Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
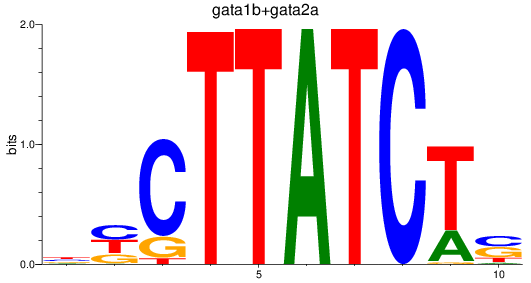
Results for gata1b+gata2a
Z-value: 2.65
Transcription factors associated with gata1b+gata2a
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
gata1b
|
ENSDARG00000059130 | GATA binding protein 1b |
|
gata2a
|
ENSDARG00000059327 | GATA binding protein 2a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| gata1b | dr11_v1_chr8_-_7474997_7474997 | -0.51 | 1.8e-07 | Click! |
| gata2a | dr11_v1_chr11_-_3865472_3865472 | -0.36 | 3.0e-04 | Click! |
Activity profile of gata1b+gata2a motif
Sorted Z-values of gata1b+gata2a motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr23_+_7379728 | 68.30 |
ENSDART00000012194
|
gata5
|
GATA binding protein 5 |
| chr5_-_64168415 | 50.35 |
ENSDART00000048395
|
cmlc1
|
cardiac myosin light chain-1 |
| chr2_-_51794472 | 43.86 |
ENSDART00000186652
|
BX908782.3
|
|
| chr2_-_24289641 | 42.06 |
ENSDART00000128784
ENSDART00000123565 ENSDART00000141922 ENSDART00000184550 ENSDART00000191469 |
myh7l
|
myosin heavy chain 7-like |
| chr16_-_36834505 | 37.15 |
ENSDART00000141275
ENSDART00000139588 ENSDART00000041993 |
pnp4b
|
purine nucleoside phosphorylase 4b |
| chr13_+_10945337 | 36.80 |
ENSDART00000091845
|
abcg5
|
ATP-binding cassette, sub-family G (WHITE), member 5 |
| chr8_-_40464935 | 36.39 |
ENSDART00000040013
|
myl7
|
myosin, light chain 7, regulatory |
| chr25_-_28674739 | 36.27 |
ENSDART00000067073
|
lrrc10
|
leucine rich repeat containing 10 |
| chr2_+_4207209 | 35.39 |
ENSDART00000157903
ENSDART00000166476 |
gata6
|
GATA binding protein 6 |
| chr18_-_42830563 | 30.70 |
ENSDART00000191488
|
ttc36
|
tetratricopeptide repeat domain 36 |
| chr13_-_20381485 | 30.68 |
ENSDART00000131351
|
si:ch211-270n8.1
|
si:ch211-270n8.1 |
| chr10_+_10801564 | 29.22 |
ENSDART00000027026
|
ambp
|
alpha-1-microglobulin/bikunin precursor |
| chr13_-_10945288 | 28.73 |
ENSDART00000114315
ENSDART00000164667 ENSDART00000159482 |
abcg8
|
ATP-binding cassette, sub-family G (WHITE), member 8 |
| chr14_+_36738069 | 27.49 |
ENSDART00000105590
|
tdo2a
|
tryptophan 2,3-dioxygenase a |
| chr4_+_1530287 | 27.40 |
ENSDART00000067446
|
slc38a4
|
solute carrier family 38, member 4 |
| chr15_-_21877726 | 26.22 |
ENSDART00000127819
ENSDART00000145646 ENSDART00000100897 ENSDART00000144739 |
zgc:162608
|
zgc:162608 |
| chr22_-_36856405 | 24.63 |
ENSDART00000029588
|
kng1
|
kininogen 1 |
| chr10_+_10801719 | 24.39 |
ENSDART00000193648
|
ambp
|
alpha-1-microglobulin/bikunin precursor |
| chr19_+_8606883 | 24.21 |
ENSDART00000054469
ENSDART00000185264 |
s100a10a
|
S100 calcium binding protein A10a |
| chr23_-_5685023 | 22.90 |
ENSDART00000148680
ENSDART00000149365 |
tnnt2a
|
troponin T type 2a (cardiac) |
| chr24_-_39858710 | 22.46 |
ENSDART00000134251
|
slc12a7b
|
solute carrier family 12 (potassium/chloride transporter), member 7b |
| chr7_-_41858513 | 21.85 |
ENSDART00000109918
|
mylk3
|
myosin light chain kinase 3 |
| chr21_-_25756119 | 21.75 |
ENSDART00000002341
|
cldnc
|
claudin c |
| chr6_-_49078263 | 21.44 |
ENSDART00000032982
|
slc5a8l
|
solute carrier family 5 (iodide transporter), member 8-like |
| chr11_-_7380674 | 20.67 |
ENSDART00000014979
ENSDART00000103418 |
vtg3
|
vitellogenin 3, phosvitinless |
| chr24_-_2843107 | 20.47 |
ENSDART00000165290
|
cyb5a
|
cytochrome b5 type A (microsomal) |
| chr7_-_41851605 | 20.35 |
ENSDART00000142981
|
mylk3
|
myosin light chain kinase 3 |
| chr5_+_21211135 | 18.98 |
ENSDART00000088492
|
bmp10
|
bone morphogenetic protein 10 |
| chr7_+_35075847 | 17.91 |
ENSDART00000193469
ENSDART00000037346 |
ctrb1
|
chymotrypsinogen B1 |
| chr6_-_609880 | 17.61 |
ENSDART00000149248
ENSDART00000148867 ENSDART00000149414 ENSDART00000148552 ENSDART00000148391 |
lgals2b
|
lectin, galactoside-binding, soluble, 2b |
| chr23_+_25856541 | 17.51 |
ENSDART00000145426
ENSDART00000028236 |
hnf4a
|
hepatocyte nuclear factor 4, alpha |
| chr20_+_40150612 | 14.71 |
ENSDART00000143680
ENSDART00000109681 ENSDART00000101041 ENSDART00000121818 |
trdn
|
triadin |
| chr25_-_7974494 | 14.69 |
ENSDART00000171446
|
hal
|
histidine ammonia-lyase |
| chr3_-_33395233 | 14.12 |
ENSDART00000167349
|
si:dkey-283b1.6
|
si:dkey-283b1.6 |
| chr12_-_32421046 | 13.79 |
ENSDART00000075567
|
enpp7.1
|
ectonucleotide pyrophosphatase/phosphodiesterase 7, tandem duplicate 1 |
| chr18_-_16795262 | 13.34 |
ENSDART00000048722
|
ampd3b
|
adenosine monophosphate deaminase 3b |
| chr11_-_1400507 | 13.33 |
ENSDART00000173029
ENSDART00000172953 ENSDART00000111140 |
rpl29
|
ribosomal protein L29 |
| chr13_-_23031803 | 13.23 |
ENSDART00000056523
|
hkdc1
|
hexokinase domain containing 1 |
| chr13_+_2908764 | 12.54 |
ENSDART00000162362
|
wu:fj16a03
|
wu:fj16a03 |
| chr20_+_15552657 | 12.27 |
ENSDART00000063912
|
jun
|
Jun proto-oncogene, AP-1 transcription factor subunit |
| chr20_+_33981946 | 12.02 |
ENSDART00000131775
|
si:dkey-51e6.1
|
si:dkey-51e6.1 |
| chr7_+_12835048 | 11.92 |
ENSDART00000016465
|
cx36.7
|
connexin 36.7 |
| chr16_+_12611362 | 11.91 |
ENSDART00000055160
|
il11a
|
interleukin 11a |
| chr13_+_22480496 | 11.64 |
ENSDART00000136863
ENSDART00000131870 ENSDART00000078720 ENSDART00000078740 ENSDART00000139218 |
ldb3a
|
LIM domain binding 3a |
| chr20_+_23625387 | 11.61 |
ENSDART00000147945
ENSDART00000150497 |
palld
|
palladin, cytoskeletal associated protein |
| chr8_-_40555340 | 11.48 |
ENSDART00000163348
|
NPC1L1
|
NPC1 like intracellular cholesterol transporter 1 |
| chr21_-_40880317 | 11.41 |
ENSDART00000100054
ENSDART00000137696 |
elnb
|
elastin b |
| chr25_+_18564266 | 11.27 |
ENSDART00000172338
|
cav1
|
caveolin 1 |
| chr5_+_23151169 | 11.15 |
ENSDART00000125638
|
tbx5b
|
T-box 5b |
| chr16_-_24668620 | 11.09 |
ENSDART00000012807
|
pon3.2
|
paraoxonase 3, tandem duplicate 2 |
| chr14_+_2487672 | 10.97 |
ENSDART00000170629
ENSDART00000123063 |
fgf18a
|
fibroblast growth factor 18a |
| chr15_-_43164591 | 10.78 |
ENSDART00000171305
|
ap1s3a
|
adaptor-related protein complex 1, sigma 3 subunit, a |
| chr5_-_30620625 | 10.74 |
ENSDART00000098273
|
tcnl
|
transcobalamin like |
| chr16_-_24815091 | 10.69 |
ENSDART00000154269
ENSDART00000131025 |
kcnn4
|
potassium intermediate/small conductance calcium-activated channel, subfamily N, member 4 |
| chr20_+_33875256 | 10.68 |
ENSDART00000002554
|
rxrgb
|
retinoid X receptor, gamma b |
| chr21_+_19547806 | 10.46 |
ENSDART00000159707
ENSDART00000184869 ENSDART00000181321 ENSDART00000058487 ENSDART00000058485 |
rai14
|
retinoic acid induced 14 |
| chr23_-_23401305 | 10.45 |
ENSDART00000078936
|
her9
|
hairy-related 9 |
| chr1_-_9109699 | 10.23 |
ENSDART00000147833
|
vap
|
vascular associated protein |
| chr25_-_13728111 | 10.07 |
ENSDART00000169865
|
lcat
|
lecithin-cholesterol acyltransferase |
| chr13_-_25774183 | 10.05 |
ENSDART00000046981
|
pdlim1
|
PDZ and LIM domain 1 (elfin) |
| chr15_+_26600611 | 10.02 |
ENSDART00000155352
|
slc47a3
|
solute carrier family 47 (multidrug and toxin extrusion), member 3 |
| chr7_-_73752955 | 9.91 |
ENSDART00000171254
ENSDART00000009888 |
casq1b
|
calsequestrin 1b |
| chr22_+_15336752 | 9.84 |
ENSDART00000139070
|
sult3st2
|
sulfotransferase family 3, cytosolic sulfotransferase 2 |
| chr8_-_18537866 | 9.66 |
ENSDART00000148802
ENSDART00000148962 ENSDART00000149506 |
nexn
|
nexilin (F actin binding protein) |
| chr16_+_25126935 | 9.42 |
ENSDART00000058945
|
zgc:92590
|
zgc:92590 |
| chr18_-_6803424 | 9.41 |
ENSDART00000142647
|
si:dkey-266m15.5
|
si:dkey-266m15.5 |
| chr8_-_50287949 | 9.33 |
ENSDART00000023639
|
nkx2.7
|
NK2 transcription factor related 7 |
| chr13_-_40282770 | 9.28 |
ENSDART00000140875
|
zgc:123010
|
zgc:123010 |
| chr17_+_39242437 | 9.23 |
ENSDART00000156138
ENSDART00000128863 |
zgc:174356
|
zgc:174356 |
| chr21_+_12010505 | 9.14 |
ENSDART00000123522
|
aqp7
|
aquaporin 7 |
| chr3_+_55105066 | 9.12 |
ENSDART00000110848
|
si:ch211-5k11.8
|
si:ch211-5k11.8 |
| chr7_+_54642005 | 9.11 |
ENSDART00000171864
|
fgf19
|
fibroblast growth factor 19 |
| chr23_-_33750135 | 9.03 |
ENSDART00000187641
|
bin2a
|
bridging integrator 2a |
| chr23_-_33750307 | 9.02 |
ENSDART00000162772
|
bin2a
|
bridging integrator 2a |
| chr20_-_33705044 | 9.00 |
ENSDART00000166573
|
rock2b
|
rho-associated, coiled-coil containing protein kinase 2b |
| chr7_+_69019851 | 8.99 |
ENSDART00000162891
|
CABZ01057488.1
|
|
| chr10_+_26612321 | 8.86 |
ENSDART00000134322
|
fhl1b
|
four and a half LIM domains 1b |
| chr19_+_20778011 | 8.83 |
ENSDART00000024208
|
nutf2l
|
nuclear transport factor 2, like |
| chr5_-_51166025 | 8.80 |
ENSDART00000097471
|
card9
|
caspase recruitment domain family, member 9 |
| chr13_+_22479988 | 8.66 |
ENSDART00000188182
ENSDART00000192972 ENSDART00000178372 |
ldb3a
|
LIM domain binding 3a |
| chr13_+_844150 | 8.39 |
ENSDART00000058260
|
gsta.1
|
glutathione S-transferase, alpha tandem duplicate 1 |
| chr16_-_24945337 | 8.38 |
ENSDART00000154004
|
has1
|
hyaluronan synthase 1 |
| chr11_+_31285127 | 8.24 |
ENSDART00000160154
|
si:dkey-238i5.2
|
si:dkey-238i5.2 |
| chr8_+_6576940 | 8.15 |
ENSDART00000138135
|
vsig8b
|
V-set and immunoglobulin domain containing 8b |
| chr13_+_835390 | 7.95 |
ENSDART00000171329
|
gsta.1
|
glutathione S-transferase, alpha tandem duplicate 1 |
| chr5_+_15203421 | 7.85 |
ENSDART00000040826
|
tbx1
|
T-box 1 |
| chr1_+_9954489 | 7.66 |
ENSDART00000005895
ENSDART00000183003 |
pdia2
|
protein disulfide isomerase family A, member 2 |
| chr7_+_44445595 | 7.60 |
ENSDART00000108766
|
cdh5
|
cadherin 5 |
| chr16_+_54829574 | 7.52 |
ENSDART00000148392
|
pabpc1a
|
poly(A) binding protein, cytoplasmic 1a |
| chr12_+_6391243 | 7.42 |
ENSDART00000152765
|
prkg1b
|
protein kinase, cGMP-dependent, type Ib |
| chr24_-_12958668 | 7.34 |
ENSDART00000178982
|
FITM1 (1 of many)
|
Danio rerio fat storage inducing transmembrane protein 1 (LOC792443), mRNA. |
| chr2_-_38261272 | 7.33 |
ENSDART00000143743
|
si:ch211-14a17.10
|
si:ch211-14a17.10 |
| chr8_+_31872992 | 7.32 |
ENSDART00000146933
|
c7a
|
complement component 7a |
| chr18_+_48428126 | 7.26 |
ENSDART00000137978
|
fli1a
|
Fli-1 proto-oncogene, ETS transcription factor a |
| chr7_-_15252540 | 7.20 |
ENSDART00000173072
ENSDART00000125258 |
si:dkey-172h23.2
|
si:dkey-172h23.2 |
| chr14_+_32839535 | 7.13 |
ENSDART00000168975
|
arr3b
|
arrestin 3b, retinal (X-arrestin) |
| chr20_+_18992679 | 6.99 |
ENSDART00000147105
ENSDART00000152811 |
tdh
|
L-threonine dehydrogenase |
| chr3_+_16762483 | 6.91 |
ENSDART00000132732
|
tmem86b
|
transmembrane protein 86B |
| chr2_+_21486529 | 6.84 |
ENSDART00000047468
|
inhbab
|
inhibin, beta Ab |
| chr17_+_23554932 | 6.81 |
ENSDART00000135814
|
pank1a
|
pantothenate kinase 1a |
| chr17_+_30894431 | 6.73 |
ENSDART00000127996
|
degs2
|
delta(4)-desaturase, sphingolipid 2 |
| chr3_+_55100045 | 6.66 |
ENSDART00000113098
|
hbaa1
|
hemoglobin, alpha adult 1 |
| chr17_+_34206167 | 6.61 |
ENSDART00000136167
|
mpp5a
|
membrane protein, palmitoylated 5a (MAGUK p55 subfamily member 5) |
| chr20_-_33704753 | 6.57 |
ENSDART00000157427
|
rock2b
|
rho-associated, coiled-coil containing protein kinase 2b |
| chr17_+_45654724 | 6.55 |
ENSDART00000098952
|
zgc:162184
|
zgc:162184 |
| chr14_-_28052474 | 6.50 |
ENSDART00000172948
ENSDART00000135337 |
TSC22D3 (1 of many)
zgc:64189
|
si:ch211-220e11.3 zgc:64189 |
| chr11_-_25418856 | 6.49 |
ENSDART00000013714
|
gata1a
|
GATA binding protein 1a |
| chr20_-_25533739 | 6.49 |
ENSDART00000063064
|
cyp2ad6
|
cytochrome P450, family 2, subfamily AD, polypeptide 6 |
| chr24_+_2843268 | 6.46 |
ENSDART00000170529
|
si:ch211-152c8.2
|
si:ch211-152c8.2 |
| chr22_-_11626014 | 6.33 |
ENSDART00000063133
ENSDART00000160085 |
gcga
|
glucagon a |
| chr14_+_17382803 | 6.21 |
ENSDART00000040383
|
si:ch211-255i20.3
|
si:ch211-255i20.3 |
| chr9_+_343062 | 6.16 |
ENSDART00000164012
ENSDART00000164365 |
bpifcl
|
BPI fold containing family C, like |
| chr22_+_5120033 | 6.16 |
ENSDART00000169200
|
mibp
|
muscle-specific beta 1 integrin binding protein |
| chr1_+_36674584 | 6.12 |
ENSDART00000186772
ENSDART00000192274 |
ednraa
|
endothelin receptor type Aa |
| chr17_-_29736019 | 6.10 |
ENSDART00000183483
|
ush2a
|
Usher syndrome 2A (autosomal recessive, mild) |
| chr5_-_26247973 | 6.09 |
ENSDART00000098527
|
erap1b
|
endoplasmic reticulum aminopeptidase 1b |
| chr21_+_39941875 | 6.04 |
ENSDART00000190414
|
slc47a1
|
solute carrier family 47 (multidrug and toxin extrusion), member 1 |
| chr12_-_20665164 | 5.90 |
ENSDART00000105352
|
gip
|
gastric inhibitory polypeptide |
| chr6_-_33129045 | 5.90 |
ENSDART00000073757
|
tspan1
|
tetraspanin 1 |
| chr18_+_33550667 | 5.77 |
ENSDART00000141246
ENSDART00000136070 |
si:dkey-47k20.1
|
si:dkey-47k20.1 |
| chr12_-_23658888 | 5.76 |
ENSDART00000088319
|
map3k8
|
mitogen-activated protein kinase kinase kinase 8 |
| chr6_-_33129312 | 5.74 |
ENSDART00000156987
|
tspan1
|
tetraspanin 1 |
| chr2_-_37312927 | 5.68 |
ENSDART00000141214
|
skila
|
SKI-like proto-oncogene a |
| chr5_+_26248380 | 5.67 |
ENSDART00000079049
|
si:ch211-214j8.1
|
si:ch211-214j8.1 |
| chr6_+_51932563 | 5.67 |
ENSDART00000181778
|
angpt4
|
angiopoietin 4 |
| chr20_-_26066020 | 5.61 |
ENSDART00000078559
|
myct1a
|
myc target 1a |
| chr4_+_18824959 | 5.60 |
ENSDART00000146141
ENSDART00000040424 |
slc26a3.1
|
solute carrier family 26 (anion exchanger), member 3 |
| chr23_+_26079467 | 5.58 |
ENSDART00000129617
|
atp6ap1b
|
ATPase H+ transporting accessory protein 1b |
| chr2_+_38261748 | 5.49 |
ENSDART00000076478
|
dhrs1
|
dehydrogenase/reductase (SDR family) member 1 |
| chr20_+_46897504 | 5.39 |
ENSDART00000158124
|
si:ch73-21k16.1
|
si:ch73-21k16.1 |
| chr9_-_42871756 | 5.39 |
ENSDART00000191396
|
ttn.1
|
titin, tandem duplicate 1 |
| chr24_+_14801844 | 5.33 |
ENSDART00000141620
|
pi15a
|
peptidase inhibitor 15a |
| chr24_+_19518570 | 5.29 |
ENSDART00000056081
|
sulf1
|
sulfatase 1 |
| chr12_+_47945664 | 5.27 |
ENSDART00000189824
ENSDART00000055743 ENSDART00000188936 |
CABZ01074994.1
|
|
| chr17_-_26867725 | 5.18 |
ENSDART00000153590
|
si:dkey-221l4.10
|
si:dkey-221l4.10 |
| chr16_+_54829293 | 5.05 |
ENSDART00000024729
|
pabpc1a
|
poly(A) binding protein, cytoplasmic 1a |
| chr3_+_17251542 | 5.02 |
ENSDART00000024832
|
stat5a
|
signal transducer and activator of transcription 5a |
| chr12_+_23850661 | 4.99 |
ENSDART00000152921
|
svila
|
supervillin a |
| chr11_-_28614608 | 4.99 |
ENSDART00000065853
|
dhrs3b
|
dehydrogenase/reductase (SDR family) member 3b |
| chr2_+_39021282 | 4.90 |
ENSDART00000056577
|
RBP1 (1 of many)
|
si:ch211-119o8.7 |
| chr24_+_10202718 | 4.82 |
ENSDART00000126668
|
pou6f2
|
POU class 6 homeobox 2 |
| chr12_-_20350629 | 4.82 |
ENSDART00000066384
|
hbbe2
|
hemoglobin beta embryonic-2 |
| chr5_-_66301142 | 4.81 |
ENSDART00000067541
|
prlrb
|
prolactin receptor b |
| chr10_-_17484762 | 4.73 |
ENSDART00000137905
ENSDART00000007961 |
nt5c2l1
|
5'-nucleotidase, cytosolic II, like 1 |
| chr14_+_46268248 | 4.63 |
ENSDART00000172979
|
cabp2b
|
calcium binding protein 2b |
| chr14_+_26546175 | 4.56 |
ENSDART00000105932
|
si:dkeyp-110e4.11
|
si:dkeyp-110e4.11 |
| chr5_-_8817841 | 4.55 |
ENSDART00000170921
|
fgf10b
|
fibroblast growth factor 10b |
| chr16_+_13424534 | 4.52 |
ENSDART00000114944
|
zgc:101640
|
zgc:101640 |
| chr16_-_17300030 | 4.48 |
ENSDART00000149267
|
kel
|
Kell blood group, metallo-endopeptidase |
| chr25_-_35102781 | 4.46 |
ENSDART00000180881
ENSDART00000153747 |
si:dkey-108k21.24
|
si:dkey-108k21.24 |
| chr12_-_29301022 | 4.43 |
ENSDART00000187826
|
sh2d4bb
|
SH2 domain containing 4Bb |
| chr20_+_25879826 | 4.36 |
ENSDART00000018519
|
zgc:153896
|
zgc:153896 |
| chr17_+_8175998 | 4.35 |
ENSDART00000131200
|
myct1b
|
myc target 1b |
| chr11_-_34577034 | 4.31 |
ENSDART00000133302
ENSDART00000184367 |
pfkfb4a
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 4a |
| chr7_+_34290051 | 4.30 |
ENSDART00000123498
|
fibinb
|
fin bud initiation factor b |
| chr3_-_34547000 | 4.29 |
ENSDART00000166623
|
sept9a
|
septin 9a |
| chr4_+_16725960 | 4.25 |
ENSDART00000034441
|
tcp11l2
|
t-complex 11, testis-specific-like 2 |
| chr19_+_7124337 | 4.24 |
ENSDART00000031380
|
tap2a
|
transporter associated with antigen processing, subunit type a |
| chr8_-_1219815 | 4.21 |
ENSDART00000016800
ENSDART00000149969 |
znf367
|
zinc finger protein 367 |
| chr21_+_19648814 | 4.15 |
ENSDART00000048581
|
fgf10a
|
fibroblast growth factor 10a |
| chr14_-_25949951 | 4.13 |
ENSDART00000141304
|
sparc
|
secreted protein, acidic, cysteine-rich (osteonectin) |
| chr2_-_17393216 | 4.09 |
ENSDART00000123137
|
st3gal3b
|
ST3 beta-galactoside alpha-2,3-sialyltransferase 3b |
| chr2_+_47927026 | 4.08 |
ENSDART00000143023
|
ftr25
|
finTRIM family, member 25 |
| chr24_+_19518303 | 4.03 |
ENSDART00000027022
ENSDART00000056080 |
sulf1
|
sulfatase 1 |
| chr17_+_51746830 | 4.03 |
ENSDART00000184230
|
odc1
|
ornithine decarboxylase 1 |
| chr3_-_26204867 | 4.02 |
ENSDART00000103748
|
gdpd3a
|
glycerophosphodiester phosphodiesterase domain containing 3a |
| chr13_+_45476181 | 3.98 |
ENSDART00000045329
|
mgst3b
|
microsomal glutathione S-transferase 3b |
| chr5_-_16425781 | 3.97 |
ENSDART00000185624
ENSDART00000180617 |
slc39a14
|
solute carrier family 39 (zinc transporter), member 14 |
| chr19_+_43523690 | 3.94 |
ENSDART00000113031
|
wasf2
|
WAS protein family, member 2 |
| chr16_+_26449615 | 3.94 |
ENSDART00000039746
|
epb41b
|
erythrocyte membrane protein band 4.1b |
| chr18_+_48428713 | 3.76 |
ENSDART00000076861
|
fli1a
|
Fli-1 proto-oncogene, ETS transcription factor a |
| chr5_-_68795063 | 3.69 |
ENSDART00000016307
|
her1
|
hairy-related 1 |
| chr7_+_5976613 | 3.65 |
ENSDART00000173105
|
si:dkey-23a13.21
|
si:dkey-23a13.21 |
| chr14_-_41556720 | 3.57 |
ENSDART00000149244
|
itga6l
|
integrin, alpha 6, like |
| chr21_+_39941184 | 3.56 |
ENSDART00000133604
|
slc47a1
|
solute carrier family 47 (multidrug and toxin extrusion), member 1 |
| chr8_+_26059677 | 3.56 |
ENSDART00000009178
|
impdh2
|
IMP (inosine 5'-monophosphate) dehydrogenase 2 |
| chr23_-_45501177 | 3.56 |
ENSDART00000150103
|
col24a1
|
collagen type XXIV alpha 1 |
| chr23_+_25172682 | 3.43 |
ENSDART00000191197
ENSDART00000183497 |
si:dkey-151g10.3
|
si:dkey-151g10.3 |
| chr18_+_17155896 | 3.43 |
ENSDART00000140654
|
trhr2
|
thyrotropin releasing hormone receptor 2 |
| chr6_+_51932267 | 3.35 |
ENSDART00000156256
|
angpt4
|
angiopoietin 4 |
| chr14_+_29780113 | 3.30 |
ENSDART00000173195
|
zgc:153146
|
zgc:153146 |
| chr6_-_9695294 | 3.29 |
ENSDART00000162728
|
nop58
|
NOP58 ribonucleoprotein homolog (yeast) |
| chr7_+_17120811 | 3.23 |
ENSDART00000147140
|
prmt3
|
protein arginine methyltransferase 3 |
| chr18_+_18000695 | 3.21 |
ENSDART00000146898
|
si:ch211-212o1.2
|
si:ch211-212o1.2 |
| chr13_+_24263049 | 3.20 |
ENSDART00000135992
ENSDART00000088005 |
abcb10
|
ATP-binding cassette, sub-family B (MDR/TAP), member 10 |
| chr19_+_37509638 | 3.20 |
ENSDART00000139999
|
si:ch211-250g4.3
|
si:ch211-250g4.3 |
| chr1_-_17570013 | 3.18 |
ENSDART00000146946
|
acsl1a
|
acyl-CoA synthetase long chain family member 1a |
| chr8_+_23709280 | 3.17 |
ENSDART00000185842
|
ppardb
|
peroxisome proliferator-activated receptor delta b |
| chr17_-_26868169 | 3.16 |
ENSDART00000157204
|
si:dkey-221l4.10
|
si:dkey-221l4.10 |
| chr12_-_36070473 | 3.12 |
ENSDART00000165720
|
gpr142
|
G protein-coupled receptor 142 |
| chr22_+_5118361 | 3.12 |
ENSDART00000168371
ENSDART00000170222 ENSDART00000158846 |
mibp
|
muscle-specific beta 1 integrin binding protein |
| chr14_+_30568961 | 3.11 |
ENSDART00000184303
|
mrpl11
|
mitochondrial ribosomal protein L11 |
| chr23_-_35483163 | 3.10 |
ENSDART00000138660
ENSDART00000113643 ENSDART00000189269 |
fbxo25
|
F-box protein 25 |
| chr10_+_19525839 | 3.09 |
ENSDART00000162912
ENSDART00000158512 |
vsig8a
|
V-set and immunoglobulin domain containing 8a |
| chr2_+_9757453 | 3.07 |
ENSDART00000168972
|
pcyt1aa
|
phosphate cytidylyltransferase 1, choline, alpha a |
| chr19_+_40250682 | 3.07 |
ENSDART00000102888
|
cdk6
|
cyclin-dependent kinase 6 |
| chr2_-_6482240 | 3.04 |
ENSDART00000132623
|
rgs13
|
regulator of G protein signaling 13 |
| chr12_+_15078124 | 3.01 |
ENSDART00000187571
|
CR456635.1
|
|
| chr20_-_27390258 | 2.93 |
ENSDART00000010584
|
prph2l
|
peripherin 2, like |
| chr6_-_39275793 | 2.93 |
ENSDART00000180477
ENSDART00000148531 |
arhgef25b
|
Rho guanine nucleotide exchange factor (GEF) 25b |
Network of associatons between targets according to the STRING database.
First level regulatory network of gata1b+gata2a
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 21.5 | 128.9 | GO:0055005 | ventricular cardiac myofibril assembly(GO:0055005) |
| 20.7 | 103.7 | GO:0001773 | myeloid dendritic cell activation(GO:0001773) myeloid dendritic cell differentiation(GO:0043011) |
| 5.7 | 22.9 | GO:0003228 | atrial cardiac muscle tissue development(GO:0003228) |
| 5.2 | 26.2 | GO:0010873 | regulation of cholesterol esterification(GO:0010872) positive regulation of cholesterol esterification(GO:0010873) |
| 4.9 | 14.7 | GO:0043606 | formate metabolic process(GO:0015942) histidine catabolic process to glutamate and formamide(GO:0019556) histidine catabolic process to glutamate and formate(GO:0019557) formamide metabolic process(GO:0043606) |
| 4.0 | 12.0 | GO:0035971 | peptidyl-histidine dephosphorylation(GO:0035971) |
| 3.7 | 11.2 | GO:0003214 | cardiac left ventricle morphogenesis(GO:0003214) cardiac left ventricle formation(GO:0003218) |
| 3.7 | 47.5 | GO:0061026 | cardiac muscle tissue regeneration(GO:0061026) |
| 3.5 | 24.6 | GO:0030195 | negative regulation of blood coagulation(GO:0030195) negative regulation of wound healing(GO:0061045) negative regulation of hemostasis(GO:1900047) |
| 3.4 | 27.5 | GO:0019441 | tryptophan catabolic process to kynurenine(GO:0019441) |
| 3.3 | 9.9 | GO:0014722 | regulation of skeletal muscle contraction by calcium ion signaling(GO:0014722) regulation of skeletal muscle contraction by regulation of release of sequestered calcium ion(GO:0014809) |
| 3.0 | 18.1 | GO:0071800 | podosome assembly(GO:0071800) |
| 2.8 | 8.4 | GO:0045226 | extracellular polysaccharide biosynthetic process(GO:0045226) extracellular polysaccharide metabolic process(GO:0046379) |
| 2.6 | 7.8 | GO:0003156 | regulation of organ formation(GO:0003156) |
| 2.4 | 65.5 | GO:0042632 | cholesterol homeostasis(GO:0042632) sterol homeostasis(GO:0055092) |
| 2.2 | 8.7 | GO:0061033 | lung growth(GO:0060437) secretion by lung epithelial cell involved in lung growth(GO:0061033) |
| 1.7 | 6.6 | GO:0016332 | establishment or maintenance of polarity of embryonic epithelium(GO:0016332) |
| 1.6 | 6.5 | GO:0030224 | monocyte differentiation(GO:0030224) |
| 1.4 | 9.7 | GO:0048739 | cardiac muscle fiber development(GO:0048739) |
| 1.4 | 13.8 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 1.3 | 4.0 | GO:0033212 | iron assimilation(GO:0033212) |
| 1.3 | 7.6 | GO:2000114 | blood vessel maturation(GO:0001955) regulation of establishment of cell polarity(GO:2000114) |
| 1.2 | 4.8 | GO:0038161 | prolactin signaling pathway(GO:0038161) |
| 1.2 | 26.4 | GO:1990798 | pancreas regeneration(GO:1990798) |
| 1.2 | 9.3 | GO:0021910 | smoothened signaling pathway involved in ventral spinal cord patterning(GO:0021910) regulation of myotome development(GO:2000290) |
| 1.1 | 5.6 | GO:1901207 | regulation of heart looping(GO:1901207) |
| 1.1 | 13.2 | GO:0001678 | cellular glucose homeostasis(GO:0001678) |
| 1.0 | 51.4 | GO:0018298 | protein-chromophore linkage(GO:0018298) |
| 1.0 | 29.2 | GO:0032474 | otolith morphogenesis(GO:0032474) |
| 1.0 | 6.2 | GO:0038065 | collagen-activated tyrosine kinase receptor signaling pathway(GO:0038063) collagen-activated signaling pathway(GO:0038065) |
| 1.0 | 6.1 | GO:0035176 | social behavior(GO:0035176) intraspecies interaction between organisms(GO:0051703) |
| 1.0 | 5.0 | GO:0060397 | JAK-STAT cascade involved in growth hormone signaling pathway(GO:0060397) |
| 1.0 | 20.6 | GO:0015671 | oxygen transport(GO:0015671) |
| 1.0 | 13.3 | GO:0032264 | IMP salvage(GO:0032264) |
| 0.9 | 9.3 | GO:0003207 | cardiac chamber formation(GO:0003207) |
| 0.9 | 16.5 | GO:0046686 | response to cadmium ion(GO:0046686) |
| 0.9 | 7.0 | GO:0006567 | threonine metabolic process(GO:0006566) threonine catabolic process(GO:0006567) |
| 0.8 | 22.5 | GO:0006884 | cell volume homeostasis(GO:0006884) |
| 0.7 | 15.6 | GO:1901888 | epithelial cilium movement involved in determination of left/right asymmetry(GO:0060287) regulation of cell junction assembly(GO:1901888) |
| 0.7 | 2.7 | GO:0061015 | RNA import into nucleus(GO:0006404) snRNA import into nucleus(GO:0061015) |
| 0.6 | 20.7 | GO:0032355 | response to estradiol(GO:0032355) |
| 0.6 | 7.1 | GO:0002031 | G-protein coupled receptor internalization(GO:0002031) |
| 0.6 | 5.9 | GO:0042304 | regulation of fatty acid biosynthetic process(GO:0042304) |
| 0.6 | 2.8 | GO:0002025 | vasodilation by norepinephrine-epinephrine involved in regulation of systemic arterial blood pressure(GO:0002025) |
| 0.6 | 8.3 | GO:0031641 | regulation of myelination(GO:0031641) |
| 0.6 | 6.1 | GO:0019885 | antigen processing and presentation of endogenous peptide antigen(GO:0002483) antigen processing and presentation of endogenous antigen(GO:0019883) antigen processing and presentation of endogenous peptide antigen via MHC class I(GO:0019885) |
| 0.5 | 1.6 | GO:0046048 | pyrimidine ribonucleoside diphosphate metabolic process(GO:0009193) UDP metabolic process(GO:0046048) |
| 0.5 | 25.8 | GO:0010862 | positive regulation of pathway-restricted SMAD protein phosphorylation(GO:0010862) |
| 0.5 | 7.2 | GO:0015937 | coenzyme A biosynthetic process(GO:0015937) |
| 0.5 | 10.2 | GO:0042541 | hemoglobin biosynthetic process(GO:0042541) |
| 0.5 | 3.6 | GO:0006177 | GMP biosynthetic process(GO:0006177) GMP metabolic process(GO:0046037) |
| 0.5 | 16.3 | GO:0006749 | glutathione metabolic process(GO:0006749) |
| 0.5 | 2.0 | GO:2001244 | positive regulation of intrinsic apoptotic signaling pathway(GO:2001244) |
| 0.5 | 27.4 | GO:0003333 | amino acid transmembrane transport(GO:0003333) |
| 0.5 | 6.7 | GO:0034311 | diol metabolic process(GO:0034311) |
| 0.5 | 6.1 | GO:0010996 | response to auditory stimulus(GO:0010996) |
| 0.4 | 5.4 | GO:0048769 | sarcomerogenesis(GO:0048769) |
| 0.4 | 4.0 | GO:0033387 | putrescine biosynthetic process from ornithine(GO:0033387) |
| 0.4 | 6.3 | GO:0003323 | type B pancreatic cell development(GO:0003323) |
| 0.4 | 5.0 | GO:0042572 | retinol metabolic process(GO:0042572) |
| 0.4 | 3.2 | GO:0010887 | negative regulation of cholesterol storage(GO:0010887) |
| 0.4 | 9.8 | GO:0051923 | sulfation(GO:0051923) |
| 0.3 | 10.7 | GO:0032526 | response to retinoic acid(GO:0032526) |
| 0.3 | 3.1 | GO:0043697 | dedifferentiation(GO:0043696) cell dedifferentiation(GO:0043697) |
| 0.3 | 31.7 | GO:0071466 | cellular response to xenobiotic stimulus(GO:0071466) |
| 0.3 | 2.3 | GO:0070584 | mitochondrion morphogenesis(GO:0070584) |
| 0.3 | 4.7 | GO:0009651 | response to salt stress(GO:0009651) |
| 0.3 | 18.0 | GO:0043123 | positive regulation of I-kappaB kinase/NF-kappaB signaling(GO:0043123) |
| 0.3 | 4.7 | GO:0010765 | positive regulation of sodium ion transport(GO:0010765) positive regulation of potassium ion transport(GO:0043268) positive regulation of potassium ion transmembrane transport(GO:1901381) positive regulation of sodium ion transmembrane transport(GO:1902307) regulation of potassium ion import(GO:1903286) positive regulation of potassium ion import(GO:1903288) positive regulation of sodium ion transmembrane transporter activity(GO:2000651) |
| 0.3 | 9.1 | GO:0048665 | neuron fate specification(GO:0048665) |
| 0.3 | 4.3 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 0.3 | 3.9 | GO:0051127 | positive regulation of actin nucleation(GO:0051127) positive regulation of Arp2/3 complex-mediated actin nucleation(GO:2000601) |
| 0.3 | 1.3 | GO:0007624 | ultradian rhythm(GO:0007624) |
| 0.3 | 7.3 | GO:0019835 | cytolysis(GO:0019835) |
| 0.2 | 4.3 | GO:0035118 | embryonic pectoral fin morphogenesis(GO:0035118) |
| 0.2 | 5.7 | GO:0030512 | negative regulation of transforming growth factor beta receptor signaling pathway(GO:0030512) negative regulation of cellular response to transforming growth factor beta stimulus(GO:1903845) |
| 0.2 | 3.2 | GO:0018216 | peptidyl-arginine methylation(GO:0018216) |
| 0.2 | 5.4 | GO:0019432 | triglyceride biosynthetic process(GO:0019432) |
| 0.2 | 25.4 | GO:0001756 | somitogenesis(GO:0001756) |
| 0.2 | 7.7 | GO:0051016 | barbed-end actin filament capping(GO:0051016) |
| 0.2 | 2.9 | GO:0002574 | thrombocyte differentiation(GO:0002574) |
| 0.2 | 1.8 | GO:0021707 | cerebellar granular layer development(GO:0021681) cerebellar granular layer morphogenesis(GO:0021683) cerebellar granular layer formation(GO:0021684) cerebellar granule cell differentiation(GO:0021707) |
| 0.2 | 0.9 | GO:0090342 | regulation of cell aging(GO:0090342) |
| 0.2 | 2.9 | GO:0036092 | phosphatidylinositol-3-phosphate biosynthetic process(GO:0036092) |
| 0.1 | 11.8 | GO:0008543 | fibroblast growth factor receptor signaling pathway(GO:0008543) |
| 0.1 | 13.3 | GO:0002181 | cytoplasmic translation(GO:0002181) |
| 0.1 | 0.6 | GO:0034729 | histone H3-K79 methylation(GO:0034729) regulation of transcription regulatory region DNA binding(GO:2000677) |
| 0.1 | 1.4 | GO:0090050 | positive regulation of blood vessel endothelial cell migration(GO:0043536) positive regulation of cell migration involved in sprouting angiogenesis(GO:0090050) |
| 0.1 | 8.8 | GO:1902593 | protein import into nucleus(GO:0006606) protein targeting to nucleus(GO:0044744) single-organism nuclear import(GO:1902593) |
| 0.1 | 2.1 | GO:0008272 | sulfate transport(GO:0008272) |
| 0.1 | 2.0 | GO:0098962 | regulation of postsynaptic neurotransmitter receptor activity(GO:0098962) |
| 0.1 | 2.5 | GO:0042476 | odontogenesis(GO:0042476) |
| 0.1 | 0.8 | GO:2000251 | positive regulation of actin cytoskeleton reorganization(GO:2000251) |
| 0.1 | 9.0 | GO:0008284 | positive regulation of cell proliferation(GO:0008284) |
| 0.1 | 23.1 | GO:0009116 | nucleoside metabolic process(GO:0009116) |
| 0.1 | 12.6 | GO:0006417 | regulation of translation(GO:0006417) |
| 0.1 | 3.9 | GO:0030866 | cortical actin cytoskeleton organization(GO:0030866) |
| 0.1 | 5.2 | GO:0007030 | Golgi organization(GO:0007030) |
| 0.1 | 7.7 | GO:0034976 | response to endoplasmic reticulum stress(GO:0034976) |
| 0.1 | 0.6 | GO:0045732 | positive regulation of protein catabolic process(GO:0045732) |
| 0.0 | 5.3 | GO:0010466 | negative regulation of peptidase activity(GO:0010466) |
| 0.0 | 8.7 | GO:0009952 | anterior/posterior pattern specification(GO:0009952) |
| 0.0 | 4.7 | GO:0002040 | sprouting angiogenesis(GO:0002040) |
| 0.0 | 4.5 | GO:0016485 | protein processing(GO:0016485) |
| 0.0 | 4.8 | GO:0031099 | regeneration(GO:0031099) |
| 0.0 | 1.4 | GO:0046856 | phosphatidylinositol dephosphorylation(GO:0046856) |
| 0.0 | 29.0 | GO:0006508 | proteolysis(GO:0006508) |
| 0.0 | 0.9 | GO:0006171 | cAMP biosynthetic process(GO:0006171) |
| 0.0 | 3.1 | GO:0043043 | translation(GO:0006412) peptide biosynthetic process(GO:0043043) |
| 0.0 | 0.7 | GO:0046513 | ceramide biosynthetic process(GO:0046513) |
| 0.0 | 3.3 | GO:0042254 | ribosome biogenesis(GO:0042254) |
| 0.0 | 3.8 | GO:0000377 | RNA splicing, via transesterification reactions(GO:0000375) RNA splicing, via transesterification reactions with bulged adenosine as nucleophile(GO:0000377) mRNA splicing, via spliceosome(GO:0000398) |
| 0.0 | 1.0 | GO:0007229 | integrin-mediated signaling pathway(GO:0007229) |
| 0.0 | 1.3 | GO:0000724 | double-strand break repair via homologous recombination(GO:0000724) |
| 0.0 | 0.2 | GO:0010811 | positive regulation of cell-substrate adhesion(GO:0010811) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.8 | 8.3 | GO:0043220 | compact myelin(GO:0043218) Schmidt-Lanterman incisure(GO:0043220) |
| 2.6 | 18.1 | GO:0001891 | phagocytic cup(GO:0001891) |
| 2.0 | 26.2 | GO:0042627 | chylomicron(GO:0042627) |
| 1.7 | 20.6 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 1.4 | 9.9 | GO:0033018 | sarcoplasmic reticulum lumen(GO:0033018) |
| 1.2 | 3.6 | GO:0005592 | collagen type XI trimer(GO:0005592) |
| 1.0 | 8.8 | GO:0044613 | nuclear pore central transport channel(GO:0044613) |
| 0.9 | 5.6 | GO:0033181 | plasma membrane proton-transporting V-type ATPase complex(GO:0033181) |
| 0.7 | 2.6 | GO:0098651 | collagen type IV trimer(GO:0005587) network-forming collagen trimer(GO:0098642) collagen network(GO:0098645) basement membrane collagen trimer(GO:0098651) |
| 0.6 | 30.3 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.6 | 14.7 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.5 | 3.3 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.5 | 7.3 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.4 | 12.6 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.3 | 55.1 | GO:0030017 | sarcomere(GO:0030017) |
| 0.3 | 40.4 | GO:0016459 | myosin complex(GO:0016459) |
| 0.3 | 6.6 | GO:0001917 | photoreceptor inner segment(GO:0001917) |
| 0.3 | 2.9 | GO:0005943 | phosphatidylinositol 3-kinase complex, class IA(GO:0005943) |
| 0.3 | 11.3 | GO:0044853 | caveola(GO:0005901) plasma membrane raft(GO:0044853) |
| 0.2 | 3.9 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.2 | 13.3 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.2 | 1.3 | GO:0030915 | Smc5-Smc6 complex(GO:0030915) |
| 0.2 | 2.9 | GO:0005885 | Arp2/3 protein complex(GO:0005885) |
| 0.2 | 0.8 | GO:0097433 | dense body(GO:0097433) |
| 0.2 | 7.7 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.1 | 66.8 | GO:0009986 | cell surface(GO:0009986) |
| 0.1 | 7.6 | GO:0016342 | catenin complex(GO:0016342) |
| 0.1 | 6.2 | GO:0005793 | endoplasmic reticulum-Golgi intermediate compartment(GO:0005793) |
| 0.1 | 5.0 | GO:0005811 | lipid particle(GO:0005811) |
| 0.1 | 33.3 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.1 | 8.1 | GO:0048475 | membrane coat(GO:0030117) coated membrane(GO:0048475) |
| 0.1 | 3.1 | GO:0005762 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.1 | 3.1 | GO:0016592 | mediator complex(GO:0016592) |
| 0.1 | 18.9 | GO:0005813 | centrosome(GO:0005813) |
| 0.1 | 74.5 | GO:0005615 | extracellular space(GO:0005615) |
| 0.1 | 4.0 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.0 | 5.4 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.0 | 4.5 | GO:0000786 | nucleosome(GO:0000786) |
| 0.0 | 2.0 | GO:0098978 | glutamatergic synapse(GO:0098978) |
| 0.0 | 6.7 | GO:0005635 | nuclear envelope(GO:0005635) |
| 0.0 | 1.0 | GO:0008305 | integrin complex(GO:0008305) |
| 0.0 | 3.8 | GO:0016607 | nuclear speck(GO:0016607) |
| 0.0 | 2.7 | GO:0034705 | voltage-gated potassium channel complex(GO:0008076) potassium channel complex(GO:0034705) |
| 0.0 | 44.6 | GO:0005829 | cytosol(GO:0005829) |
| 0.0 | 2.6 | GO:0014069 | postsynaptic density(GO:0014069) postsynaptic specialization(GO:0099572) |
| 0.0 | 1.0 | GO:0005930 | axoneme(GO:0005930) |
| 0.0 | 7.7 | GO:0005789 | endoplasmic reticulum membrane(GO:0005789) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 9.2 | 27.5 | GO:0004833 | tryptophan 2,3-dioxygenase activity(GO:0004833) |
| 5.2 | 26.2 | GO:0060228 | phosphatidylcholine-sterol O-acyltransferase activator activity(GO:0060228) |
| 5.1 | 20.5 | GO:0015355 | secondary active monocarboxylate transmembrane transporter activity(GO:0015355) |
| 4.1 | 37.1 | GO:0004731 | purine-nucleoside phosphorylase activity(GO:0004731) |
| 3.7 | 14.7 | GO:0016841 | ammonia-lyase activity(GO:0016841) |
| 3.2 | 42.2 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 3.0 | 12.0 | GO:0101006 | protein histidine phosphatase activity(GO:0101006) |
| 3.0 | 20.7 | GO:0045735 | nutrient reservoir activity(GO:0045735) |
| 2.8 | 8.4 | GO:0050501 | hyaluronan synthase activity(GO:0050501) |
| 2.8 | 11.1 | GO:0004064 | arylesterase activity(GO:0004064) |
| 2.4 | 12.2 | GO:0031769 | glucagon receptor binding(GO:0031769) |
| 2.2 | 13.2 | GO:0005536 | glucokinase activity(GO:0004340) hexokinase activity(GO:0004396) glucose binding(GO:0005536) fructokinase activity(GO:0008865) mannokinase activity(GO:0019158) |
| 1.9 | 15.6 | GO:0072518 | Rho-dependent protein serine/threonine kinase activity(GO:0072518) |
| 1.8 | 17.6 | GO:0016936 | galactoside binding(GO:0016936) |
| 1.7 | 20.6 | GO:0031720 | haptoglobin binding(GO:0031720) |
| 1.6 | 19.7 | GO:0005111 | type 2 fibroblast growth factor receptor binding(GO:0005111) |
| 1.5 | 30.3 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) alpha-actinin binding(GO:0051393) |
| 1.5 | 16.4 | GO:0031013 | troponin C binding(GO:0030172) troponin I binding(GO:0031013) |
| 1.4 | 7.0 | GO:0008743 | L-threonine 3-dehydrogenase activity(GO:0008743) |
| 1.3 | 9.3 | GO:0050262 | ribosylnicotinamide kinase activity(GO:0050262) |
| 1.3 | 4.0 | GO:0015086 | cadmium ion transmembrane transporter activity(GO:0015086) |
| 1.3 | 21.1 | GO:0042910 | xenobiotic transporter activity(GO:0042910) |
| 1.2 | 7.4 | GO:0004692 | cGMP-dependent protein kinase activity(GO:0004692) |
| 1.2 | 13.3 | GO:0003876 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 1.2 | 4.8 | GO:0004925 | prolactin receptor activity(GO:0004925) |
| 1.2 | 7.2 | GO:0004594 | pantothenate kinase activity(GO:0004594) |
| 1.2 | 10.7 | GO:0016286 | small conductance calcium-activated potassium channel activity(GO:0016286) |
| 1.2 | 10.7 | GO:0044323 | retinoic acid-responsive element binding(GO:0044323) |
| 1.1 | 9.1 | GO:0015168 | glycerol transmembrane transporter activity(GO:0015168) |
| 1.0 | 14.5 | GO:0008266 | poly(U) RNA binding(GO:0008266) |
| 1.0 | 13.8 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.9 | 5.6 | GO:0004997 | thyrotropin-releasing hormone receptor activity(GO:0004997) |
| 0.9 | 22.5 | GO:0015377 | cation:chloride symporter activity(GO:0015377) potassium:chloride symporter activity(GO:0015379) |
| 0.9 | 8.8 | GO:0050700 | CARD domain binding(GO:0050700) |
| 0.9 | 6.1 | GO:0004962 | endothelin receptor activity(GO:0004962) |
| 0.9 | 6.9 | GO:0016803 | ether hydrolase activity(GO:0016803) |
| 0.8 | 7.2 | GO:0004690 | cyclic nucleotide-dependent protein kinase activity(GO:0004690) |
| 0.7 | 2.8 | GO:0051380 | norepinephrine binding(GO:0051380) |
| 0.7 | 19.4 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.6 | 5.0 | GO:0004745 | retinol dehydrogenase activity(GO:0004745) |
| 0.6 | 24.6 | GO:0004869 | cysteine-type endopeptidase inhibitor activity(GO:0004869) |
| 0.6 | 3.1 | GO:0004105 | choline-phosphate cytidylyltransferase activity(GO:0004105) |
| 0.6 | 20.3 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.6 | 3.6 | GO:0003938 | IMP dehydrogenase activity(GO:0003938) |
| 0.6 | 7.7 | GO:0016864 | protein disulfide isomerase activity(GO:0003756) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
| 0.6 | 9.9 | GO:0005104 | fibroblast growth factor receptor binding(GO:0005104) |
| 0.6 | 4.1 | GO:0008118 | N-acetyllactosaminide alpha-2,3-sialyltransferase activity(GO:0008118) |
| 0.6 | 8.3 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.5 | 9.3 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 0.5 | 1.6 | GO:0009041 | uridylate kinase activity(GO:0009041) |
| 0.5 | 3.1 | GO:0070180 | large ribosomal subunit rRNA binding(GO:0070180) |
| 0.5 | 53.6 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.5 | 71.1 | GO:0016820 | hydrolase activity, acting on acid anhydrides, catalyzing transmembrane movement of substances(GO:0016820) ATPase activity, coupled to transmembrane movement of substances(GO:0042626) ATPase activity, coupled to movement of substances(GO:0043492) |
| 0.5 | 23.5 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.4 | 4.0 | GO:0004586 | ornithine decarboxylase activity(GO:0004586) |
| 0.4 | 3.9 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 0.4 | 5.6 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.4 | 10.1 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.3 | 20.5 | GO:0009055 | electron carrier activity(GO:0009055) |
| 0.3 | 4.3 | GO:0003873 | 6-phosphofructo-2-kinase activity(GO:0003873) |
| 0.3 | 4.7 | GO:0019869 | chloride channel inhibitor activity(GO:0019869) potassium channel inhibitor activity(GO:0019870) |
| 0.3 | 9.0 | GO:0030971 | receptor tyrosine kinase binding(GO:0030971) |
| 0.3 | 5.8 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
| 0.2 | 27.4 | GO:0015171 | amino acid transmembrane transporter activity(GO:0015171) |
| 0.2 | 4.7 | GO:0008253 | 5'-nucleotidase activity(GO:0008253) |
| 0.2 | 3.1 | GO:0008353 | RNA polymerase II carboxy-terminal domain kinase activity(GO:0008353) |
| 0.2 | 6.6 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.2 | 5.3 | GO:0030414 | peptidase inhibitor activity(GO:0030414) |
| 0.2 | 3.2 | GO:0016274 | arginine N-methyltransferase activity(GO:0016273) protein-arginine N-methyltransferase activity(GO:0016274) |
| 0.2 | 37.8 | GO:0005125 | cytokine activity(GO:0005125) |
| 0.2 | 20.7 | GO:0004879 | RNA polymerase II transcription factor activity, ligand-activated sequence-specific DNA binding(GO:0004879) |
| 0.2 | 5.3 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
| 0.2 | 2.0 | GO:0070530 | K63-linked polyubiquitin binding(GO:0070530) |
| 0.2 | 3.3 | GO:0030515 | snoRNA binding(GO:0030515) |
| 0.2 | 5.4 | GO:0008195 | phosphatidate phosphatase activity(GO:0008195) |
| 0.2 | 8.9 | GO:0044325 | ion channel binding(GO:0044325) |
| 0.2 | 0.8 | GO:0098973 | structural constituent of postsynaptic actin cytoskeleton(GO:0098973) |
| 0.2 | 2.9 | GO:0035005 | 1-phosphatidylinositol-4-phosphate 3-kinase activity(GO:0035005) |
| 0.2 | 40.4 | GO:0003774 | motor activity(GO:0003774) |
| 0.1 | 0.6 | GO:0031151 | histone methyltransferase activity (H3-K79 specific)(GO:0031151) |
| 0.1 | 17.6 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.1 | 5.7 | GO:0004180 | carboxypeptidase activity(GO:0004180) |
| 0.1 | 1.4 | GO:0052629 | phosphatidylinositol-3,5-bisphosphate 3-phosphatase activity(GO:0052629) |
| 0.1 | 128.5 | GO:0008270 | zinc ion binding(GO:0008270) |
| 0.1 | 2.2 | GO:0002039 | p53 binding(GO:0002039) |
| 0.1 | 30.1 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.1 | 9.8 | GO:0008146 | sulfotransferase activity(GO:0008146) |
| 0.1 | 5.0 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.1 | 6.5 | GO:0016712 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced flavin or flavoprotein as one donor, and incorporation of one atom of oxygen(GO:0016712) |
| 0.1 | 1.3 | GO:0008026 | ATP-dependent DNA helicase activity(GO:0004003) ATP-dependent helicase activity(GO:0008026) purine NTP-dependent helicase activity(GO:0070035) |
| 0.1 | 3.0 | GO:0005518 | collagen binding(GO:0005518) |
| 0.1 | 5.7 | GO:0046332 | SMAD binding(GO:0046332) |
| 0.1 | 7.6 | GO:0045296 | cadherin binding(GO:0045296) |
| 0.1 | 40.0 | GO:0003779 | actin binding(GO:0003779) |
| 0.1 | 2.1 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.1 | 1.2 | GO:0019531 | oxalate transmembrane transporter activity(GO:0019531) |
| 0.1 | 13.3 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.1 | 2.4 | GO:0004707 | MAP kinase activity(GO:0004707) |
| 0.1 | 2.7 | GO:0005251 | delayed rectifier potassium channel activity(GO:0005251) |
| 0.1 | 0.2 | GO:0015349 | thyroid hormone transmembrane transporter activity(GO:0015349) |
| 0.1 | 43.2 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.1 | 4.5 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.1 | 19.3 | GO:0005543 | phospholipid binding(GO:0005543) |
| 0.1 | 3.4 | GO:0042803 | protein homodimerization activity(GO:0042803) |
| 0.0 | 0.7 | GO:0050291 | sphingosine N-acyltransferase activity(GO:0050291) |
| 0.0 | 8.5 | GO:0046982 | protein heterodimerization activity(GO:0046982) |
| 0.0 | 6.7 | GO:0016705 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen(GO:0016705) |
| 0.0 | 0.6 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 0.0 | 1.4 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 0.1 | GO:0047676 | arachidonate-CoA ligase activity(GO:0047676) |
| 0.0 | 5.0 | GO:0008092 | cytoskeletal protein binding(GO:0008092) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.3 | 18.0 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 1.4 | 24.6 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.7 | 30.0 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.7 | 11.3 | PID WNT CANONICAL PATHWAY | Canonical Wnt signaling pathway |
| 0.5 | 7.6 | PID S1P S1P2 PATHWAY | S1P2 pathway |
| 0.4 | 17.5 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.4 | 14.0 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.4 | 62.9 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.3 | 2.6 | ST IL 13 PATHWAY | Interleukin 13 (IL-13) Pathway |
| 0.2 | 3.1 | PID IL2 STAT5 PATHWAY | IL2 signaling events mediated by STAT5 |
| 0.2 | 2.7 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
| 0.2 | 3.6 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.2 | 9.1 | PID FGF PATHWAY | FGF signaling pathway |
| 0.2 | 2.2 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.2 | 6.9 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.2 | 8.1 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.1 | 7.7 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.1 | 2.0 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.1 | 15.0 | NABA SECRETED FACTORS | Genes encoding secreted soluble factors |
| 0.1 | 2.0 | PID ATF2 PATHWAY | ATF-2 transcription factor network |
| 0.1 | 1.8 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.1 | 4.0 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.0 | 1.4 | ST T CELL SIGNAL TRANSDUCTION | T Cell Signal Transduction |
| 0.0 | 0.9 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.5 | 65.5 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 3.0 | 9.1 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 2.6 | 17.9 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 1.6 | 24.6 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 1.3 | 11.3 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 1.2 | 12.3 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 1.1 | 27.4 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 1.1 | 103.7 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 1.0 | 9.1 | REACTOME FGFR4 LIGAND BINDING AND ACTIVATION | Genes involved in FGFR4 ligand binding and activation |
| 0.9 | 17.5 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.9 | 10.1 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.7 | 11.1 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.6 | 8.4 | REACTOME HYALURONAN METABOLISM | Genes involved in Hyaluronan metabolism |
| 0.6 | 5.0 | REACTOME PROLACTIN RECEPTOR SIGNALING | Genes involved in Prolactin receptor signaling |
| 0.5 | 5.9 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GIP | Genes involved in Synthesis, Secretion, and Inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) |
| 0.5 | 5.8 | REACTOME CD28 DEPENDENT PI3K AKT SIGNALING | Genes involved in CD28 dependent PI3K/Akt signaling |
| 0.5 | 1.4 | REACTOME TRANSLOCATION OF ZAP 70 TO IMMUNOLOGICAL SYNAPSE | Genes involved in Translocation of ZAP-70 to Immunological synapse |
| 0.4 | 20.9 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.3 | 9.0 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.3 | 7.6 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.3 | 0.8 | REACTOME FGFR1 LIGAND BINDING AND ACTIVATION | Genes involved in FGFR1 ligand binding and activation |
| 0.3 | 10.8 | REACTOME NOD1 2 SIGNALING PATHWAY | Genes involved in NOD1/2 Signaling Pathway |
| 0.3 | 3.6 | REACTOME PURINE RIBONUCLEOSIDE MONOPHOSPHATE BIOSYNTHESIS | Genes involved in Purine ribonucleoside monophosphate biosynthesis |
| 0.2 | 3.2 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.2 | 6.7 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.2 | 2.0 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.2 | 2.7 | REACTOME MICRORNA MIRNA BIOGENESIS | Genes involved in MicroRNA (miRNA) Biogenesis |
| 0.1 | 10.7 | REACTOME POTASSIUM CHANNELS | Genes involved in Potassium Channels |
| 0.1 | 6.2 | REACTOME EXTRACELLULAR MATRIX ORGANIZATION | Genes involved in Extracellular matrix organization |
| 0.1 | 13.3 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.1 | 4.0 | REACTOME REGULATION OF ORNITHINE DECARBOXYLASE ODC | Genes involved in Regulation of ornithine decarboxylase (ODC) |
| 0.1 | 3.1 | REACTOME G1 PHASE | Genes involved in G1 Phase |
| 0.1 | 1.4 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.1 | 6.6 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.0 | 0.9 | REACTOME APOPTOTIC CLEAVAGE OF CELLULAR PROTEINS | Genes involved in Apoptotic cleavage of cellular proteins |
| 0.0 | 2.8 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |