Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
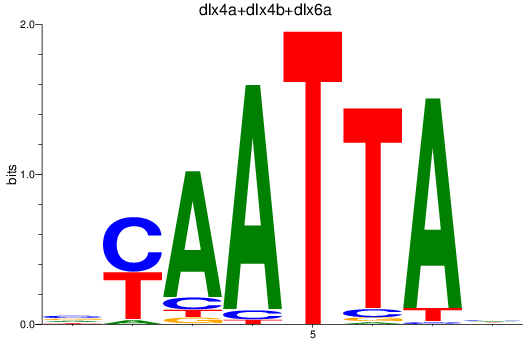
Results for dlx2a+dlx2b_dlx4a+dlx4b+dlx6a
Z-value: 0.68
Transcription factors associated with dlx2a+dlx2b_dlx4a+dlx4b+dlx6a
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
dlx2b
|
ENSDARG00000017174 | distal-less homeobox 2b |
|
dlx2a
|
ENSDARG00000079964 | distal-less homeobox 2a |
|
dlx2b
|
ENSDARG00000116017 | distal-less homeobox 2b |
|
dlx4a
|
ENSDARG00000011956 | distal-less homeobox 4a |
|
dlx6a
|
ENSDARG00000042291 | distal-less homeobox 6a |
|
dlx4b
|
ENSDARG00000071560 | distal-less homeobox 4b |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| dlx4b | dr11_v1_chr12_-_5728755_5728755 | -0.35 | 4.9e-04 | Click! |
| dlx4a | dr11_v1_chr3_+_34670076_34670076 | -0.35 | 5.5e-04 | Click! |
| dlx6a | dr11_v1_chr19_+_41419519_41419519 | -0.28 | 6.7e-03 | Click! |
| dlx2a | dr11_v1_chr9_-_3400727_3400729 | -0.12 | 2.4e-01 | Click! |
| dlx2b | dr11_v1_chr1_-_30979707_30979707 | -0.12 | 2.6e-01 | Click! |
Activity profile of dlx2a+dlx2b_dlx4a+dlx4b+dlx6a motif
Sorted Z-values of dlx2a+dlx2b_dlx4a+dlx4b+dlx6a motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr20_-_9462433 | 8.07 |
ENSDART00000152674
ENSDART00000040557 |
zgc:101840
|
zgc:101840 |
| chr6_-_54826061 | 7.68 |
ENSDART00000149982
|
tnni1b
|
troponin I type 1b (skeletal, slow) |
| chr23_+_28731379 | 6.69 |
ENSDART00000047378
|
cort
|
cortistatin |
| chr20_-_29420713 | 6.63 |
ENSDART00000147464
|
ryr3
|
ryanodine receptor 3 |
| chr25_+_29160102 | 5.84 |
ENSDART00000162854
|
pkmb
|
pyruvate kinase M1/2b |
| chr4_+_7391110 | 5.49 |
ENSDART00000160708
ENSDART00000187823 |
tnni4a
|
troponin I4a |
| chr24_-_24849091 | 5.15 |
ENSDART00000133649
ENSDART00000038290 |
crhb
|
corticotropin releasing hormone b |
| chr13_-_31435137 | 5.01 |
ENSDART00000057441
|
rtn1a
|
reticulon 1a |
| chr17_-_20717845 | 4.91 |
ENSDART00000150037
|
ank3b
|
ankyrin 3b |
| chr20_-_28800999 | 4.72 |
ENSDART00000049462
|
rab15
|
RAB15, member RAS oncogene family |
| chr19_-_32804535 | 4.40 |
ENSDART00000175613
ENSDART00000052098 |
nt5c1aa
|
5'-nucleotidase, cytosolic IAa |
| chr12_+_24342303 | 4.08 |
ENSDART00000111239
|
nrxn1a
|
neurexin 1a |
| chr4_+_12615836 | 3.91 |
ENSDART00000003583
|
lmo3
|
LIM domain only 3 |
| chr16_+_5774977 | 3.64 |
ENSDART00000134202
|
ccka
|
cholecystokinin a |
| chr7_+_30787903 | 3.42 |
ENSDART00000174000
|
apba2b
|
amyloid beta (A4) precursor protein-binding, family A, member 2b |
| chr1_+_16397063 | 3.42 |
ENSDART00000159794
|
micu3a
|
mitochondrial calcium uptake family, member 3a |
| chr9_-_21918963 | 3.39 |
ENSDART00000090782
|
lmo7a
|
LIM domain 7a |
| chr13_+_38430466 | 3.34 |
ENSDART00000132691
|
adgrb3
|
adhesion G protein-coupled receptor B3 |
| chr1_+_16396870 | 3.23 |
ENSDART00000189245
ENSDART00000113266 |
micu3a
|
mitochondrial calcium uptake family, member 3a |
| chr25_+_29662411 | 3.16 |
ENSDART00000077445
|
pim3
|
Pim-3 proto-oncogene, serine/threonine kinase |
| chr6_-_14292307 | 3.12 |
ENSDART00000177852
ENSDART00000061745 |
inpp4ab
|
inositol polyphosphate-4-phosphatase type I Ab |
| chr24_+_38306010 | 3.11 |
ENSDART00000143184
|
mybpc2b
|
myosin binding protein C, fast type b |
| chr10_-_34871737 | 3.06 |
ENSDART00000138755
|
dclk1a
|
doublecortin-like kinase 1a |
| chr11_-_6974022 | 3.04 |
ENSDART00000172851
|
COMP
|
si:ch211-43f4.1 |
| chr6_+_52804267 | 2.95 |
ENSDART00000065681
|
matn4
|
matrilin 4 |
| chr16_+_20161805 | 2.94 |
ENSDART00000192146
|
c16h2orf66
|
chromosome 16 C2orf66 homolog |
| chr8_+_21353878 | 2.91 |
ENSDART00000056420
|
alas2
|
aminolevulinate, delta-, synthase 2 |
| chr22_+_16535575 | 2.90 |
ENSDART00000083063
|
tal1
|
T-cell acute lymphocytic leukemia 1 |
| chr2_-_16380283 | 2.90 |
ENSDART00000149992
|
si:dkey-231j24.3
|
si:dkey-231j24.3 |
| chr21_+_34088110 | 2.75 |
ENSDART00000145123
ENSDART00000029599 ENSDART00000147519 |
mtmr1b
|
myotubularin related protein 1b |
| chr20_+_34933183 | 2.73 |
ENSDART00000062738
|
snap25a
|
synaptosomal-associated protein, 25a |
| chr15_-_6247775 | 2.70 |
ENSDART00000148350
|
dscamb
|
Down syndrome cell adhesion molecule b |
| chr2_-_33645411 | 2.65 |
ENSDART00000114663
|
b4galt2
|
UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 2 |
| chr4_+_11384891 | 2.63 |
ENSDART00000092381
ENSDART00000186577 ENSDART00000191054 ENSDART00000191584 |
pcloa
|
piccolo presynaptic cytomatrix protein a |
| chrM_+_9052 | 2.57 |
ENSDART00000093612
|
mt-atp6
|
ATP synthase 6, mitochondrial |
| chr19_+_46158078 | 2.56 |
ENSDART00000183933
ENSDART00000164055 |
cap2
|
CAP, adenylate cyclase-associated protein, 2 (yeast) |
| chr18_+_9637744 | 2.53 |
ENSDART00000190171
|
pclob
|
piccolo presynaptic cytomatrix protein b |
| chr4_+_26629368 | 2.50 |
ENSDART00000183575
|
iqsec3a
|
IQ motif and Sec7 domain 3a |
| chr19_-_7358184 | 2.49 |
ENSDART00000092379
|
oxr1b
|
oxidation resistance 1b |
| chr6_-_41229787 | 2.41 |
ENSDART00000065013
|
synpr
|
synaptoporin |
| chr6_+_39232245 | 2.39 |
ENSDART00000187351
|
b4galnt1b
|
beta-1,4-N-acetyl-galactosaminyl transferase 1b |
| chr21_+_34088377 | 2.24 |
ENSDART00000170070
|
mtmr1b
|
myotubularin related protein 1b |
| chr13_-_36911118 | 2.22 |
ENSDART00000048739
|
trim9
|
tripartite motif containing 9 |
| chr17_+_23298928 | 2.20 |
ENSDART00000153652
|
zgc:165461
|
zgc:165461 |
| chr16_-_17197546 | 2.13 |
ENSDART00000139939
ENSDART00000135146 ENSDART00000063800 ENSDART00000163606 |
gapdh
|
glyceraldehyde-3-phosphate dehydrogenase |
| chr16_-_12953739 | 2.11 |
ENSDART00000103894
|
cacng8b
|
calcium channel, voltage-dependent, gamma subunit 8b |
| chr14_+_23184517 | 2.07 |
ENSDART00000181410
|
enox2
|
ecto-NOX disulfide-thiol exchanger 2 |
| chr11_+_30244356 | 2.06 |
ENSDART00000036050
ENSDART00000150080 |
rs1a
|
retinoschisin 1a |
| chr15_-_22074315 | 2.04 |
ENSDART00000149830
|
drd2a
|
dopamine receptor D2a |
| chr12_+_34953038 | 1.98 |
ENSDART00000187022
ENSDART00000123988 ENSDART00000027034 |
qki2
|
QKI, KH domain containing, RNA binding 2 |
| chr16_+_17389116 | 1.97 |
ENSDART00000103750
ENSDART00000173448 |
fam131bb
|
family with sequence similarity 131, member Bb |
| chr20_-_45060241 | 1.93 |
ENSDART00000185227
|
klhl29
|
kelch-like family member 29 |
| chr20_-_32270866 | 1.92 |
ENSDART00000153140
|
armc2
|
armadillo repeat containing 2 |
| chr8_-_46894362 | 1.89 |
ENSDART00000111124
|
acot7
|
acyl-CoA thioesterase 7 |
| chr7_-_25895189 | 1.87 |
ENSDART00000173599
ENSDART00000079235 ENSDART00000173786 ENSDART00000173602 ENSDART00000079245 ENSDART00000187568 ENSDART00000173505 |
cd99l2
|
CD99 molecule-like 2 |
| chr16_+_13818500 | 1.87 |
ENSDART00000135245
|
flcn
|
folliculin |
| chr20_+_19238382 | 1.84 |
ENSDART00000136757
|
fndc4a
|
fibronectin type III domain containing 4a |
| chr11_-_37509001 | 1.83 |
ENSDART00000109753
|
bsnb
|
bassoon (presynaptic cytomatrix protein) b |
| chr18_-_1185772 | 1.80 |
ENSDART00000143245
|
nptnb
|
neuroplastin b |
| chr1_-_50859053 | 1.76 |
ENSDART00000132779
ENSDART00000137648 |
si:dkeyp-123h10.2
|
si:dkeyp-123h10.2 |
| chr2_+_50608099 | 1.73 |
ENSDART00000185805
ENSDART00000111135 |
neurod6b
|
neuronal differentiation 6b |
| chr19_-_5103141 | 1.73 |
ENSDART00000150952
|
tpi1a
|
triosephosphate isomerase 1a |
| chr11_+_30057762 | 1.71 |
ENSDART00000164139
|
nhsb
|
Nance-Horan syndrome b (congenital cataracts and dental anomalies) |
| chr17_-_12389259 | 1.71 |
ENSDART00000185724
|
snap25b
|
synaptosomal-associated protein, 25b |
| chr10_+_29698467 | 1.70 |
ENSDART00000163402
|
dlg2
|
discs, large homolog 2 (Drosophila) |
| chr22_+_16308450 | 1.68 |
ENSDART00000105678
|
lrrc39
|
leucine rich repeat containing 39 |
| chr25_+_11456696 | 1.66 |
ENSDART00000171408
|
AGBL1
|
si:ch73-141f14.1 |
| chr22_+_16308806 | 1.64 |
ENSDART00000162685
|
lrrc39
|
leucine rich repeat containing 39 |
| chr21_+_28478663 | 1.63 |
ENSDART00000077887
ENSDART00000134150 |
slc22a6l
|
solute carrier family 22 (organic anion transporter), member 6, like |
| chr17_-_16965809 | 1.59 |
ENSDART00000153697
|
nrxn3a
|
neurexin 3a |
| chr14_-_30587814 | 1.57 |
ENSDART00000144912
ENSDART00000149714 |
tmem265
|
transmembrane protein 265 |
| chr5_+_19314574 | 1.52 |
ENSDART00000133247
|
rusc2
|
RUN and SH3 domain containing 2 |
| chrM_+_9735 | 1.49 |
ENSDART00000093613
|
mt-co3
|
cytochrome c oxidase III, mitochondrial |
| chr3_+_34919810 | 1.48 |
ENSDART00000055264
|
ca10b
|
carbonic anhydrase Xb |
| chr20_+_30445971 | 1.48 |
ENSDART00000153150
|
myt1la
|
myelin transcription factor 1-like, a |
| chr12_-_33579873 | 1.47 |
ENSDART00000184661
|
tdrkh
|
tudor and KH domain containing |
| chr16_+_13818743 | 1.46 |
ENSDART00000090191
|
flcn
|
folliculin |
| chr15_+_15856178 | 1.44 |
ENSDART00000080338
|
dusp14
|
dual specificity phosphatase 14 |
| chr23_+_20563779 | 1.42 |
ENSDART00000146008
|
camkvl
|
CaM kinase-like vesicle-associated, like |
| chr20_-_46362606 | 1.41 |
ENSDART00000153087
|
bmf2
|
BCL2 modifying factor 2 |
| chr22_+_5176255 | 1.40 |
ENSDART00000092647
|
cers1
|
ceramide synthase 1 |
| chr6_-_6487876 | 1.40 |
ENSDART00000137642
|
cep170ab
|
centrosomal protein 170Ab |
| chr13_-_44630111 | 1.39 |
ENSDART00000110092
|
mdga1
|
MAM domain containing glycosylphosphatidylinositol anchor 1 |
| chr17_+_21295132 | 1.35 |
ENSDART00000103845
|
eno4
|
enolase family member 4 |
| chr21_-_42100471 | 1.34 |
ENSDART00000166148
|
gabra1
|
gamma-aminobutyric acid (GABA) A receptor, alpha 1 |
| chr16_-_47483142 | 1.32 |
ENSDART00000147072
|
cthrc1b
|
collagen triple helix repeat containing 1b |
| chr1_+_45323400 | 1.32 |
ENSDART00000148906
ENSDART00000132366 |
emp1
|
epithelial membrane protein 1 |
| chr3_-_34296361 | 1.28 |
ENSDART00000181485
ENSDART00000113726 |
tnrc6c1
|
trinucleotide repeat containing 6C1 |
| chr21_+_29077509 | 1.26 |
ENSDART00000128561
|
ecscr
|
endothelial cell surface expressed chemotaxis and apoptosis regulator |
| chr1_+_11977426 | 1.26 |
ENSDART00000103399
|
tspan5b
|
tetraspanin 5b |
| chr20_-_10120442 | 1.26 |
ENSDART00000144970
|
meis2b
|
Meis homeobox 2b |
| chr15_+_42397125 | 1.25 |
ENSDART00000169751
|
tiam1b
|
T cell lymphoma invasion and metastasis 1b |
| chr15_-_20024205 | 1.24 |
ENSDART00000161379
|
auts2b
|
autism susceptibility candidate 2b |
| chr1_-_44701313 | 1.21 |
ENSDART00000193926
|
si:dkey-28b4.8
|
si:dkey-28b4.8 |
| chr8_-_19051906 | 1.21 |
ENSDART00000089024
|
sema6bb
|
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6Bb |
| chr22_-_7129631 | 1.21 |
ENSDART00000171359
|
asic1b
|
acid-sensing (proton-gated) ion channel 1b |
| chr11_-_8126223 | 1.19 |
ENSDART00000091617
ENSDART00000192391 ENSDART00000101561 |
ttll7
|
tubulin tyrosine ligase-like family, member 7 |
| chr15_+_36309070 | 1.19 |
ENSDART00000157034
|
gmnc
|
geminin coiled-coil domain containing |
| chr15_+_31344472 | 1.18 |
ENSDART00000146695
ENSDART00000159182 ENSDART00000060125 |
or107-1
|
odorant receptor, family D, subfamily 107, member 1 |
| chr5_-_23117078 | 1.14 |
ENSDART00000051529
|
uprt
|
uracil phosphoribosyltransferase (FUR1) homolog (S. cerevisiae) |
| chr6_+_23887314 | 1.14 |
ENSDART00000163188
|
znf648
|
zinc finger protein 648 |
| chr18_+_19006063 | 1.11 |
ENSDART00000135729
|
slc24a1
|
solute carrier family 24 (sodium/potassium/calcium exchanger), member 1 |
| chr10_+_3049636 | 1.07 |
ENSDART00000081794
ENSDART00000183167 ENSDART00000191634 ENSDART00000183514 |
rasgrf2a
|
Ras protein-specific guanine nucleotide-releasing factor 2a |
| chr3_+_33341640 | 1.05 |
ENSDART00000186352
|
pyya
|
peptide YYa |
| chr21_+_28958471 | 1.03 |
ENSDART00000144331
ENSDART00000005929 |
ppp3ca
|
protein phosphatase 3, catalytic subunit, alpha isozyme |
| chr7_-_28148310 | 1.03 |
ENSDART00000044208
|
lmo1
|
LIM domain only 1 |
| chr24_+_9412450 | 1.03 |
ENSDART00000132724
|
si:ch211-285f17.1
|
si:ch211-285f17.1 |
| chr22_+_5176693 | 1.02 |
ENSDART00000160927
|
cers1
|
ceramide synthase 1 |
| chr14_+_34971554 | 1.00 |
ENSDART00000184271
|
rnf145a
|
ring finger protein 145a |
| chr3_-_46817499 | 0.98 |
ENSDART00000013717
|
elavl3
|
ELAV like neuron-specific RNA binding protein 3 |
| chr14_+_36220479 | 0.94 |
ENSDART00000148319
|
pitx2
|
paired-like homeodomain 2 |
| chr17_-_44247707 | 0.93 |
ENSDART00000126097
|
otx2b
|
orthodenticle homeobox 2b |
| chr18_+_9362455 | 0.93 |
ENSDART00000187025
|
sema3ab
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3Ab |
| chr17_+_24687338 | 0.91 |
ENSDART00000135794
|
selenon
|
selenoprotein N |
| chr9_+_17983463 | 0.91 |
ENSDART00000182150
|
akap11
|
A kinase (PRKA) anchor protein 11 |
| chr13_+_25449681 | 0.88 |
ENSDART00000101328
|
atoh7
|
atonal bHLH transcription factor 7 |
| chr21_-_10488379 | 0.86 |
ENSDART00000163878
|
hcn1
|
hyperpolarization activated cyclic nucleotide-gated potassium channel 1 |
| chr20_-_27711970 | 0.85 |
ENSDART00000139637
|
zbtb25
|
zinc finger and BTB domain containing 25 |
| chr5_+_20147830 | 0.84 |
ENSDART00000098727
|
svopa
|
SV2 related protein a |
| chr19_-_19556778 | 0.84 |
ENSDART00000164060
|
tax1bp1a
|
Tax1 (human T-cell leukemia virus type I) binding protein 1a |
| chr13_+_38990939 | 0.84 |
ENSDART00000145979
|
col19a1
|
collagen, type XIX, alpha 1 |
| chr16_-_30434279 | 0.83 |
ENSDART00000018504
|
zgc:77086
|
zgc:77086 |
| chr10_+_42690374 | 0.81 |
ENSDART00000123496
|
rhobtb2b
|
Rho-related BTB domain containing 2b |
| chr5_-_50781623 | 0.79 |
ENSDART00000114950
|
zgc:194908
|
zgc:194908 |
| chr23_+_38957472 | 0.79 |
ENSDART00000193836
|
ATP9A
|
ATPase phospholipid transporting 9A (putative) |
| chr16_+_20910186 | 0.78 |
ENSDART00000046766
|
hoxa10b
|
homeobox A10b |
| chr21_-_11654422 | 0.78 |
ENSDART00000081614
ENSDART00000132699 |
cast
|
calpastatin |
| chr11_+_38280454 | 0.77 |
ENSDART00000171496
|
CDK18
|
si:dkey-166c18.1 |
| chr21_-_17296789 | 0.77 |
ENSDART00000192180
|
gfi1b
|
growth factor independent 1B transcription repressor |
| chr13_+_21768447 | 0.74 |
ENSDART00000100941
|
chchd1
|
coiled-coil-helix-coiled-coil-helix domain containing 1 |
| chr10_-_35236949 | 0.74 |
ENSDART00000145804
|
ypel2a
|
yippee-like 2a |
| chr19_-_205104 | 0.73 |
ENSDART00000011890
|
zbtb22a
|
zinc finger and BTB domain containing 22a |
| chr6_-_44044385 | 0.73 |
ENSDART00000075497
|
rybpb
|
RING1 and YY1 binding protein b |
| chr4_+_12612723 | 0.73 |
ENSDART00000133767
|
lmo3
|
LIM domain only 3 |
| chr14_-_17622080 | 0.72 |
ENSDART00000112149
|
si:ch211-159i8.4
|
si:ch211-159i8.4 |
| chr10_+_41765944 | 0.71 |
ENSDART00000171484
|
rnf34b
|
ring finger protein 34b |
| chr6_-_46782176 | 0.69 |
ENSDART00000185559
|
igfn1.4
|
immunoglobulin-like and fibronectin type III domain containing 1, tandem duplicate 4 |
| chr7_+_42460302 | 0.69 |
ENSDART00000004120
|
adamts18
|
ADAM metallopeptidase with thrombospondin type 1 motif, 18 |
| chr12_-_8504278 | 0.69 |
ENSDART00000135865
|
egr2b
|
early growth response 2b |
| chr6_+_52880292 | 0.68 |
ENSDART00000077848
|
or137-4
|
odorant receptor, family H, subfamily 137, member 4 |
| chr17_-_19963718 | 0.67 |
ENSDART00000154251
|
chrm3a
|
cholinergic receptor, muscarinic 3a |
| chr18_+_34478959 | 0.67 |
ENSDART00000059394
|
kcnab1a
|
potassium voltage-gated channel, shaker-related subfamily, beta member 1 a |
| chr2_-_15324837 | 0.66 |
ENSDART00000015655
|
tecrl2b
|
trans-2,3-enoyl-CoA reductase-like 2b |
| chr10_+_43039947 | 0.66 |
ENSDART00000193434
|
atg10
|
ATG10 autophagy related 10 homolog (S. cerevisiae) |
| chr21_-_3853204 | 0.66 |
ENSDART00000188829
|
st6galnac4
|
ST6 (alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3)-N-acetylgalactosaminide alpha-2,6-sialyltransferase 4 |
| chr25_+_26798673 | 0.64 |
ENSDART00000157235
|
ca12
|
carbonic anhydrase XII |
| chr1_-_22512063 | 0.63 |
ENSDART00000031546
ENSDART00000190987 |
chrna6
|
cholinergic receptor, nicotinic, alpha 6 |
| chr22_-_20166660 | 0.62 |
ENSDART00000085913
ENSDART00000188241 |
btbd2a
|
BTB (POZ) domain containing 2a |
| chr11_+_35050253 | 0.62 |
ENSDART00000124800
|
fam212aa
|
family with sequence similarity 212, member Aa |
| chr21_-_5856050 | 0.61 |
ENSDART00000115367
|
CABZ01071020.1
|
|
| chr5_-_28109416 | 0.60 |
ENSDART00000042002
|
si:ch211-48m9.1
|
si:ch211-48m9.1 |
| chr3_+_33440615 | 0.60 |
ENSDART00000146005
|
gtpbp1
|
GTP binding protein 1 |
| chr16_+_54263921 | 0.59 |
ENSDART00000002856
|
drd2l
|
dopamine receptor D2 like |
| chr23_-_31974060 | 0.54 |
ENSDART00000168087
|
BX927210.1
|
|
| chr8_-_19467011 | 0.54 |
ENSDART00000162010
|
zgc:92140
|
zgc:92140 |
| chr5_-_30481263 | 0.53 |
ENSDART00000086734
|
phldb1a
|
pleckstrin homology-like domain, family B, member 1a |
| chr4_-_15420452 | 0.53 |
ENSDART00000016230
|
plxna4
|
plexin A4 |
| chr9_-_20372977 | 0.52 |
ENSDART00000113418
|
igsf3
|
immunoglobulin superfamily, member 3 |
| chr14_-_41075262 | 0.51 |
ENSDART00000180518
|
drp2
|
dystrophin related protein 2 |
| chr2_+_4146299 | 0.51 |
ENSDART00000173418
|
mib1
|
mindbomb E3 ubiquitin protein ligase 1 |
| chr10_+_41765616 | 0.51 |
ENSDART00000170682
|
rnf34b
|
ring finger protein 34b |
| chr1_+_34696503 | 0.49 |
ENSDART00000186106
|
CR339054.2
|
|
| chr10_+_36178713 | 0.48 |
ENSDART00000140816
|
or108-3
|
odorant receptor, family D, subfamily 108, member 3 |
| chr1_+_52929185 | 0.47 |
ENSDART00000147683
|
inpp4b
|
inositol polyphosphate-4-phosphatase type II B |
| chr20_-_38617766 | 0.47 |
ENSDART00000050474
|
slc30a2
|
solute carrier family 30 (zinc transporter), member 2 |
| chr15_+_40792667 | 0.47 |
ENSDART00000193186
ENSDART00000184559 |
fat3a
|
FAT atypical cadherin 3a |
| chr7_-_12464412 | 0.45 |
ENSDART00000178723
|
adamtsl3
|
ADAMTS-like 3 |
| chr7_-_48263516 | 0.44 |
ENSDART00000006619
ENSDART00000142370 ENSDART00000148273 ENSDART00000147968 |
rbpms2b
|
RNA binding protein with multiple splicing 2b |
| chr13_+_38817871 | 0.44 |
ENSDART00000187708
|
col19a1
|
collagen, type XIX, alpha 1 |
| chr4_+_12612145 | 0.44 |
ENSDART00000181201
|
lmo3
|
LIM domain only 3 |
| chr17_-_31659670 | 0.43 |
ENSDART00000030448
|
vsx2
|
visual system homeobox 2 |
| chr23_+_5499307 | 0.42 |
ENSDART00000170361
|
tead3a
|
TEA domain family member 3 a |
| chr1_+_513986 | 0.42 |
ENSDART00000109083
ENSDART00000081945 |
txnl4b
|
thioredoxin-like 4B |
| chr6_-_29515709 | 0.42 |
ENSDART00000180205
|
pex5la
|
peroxisomal biogenesis factor 5-like a |
| chr2_+_21229909 | 0.41 |
ENSDART00000046127
|
vipr1a
|
vasoactive intestinal peptide receptor 1a |
| chr21_-_15046065 | 0.41 |
ENSDART00000178507
|
mmp17a
|
matrix metallopeptidase 17a |
| chr17_-_29119362 | 0.40 |
ENSDART00000104204
|
foxg1a
|
forkhead box G1a |
| chr19_+_2631565 | 0.40 |
ENSDART00000171487
|
fam126a
|
family with sequence similarity 126, member A |
| chr16_-_13623928 | 0.39 |
ENSDART00000164344
|
si:dkeyp-69b9.6
|
si:dkeyp-69b9.6 |
| chr20_+_4060839 | 0.39 |
ENSDART00000178565
|
TRIM67
|
tripartite motif containing 67 |
| chr14_+_30568961 | 0.38 |
ENSDART00000184303
|
mrpl11
|
mitochondrial ribosomal protein L11 |
| chr4_+_22343093 | 0.37 |
ENSDART00000023588
|
guca1a
|
guanylate cyclase activator 1A |
| chr13_+_2625150 | 0.37 |
ENSDART00000164177
|
plpp4
|
phospholipid phosphatase 4 |
| chr18_-_2433011 | 0.36 |
ENSDART00000181922
ENSDART00000193276 |
CR769778.1
|
|
| chr21_-_40676224 | 0.35 |
ENSDART00000162623
|
arxb
|
aristaless related homeobox b |
| chr7_-_44957503 | 0.34 |
ENSDART00000165077
|
cdh16
|
cadherin 16, KSP-cadherin |
| chr24_-_39518599 | 0.34 |
ENSDART00000145606
ENSDART00000031486 |
lyrm1
|
LYR motif containing 1 |
| chr15_-_2184638 | 0.34 |
ENSDART00000135460
|
shox2
|
short stature homeobox 2 |
| chr8_+_50953776 | 0.34 |
ENSDART00000013870
|
zgc:56596
|
zgc:56596 |
| chr9_+_20557793 | 0.33 |
ENSDART00000178434
|
man1a2
|
mannosidase, alpha, class 1A, member 2 |
| chr5_+_38837429 | 0.33 |
ENSDART00000160236
|
fras1
|
Fraser extracellular matrix complex subunit 1 |
| chr19_-_38830582 | 0.32 |
ENSDART00000189966
ENSDART00000183055 |
adgrb2
|
adhesion G protein-coupled receptor B2 |
| chr1_-_54765262 | 0.32 |
ENSDART00000150362
|
si:ch211-197k17.3
|
si:ch211-197k17.3 |
| chr21_-_11655010 | 0.31 |
ENSDART00000144370
ENSDART00000139814 ENSDART00000139289 |
cast
|
calpastatin |
| chr19_+_22062202 | 0.31 |
ENSDART00000100181
|
sall3b
|
spalt-like transcription factor 3b |
| chr10_+_16584382 | 0.30 |
ENSDART00000112039
|
CR790388.1
|
|
| chr16_+_35661771 | 0.30 |
ENSDART00000161393
|
map7d1a
|
MAP7 domain containing 1a |
| chr13_+_22295905 | 0.30 |
ENSDART00000180133
ENSDART00000181125 |
usp54a
|
ubiquitin specific peptidase 54a |
| chr6_+_36807861 | 0.29 |
ENSDART00000161708
|
si:ch73-29l19.1
|
si:ch73-29l19.1 |
| chr23_-_17484555 | 0.29 |
ENSDART00000187181
|
dnajc5ab
|
DnaJ (Hsp40) homolog, subfamily C, member 5ab |
Network of associatons between targets according to the STRING database.
First level regulatory network of dlx2a+dlx2b_dlx4a+dlx4b+dlx6a
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.7 | 5.2 | GO:0035902 | response to immobilization stress(GO:0035902) |
| 1.0 | 2.9 | GO:0045601 | regulation of endothelial cell differentiation(GO:0045601) |
| 0.7 | 7.0 | GO:1904071 | presynaptic active zone assembly(GO:1904071) |
| 0.7 | 3.3 | GO:0016322 | neuron remodeling(GO:0016322) |
| 0.5 | 3.3 | GO:0030511 | positive regulation of transforming growth factor beta receptor signaling pathway(GO:0030511) positive regulation of cellular response to transforming growth factor beta stimulus(GO:1903846) |
| 0.4 | 1.7 | GO:0046166 | methylglyoxal biosynthetic process(GO:0019242) glyceraldehyde-3-phosphate biosynthetic process(GO:0046166) |
| 0.4 | 2.1 | GO:0007624 | ultradian rhythm(GO:0007624) |
| 0.4 | 1.1 | GO:0060292 | long term synaptic depression(GO:0060292) |
| 0.4 | 6.6 | GO:0036444 | calcium ion transmembrane import into mitochondrion(GO:0036444) |
| 0.3 | 4.3 | GO:0046085 | adenosine metabolic process(GO:0046085) |
| 0.3 | 1.9 | GO:0007288 | sperm axoneme assembly(GO:0007288) |
| 0.3 | 7.5 | GO:0051209 | release of sequestered calcium ion into cytosol(GO:0051209) |
| 0.3 | 1.3 | GO:0070208 | protein heterotrimerization(GO:0070208) |
| 0.3 | 13.2 | GO:0003009 | skeletal muscle contraction(GO:0003009) |
| 0.2 | 1.2 | GO:1903251 | multi-ciliated epithelial cell differentiation(GO:1903251) |
| 0.2 | 3.1 | GO:0001574 | ganglioside biosynthetic process(GO:0001574) |
| 0.2 | 0.7 | GO:0045987 | positive regulation of smooth muscle contraction(GO:0045987) |
| 0.2 | 1.3 | GO:0003209 | cardiac atrium morphogenesis(GO:0003209) |
| 0.2 | 0.6 | GO:0021519 | optic cup formation involved in camera-type eye development(GO:0003408) spinal cord association neuron specification(GO:0021519) |
| 0.2 | 0.9 | GO:0061072 | iris morphogenesis(GO:0061072) |
| 0.2 | 2.6 | GO:0051967 | negative regulation of cytosolic calcium ion concentration(GO:0051481) negative regulation of synaptic transmission, glutamatergic(GO:0051967) |
| 0.2 | 2.9 | GO:0006782 | protoporphyrinogen IX biosynthetic process(GO:0006782) protoporphyrinogen IX metabolic process(GO:0046501) |
| 0.2 | 0.5 | GO:0007414 | axonal defasciculation(GO:0007414) |
| 0.2 | 1.2 | GO:1900118 | negative regulation of execution phase of apoptosis(GO:1900118) regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001270) negative regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001271) |
| 0.2 | 1.2 | GO:0070207 | protein homotrimerization(GO:0070207) |
| 0.2 | 1.3 | GO:0032889 | regulation of vacuole fusion, non-autophagic(GO:0032889) |
| 0.2 | 0.7 | GO:1901380 | negative regulation of potassium ion transmembrane transporter activity(GO:1901017) negative regulation of potassium ion transmembrane transport(GO:1901380) negative regulation of delayed rectifier potassium channel activity(GO:1902260) negative regulation of voltage-gated potassium channel activity(GO:1903817) |
| 0.2 | 1.1 | GO:0097341 | inhibition of cysteine-type endopeptidase activity(GO:0097340) zymogen inhibition(GO:0097341) |
| 0.2 | 4.5 | GO:0070593 | dendrite self-avoidance(GO:0070593) |
| 0.2 | 0.9 | GO:0048755 | branching morphogenesis of a nerve(GO:0048755) |
| 0.2 | 8.0 | GO:0006096 | glycolytic process(GO:0006096) |
| 0.1 | 4.7 | GO:0032482 | Rab protein signal transduction(GO:0032482) |
| 0.1 | 4.4 | GO:0016082 | synaptic vesicle priming(GO:0016082) |
| 0.1 | 2.6 | GO:0030817 | regulation of cAMP metabolic process(GO:0030814) regulation of cAMP biosynthetic process(GO:0030817) regulation of adenylate cyclase activity(GO:0045761) |
| 0.1 | 1.3 | GO:0035479 | angioblast cell migration from lateral mesoderm to midline(GO:0035479) |
| 0.1 | 2.1 | GO:0098943 | neurotransmitter receptor transport, postsynaptic endosome to lysosome(GO:0098943) |
| 0.1 | 1.5 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.1 | 3.3 | GO:0055008 | cardiac muscle tissue morphogenesis(GO:0055008) |
| 0.1 | 2.6 | GO:0015985 | energy coupled proton transport, down electrochemical gradient(GO:0015985) ATP synthesis coupled proton transport(GO:0015986) |
| 0.1 | 0.5 | GO:0061088 | sequestering of zinc ion(GO:0032119) regulation of sequestering of zinc ion(GO:0061088) |
| 0.1 | 3.6 | GO:0007586 | digestion(GO:0007586) |
| 0.1 | 5.1 | GO:0046856 | phosphatidylinositol dephosphorylation(GO:0046856) |
| 0.1 | 0.4 | GO:0051145 | smooth muscle cell differentiation(GO:0051145) negative regulation of muscle cell differentiation(GO:0051148) |
| 0.1 | 1.9 | GO:0042759 | long-chain fatty acid biosynthetic process(GO:0042759) |
| 0.1 | 1.3 | GO:0035278 | miRNA mediated inhibition of translation(GO:0035278) negative regulation of translation, ncRNA-mediated(GO:0040033) regulation of translation, ncRNA-mediated(GO:0045974) |
| 0.1 | 0.9 | GO:0035677 | posterior lateral line neuromast hair cell development(GO:0035677) |
| 0.1 | 1.7 | GO:0010923 | negative regulation of phosphatase activity(GO:0010923) |
| 0.1 | 1.0 | GO:0048016 | calcineurin-NFAT signaling cascade(GO:0033173) inositol phosphate-mediated signaling(GO:0048016) |
| 0.1 | 1.2 | GO:0018095 | protein polyglutamylation(GO:0018095) |
| 0.1 | 0.9 | GO:0003406 | retinal pigment epithelium development(GO:0003406) |
| 0.1 | 1.2 | GO:0010507 | negative regulation of autophagy(GO:0010507) |
| 0.1 | 2.0 | GO:0014866 | skeletal myofibril assembly(GO:0014866) |
| 0.1 | 2.5 | GO:0032012 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.1 | 0.4 | GO:0043584 | nose development(GO:0043584) |
| 0.0 | 1.1 | GO:0007631 | feeding behavior(GO:0007631) |
| 0.0 | 2.4 | GO:0046513 | ceramide biosynthetic process(GO:0046513) |
| 0.0 | 0.7 | GO:0048931 | myelination of lateral line nerve axons(GO:0048897) posterior lateral line nerve glial cell differentiation(GO:0048931) myelination of posterior lateral line nerve axons(GO:0048932) lateral line nerve glial cell morphogenesis involved in differentiation(GO:0048938) posterior lateral line nerve glial cell development(GO:0048941) posterior lateral line nerve glial cell morphogenesis involved in differentiation(GO:0048942) |
| 0.0 | 0.4 | GO:0016560 | protein import into peroxisome matrix, docking(GO:0016560) protein to membrane docking(GO:0022615) |
| 0.0 | 1.8 | GO:0036269 | swimming behavior(GO:0036269) |
| 0.0 | 0.8 | GO:0043652 | engulfment of apoptotic cell(GO:0043652) |
| 0.0 | 0.3 | GO:0071816 | tail-anchored membrane protein insertion into ER membrane(GO:0071816) |
| 0.0 | 2.5 | GO:0048846 | axon extension involved in axon guidance(GO:0048846) neuron projection extension involved in neuron projection guidance(GO:1902284) |
| 0.0 | 6.6 | GO:0030334 | regulation of cell migration(GO:0030334) |
| 0.0 | 1.1 | GO:0050772 | positive regulation of axonogenesis(GO:0050772) |
| 0.0 | 4.1 | GO:0002040 | sprouting angiogenesis(GO:0002040) |
| 0.0 | 0.7 | GO:0042761 | very long-chain fatty acid biosynthetic process(GO:0042761) |
| 0.0 | 0.3 | GO:0010669 | epithelial structure maintenance(GO:0010669) |
| 0.0 | 0.7 | GO:0032543 | mitochondrial translation(GO:0032543) |
| 0.0 | 0.3 | GO:0098887 | neurotransmitter receptor transport, endosome to postsynaptic membrane(GO:0098887) endosome to plasma membrane protein transport(GO:0099638) neurotransmitter receptor transport, endosome to plasma membrane(GO:0099639) |
| 0.0 | 2.6 | GO:0006979 | response to oxidative stress(GO:0006979) |
| 0.0 | 0.4 | GO:0035329 | hippo signaling(GO:0035329) |
| 0.0 | 0.1 | GO:1990511 | piRNA biosynthetic process(GO:1990511) |
| 0.0 | 0.1 | GO:0036149 | phosphatidylinositol acyl-chain remodeling(GO:0036149) |
| 0.0 | 1.7 | GO:0002088 | lens development in camera-type eye(GO:0002088) |
| 0.0 | 0.3 | GO:0016525 | negative regulation of angiogenesis(GO:0016525) |
| 0.0 | 0.5 | GO:0060536 | cartilage morphogenesis(GO:0060536) |
| 0.0 | 0.1 | GO:0021794 | thalamus development(GO:0021794) |
| 0.0 | 0.6 | GO:0006730 | one-carbon metabolic process(GO:0006730) |
| 0.0 | 0.4 | GO:0044243 | collagen catabolic process(GO:0030574) multicellular organism catabolic process(GO:0044243) |
| 0.0 | 0.1 | GO:0045670 | regulation of osteoclast differentiation(GO:0045670) |
| 0.0 | 0.4 | GO:0046839 | phospholipid dephosphorylation(GO:0046839) |
| 0.0 | 0.2 | GO:0036353 | histone H2A-K119 monoubiquitination(GO:0036353) |
| 0.0 | 2.4 | GO:0001525 | angiogenesis(GO:0001525) |
| 0.0 | 2.6 | GO:0060537 | muscle tissue development(GO:0060537) |
| 0.0 | 1.6 | GO:0030155 | regulation of cell adhesion(GO:0030155) |
| 0.0 | 1.5 | GO:0070588 | calcium ion transmembrane transport(GO:0070588) |
| 0.0 | 0.6 | GO:0060216 | definitive hemopoiesis(GO:0060216) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 6.6 | GO:0005790 | smooth endoplasmic reticulum(GO:0005790) |
| 0.7 | 4.4 | GO:0070032 | synaptobrevin 2-SNAP-25-syntaxin-1a-complexin I complex(GO:0070032) |
| 0.7 | 7.0 | GO:0098982 | GABA-ergic synapse(GO:0098982) |
| 0.6 | 3.3 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.5 | 6.6 | GO:1990246 | uniplex complex(GO:1990246) |
| 0.5 | 4.7 | GO:0032593 | insulin-responsive compartment(GO:0032593) |
| 0.3 | 13.2 | GO:0005861 | troponin complex(GO:0005861) |
| 0.2 | 1.0 | GO:0005955 | calcineurin complex(GO:0005955) |
| 0.2 | 1.3 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.2 | 3.3 | GO:0031430 | M band(GO:0031430) |
| 0.1 | 2.6 | GO:0045263 | proton-transporting ATP synthase complex, coupling factor F(o)(GO:0045263) |
| 0.1 | 2.6 | GO:0098978 | glutamatergic synapse(GO:0098978) |
| 0.1 | 1.5 | GO:0045277 | respiratory chain complex IV(GO:0045277) |
| 0.1 | 1.5 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.1 | 3.8 | GO:0098839 | postsynaptic density membrane(GO:0098839) postsynaptic specialization membrane(GO:0099634) |
| 0.1 | 0.7 | GO:0044224 | juxtaparanode region of axon(GO:0044224) |
| 0.0 | 0.3 | GO:0071818 | BAT3 complex(GO:0071818) |
| 0.0 | 2.6 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.0 | 1.3 | GO:1902711 | GABA receptor complex(GO:1902710) GABA-A receptor complex(GO:1902711) |
| 0.0 | 2.4 | GO:0099501 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 0.0 | 8.1 | GO:0030424 | axon(GO:0030424) |
| 0.0 | 2.1 | GO:0043197 | dendritic spine(GO:0043197) neuron spine(GO:0044309) |
| 0.0 | 0.5 | GO:0002116 | semaphorin receptor complex(GO:0002116) |
| 0.0 | 3.3 | GO:0098852 | lysosomal membrane(GO:0005765) lytic vacuole membrane(GO:0098852) |
| 0.0 | 0.4 | GO:0005682 | U5 snRNP(GO:0005682) |
| 0.0 | 3.8 | GO:0005759 | mitochondrial matrix(GO:0005759) |
| 0.0 | 0.5 | GO:0045180 | basal cortex(GO:0045180) |
| 0.0 | 1.8 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.0 | 1.4 | GO:0016459 | myosin complex(GO:0016459) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.3 | 6.6 | GO:0005219 | ryanodine-sensitive calcium-release channel activity(GO:0005219) calcium-induced calcium release activity(GO:0048763) |
| 1.0 | 5.8 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 1.0 | 2.9 | GO:0016748 | succinyltransferase activity(GO:0016748) |
| 0.9 | 2.6 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 0.5 | 2.1 | GO:0004365 | glyceraldehyde-3-phosphate dehydrogenase (NAD+) (phosphorylating) activity(GO:0004365) glyceraldehyde-3-phosphate dehydrogenase (NAD(P)+) (phosphorylating) activity(GO:0043891) |
| 0.5 | 3.6 | GO:0016316 | phosphatidylinositol-3,4-bisphosphate 4-phosphatase activity(GO:0016316) |
| 0.5 | 5.0 | GO:0052629 | phosphatidylinositol-3,5-bisphosphate 3-phosphatase activity(GO:0052629) |
| 0.4 | 1.7 | GO:0008929 | triose-phosphate isomerase activity(GO:0004807) methylglyoxal synthase activity(GO:0008929) |
| 0.4 | 7.0 | GO:0098882 | structural constituent of presynaptic active zone(GO:0098882) |
| 0.3 | 1.9 | GO:0036042 | long-chain fatty acyl-CoA binding(GO:0036042) |
| 0.2 | 1.2 | GO:0044736 | acid-sensing ion channel activity(GO:0044736) |
| 0.2 | 0.7 | GO:0019777 | Atg12 transferase activity(GO:0019777) |
| 0.2 | 1.1 | GO:0031841 | neuropeptide Y receptor binding(GO:0031841) type 2 neuropeptide Y receptor binding(GO:0031843) |
| 0.2 | 4.4 | GO:0008253 | 5'-nucleotidase activity(GO:0008253) |
| 0.2 | 2.6 | GO:0001591 | dopamine neurotransmitter receptor activity, coupled via Gi/Go(GO:0001591) |
| 0.2 | 1.0 | GO:0004723 | calcium-dependent protein serine/threonine phosphatase activity(GO:0004723) calmodulin-dependent protein phosphatase activity(GO:0033192) |
| 0.2 | 6.8 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.2 | 1.3 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.2 | 2.4 | GO:0050291 | sphingosine N-acyltransferase activity(GO:0050291) |
| 0.2 | 1.1 | GO:0010859 | calcium-dependent cysteine-type endopeptidase inhibitor activity(GO:0010859) |
| 0.1 | 3.4 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.1 | 4.5 | GO:0098632 | protein binding involved in cell-cell adhesion(GO:0098632) |
| 0.1 | 1.1 | GO:0008273 | calcium, potassium:sodium antiporter activity(GO:0008273) |
| 0.1 | 3.6 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.1 | 2.6 | GO:0008378 | galactosyltransferase activity(GO:0008378) |
| 0.1 | 1.3 | GO:0008503 | benzodiazepine receptor activity(GO:0008503) GABA-gated chloride ion channel activity(GO:0022851) |
| 0.1 | 1.5 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.1 | 1.2 | GO:0070739 | protein-glutamic acid ligase activity(GO:0070739) tubulin-glutamic acid ligase activity(GO:0070740) |
| 0.1 | 0.7 | GO:0001665 | alpha-N-acetylgalactosaminide alpha-2,6-sialyltransferase activity(GO:0001665) |
| 0.1 | 2.1 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.1 | 8.2 | GO:0005179 | hormone activity(GO:0005179) |
| 0.1 | 1.4 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) MAP kinase phosphatase activity(GO:0033549) |
| 0.1 | 0.4 | GO:0004999 | vasoactive intestinal polypeptide receptor activity(GO:0004999) |
| 0.1 | 0.4 | GO:0070180 | large ribosomal subunit rRNA binding(GO:0070180) |
| 0.1 | 0.9 | GO:0038191 | neuropilin binding(GO:0038191) |
| 0.1 | 0.7 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 0.1 | 2.4 | GO:0008376 | acetylgalactosaminyltransferase activity(GO:0008376) |
| 0.1 | 1.5 | GO:0016675 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors(GO:0016675) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.1 | 0.4 | GO:0005052 | peroxisome matrix targeting signal-1 binding(GO:0005052) |
| 0.0 | 0.9 | GO:0030552 | cAMP binding(GO:0030552) |
| 0.0 | 0.4 | GO:0008048 | calcium sensitive guanylate cyclase activator activity(GO:0008048) |
| 0.0 | 3.4 | GO:0004722 | protein serine/threonine phosphatase activity(GO:0004722) |
| 0.0 | 2.0 | GO:0017124 | SH3 domain binding(GO:0017124) |
| 0.0 | 2.6 | GO:0015078 | hydrogen ion transmembrane transporter activity(GO:0015078) |
| 0.0 | 0.7 | GO:0004033 | aldo-keto reductase (NADP) activity(GO:0004033) |
| 0.0 | 1.4 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.0 | 0.9 | GO:0051018 | protein kinase A binding(GO:0051018) |
| 0.0 | 1.2 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.0 | 0.6 | GO:0022848 | acetylcholine-gated cation channel activity(GO:0022848) |
| 0.0 | 2.1 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.0 | 0.5 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.0 | 0.5 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 0.6 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.0 | 5.5 | GO:0005085 | guanyl-nucleotide exchange factor activity(GO:0005085) |
| 0.0 | 0.1 | GO:0004366 | glycerol-3-phosphate O-acyltransferase activity(GO:0004366) |
| 0.0 | 0.1 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.0 | 0.4 | GO:0008195 | phosphatidate phosphatase activity(GO:0008195) |
| 0.0 | 0.3 | GO:0030159 | receptor signaling complex scaffold activity(GO:0030159) |
| 0.0 | 1.5 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 0.1 | GO:0031078 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.0 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.1 | 1.5 | PID RHODOPSIN PATHWAY | Visual signal transduction: Rods |
| 0.0 | 6.2 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 2.1 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.0 | 1.3 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.0 | 0.9 | SIG CHEMOTAXIS | Genes related to chemotaxis |
| 0.0 | 0.6 | PID NOTCH PATHWAY | Notch signaling pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.2 | 2.9 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.1 | 2.6 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.1 | 2.1 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.1 | 1.3 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.1 | 1.0 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.1 | 0.6 | REACTOME PRESYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Presynaptic nicotinic acetylcholine receptors |
| 0.1 | 2.6 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 0.1 | 0.6 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.0 | 0.5 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.0 | 0.5 | REACTOME SYNTHESIS OF PIPS AT THE EARLY ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the early endosome membrane |
| 0.0 | 1.3 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 0.6 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.0 | 0.1 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 0.0 | 0.9 | REACTOME POTASSIUM CHANNELS | Genes involved in Potassium Channels |
| 0.0 | 2.2 | REACTOME ANTIGEN PROCESSING UBIQUITINATION PROTEASOME DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |