO:0016197) |
| 0.0 | 0.4 | GO:0043171 | peptide catabolic process(GO:0043171) |
| 0.0 | 8.1 | GO:0006955 | immune response(GO:0006955) |
| 0.0 | 3.1 | GO:0007018 | microtubule-based movement(GO:0007018) |
| 0.0 | 0.2 | GO:0051443 | protein neddylation(GO:0045116) positive regulation of ubiquitin-protein transferase activity(GO:0051443) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 9.6 | GO:0033018 | sarcoplasmic reticulum lumen(GO:0033018) |
| 0.6 | 9.5 | GO:0030663 | COPI-coated vesicle membrane(GO:0030663) |
| 0.4 | 27.0 | GO:0030134 | ER to Golgi transport vesicle(GO:0030134) |
| 0.4 | 1.8 | GO:0000839 | Hrd1p ubiquitin ligase ERAD-L complex(GO:0000839) |
| 0.3 | 2.7 | GO:0070938 | contractile ring(GO:0070938) |
| 0.2 | 9.3 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.2 | 1.5 | GO:0071005 | U2-type precatalytic spliceosome(GO:0071005) |
| 0.2 | 1.7 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.0 | 1.0 | GO:0030667 | secretory granule membrane(GO:0030667) |
| 0.0 | 0.4 | GO:0017119 | Golgi transport complex(GO:0017119) |
| 0.0 | 2.1 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.0 | 3.0 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.0 | 0.3 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 0.0 | 7.9 | GO:0005789 | endoplasmic reticulum membrane(GO:0005789) |
| 0.0 | 3.5 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.0 | 1.8 | GO:0048471 | perinuclear region of cytoplasm(GO:0048471) |
| 0.0 | 5.3 | GO:0005783 | endoplasmic reticulum(GO:0005783) |
| 0.0 | 0.9 | GO:0031201 | SNARE complex(GO:0031201) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 4.7 | GO:0004779 | adenylylsulfate kinase activity(GO:0004020) sulfate adenylyltransferase activity(GO:0004779) sulfate adenylyltransferase (ATP) activity(GO:0004781) |
| 1.4 | 5.4 | GO:0008905 | mannose-1-phosphate guanylyltransferase activity(GO:0004475) mannose-phosphate guanylyltransferase activity(GO:0008905) |
| 1.2 | 15.7 | GO:0005537 | mannose binding(GO:0005537) |
| 1.2 | 3.6 | GO:0005093 | Rab GDP-dissociation inhibitor activity(GO:0005093) |
| 0.7 | 2.7 | GO:0048407 | platelet-derived growth factor-activated receptor activity(GO:0005017) platelet-derived growth factor binding(GO:0048407) |
| 0.7 | 2.0 | GO:0046923 | ER retention sequence binding(GO:0046923) |
| 0.7 | 4.0 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.6 | 1.8 | GO:1990381 | ubiquitin-specific protease binding(GO:1990381) |
| 0.4 | 3.0 | GO:0019992 | diacylglycerol binding(GO:0019992) |
| 0.3 | 1.7 | GO:0016774 | phosphotransferase activity, carboxyl group as acceptor(GO:0016774) |
| 0.3 | 1.0 | GO:0004500 | dopamine beta-monooxygenase activity(GO:0004500) |
| 0.3 | 1.5 | GO:0030620 | U2 snRNA binding(GO:0030620) |
| 0.2 | 2.1 | GO:0031826 | type 2A serotonin receptor binding(GO:0031826) |
| 0.2 | 5.2 | GO:0047617 | acyl-CoA hydrolase activity(GO:0047617) |
| 0.2 | 0.9 | GO:0004925 | prolactin receptor activity(GO:0004925) |
| 0.1 | 1.7 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.1 | 2.7 | GO:0031267 | small GTPase binding(GO:0031267) |
| 0.1 | 2.3 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.1 | 4.2 | GO:0051087 | chaperone binding(GO:0051087) |
| 0.1 | 3.1 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.1 | 1.1 | GO:0016868 | intramolecular transferase activity, phosphotransferases(GO:0016868) |
| 0.1 | 2.2 | GO:0016646 | oxidoreductase activity, acting on the CH-NH group of donors, NAD or NADP as acceptor(GO:0016646) |
| 0.1 | 0.4 | GO:0017150 | tRNA dihydrouridine synthase activity(GO:0017150) |
| 0.0 | 8.4 | GO:0000149 | SNARE binding(GO:0000149) |
| 0.0 | 1.4 | GO:0022833 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 0.0 | 0.1 | GO:0047453 | ATP-dependent NAD(P)H-hydrate dehydratase activity(GO:0047453) ADP-dependent NAD(P)H-hydrate dehydratase activity(GO:0052855) |
| 0.0 | 1.1 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 13.4 | GO:0005525 | GTP binding(GO:0005525) |
| 0.0 | 0.4 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 15.7 | PID TCPTP PATHWAY | Signaling events mediated by TCPTP |
| 0.2 | 2.7 | PID PDGFRA PATHWAY | PDGFR-alpha signaling pathway |
| 0.1 | 3.6 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.1 | 4.8 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.1 | 3.9 | PID ARF6 TRAFFICKING PATHWAY | Arf6 trafficking events |
| 0.1 | 3.1 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.0 | 1.2 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 2.7 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 20.9 | REACTOME TRANSPORT TO THE GOLGI AND SUBSEQUENT MODIFICATION | Genes involved in Transport to the Golgi and subsequent modification |
| 0.3 | 1.8 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.2 | 1.7 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.2 | 2.6 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.1 | 1.1 | REACTOME SYNTHESIS OF SUBSTRATES IN N GLYCAN BIOSYTHESIS | Genes involved in Synthesis of substrates in N-glycan biosythesis |
| 0.1 | 8.1 | REACTOME SRP DEPENDENT COTRANSLATIONAL PROTEIN TARGETING TO MEMBRANE | Genes involved in SRP-dependent cotranslational protein targeting to membrane |
| 0.1 | 2.0 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.0 | 1.5 | REACTOME MRNA SPLICING MINOR PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |
| 0.0 | 2.2 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.0 | 2.7 | REACTOME DOWNSTREAM SIGNAL TRANSDUCTION | Genes involved in Downstream signal transduction |
| 0.0 | 3.6 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |