Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
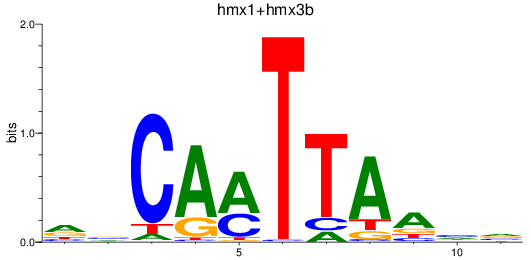
Results for hmx1+hmx3b
Z-value: 0.31
Transcription factors associated with hmx1+hmx3b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
hmx1
|
ENSDARG00000095651 | H6 family homeobox 1 |
|
hmx3b
|
ENSDARG00000096716 | H6 family homeobox 3b |
|
hmx1
|
ENSDARG00000109725 | H6 family homeobox 1 |
|
hmx1
|
ENSDARG00000111765 | H6 family homeobox 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| hmx3b | dr11_v1_chr12_-_49154934_49154934 | -0.25 | 1.7e-02 | Click! |
| hmx1 | dr11_v1_chr1_-_40914752_40914752 | -0.08 | 4.3e-01 | Click! |
Activity profile of hmx1+hmx3b motif
Sorted Z-values of hmx1+hmx3b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_+_9295244 | 1.79 |
ENSDART00000132255
ENSDART00000144299 |
si:ch73-15n24.1
|
si:ch73-15n24.1 |
| chr20_-_23426339 | 0.96 |
ENSDART00000004625
|
zar1
|
zygote arrest 1 |
| chr11_+_18183220 | 0.95 |
ENSDART00000113468
|
LO018315.10
|
|
| chr13_-_50108337 | 0.91 |
ENSDART00000133308
|
nid1a
|
nidogen 1a |
| chr5_-_69620722 | 0.90 |
ENSDART00000097248
|
aldh2.2
|
aldehyde dehydrogenase 2 family (mitochondrial), tandem duplicate 2 |
| chr25_+_3326885 | 0.82 |
ENSDART00000104866
|
ldhbb
|
lactate dehydrogenase Bb |
| chr11_+_18157260 | 0.82 |
ENSDART00000144659
|
zgc:173545
|
zgc:173545 |
| chr18_-_5595546 | 0.81 |
ENSDART00000191825
|
cyp1a
|
cytochrome P450, family 1, subfamily A |
| chr10_-_44560165 | 0.81 |
ENSDART00000181217
ENSDART00000076084 |
npm2b
|
nucleophosmin/nucleoplasmin, 2b |
| chr11_+_18130300 | 0.79 |
ENSDART00000169146
|
zgc:175135
|
zgc:175135 |
| chr11_-_37691449 | 0.75 |
ENSDART00000185340
|
zgc:158258
|
zgc:158258 |
| chr25_+_3327071 | 0.69 |
ENSDART00000136131
ENSDART00000133243 |
ldhbb
|
lactate dehydrogenase Bb |
| chr3_+_22327738 | 0.69 |
ENSDART00000055675
|
gh1
|
growth hormone 1 |
| chr22_-_20695237 | 0.67 |
ENSDART00000112722
|
org
|
oogenesis-related gene |
| chr22_+_16293071 | 0.64 |
ENSDART00000170960
|
dbt
|
dihydrolipoamide branched chain transacylase E2 |
| chr5_+_37032038 | 0.56 |
ENSDART00000045036
|
nfkbib
|
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, beta |
| chr2_-_17115256 | 0.52 |
ENSDART00000190488
|
pif1
|
PIF1 5'-to-3' DNA helicase homolog (S. cerevisiae) |
| chr22_+_1911269 | 0.47 |
ENSDART00000164158
ENSDART00000168205 |
znf1156
|
zinc finger protein 1156 |
| chrM_+_8130 | 0.46 |
ENSDART00000093609
|
mt-co2
|
cytochrome c oxidase II, mitochondrial |
| chr2_+_16652238 | 0.46 |
ENSDART00000091351
|
gk5
|
glycerol kinase 5 (putative) |
| chr1_-_56032619 | 0.45 |
ENSDART00000143793
|
c3a.4
|
complement component c3a, duplicate 4 |
| chr5_+_32835219 | 0.40 |
ENSDART00000140832
ENSDART00000186055 |
si:ch211-208h16.4
|
si:ch211-208h16.4 |
| chr6_+_11681011 | 0.39 |
ENSDART00000151447
ENSDART00000151618 |
calcrlb
|
calcitonin receptor-like b |
| chr5_+_24287927 | 0.39 |
ENSDART00000143563
|
zdhhc23a
|
zinc finger, DHHC-type containing 23a |
| chr7_+_17443567 | 0.39 |
ENSDART00000060383
|
nitr2b
|
novel immune-type receptor 2b |
| chr10_+_16069987 | 0.37 |
ENSDART00000043936
|
megf10
|
multiple EGF-like-domains 10 |
| chr6_-_37422841 | 0.36 |
ENSDART00000138351
|
cth
|
cystathionase (cystathionine gamma-lyase) |
| chr5_-_25645593 | 0.36 |
ENSDART00000044471
|
tmc2b
|
transmembrane channel-like 2b |
| chr10_+_40660772 | 0.35 |
ENSDART00000148007
|
taar19l
|
trace amine associated receptor 19l |
| chr6_-_40352215 | 0.35 |
ENSDART00000103992
|
ttll3
|
tubulin tyrosine ligase-like family, member 3 |
| chr13_+_30169681 | 0.35 |
ENSDART00000138326
|
pald1b
|
phosphatase domain containing, paladin 1b |
| chr23_+_9353552 | 0.34 |
ENSDART00000163298
|
BX511246.1
|
|
| chrM_+_4993 | 0.33 |
ENSDART00000093600
|
mt-nd2
|
NADH dehydrogenase 2, mitochondrial |
| chr20_+_37295006 | 0.32 |
ENSDART00000153137
|
cx23
|
connexin 23 |
| chr14_-_9128919 | 0.31 |
ENSDART00000108641
|
sh2d1ab
|
SH2 domain containing 1A duplicate b |
| chr19_+_43604256 | 0.31 |
ENSDART00000151080
ENSDART00000110305 |
si:ch211-199g17.9
|
si:ch211-199g17.9 |
| chr18_-_25568994 | 0.31 |
ENSDART00000133029
|
si:ch211-13k12.2
|
si:ch211-13k12.2 |
| chr2_+_24781026 | 0.30 |
ENSDART00000145692
|
pde4ca
|
phosphodiesterase 4C, cAMP-specific a |
| chr21_+_21702623 | 0.30 |
ENSDART00000136840
|
or125-2
|
odorant receptor, family E, subfamily 125, member 2 |
| chr17_+_7510220 | 0.28 |
ENSDART00000192648
|
klhl10b.1
|
kelch-like family member 10b, tandem duplicate 1 |
| chr18_+_44080948 | 0.28 |
ENSDART00000191036
|
BX908388.2
|
|
| chr2_-_25159309 | 0.28 |
ENSDART00000137290
|
si:dkey-223d7.6
|
si:dkey-223d7.6 |
| chr25_-_10503043 | 0.27 |
ENSDART00000155404
|
cox8b
|
cytochrome c oxidase subunit 8b |
| chr4_-_72609735 | 0.26 |
ENSDART00000174299
ENSDART00000159227 |
si:cabz01054394.6
|
si:cabz01054394.6 |
| chr11_+_30244356 | 0.26 |
ENSDART00000036050
ENSDART00000150080 |
rs1a
|
retinoschisin 1a |
| chr15_+_33696559 | 0.26 |
ENSDART00000188248
ENSDART00000161951 |
hdac12
|
histone deacetylase 12 |
| chr7_+_71683853 | 0.25 |
ENSDART00000163002
|
emilin2b
|
elastin microfibril interfacer 2b |
| chr22_-_4649238 | 0.25 |
ENSDART00000125302
|
fbn2b
|
fibrillin 2b |
| chr10_-_39154594 | 0.24 |
ENSDART00000148825
|
slc37a4b
|
solute carrier family 37 (glucose-6-phosphate transporter), member 4b |
| chr1_-_10577945 | 0.23 |
ENSDART00000179237
ENSDART00000040502 ENSDART00000186876 |
trpc5a
|
transient receptor potential cation channel, subfamily C, member 5a |
| chr15_+_1134870 | 0.23 |
ENSDART00000155392
|
p2ry13
|
purinergic receptor P2Y13 |
| chr10_+_40690411 | 0.23 |
ENSDART00000146865
|
taar19m
|
trace amine associated receptor 19m |
| chr21_-_37973819 | 0.22 |
ENSDART00000133405
|
ripply1
|
ripply transcriptional repressor 1 |
| chr24_+_26134209 | 0.22 |
ENSDART00000038824
|
tmtopsb
|
teleost multiple tissue opsin b |
| chr8_+_22472584 | 0.22 |
ENSDART00000138303
|
si:dkey-23c22.9
|
si:dkey-23c22.9 |
| chr19_-_6239248 | 0.21 |
ENSDART00000014127
|
pou2f2a
|
POU class 2 homeobox 2a |
| chr5_+_43782267 | 0.19 |
ENSDART00000130355
|
nos2a
|
nitric oxide synthase 2a, inducible |
| chr18_-_33080454 | 0.19 |
ENSDART00000191907
|
v2ra18
|
vomeronasal 2 receptor, a18 |
| chr8_-_19649617 | 0.18 |
ENSDART00000189033
|
fam78bb
|
family with sequence similarity 78, member B b |
| chr4_+_20576857 | 0.17 |
ENSDART00000125340
|
BX248410.2
|
|
| chr21_-_14966718 | 0.16 |
ENSDART00000151200
|
mmp17a
|
matrix metallopeptidase 17a |
| chr23_-_36857964 | 0.16 |
ENSDART00000188822
ENSDART00000134061 ENSDART00000093061 |
hipk1a
|
homeodomain interacting protein kinase 1a |
| chr20_+_28861629 | 0.16 |
ENSDART00000187274
ENSDART00000047826 |
slc10a1
|
solute carrier family 10 (sodium/bile acid cotransporter), member 1 |
| chr23_-_18913032 | 0.16 |
ENSDART00000136678
|
si:ch211-209j10.6
|
si:ch211-209j10.6 |
| chr22_-_6988102 | 0.15 |
ENSDART00000185618
|
fgfr1bl
|
fibroblast growth factor receptor 1b, like |
| chr10_-_39153959 | 0.15 |
ENSDART00000150193
ENSDART00000111362 |
slc37a4b
|
solute carrier family 37 (glucose-6-phosphate transporter), member 4b |
| chr6_-_40697585 | 0.15 |
ENSDART00000113196
|
si:ch211-157b11.14
|
si:ch211-157b11.14 |
| chr20_+_28861435 | 0.14 |
ENSDART00000142678
|
slc10a1
|
solute carrier family 10 (sodium/bile acid cotransporter), member 1 |
| chr21_-_14803366 | 0.14 |
ENSDART00000190872
|
si:dkey-11o18.5
|
si:dkey-11o18.5 |
| chr4_-_4751981 | 0.13 |
ENSDART00000147436
ENSDART00000092984 ENSDART00000158466 |
creb3l2
|
cAMP responsive element binding protein 3-like 2 |
| chr5_+_32009542 | 0.13 |
ENSDART00000182025
ENSDART00000179879 |
scai
|
suppressor of cancer cell invasion |
| chr15_-_14884332 | 0.12 |
ENSDART00000165237
|
si:ch211-24o8.4
|
si:ch211-24o8.4 |
| chr8_+_7097740 | 0.12 |
ENSDART00000159670
|
abtb1
|
ankyrin repeat and BTB (POZ) domain containing 1 |
| chr18_-_20458412 | 0.12 |
ENSDART00000012241
|
kif23
|
kinesin family member 23 |
| chr6_+_4528631 | 0.11 |
ENSDART00000122042
|
RNF219
|
ring finger protein 219 |
| chr23_+_38953545 | 0.11 |
ENSDART00000184621
|
ATP9A
|
ATPase phospholipid transporting 9A (putative) |
| chr4_-_837768 | 0.09 |
ENSDART00000185280
ENSDART00000135618 |
sobpb
|
sine oculis binding protein homolog (Drosophila) b |
| chr1_-_23596391 | 0.09 |
ENSDART00000155184
|
lcorl
|
ligand dependent nuclear receptor corepressor-like |
| chr2_-_42827336 | 0.09 |
ENSDART00000140913
|
adcy8
|
adenylate cyclase 8 (brain) |
| chr22_+_1313046 | 0.08 |
ENSDART00000170421
|
si:ch73-138e16.6
|
si:ch73-138e16.6 |
| chr16_-_42965192 | 0.07 |
ENSDART00000113714
|
mtx1a
|
metaxin 1a |
| chr8_-_23194408 | 0.06 |
ENSDART00000141090
|
si:ch211-196c10.15
|
si:ch211-196c10.15 |
| chr5_-_42272517 | 0.04 |
ENSDART00000137692
ENSDART00000164363 |
si:ch211-207c6.2
|
si:ch211-207c6.2 |
| chr16_-_6391275 | 0.04 |
ENSDART00000060550
|
ndufb9
|
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 9 |
| chr18_+_11970987 | 0.04 |
ENSDART00000144111
|
si:dkeyp-2c8.3
|
si:dkeyp-2c8.3 |
| chr9_-_36924388 | 0.03 |
ENSDART00000059756
|
ralba
|
v-ral simian leukemia viral oncogene homolog Ba (ras related) |
| chr4_+_13931733 | 0.02 |
ENSDART00000141742
ENSDART00000067175 |
pphln1
|
periphilin 1 |
| chr8_+_23194212 | 0.02 |
ENSDART00000028946
ENSDART00000186842 ENSDART00000137815 ENSDART00000136914 ENSDART00000144855 ENSDART00000182689 |
tpd52l2a
|
tumor protein D52-like 2a |
| chr13_+_23176330 | 0.02 |
ENSDART00000168351
|
sorbs1
|
sorbin and SH3 domain containing 1 |
| chr15_-_1822548 | 0.01 |
ENSDART00000082026
ENSDART00000180230 |
mmp28
|
matrix metallopeptidase 28 |
| chr22_-_7129631 | 0.00 |
ENSDART00000171359
|
asic1b
|
acid-sensing (proton-gated) ion channel 1b |
Network of associatons between targets according to the STRING database.
First level regulatory network of hmx1+hmx3b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.7 | GO:0050996 | positive regulation of lipid catabolic process(GO:0050996) |
| 0.1 | 0.2 | GO:0060958 | cell proliferation involved in heart valve morphogenesis(GO:0003249) regulation of cell proliferation involved in heart valve morphogenesis(GO:0003250) endocardial cell development(GO:0060958) cell proliferation involved in heart valve development(GO:2000793) |
| 0.1 | 0.5 | GO:0019563 | glycerol catabolic process(GO:0019563) |
| 0.1 | 0.8 | GO:0045740 | positive regulation of DNA replication(GO:0045740) |
| 0.1 | 0.4 | GO:0019344 | homoserine metabolic process(GO:0009092) cysteine biosynthetic process via cystathionine(GO:0019343) cysteine biosynthetic process(GO:0019344) transsulfuration(GO:0019346) |
| 0.0 | 0.5 | GO:0000002 | mitochondrial genome maintenance(GO:0000002) |
| 0.0 | 0.1 | GO:0070278 | extracellular matrix constituent secretion(GO:0070278) |
| 0.0 | 0.4 | GO:0060005 | reflex(GO:0060004) vestibular reflex(GO:0060005) |
| 0.0 | 0.2 | GO:0030823 | regulation of cGMP metabolic process(GO:0030823) positive regulation of cGMP metabolic process(GO:0030825) regulation of cGMP biosynthetic process(GO:0030826) positive regulation of cGMP biosynthetic process(GO:0030828) regulation of guanylate cyclase activity(GO:0031282) positive regulation of guanylate cyclase activity(GO:0031284) |
| 0.0 | 0.6 | GO:0009081 | branched-chain amino acid metabolic process(GO:0009081) |
| 0.0 | 0.3 | GO:0015721 | bile acid and bile salt transport(GO:0015721) |
| 0.0 | 0.4 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
| 0.0 | 0.3 | GO:0007130 | synaptonemal complex assembly(GO:0007130) synaptonemal complex organization(GO:0070193) |
| 0.0 | 0.1 | GO:0040016 | embryonic cleavage(GO:0040016) |
| 0.0 | 0.3 | GO:0006198 | cAMP catabolic process(GO:0006198) |
| 0.0 | 0.7 | GO:0006119 | oxidative phosphorylation(GO:0006119) |
| 0.0 | 0.2 | GO:0032525 | somite rostral/caudal axis specification(GO:0032525) |
| 0.0 | 0.9 | GO:0042493 | response to drug(GO:0042493) |
| 0.0 | 0.6 | GO:0071466 | cellular response to xenobiotic stimulus(GO:0071466) |
| 0.0 | 0.4 | GO:0018230 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 0.0 | 0.6 | GO:0070098 | chemokine-mediated signaling pathway(GO:0070098) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.7 | GO:0045277 | respiratory chain complex IV(GO:0045277) |
| 0.0 | 0.3 | GO:0000795 | synaptonemal complex(GO:0000795) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:0005131 | growth hormone receptor binding(GO:0005131) growth hormone activity(GO:0070186) |
| 0.2 | 1.5 | GO:0004459 | L-lactate dehydrogenase activity(GO:0004459) |
| 0.1 | 0.8 | GO:0070330 | aromatase activity(GO:0070330) |
| 0.1 | 0.4 | GO:0001605 | adrenomedullin receptor activity(GO:0001605) calcitonin gene-related peptide receptor activity(GO:0001635) |
| 0.1 | 0.4 | GO:0070738 | protein-glycine ligase activity(GO:0070735) tubulin-glycine ligase activity(GO:0070738) |
| 0.1 | 0.5 | GO:0043139 | 5'-3' DNA helicase activity(GO:0043139) |
| 0.1 | 0.9 | GO:0004029 | aldehyde dehydrogenase (NAD) activity(GO:0004029) |
| 0.1 | 0.3 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.0 | 0.8 | GO:0031726 | CCR1 chemokine receptor binding(GO:0031726) CCR4 chemokine receptor binding(GO:0031729) |
| 0.0 | 0.6 | GO:0005504 | fatty acid binding(GO:0005504) |
| 0.0 | 0.2 | GO:0004517 | nitric-oxide synthase activity(GO:0004517) |
| 0.0 | 0.7 | GO:0016675 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors(GO:0016675) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.0 | 0.4 | GO:0016846 | carbon-sulfur lyase activity(GO:0016846) |
| 0.0 | 0.3 | GO:0008137 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.0 | 0.2 | GO:0001608 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.0 | 0.3 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.0 | 0.4 | GO:0008381 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 0.0 | 0.2 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | ST TUMOR NECROSIS FACTOR PATHWAY | Tumor Necrosis Factor Pathway. |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.7 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.1 | 0.3 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 0.0 | 0.6 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.0 | 0.6 | REACTOME RIP MEDIATED NFKB ACTIVATION VIA DAI | Genes involved in RIP-mediated NFkB activation via DAI |
| 0.0 | 0.2 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.0 | 0.4 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |