Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
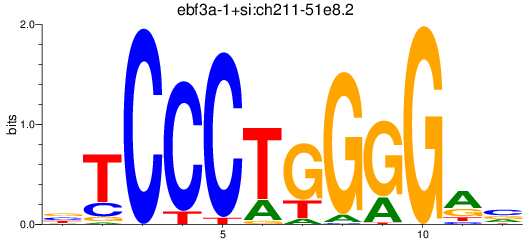
Results for ebf3a-1+si:ch211-51e8.2
Z-value: 0.73
Transcription factors associated with ebf3a-1+si:ch211-51e8.2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
si_ch211-51e8.2
|
ENSDARG00000092635 | si_ch211-51e8.2 |
|
ebf3a-1
|
ENSDARG00000099849 | EBF transcription factor 1a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| EBF1 (1 of many) | dr11_v1_chr21_-_33995213_33995213 | -0.28 | 6.5e-03 | Click! |
| ebf3a | dr11_v1_chr14_+_35023923_35023923 | 0.17 | 1.0e-01 | Click! |
Activity profile of ebf3a-1+si:ch211-51e8.2 motif
Sorted Z-values of ebf3a-1+si:ch211-51e8.2 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr22_+_11756040 | 15.44 |
ENSDART00000105808
|
krt97
|
keratin 97 |
| chr22_+_11775269 | 10.71 |
ENSDART00000140272
|
krt96
|
keratin 96 |
| chr9_-_22831836 | 8.60 |
ENSDART00000142585
|
neb
|
nebulin |
| chr7_+_66822229 | 8.09 |
ENSDART00000112109
|
lyve1a
|
lymphatic vessel endothelial hyaluronic receptor 1a |
| chr8_-_1051438 | 5.97 |
ENSDART00000067093
ENSDART00000170737 |
smyd1b
|
SET and MYND domain containing 1b |
| chr19_-_35596207 | 5.89 |
ENSDART00000136811
|
col8a2
|
collagen, type VIII, alpha 2 |
| chr10_-_25769334 | 5.64 |
ENSDART00000134176
|
postna
|
periostin, osteoblast specific factor a |
| chr10_+_158590 | 5.41 |
ENSDART00000081982
|
KCNJ15
|
potassium voltage-gated channel subfamily J member 15 |
| chr14_-_21959712 | 5.40 |
ENSDART00000021417
|
p2rx3a
|
purinergic receptor P2X, ligand-gated ion channel, 3a |
| chr12_-_46176115 | 4.88 |
ENSDART00000152848
|
si:ch211-226h7.8
|
si:ch211-226h7.8 |
| chr22_+_28446365 | 4.66 |
ENSDART00000189359
|
abi3bpb
|
ABI family, member 3 (NESH) binding protein b |
| chr2_-_37465517 | 4.40 |
ENSDART00000139983
|
si:dkey-57k2.6
|
si:dkey-57k2.6 |
| chr3_+_26081343 | 4.19 |
ENSDART00000134647
|
atp2a1
|
ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 1 |
| chr2_-_30987538 | 4.12 |
ENSDART00000148157
|
emilin2a
|
elastin microfibril interfacer 2a |
| chr20_-_29471577 | 4.12 |
ENSDART00000043382
|
grem1a
|
gremlin 1a, DAN family BMP antagonist |
| chr3_+_32416948 | 3.83 |
ENSDART00000157324
ENSDART00000154267 ENSDART00000186094 ENSDART00000155860 ENSDART00000156986 |
prrg2
|
proline rich Gla (G-carboxyglutamic acid) 2 |
| chr9_+_44431174 | 3.75 |
ENSDART00000149726
|
ppp1r1c
|
protein phosphatase 1, regulatory (inhibitor) subunit 1C |
| chr3_+_13559199 | 3.65 |
ENSDART00000166547
|
si:ch73-106n3.1
|
si:ch73-106n3.1 |
| chr9_-_33081781 | 3.64 |
ENSDART00000165748
|
zgc:172053
|
zgc:172053 |
| chr4_+_5193198 | 3.38 |
ENSDART00000067388
|
fgf23
|
fibroblast growth factor 23 |
| chr2_-_23172708 | 3.32 |
ENSDART00000041365
|
prrx1a
|
paired related homeobox 1a |
| chr15_+_20239141 | 3.29 |
ENSDART00000101152
ENSDART00000152473 |
spint2
|
serine peptidase inhibitor, Kunitz type, 2 |
| chr13_-_13751017 | 3.06 |
ENSDART00000180182
|
ky
|
kyphoscoliosis peptidase |
| chr19_+_5651325 | 2.89 |
ENSDART00000082067
|
ft1
|
alpha(1,3)fucosyltransferase gene 1 |
| chr2_+_5371492 | 2.85 |
ENSDART00000139762
|
si:ch1073-184j22.1
|
si:ch1073-184j22.1 |
| chr9_+_56443416 | 2.79 |
ENSDART00000167947
|
CABZ01079480.1
|
|
| chr4_-_20115570 | 2.73 |
ENSDART00000134528
|
si:dkey-159a18.1
|
si:dkey-159a18.1 |
| chr15_+_37559570 | 2.71 |
ENSDART00000085522
|
hspb6
|
heat shock protein, alpha-crystallin-related, b6 |
| chr7_+_39410180 | 2.66 |
ENSDART00000168641
|
CT030188.1
|
|
| chr10_-_35149513 | 2.65 |
ENSDART00000063434
ENSDART00000131291 |
ripk4
|
receptor-interacting serine-threonine kinase 4 |
| chr12_-_26406323 | 2.64 |
ENSDART00000131896
|
myoz1b
|
myozenin 1b |
| chr14_-_6727717 | 2.61 |
ENSDART00000166979
|
si:dkeyp-44a8.4
|
si:dkeyp-44a8.4 |
| chr19_-_47523793 | 2.57 |
ENSDART00000124234
|
grem1a
|
gremlin 1a, DAN family BMP antagonist |
| chr15_+_59417 | 2.57 |
ENSDART00000187260
|
AXL
|
AXL receptor tyrosine kinase |
| chr20_-_2725594 | 2.53 |
ENSDART00000152120
|
akirin2
|
akirin 2 |
| chr23_+_36601984 | 2.48 |
ENSDART00000128598
|
igfbp6b
|
insulin-like growth factor binding protein 6b |
| chr12_-_43693192 | 2.48 |
ENSDART00000172284
|
CU914622.1
|
|
| chr1_-_59411901 | 2.45 |
ENSDART00000167015
|
si:ch211-188p14.4
|
si:ch211-188p14.4 |
| chr19_+_19786117 | 2.45 |
ENSDART00000167757
ENSDART00000163546 |
hoxa1a
|
homeobox A1a |
| chr2_-_4868002 | 2.38 |
ENSDART00000189835
|
tfr1a
|
transferrin receptor 1a |
| chr13_+_31297016 | 2.33 |
ENSDART00000141786
ENSDART00000086143 |
antxr1d
|
anthrax toxin receptor 1d |
| chr5_-_5831037 | 2.31 |
ENSDART00000112856
|
zmp:0000000846
|
zmp:0000000846 |
| chr7_+_39410393 | 2.31 |
ENSDART00000158561
ENSDART00000185173 |
CT030188.1
|
|
| chr22_+_20710434 | 2.29 |
ENSDART00000135521
|
peak3
|
PEAK family member 3 |
| chr25_+_25453571 | 2.28 |
ENSDART00000112330
|
rassf7a
|
Ras association (RalGDS/AF-6) domain family (N-terminal) member 7a |
| chr24_+_3050020 | 2.15 |
ENSDART00000004241
|
inhbaa
|
inhibin, beta Aa |
| chr22_+_661711 | 2.13 |
ENSDART00000113795
|
elf3
|
E74-like factor 3 (ets domain transcription factor, epithelial-specific ) |
| chr19_+_41464870 | 2.12 |
ENSDART00000102778
|
dlx6a
|
distal-less homeobox 6a |
| chr19_+_19756425 | 2.07 |
ENSDART00000167606
|
hoxa3a
|
homeobox A3a |
| chr23_+_19790962 | 2.04 |
ENSDART00000142228
|
flna
|
filamin A, alpha (actin binding protein 280) |
| chr1_-_191971 | 1.94 |
ENSDART00000164091
ENSDART00000159237 |
grtp1a
|
growth hormone regulated TBC protein 1a |
| chr4_-_58964138 | 1.91 |
ENSDART00000150259
|
si:ch211-64i20.3
|
si:ch211-64i20.3 |
| chr12_+_30360579 | 1.89 |
ENSDART00000152900
|
si:ch211-225b10.3
|
si:ch211-225b10.3 |
| chr14_-_41369629 | 1.88 |
ENSDART00000173040
|
cstf2
|
cleavage stimulation factor, 3' pre-RNA, subunit 2 |
| chr3_-_11203689 | 1.83 |
ENSDART00000166328
|
BX323882.1
|
|
| chr23_-_35064785 | 1.80 |
ENSDART00000172240
|
BX294434.1
|
|
| chr7_+_34794829 | 1.80 |
ENSDART00000009698
ENSDART00000075089 ENSDART00000173456 |
esrp2
|
epithelial splicing regulatory protein 2 |
| chr10_+_35422802 | 1.78 |
ENSDART00000147303
|
hhla2a.1
|
HERV-H LTR-associating 2a, tandem duplicate 1 |
| chr5_-_38063585 | 1.78 |
ENSDART00000164947
|
si:dkey-111e8.5
|
si:dkey-111e8.5 |
| chr6_+_42475730 | 1.56 |
ENSDART00000150226
|
mst1ra
|
macrophage stimulating 1 receptor a |
| chr20_-_49657134 | 1.55 |
ENSDART00000151248
|
col12a1b
|
collagen, type XII, alpha 1b |
| chr20_-_33966148 | 1.55 |
ENSDART00000148111
|
selp
|
selectin P |
| chr7_-_52558495 | 1.54 |
ENSDART00000138263
ENSDART00000009938 ENSDART00000174292 ENSDART00000174218 ENSDART00000174335 |
tcf12
|
transcription factor 12 |
| chr22_+_26290209 | 1.52 |
ENSDART00000060898
|
mrps28
|
mitochondrial ribosomal protein S28 |
| chr24_+_7800486 | 1.49 |
ENSDART00000145504
|
ptprh
|
protein tyrosine phosphatase, receptor type, h |
| chr2_+_22495274 | 1.45 |
ENSDART00000167915
|
lrrc8da
|
leucine rich repeat containing 8 VRAC subunit Da |
| chr20_-_2725930 | 1.45 |
ENSDART00000081643
|
akirin2
|
akirin 2 |
| chr14_-_858985 | 1.44 |
ENSDART00000148687
ENSDART00000149375 |
slc34a1a
|
solute carrier family 34 (type II sodium/phosphate cotransporter), member 1a |
| chr17_+_30591287 | 1.44 |
ENSDART00000154243
|
si:dkey-190l8.2
|
si:dkey-190l8.2 |
| chr22_+_21549419 | 1.43 |
ENSDART00000139411
|
plpp2b
|
phospholipid phosphatase 2b |
| chr22_+_3303671 | 1.41 |
ENSDART00000075049
|
gipc3
|
GIPC PDZ domain containing family, member 3 |
| chr6_-_8264751 | 1.39 |
ENSDART00000091628
|
ccdc151
|
coiled-coil domain containing 151 |
| chr18_-_21177674 | 1.36 |
ENSDART00000060175
|
si:dkey-12e7.4
|
si:dkey-12e7.4 |
| chr2_+_36620011 | 1.33 |
ENSDART00000177428
|
pak2a
|
p21 protein (Cdc42/Rac)-activated kinase 2a |
| chr13_-_47800342 | 1.31 |
ENSDART00000121817
|
fbln7
|
fibulin 7 |
| chr15_+_28318005 | 1.30 |
ENSDART00000175860
|
myo1cb
|
myosin Ic, paralog b |
| chr11_+_7214353 | 1.27 |
ENSDART00000156764
|
nwd1
|
NACHT and WD repeat domain containing 1 |
| chr25_-_1235457 | 1.25 |
ENSDART00000093093
|
coro2bb
|
coronin, actin binding protein, 2Bb |
| chr3_+_16722014 | 1.24 |
ENSDART00000008711
|
gys1
|
glycogen synthase 1 (muscle) |
| chr16_+_23796612 | 1.23 |
ENSDART00000131698
|
rab13
|
RAB13, member RAS oncogene family |
| chr7_-_18554603 | 1.22 |
ENSDART00000108938
|
msantd1
|
Myb/SANT-like DNA-binding domain containing 1 |
| chr15_-_1941042 | 1.21 |
ENSDART00000112946
ENSDART00000155504 |
dock10
|
dedicator of cytokinesis 10 |
| chr25_+_35212919 | 1.20 |
ENSDART00000180127
|
ano5a
|
anoctamin 5a |
| chr1_-_8963484 | 1.18 |
ENSDART00000188035
|
socs1b
|
suppressor of cytokine signaling 1b |
| chr4_+_20263097 | 1.18 |
ENSDART00000138820
|
lrtm2a
|
leucine-rich repeats and transmembrane domains 2a |
| chr8_-_15182232 | 1.14 |
ENSDART00000138855
|
bcar3
|
BCAR3, NSP family adaptor protein |
| chr10_-_24753715 | 1.12 |
ENSDART00000192401
|
ilk
|
integrin-linked kinase |
| chr16_-_47483142 | 1.11 |
ENSDART00000147072
|
cthrc1b
|
collagen triple helix repeat containing 1b |
| chr12_-_28548789 | 1.10 |
ENSDART00000153062
|
si:ch73-180n10.1
|
si:ch73-180n10.1 |
| chr10_-_27199135 | 1.09 |
ENSDART00000189511
ENSDART00000180314 |
auts2a
|
autism susceptibility candidate 2a |
| chr19_+_58954 | 1.02 |
ENSDART00000162379
|
col14a1b
|
collagen, type XIV, alpha 1b |
| chr5_+_38900249 | 1.02 |
ENSDART00000097856
|
fras1
|
Fraser extracellular matrix complex subunit 1 |
| chr25_+_9003879 | 1.01 |
ENSDART00000186228
|
rag1
|
recombination activating gene 1 |
| chr7_+_73807504 | 1.01 |
ENSDART00000166244
|
si:ch73-252p3.1
|
si:ch73-252p3.1 |
| chr7_-_3792972 | 0.96 |
ENSDART00000146304
|
si:dkey-28d5.12
|
si:dkey-28d5.12 |
| chr3_+_33745014 | 0.96 |
ENSDART00000159966
|
nacc1a
|
nucleus accumbens associated 1, BEN and BTB (POZ) domain containing a |
| chr2_-_37896646 | 0.95 |
ENSDART00000075931
|
hbl1
|
hexose-binding lectin 1 |
| chr15_+_28268135 | 0.92 |
ENSDART00000152536
ENSDART00000188550 |
myo1cb
|
myosin Ic, paralog b |
| chr20_+_46371458 | 0.88 |
ENSDART00000152912
|
adgrg11
|
adhesion G protein-coupled receptor G11 |
| chr23_-_33775145 | 0.86 |
ENSDART00000132147
ENSDART00000027959 ENSDART00000160116 |
racgap1
|
Rac GTPase activating protein 1 |
| chr16_-_7228276 | 0.84 |
ENSDART00000149030
|
nt5c3a
|
5'-nucleotidase, cytosolic IIIA |
| chr8_+_48858132 | 0.82 |
ENSDART00000124737
ENSDART00000079644 |
tp73
|
tumor protein p73 |
| chr25_+_4954220 | 0.79 |
ENSDART00000156034
|
si:ch73-265h17.2
|
si:ch73-265h17.2 |
| chr20_-_2619316 | 0.78 |
ENSDART00000185777
|
bub1
|
BUB1 mitotic checkpoint serine/threonine kinase |
| chr22_-_27136451 | 0.77 |
ENSDART00000153589
|
si:dkey-16m19.1
|
si:dkey-16m19.1 |
| chr12_+_6195191 | 0.77 |
ENSDART00000043236
ENSDART00000186420 |
prkg1b
|
protein kinase, cGMP-dependent, type Ib |
| chr25_+_9003230 | 0.77 |
ENSDART00000180330
ENSDART00000142917 |
rag1
|
recombination activating gene 1 |
| chr16_-_43041324 | 0.77 |
ENSDART00000155445
ENSDART00000156836 ENSDART00000154945 |
si:dkey-7j14.6
|
si:dkey-7j14.6 |
| chr6_+_40794015 | 0.76 |
ENSDART00000144479
|
gata2b
|
GATA binding protein 2b |
| chr8_+_25342896 | 0.76 |
ENSDART00000129032
|
CR847543.1
|
|
| chr11_-_3308569 | 0.73 |
ENSDART00000036581
|
cdk2
|
cyclin-dependent kinase 2 |
| chr12_+_25945560 | 0.73 |
ENSDART00000109799
|
mmrn2b
|
multimerin 2b |
| chr8_-_33154677 | 0.72 |
ENSDART00000133300
|
zbtb34
|
zinc finger and BTB domain containing 34 |
| chr21_-_45186431 | 0.72 |
ENSDART00000193449
|
si:ch73-269m14.2
|
si:ch73-269m14.2 |
| chr6_+_11850359 | 0.71 |
ENSDART00000109552
ENSDART00000188139 ENSDART00000181499 ENSDART00000178269 |
baz2ba
|
bromodomain adjacent to zinc finger domain, 2Ba |
| chr17_-_24439672 | 0.67 |
ENSDART00000155020
|
wdpcp
|
WD repeat containing planar cell polarity effector |
| chr21_-_11820379 | 0.62 |
ENSDART00000126640
|
RHOBTB3
|
si:dkey-6b12.5 |
| chr10_-_29101715 | 0.62 |
ENSDART00000149674
ENSDART00000171194 ENSDART00000192019 |
si:ch211-103f14.3
|
si:ch211-103f14.3 |
| chr6_+_54711306 | 0.61 |
ENSDART00000074605
|
pkp1b
|
plakophilin 1b |
| chr12_-_7807298 | 0.60 |
ENSDART00000191355
|
ank3b
|
ankyrin 3b |
| chr24_+_27023616 | 0.59 |
ENSDART00000089541
|
dip2ca
|
disco-interacting protein 2 homolog Ca |
| chr15_+_16521785 | 0.58 |
ENSDART00000062191
|
galnt17
|
polypeptide N-acetylgalactosaminyltransferase 17 |
| chr7_+_11459235 | 0.57 |
ENSDART00000159611
|
il16
|
interleukin 16 |
| chr4_-_16345227 | 0.57 |
ENSDART00000079521
|
kera
|
keratocan |
| chr1_-_35924495 | 0.56 |
ENSDART00000184424
|
smad1
|
SMAD family member 1 |
| chr19_+_56351 | 0.54 |
ENSDART00000168334
|
col14a1b
|
collagen, type XIV, alpha 1b |
| chr1_-_54191626 | 0.53 |
ENSDART00000062941
|
nthl1
|
nth-like DNA glycosylase 1 |
| chr4_+_16323970 | 0.53 |
ENSDART00000190651
|
BX322608.1
|
|
| chr16_+_8677112 | 0.52 |
ENSDART00000173153
|
si:ch211-207g17.3
|
si:ch211-207g17.3 |
| chr6_+_2951645 | 0.50 |
ENSDART00000183181
ENSDART00000181271 |
ptprfa
|
protein tyrosine phosphatase, receptor type, f, a |
| chr1_-_58880613 | 0.50 |
ENSDART00000188136
|
CABZ01084501.2
|
|
| chr5_+_43870389 | 0.49 |
ENSDART00000141002
|
zgc:112966
|
zgc:112966 |
| chr3_+_32118670 | 0.48 |
ENSDART00000055287
ENSDART00000111688 |
zgc:109934
|
zgc:109934 |
| chr14_-_16328331 | 0.47 |
ENSDART00000111262
|
pcdh12
|
protocadherin 12 |
| chr6_-_56111829 | 0.47 |
ENSDART00000154397
|
gcnt7
|
glucosaminyl (N-acetyl) transferase family member 7 |
| chr11_+_807153 | 0.46 |
ENSDART00000173289
|
vgll4b
|
vestigial-like family member 4b |
| chr21_+_26389391 | 0.46 |
ENSDART00000077197
|
tmsb
|
thymosin, beta |
| chr22_-_9618867 | 0.46 |
ENSDART00000131246
|
si:ch73-286h23.4
|
si:ch73-286h23.4 |
| chr10_-_5022176 | 0.45 |
ENSDART00000148277
|
hnrnpdl
|
heterogeneous nuclear ribonucleoprotein D-like |
| chr15_+_44250335 | 0.43 |
ENSDART00000186162
ENSDART00000193503 ENSDART00000180275 |
zgc:162962
|
zgc:162962 |
| chr18_+_33468099 | 0.42 |
ENSDART00000131769
|
v2rh9
|
vomeronasal 2 receptor, h9 |
| chr15_-_3736149 | 0.41 |
ENSDART00000182986
|
lpar6a
|
lysophosphatidic acid receptor 6a |
| chr13_-_16226312 | 0.41 |
ENSDART00000163952
|
zgc:110045
|
zgc:110045 |
| chr16_-_19254323 | 0.39 |
ENSDART00000138744
|
dnah11
|
dynein, axonemal, heavy chain 11 |
| chr15_+_1372343 | 0.38 |
ENSDART00000152285
|
schip1
|
schwannomin interacting protein 1 |
| chr2_-_10563576 | 0.37 |
ENSDART00000185818
ENSDART00000190887 |
ccdc18
|
coiled-coil domain containing 18 |
| chr21_-_26558268 | 0.35 |
ENSDART00000065390
|
CR631122.1
|
|
| chr4_+_58240778 | 0.35 |
ENSDART00000181184
ENSDART00000192348 ENSDART00000189444 ENSDART00000164424 |
si:dkey-16p6.1
|
si:dkey-16p6.1 |
| chr2_-_56649883 | 0.34 |
ENSDART00000191786
|
gpx4b
|
glutathione peroxidase 4b |
| chr15_+_31515976 | 0.33 |
ENSDART00000156471
|
tex26
|
testis expressed 26 |
| chr15_-_4596623 | 0.33 |
ENSDART00000132227
|
eif4h
|
eukaryotic translation initiation factor 4h |
| chr3_-_55649319 | 0.32 |
ENSDART00000176127
|
axin2
|
axin 2 (conductin, axil) |
| chr8_+_24740013 | 0.32 |
ENSDART00000126897
|
lamtor5
|
late endosomal/lysosomal adaptor, MAPK and MTOR activator 5 |
| chr4_+_5318901 | 0.31 |
ENSDART00000067378
|
si:ch211-214j24.10
|
si:ch211-214j24.10 |
| chr1_+_54835131 | 0.30 |
ENSDART00000145070
|
si:ch211-196h16.4
|
si:ch211-196h16.4 |
| chr12_+_34763795 | 0.30 |
ENSDART00000153097
|
slc38a10
|
solute carrier family 38, member 10 |
| chr11_+_3308656 | 0.30 |
ENSDART00000082458
|
sarnp
|
SAP domain containing ribonucleoprotein |
| chr9_-_21268576 | 0.28 |
ENSDART00000080604
|
sap18
|
Sin3A-associated protein |
| chr12_+_48744016 | 0.28 |
ENSDART00000175518
|
LO017648.1
|
|
| chr13_-_49819027 | 0.28 |
ENSDART00000067824
|
b3galnt2
|
beta-1,3-N-acetylgalactosaminyltransferase 2 |
| chr13_+_29193649 | 0.27 |
ENSDART00000045951
|
unc5b
|
unc-5 netrin receptor B |
| chr20_-_53321499 | 0.27 |
ENSDART00000179894
ENSDART00000127427 ENSDART00000084952 |
wasf1
|
WAS protein family, member 1 |
| chr5_+_37343643 | 0.26 |
ENSDART00000163993
ENSDART00000164459 ENSDART00000167006 ENSDART00000128122 ENSDART00000183720 |
klhl13
|
kelch-like family member 13 |
| chr11_+_19060079 | 0.26 |
ENSDART00000158123
|
magi1b
|
membrane associated guanylate kinase, WW and PDZ domain containing 1b |
| chr9_-_21912227 | 0.25 |
ENSDART00000145576
|
lmo7a
|
LIM domain 7a |
| chr10_-_31563049 | 0.23 |
ENSDART00000023575
|
robo3
|
roundabout, axon guidance receptor, homolog 3 (Drosophila) |
| chr22_-_9677264 | 0.23 |
ENSDART00000181737
|
si:dkey-286j17.4
|
si:dkey-286j17.4 |
| chr16_-_30927764 | 0.20 |
ENSDART00000156804
|
slc45a4
|
solute carrier family 45, member 4 |
| chr14_-_27289042 | 0.19 |
ENSDART00000159727
|
pcdh11
|
protocadherin 11 |
| chr25_+_30460378 | 0.18 |
ENSDART00000016310
|
banp
|
BTG3 associated nuclear protein |
| chr23_+_26946429 | 0.18 |
ENSDART00000185564
|
cacnb3b
|
calcium channel, voltage-dependent, beta 3b |
| chr9_+_41612642 | 0.17 |
ENSDART00000138473
|
spegb
|
SPEG complex locus b |
| chr11_-_8271374 | 0.17 |
ENSDART00000168253
|
pimr202
|
Pim proto-oncogene, serine/threonine kinase, related 202 |
| chr23_-_36418059 | 0.17 |
ENSDART00000135232
|
znf740b
|
zinc finger protein 740b |
| chr11_-_8288925 | 0.16 |
ENSDART00000168524
|
pimr202
|
Pim proto-oncogene, serine/threonine kinase, related 202 |
| chr5_+_27609014 | 0.16 |
ENSDART00000184912
ENSDART00000189071 ENSDART00000181080 |
zmat4a
|
zinc finger, matrin-type 4a |
| chr16_-_49846359 | 0.15 |
ENSDART00000184246
|
znf385d
|
zinc finger protein 385D |
| chr18_-_210478 | 0.15 |
ENSDART00000136693
|
tm2d3
|
TM2 domain containing 3 |
| chr18_+_34225520 | 0.14 |
ENSDART00000126115
|
v2rl1
|
vomeronasal 2 receptor, l1 |
| chr2_+_37110504 | 0.12 |
ENSDART00000132794
ENSDART00000042974 |
slc1a8b
|
solute carrier family 1 (glutamate transporter), member 8b |
| chr3_-_54846444 | 0.11 |
ENSDART00000074010
|
ubald1b
|
UBA-like domain containing 1b |
| chr2_-_52020049 | 0.11 |
ENSDART00000128198
ENSDART00000135331 |
scn12aa
|
sodium channel, voltage gated, type XII, alpha a |
| chr18_+_33340868 | 0.10 |
ENSDART00000099139
|
v2rh14
|
vomeronasal 2 receptor, h14 |
| chr17_+_45454943 | 0.09 |
ENSDART00000074838
|
kcnk3b
|
potassium channel, subfamily K, member 3b |
| chr3_+_19248973 | 0.08 |
ENSDART00000174668
|
smarca4a
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 4a |
| chr7_-_24005268 | 0.07 |
ENSDART00000173608
|
si:dkey-183c6.9
|
si:dkey-183c6.9 |
| chr15_-_2184638 | 0.05 |
ENSDART00000135460
|
shox2
|
short stature homeobox 2 |
| chr22_-_18387059 | 0.04 |
ENSDART00000007769
|
tssk6
|
testis-specific serine kinase 6 |
| chr15_-_5207805 | 0.03 |
ENSDART00000174046
|
or128-7
|
odorant receptor, family E, subfamily 128, member 7 |
| chr18_+_33818577 | 0.02 |
ENSDART00000131788
|
olfcq19
|
olfactory receptor C family, q19 |
| chr19_+_10979983 | 0.01 |
ENSDART00000142975
|
si:ch1073-70f20.1
|
si:ch1073-70f20.1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of ebf3a-1+si:ch211-51e8.2
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 8.6 | GO:0071691 | cardiac muscle thin filament assembly(GO:0071691) |
| 1.3 | 6.7 | GO:0038098 | sequestering of BMP from receptor via BMP binding(GO:0038098) |
| 1.0 | 4.2 | GO:0031448 | regulation of twitch skeletal muscle contraction(GO:0014724) regulation of fast-twitch skeletal muscle fiber contraction(GO:0031446) positive regulation of fast-twitch skeletal muscle fiber contraction(GO:0031448) negative regulation of striated muscle contraction(GO:0045988) relaxation of skeletal muscle(GO:0090076) |
| 0.6 | 4.8 | GO:0030643 | cellular anion homeostasis(GO:0030002) cellular monovalent inorganic anion homeostasis(GO:0030320) cellular phosphate ion homeostasis(GO:0030643) cellular divalent inorganic anion homeostasis(GO:0072501) cellular trivalent inorganic anion homeostasis(GO:0072502) |
| 0.6 | 2.4 | GO:0015682 | ferric iron transport(GO:0015682) transferrin transport(GO:0033572) trivalent inorganic cation transport(GO:0072512) |
| 0.5 | 6.0 | GO:0030240 | skeletal muscle thin filament assembly(GO:0030240) |
| 0.4 | 5.4 | GO:0033198 | response to ATP(GO:0033198) |
| 0.4 | 3.3 | GO:0086002 | cardiac muscle cell action potential involved in contraction(GO:0086002) |
| 0.3 | 1.2 | GO:0048210 | Golgi vesicle fusion to target membrane(GO:0048210) |
| 0.3 | 2.1 | GO:0060017 | parathyroid gland development(GO:0060017) |
| 0.3 | 2.3 | GO:1901998 | toxin transport(GO:1901998) |
| 0.3 | 1.5 | GO:0097241 | hematopoietic stem cell migration to bone marrow(GO:0097241) |
| 0.2 | 2.5 | GO:0043567 | regulation of insulin-like growth factor receptor signaling pathway(GO:0043567) |
| 0.2 | 1.9 | GO:0098787 | mRNA cleavage involved in mRNA processing(GO:0098787) pre-mRNA cleavage required for polyadenylation(GO:0098789) |
| 0.2 | 1.2 | GO:0060330 | regulation of response to interferon-gamma(GO:0060330) regulation of interferon-gamma-mediated signaling pathway(GO:0060334) |
| 0.2 | 1.1 | GO:0035989 | tendon development(GO:0035989) |
| 0.2 | 2.9 | GO:0036065 | fucosylation(GO:0036065) |
| 0.2 | 1.8 | GO:0033151 | V(D)J recombination(GO:0033151) |
| 0.2 | 1.8 | GO:0031294 | lymphocyte costimulation(GO:0031294) T cell costimulation(GO:0031295) |
| 0.1 | 0.5 | GO:0006296 | nucleotide-excision repair, DNA incision, 5'-to lesion(GO:0006296) |
| 0.1 | 8.1 | GO:0001946 | lymphangiogenesis(GO:0001946) |
| 0.1 | 0.5 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 0.1 | 0.8 | GO:0051754 | meiotic sister chromatid cohesion, centromeric(GO:0051754) |
| 0.1 | 2.1 | GO:0035141 | medial fin morphogenesis(GO:0035141) |
| 0.1 | 1.6 | GO:0051897 | positive regulation of protein kinase B signaling(GO:0051897) |
| 0.1 | 0.6 | GO:0045110 | intermediate filament bundle assembly(GO:0045110) |
| 0.1 | 1.5 | GO:0048679 | regulation of axon regeneration(GO:0048679) regulation of neuron projection regeneration(GO:0070570) |
| 0.1 | 0.4 | GO:0035332 | positive regulation of hippo signaling(GO:0035332) |
| 0.1 | 0.5 | GO:0097374 | sensory neuron axon guidance(GO:0097374) |
| 0.1 | 4.0 | GO:0045089 | positive regulation of innate immune response(GO:0045089) |
| 0.1 | 1.1 | GO:0033138 | positive regulation of peptidyl-serine phosphorylation(GO:0033138) |
| 0.1 | 1.1 | GO:0021884 | forebrain neuron development(GO:0021884) |
| 0.0 | 2.1 | GO:0010862 | positive regulation of pathway-restricted SMAD protein phosphorylation(GO:0010862) |
| 0.0 | 2.2 | GO:0030050 | vesicle transport along actin filament(GO:0030050) actin filament-based transport(GO:0099515) |
| 0.0 | 1.9 | GO:0006890 | retrograde vesicle-mediated transport, Golgi to ER(GO:0006890) |
| 0.0 | 12.5 | GO:0030198 | extracellular matrix organization(GO:0030198) extracellular structure organization(GO:0043062) |
| 0.0 | 0.6 | GO:0050930 | regulation of positive chemotaxis(GO:0050926) positive regulation of positive chemotaxis(GO:0050927) induction of positive chemotaxis(GO:0050930) |
| 0.0 | 4.7 | GO:1990266 | neutrophil migration(GO:1990266) |
| 0.0 | 0.3 | GO:1904263 | positive regulation of TORC1 signaling(GO:1904263) |
| 0.0 | 0.6 | GO:0060395 | SMAD protein signal transduction(GO:0060395) |
| 0.0 | 1.2 | GO:0005978 | glycogen biosynthetic process(GO:0005978) glucan biosynthetic process(GO:0009250) |
| 0.0 | 0.3 | GO:2000054 | negative regulation of Wnt signaling pathway involved in dorsal/ventral axis specification(GO:2000054) |
| 0.0 | 0.3 | GO:0038007 | netrin-activated signaling pathway(GO:0038007) |
| 0.0 | 0.3 | GO:0016973 | poly(A)+ mRNA export from nucleus(GO:0016973) |
| 0.0 | 0.3 | GO:2000601 | positive regulation of actin nucleation(GO:0051127) positive regulation of Arp2/3 complex-mediated actin nucleation(GO:2000601) |
| 0.0 | 0.7 | GO:0010389 | regulation of G2/M transition of mitotic cell cycle(GO:0010389) |
| 0.0 | 0.2 | GO:0071679 | commissural neuron axon guidance(GO:0071679) |
| 0.0 | 1.4 | GO:0046839 | phospholipid dephosphorylation(GO:0046839) |
| 0.0 | 2.5 | GO:0007160 | cell-matrix adhesion(GO:0007160) |
| 0.0 | 0.5 | GO:0035329 | hippo signaling(GO:0035329) |
| 0.0 | 0.8 | GO:0007050 | cell cycle arrest(GO:0007050) |
| 0.0 | 0.9 | GO:0000281 | mitotic cytokinesis(GO:0000281) |
| 0.0 | 2.3 | GO:0070507 | regulation of microtubule cytoskeleton organization(GO:0070507) |
| 0.0 | 1.9 | GO:0090630 | activation of GTPase activity(GO:0090630) |
| 0.0 | 1.3 | GO:0031098 | stress-activated protein kinase signaling cascade(GO:0031098) |
| 0.0 | 0.1 | GO:0097010 | eukaryotic translation initiation factor 4F complex assembly(GO:0097010) |
| 0.0 | 0.1 | GO:0021634 | optic nerve formation(GO:0021634) |
| 0.0 | 1.7 | GO:0006954 | inflammatory response(GO:0006954) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 4.2 | GO:0031673 | H zone(GO:0031673) |
| 0.4 | 1.8 | GO:0097519 | DNA recombinase complex(GO:0097519) |
| 0.3 | 6.1 | GO:0031430 | M band(GO:0031430) |
| 0.3 | 5.4 | GO:0031229 | integral component of nuclear inner membrane(GO:0005639) intrinsic component of nuclear inner membrane(GO:0031229) nuclear membrane part(GO:0044453) |
| 0.2 | 0.7 | GO:0097541 | axonemal basal plate(GO:0097541) |
| 0.2 | 22.1 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.1 | 12.4 | GO:0030018 | Z disc(GO:0030018) |
| 0.1 | 1.2 | GO:0032593 | insulin-responsive compartment(GO:0032593) |
| 0.1 | 1.9 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.1 | 1.4 | GO:0005903 | brush border(GO:0005903) |
| 0.1 | 1.5 | GO:0000314 | organellar small ribosomal subunit(GO:0000314) mitochondrial small ribosomal subunit(GO:0005763) |
| 0.1 | 1.4 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.1 | 2.4 | GO:0048770 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.1 | 0.3 | GO:0071986 | Ragulator complex(GO:0071986) |
| 0.0 | 2.9 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.0 | 2.2 | GO:0005902 | microvillus(GO:0005902) |
| 0.0 | 4.4 | GO:0016324 | apical plasma membrane(GO:0016324) |
| 0.0 | 0.6 | GO:0030057 | desmosome(GO:0030057) |
| 0.0 | 1.5 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.0 | 0.8 | GO:0000778 | condensed nuclear chromosome kinetochore(GO:0000778) condensed nuclear chromosome, centromeric region(GO:0000780) |
| 0.0 | 0.3 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 0.0 | 0.6 | GO:0071144 | SMAD2-SMAD3 protein complex(GO:0071144) |
| 0.0 | 1.9 | GO:0030134 | ER to Golgi transport vesicle(GO:0030134) |
| 0.0 | 10.5 | GO:0031012 | extracellular matrix(GO:0031012) |
| 0.0 | 1.2 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.0 | 18.9 | GO:0005615 | extracellular space(GO:0005615) |
| 0.0 | 0.3 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.0 | 0.7 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.0 | 4.0 | GO:0000785 | chromatin(GO:0000785) |
| 0.0 | 3.9 | GO:0009986 | cell surface(GO:0009986) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 6.7 | GO:0036122 | BMP binding(GO:0036122) |
| 0.6 | 2.4 | GO:0004998 | transferrin receptor activity(GO:0004998) |
| 0.4 | 5.4 | GO:0004931 | extracellular ATP-gated cation channel activity(GO:0004931) ATP-gated ion channel activity(GO:0035381) |
| 0.4 | 1.8 | GO:1990238 | double-stranded DNA endodeoxyribonuclease activity(GO:1990238) |
| 0.4 | 3.4 | GO:0005105 | type 1 fibroblast growth factor receptor binding(GO:0005105) |
| 0.3 | 1.2 | GO:0004373 | glycogen (starch) synthase activity(GO:0004373) |
| 0.3 | 2.6 | GO:0051373 | telethonin binding(GO:0031433) FATZ binding(GO:0051373) |
| 0.3 | 6.0 | GO:0042800 | histone methyltransferase activity (H3-K4 specific)(GO:0042800) |
| 0.2 | 2.9 | GO:0046920 | alpha-(1->3)-fucosyltransferase activity(GO:0046920) |
| 0.2 | 1.4 | GO:0005436 | sodium:phosphate symporter activity(GO:0005436) |
| 0.2 | 2.5 | GO:0031995 | insulin-like growth factor I binding(GO:0031994) insulin-like growth factor II binding(GO:0031995) |
| 0.2 | 8.1 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.2 | 0.5 | GO:0000703 | oxidized pyrimidine nucleobase lesion DNA N-glycosylase activity(GO:0000703) |
| 0.1 | 0.8 | GO:0004692 | cGMP-dependent protein kinase activity(GO:0004692) |
| 0.1 | 2.1 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.1 | 0.6 | GO:0042609 | CD4 receptor binding(GO:0042609) |
| 0.1 | 0.5 | GO:0001223 | transcription coactivator binding(GO:0001223) |
| 0.1 | 0.4 | GO:0034046 | poly(G) binding(GO:0034046) |
| 0.1 | 10.9 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.1 | 1.2 | GO:0048495 | Roundabout binding(GO:0048495) |
| 0.1 | 0.7 | GO:0030332 | cyclin binding(GO:0030332) |
| 0.1 | 0.3 | GO:0047066 | phospholipid-hydroperoxide glutathione peroxidase activity(GO:0047066) |
| 0.1 | 2.4 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.1 | 1.4 | GO:0042577 | lipid phosphatase activity(GO:0042577) |
| 0.0 | 2.2 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.0 | 0.8 | GO:0008253 | 5'-nucleotidase activity(GO:0008253) |
| 0.0 | 1.2 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.0 | 19.9 | GO:0005198 | structural molecule activity(GO:0005198) |
| 0.0 | 10.0 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.0 | 0.1 | GO:0005252 | open rectifier potassium channel activity(GO:0005252) |
| 0.0 | 3.3 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 0.3 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 0.0 | 0.6 | GO:0070411 | I-SMAD binding(GO:0070411) |
| 0.0 | 0.3 | GO:0005042 | netrin receptor activity(GO:0005042) |
| 0.0 | 0.5 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.0 | 5.1 | GO:0050839 | cell adhesion molecule binding(GO:0050839) |
| 0.0 | 2.8 | GO:0005179 | hormone activity(GO:0005179) |
| 0.0 | 0.2 | GO:0070915 | lysophosphatidic acid receptor activity(GO:0070915) |
| 0.0 | 2.1 | GO:0008083 | growth factor activity(GO:0008083) |
| 0.0 | 0.4 | GO:0045505 | dynein intermediate chain binding(GO:0045505) |
| 0.0 | 0.1 | GO:0034057 | RNA strand annealing activity(GO:0033592) RNA strand-exchange activity(GO:0034057) |
| 0.0 | 1.4 | GO:0004714 | transmembrane receptor protein tyrosine kinase activity(GO:0004714) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 5.4 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.2 | 5.9 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.1 | 1.1 | SA PTEN PATHWAY | PTEN is a tumor suppressor that dephosphorylates the lipid messenger phosphatidylinositol triphosphate. |
| 0.1 | 2.3 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.1 | 3.2 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.1 | 1.2 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.1 | 3.4 | PID FGF PATHWAY | FGF signaling pathway |
| 0.1 | 0.7 | SA G1 AND S PHASES | Cdk2, 4, and 6 bind cyclin D in G1, while cdk2/cyclin E promotes the G1/S transition. |
| 0.1 | 2.6 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.0 | 1.1 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.0 | 0.6 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.0 | 1.0 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.0 | 3.1 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 0.3 | PID BETA CATENIN DEG PATHWAY | Degradation of beta catenin |
| 0.0 | 0.3 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.0 | 0.8 | PID P73PATHWAY | p73 transcription factor network |
| 0.0 | 2.1 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 3.4 | REACTOME SIGNALING BY ACTIVATED POINT MUTANTS OF FGFR1 | Genes involved in Signaling by activated point mutants of FGFR1 |
| 0.3 | 8.6 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.2 | 4.9 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.2 | 4.2 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.1 | 5.9 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.1 | 1.9 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.1 | 1.1 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.1 | 0.6 | REACTOME KERATAN SULFATE DEGRADATION | Genes involved in Keratan sulfate degradation |
| 0.0 | 0.5 | REACTOME BASE FREE SUGAR PHOSPHATE REMOVAL VIA THE SINGLE NUCLEOTIDE REPLACEMENT PATHWAY | Genes involved in Base-free sugar-phosphate removal via the single-nucleotide replacement pathway |
| 0.0 | 0.6 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.0 | 1.5 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 0.9 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.0 | 1.9 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.0 | 1.2 | REACTOME GLUCOSE METABOLISM | Genes involved in Glucose metabolism |
| 0.0 | 0.8 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.0 | 0.3 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |