Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
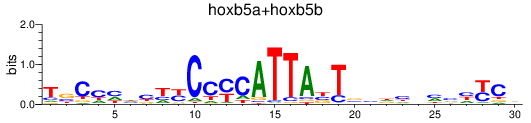
Results for hoxb5a+hoxb5b
Z-value: 0.62
Transcription factors associated with hoxb5a+hoxb5b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
hoxb5a
|
ENSDARG00000013057 | homeobox B5a |
|
hoxb5b
|
ENSDARG00000054030 | homeobox B5b |
|
hoxb5b
|
ENSDARG00000113174 | homeobox B5b |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| hoxb5b | dr11_v1_chr12_+_27129659_27129659 | 0.43 | 1.9e-05 | Click! |
| hoxb5a | dr11_v1_chr3_+_23707691_23707691 | 0.18 | 7.7e-02 | Click! |
Activity profile of hoxb5a+hoxb5b motif
Sorted Z-values of hoxb5a+hoxb5b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr23_+_22656477 | 5.98 |
ENSDART00000009337
ENSDART00000133322 |
eno1a
|
enolase 1a, (alpha) |
| chr1_+_12766351 | 5.69 |
ENSDART00000165785
|
pcdh10a
|
protocadherin 10a |
| chr16_-_27640995 | 5.58 |
ENSDART00000019658
|
nacad
|
NAC alpha domain containing |
| chr9_-_12034444 | 5.37 |
ENSDART00000038651
|
znf804a
|
zinc finger protein 804A |
| chr15_-_12500938 | 5.11 |
ENSDART00000159627
|
scn4ba
|
sodium channel, voltage-gated, type IV, beta a |
| chr15_-_12319065 | 4.90 |
ENSDART00000162973
ENSDART00000170543 |
fxyd6
|
FXYD domain containing ion transport regulator 6 |
| chr6_-_52156427 | 4.73 |
ENSDART00000082821
|
rims4
|
regulating synaptic membrane exocytosis 4 |
| chr2_+_44426609 | 4.52 |
ENSDART00000112711
ENSDART00000154188 |
kcnab1b
|
potassium voltage-gated channel, shaker-related subfamily, beta member 1 b |
| chr7_+_30867008 | 4.44 |
ENSDART00000193106
|
apba2b
|
amyloid beta (A4) precursor protein-binding, family A, member 2b |
| chr22_+_11857356 | 4.42 |
ENSDART00000179540
|
mras
|
muscle RAS oncogene homolog |
| chr19_+_46259619 | 4.10 |
ENSDART00000158032
|
grinaa
|
glutamate receptor, ionotropic, N-methyl D-aspartate-associated protein 1a (glutamate binding) |
| chr20_-_29418620 | 4.09 |
ENSDART00000172634
|
ryr3
|
ryanodine receptor 3 |
| chr11_-_44030962 | 4.03 |
ENSDART00000171910
|
FP016005.1
|
|
| chr7_+_33314925 | 4.03 |
ENSDART00000148590
|
coro2ba
|
coronin, actin binding protein, 2Ba |
| chr14_-_25078569 | 3.99 |
ENSDART00000172802
ENSDART00000173345 ENSDART00000135004 |
matr3l1.1
|
matrin 3-like 1.1 |
| chr12_+_7865470 | 3.90 |
ENSDART00000161683
|
BX548028.1
|
|
| chr4_+_3482312 | 3.77 |
ENSDART00000109044
|
grm8a
|
glutamate receptor, metabotropic 8a |
| chr3_+_28831450 | 3.74 |
ENSDART00000055422
|
flr
|
fleer |
| chr8_-_15398760 | 3.69 |
ENSDART00000162011
|
agbl4
|
ATP/GTP binding protein-like 4 |
| chr22_-_15385442 | 3.66 |
ENSDART00000090975
|
tmem264
|
transmembrane protein 264 |
| chr16_-_9980402 | 3.54 |
ENSDART00000066372
|
id4
|
inhibitor of DNA binding 4 |
| chr19_-_8600383 | 3.46 |
ENSDART00000081546
|
trim46b
|
tripartite motif containing 46b |
| chr6_+_54888493 | 3.44 |
ENSDART00000113331
|
nav1b
|
neuron navigator 1b |
| chr1_-_8192418 | 3.02 |
ENSDART00000136654
|
grapb
|
GRB2-related adaptor protein b |
| chr9_+_6082793 | 2.92 |
ENSDART00000192045
|
st6gal2a
|
ST6 beta-galactosamide alpha-2,6-sialyltranferase 2a |
| chr5_-_26284276 | 2.71 |
ENSDART00000148608
|
arvcfb
|
ARVCF, delta catenin family member b |
| chr23_-_637347 | 2.69 |
ENSDART00000132175
|
l1camb
|
L1 cell adhesion molecule, paralog b |
| chr1_-_46981134 | 2.54 |
ENSDART00000130607
|
pknox1.2
|
pbx/knotted 1 homeobox 1.2 |
| chr2_+_24317465 | 2.48 |
ENSDART00000139339
ENSDART00000136284 |
mrpl34
|
mitochondrial ribosomal protein L34 |
| chr6_-_15603675 | 2.33 |
ENSDART00000143502
|
lrrfip1b
|
leucine rich repeat (in FLII) interacting protein 1b |
| chr14_-_31893996 | 2.22 |
ENSDART00000173222
|
gpr101
|
G protein-coupled receptor 101 |
| chr2_-_42065069 | 2.19 |
ENSDART00000140188
|
cspp1b
|
centrosome and spindle pole associated protein 1b |
| chr5_-_41142467 | 2.09 |
ENSDART00000129415
|
zfr
|
zinc finger RNA binding protein |
| chr23_+_39331216 | 2.08 |
ENSDART00000160957
|
kcng1
|
potassium voltage-gated channel, subfamily G, member 1 |
| chr18_+_33580391 | 2.04 |
ENSDART00000189300
|
si:dkey-47k20.4
|
si:dkey-47k20.4 |
| chr10_+_2684958 | 1.82 |
ENSDART00000112019
|
setd9
|
SET domain containing 9 |
| chr3_-_5413018 | 1.76 |
ENSDART00000063138
|
vmo1a
|
vitelline membrane outer layer 1 homolog a |
| chr7_+_36035432 | 1.72 |
ENSDART00000179004
|
irx3a
|
iroquois homeobox 3a |
| chr6_-_18992896 | 1.71 |
ENSDART00000170228
|
sept9b
|
septin 9b |
| chr24_-_29822913 | 1.68 |
ENSDART00000160929
|
aglb
|
amylo-alpha-1, 6-glucosidase, 4-alpha-glucanotransferase b |
| chr1_-_49498116 | 1.63 |
ENSDART00000137357
|
zgc:175214
|
zgc:175214 |
| chr15_-_25099679 | 1.62 |
ENSDART00000154628
|
rflnb
|
refilin B |
| chr7_-_72426484 | 1.60 |
ENSDART00000190063
|
rph3ab
|
rabphilin 3A homolog (mouse), b |
| chr22_+_696931 | 1.42 |
ENSDART00000149712
ENSDART00000009756 |
gpr37l1a
|
G protein-coupled receptor 37 like 1a |
| chr11_-_2297832 | 1.41 |
ENSDART00000158266
|
znf740a
|
zinc finger protein 740a |
| chr4_-_59094319 | 1.39 |
ENSDART00000162560
|
si:dkey-28i19.1
|
si:dkey-28i19.1 |
| chr8_+_25962833 | 1.38 |
ENSDART00000086583
|
si:dkey-72l14.4
|
si:dkey-72l14.4 |
| chr17_+_49081828 | 1.34 |
ENSDART00000156492
|
tiam2a
|
T cell lymphoma invasion and metastasis 2a |
| chr17_-_39772999 | 1.28 |
ENSDART00000155727
|
pimr60
|
Pim proto-oncogene, serine/threonine kinase, related 60 |
| chr22_-_7461603 | 1.26 |
ENSDART00000170630
|
BX511034.6
|
|
| chr6_-_19340889 | 1.24 |
ENSDART00000181407
|
mif4gda
|
MIF4G domain containing a |
| chr10_+_35439952 | 1.14 |
ENSDART00000006284
|
hhla2a.2
|
HERV-H LTR-associating 2a, tandem duplicate 2 |
| chr11_+_24957858 | 0.95 |
ENSDART00000145647
|
sulf2a
|
sulfatase 2a |
| chr5_+_59392183 | 0.94 |
ENSDART00000082983
ENSDART00000180882 |
clip2
|
CAP-GLY domain containing linker protein 2 |
| chr14_+_33723309 | 0.89 |
ENSDART00000132488
|
apln
|
apelin |
| chr22_+_1680628 | 0.81 |
ENSDART00000166672
|
si:dkey-1b17.6
|
si:dkey-1b17.6 |
| chr7_+_56889879 | 0.80 |
ENSDART00000039810
|
myo9aa
|
myosin IXAa |
| chr22_+_12477996 | 0.75 |
ENSDART00000177704
|
CR847870.3
|
|
| chr22_+_17536989 | 0.75 |
ENSDART00000149531
|
hnrnpm
|
heterogeneous nuclear ribonucleoprotein M |
| chr18_-_14337065 | 0.73 |
ENSDART00000135703
|
hsbp1b
|
heat shock factor binding protein 1b |
| chr9_+_30475563 | 0.72 |
ENSDART00000133118
|
gja5a
|
gap junction protein, alpha 5a |
| chr19_-_27827744 | 0.67 |
ENSDART00000181620
|
papd7
|
PAP associated domain containing 7 |
| chr19_-_30904590 | 0.66 |
ENSDART00000137633
|
si:ch211-194e15.5
|
si:ch211-194e15.5 |
| chr23_+_3513201 | 0.63 |
ENSDART00000092258
|
spata2
|
spermatogenesis associated 2 |
| chr2_-_3038904 | 0.60 |
ENSDART00000186795
|
guk1a
|
guanylate kinase 1a |
| chr15_-_35031523 | 0.59 |
ENSDART00000154848
|
si:ch211-272b8.6
|
si:ch211-272b8.6 |
| chr20_+_33708318 | 0.54 |
ENSDART00000135845
|
ak9
|
adenylate kinase 9 |
| chr14_+_8476769 | 0.52 |
ENSDART00000131773
|
dnajc4
|
DnaJ (Hsp40) homolog, subfamily C, member 4 |
| chr13_-_11986754 | 0.50 |
ENSDART00000164214
|
npm3
|
nucleophosmin/nucleoplasmin, 3 |
| chr12_+_17436904 | 0.50 |
ENSDART00000079130
|
atad1b
|
ATPase family, AAA domain containing 1b |
| chr9_+_16854121 | 0.43 |
ENSDART00000110866
|
cln5
|
CLN5, intracellular trafficking protein |
| chr20_+_51730658 | 0.31 |
ENSDART00000010271
|
aida
|
axin interactor, dorsalization associated |
| chr9_+_48755638 | 0.29 |
ENSDART00000047401
|
abcb11a
|
ATP-binding cassette, sub-family B (MDR/TAP), member 11a |
| chr8_+_247163 | 0.27 |
ENSDART00000122378
|
cep120
|
centrosomal protein 120 |
| chr5_-_16791223 | 0.25 |
ENSDART00000192744
|
atad1a
|
ATPase family, AAA domain containing 1a |
| chr2_+_22823952 | 0.22 |
ENSDART00000147935
|
aire
|
autoimmune regulator |
| chr20_+_48543492 | 0.20 |
ENSDART00000175480
|
LO018254.1
|
|
| chr1_+_42224769 | 0.19 |
ENSDART00000177496
ENSDART00000184778 ENSDART00000110860 |
ctnna2
|
catenin (cadherin-associated protein), alpha 2 |
| chr15_-_41408148 | 0.18 |
ENSDART00000133459
|
CR854897.1
|
|
| chr15_-_684911 | 0.18 |
ENSDART00000161685
|
zgc:171901
|
zgc:171901 |
| chr16_+_16826504 | 0.17 |
ENSDART00000108691
|
kcnj14
|
potassium inwardly-rectifying channel, subfamily J, member 14 |
| chr4_-_76154252 | 0.13 |
ENSDART00000161269
|
BX957252.1
|
|
| chr4_+_41789497 | 0.11 |
ENSDART00000126634
|
si:dkey-237m9.2
|
si:dkey-237m9.2 |
| chr3_+_39579393 | 0.10 |
ENSDART00000055170
|
cln3
|
ceroid-lipofuscinosis, neuronal 3 |
| chr20_+_53181017 | 0.10 |
ENSDART00000189692
ENSDART00000177109 |
FIG4
|
FIG4 phosphoinositide 5-phosphatase |
| chr11_-_6070192 | 0.07 |
ENSDART00000162776
|
babam1
|
BRISC and BRCA1 A complex member 1 |
| chr3_+_54776475 | 0.06 |
ENSDART00000142710
|
si:ch211-74m13.1
|
si:ch211-74m13.1 |
| chr4_+_77966055 | 0.06 |
ENSDART00000174203
ENSDART00000130100 ENSDART00000080665 ENSDART00000174317 ENSDART00000190123 |
zgc:113921
|
zgc:113921 |
| chr9_+_51147380 | 0.05 |
ENSDART00000003913
|
ifih1
|
interferon induced with helicase C domain 1 |
| chr4_+_42295479 | 0.02 |
ENSDART00000162279
|
znf1118
|
zinc finger protein 1118 |
| chr23_+_22293682 | 0.00 |
ENSDART00000187304
|
CR354612.1
|
|
Network of associatons between targets according to the STRING database.
First level regulatory network of hoxb5a+hoxb5b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 4.5 | GO:1902260 | negative regulation of potassium ion transmembrane transporter activity(GO:1901017) negative regulation of potassium ion transmembrane transport(GO:1901380) negative regulation of delayed rectifier potassium channel activity(GO:1902260) negative regulation of voltage-gated potassium channel activity(GO:1903817) |
| 0.7 | 6.0 | GO:0032889 | regulation of vacuole fusion, non-autophagic(GO:0032889) |
| 0.7 | 2.1 | GO:1902259 | regulation of delayed rectifier potassium channel activity(GO:1902259) |
| 0.5 | 1.6 | GO:1900157 | negative regulation of chondrocyte differentiation(GO:0032331) negative regulation of cartilage development(GO:0061037) negative regulation of chondrocyte development(GO:0061182) regulation of bone mineralization involved in bone maturation(GO:1900157) negative regulation of bone mineralization involved in bone maturation(GO:1900158) negative regulation of bone development(GO:1903011) |
| 0.5 | 3.7 | GO:0036372 | opsin transport(GO:0036372) |
| 0.4 | 4.1 | GO:1902108 | regulation of mitochondrial membrane permeability involved in apoptotic process(GO:1902108) |
| 0.3 | 5.7 | GO:0097324 | melanocyte migration(GO:0097324) |
| 0.3 | 3.5 | GO:0003307 | regulation of Wnt signaling pathway involved in heart development(GO:0003307) |
| 0.2 | 4.7 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.2 | 0.7 | GO:0010610 | regulation of mRNA stability involved in response to stress(GO:0010610) regulation of mRNA stability involved in response to oxidative stress(GO:2000815) |
| 0.2 | 0.8 | GO:0098529 | neuromuscular junction development, skeletal muscle fiber(GO:0098529) |
| 0.1 | 0.9 | GO:0090134 | mesendoderm migration(GO:0090133) cell migration involved in mesendoderm migration(GO:0090134) |
| 0.1 | 0.5 | GO:0009182 | purine deoxyribonucleoside diphosphate metabolic process(GO:0009182) deoxyribonucleoside triphosphate biosynthetic process(GO:0009202) dGDP metabolic process(GO:0046066) |
| 0.1 | 0.6 | GO:0010939 | regulation of necrotic cell death(GO:0010939) regulation of necroptotic process(GO:0060544) |
| 0.1 | 2.2 | GO:0032467 | positive regulation of cytokinesis(GO:0032467) |
| 0.1 | 1.7 | GO:0044247 | polysaccharide catabolic process(GO:0000272) glycogen catabolic process(GO:0005980) glucan catabolic process(GO:0009251) cellular polysaccharide catabolic process(GO:0044247) |
| 0.1 | 2.7 | GO:0045773 | positive regulation of axon extension(GO:0045773) |
| 0.1 | 4.1 | GO:0051209 | release of sequestered calcium ion into cytosol(GO:0051209) |
| 0.1 | 3.8 | GO:0007216 | G-protein coupled glutamate receptor signaling pathway(GO:0007216) |
| 0.1 | 0.2 | GO:0002513 | tolerance induction(GO:0002507) tolerance induction to self antigen(GO:0002513) |
| 0.1 | 1.1 | GO:0031295 | lymphocyte costimulation(GO:0031294) T cell costimulation(GO:0031295) |
| 0.1 | 4.9 | GO:2000649 | regulation of sodium ion transmembrane transporter activity(GO:2000649) |
| 0.1 | 1.4 | GO:0032418 | lysosome localization(GO:0032418) |
| 0.1 | 2.7 | GO:0016339 | calcium-dependent cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0016339) |
| 0.1 | 0.3 | GO:0015722 | canalicular bile acid transport(GO:0015722) bile acid secretion(GO:0032782) |
| 0.1 | 0.9 | GO:0035860 | esophagus smooth muscle contraction(GO:0014846) glial cell-derived neurotrophic factor receptor signaling pathway(GO:0035860) |
| 0.1 | 5.6 | GO:0006612 | protein targeting to membrane(GO:0006612) |
| 0.1 | 0.3 | GO:0010825 | positive regulation of centrosome duplication(GO:0010825) |
| 0.1 | 2.5 | GO:0060037 | pharyngeal system development(GO:0060037) |
| 0.1 | 0.3 | GO:0043508 | negative regulation of JUN kinase activity(GO:0043508) |
| 0.1 | 2.2 | GO:0071880 | adenylate cyclase-activating adrenergic receptor signaling pathway(GO:0071880) |
| 0.1 | 3.4 | GO:0001764 | neuron migration(GO:0001764) |
| 0.0 | 0.2 | GO:0051823 | regulation of synapse structural plasticity(GO:0051823) |
| 0.0 | 1.2 | GO:0006446 | regulation of translational initiation(GO:0006446) |
| 0.0 | 1.3 | GO:0050772 | positive regulation of axonogenesis(GO:0050772) |
| 0.0 | 1.4 | GO:0007193 | adenylate cyclase-inhibiting G-protein coupled receptor signaling pathway(GO:0007193) |
| 0.0 | 0.7 | GO:0014029 | neural crest formation(GO:0014029) |
| 0.0 | 1.7 | GO:0031018 | endocrine pancreas development(GO:0031018) |
| 0.0 | 1.7 | GO:0061640 | cytoskeleton-dependent cytokinesis(GO:0061640) |
| 0.0 | 0.5 | GO:0007040 | lysosome organization(GO:0007040) lytic vacuole organization(GO:0080171) |
| 0.0 | 0.9 | GO:0031122 | cytoplasmic microtubule organization(GO:0031122) |
| 0.0 | 3.9 | GO:0007265 | Ras protein signal transduction(GO:0007265) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 5.6 | GO:0005854 | nascent polypeptide-associated complex(GO:0005854) |
| 0.7 | 6.0 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.7 | 3.7 | GO:0005879 | axonemal microtubule(GO:0005879) |
| 0.6 | 4.1 | GO:0005790 | smooth endoplasmic reticulum(GO:0005790) |
| 0.5 | 3.4 | GO:0043194 | axon initial segment(GO:0043194) |
| 0.3 | 4.5 | GO:0044224 | juxtaparanode region of axon(GO:0044224) |
| 0.2 | 2.7 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.2 | 4.7 | GO:0048788 | cytoskeleton of presynaptic active zone(GO:0048788) |
| 0.1 | 2.5 | GO:0005762 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.1 | 0.7 | GO:0005922 | connexon complex(GO:0005922) |
| 0.0 | 0.7 | GO:0071014 | post-mRNA release spliceosomal complex(GO:0071014) |
| 0.0 | 2.9 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.0 | 1.7 | GO:0031105 | septin ring(GO:0005940) septin complex(GO:0031105) septin cytoskeleton(GO:0032156) |
| 0.0 | 4.4 | GO:0044309 | dendritic spine(GO:0043197) neuron spine(GO:0044309) |
| 0.0 | 2.1 | GO:0008076 | voltage-gated potassium channel complex(GO:0008076) potassium channel complex(GO:0034705) |
| 0.0 | 0.9 | GO:0035371 | microtubule plus-end(GO:0035371) |
| 0.0 | 1.6 | GO:0032432 | actin filament bundle(GO:0032432) |
| 0.0 | 2.2 | GO:0000922 | spindle pole(GO:0000922) |
| 0.0 | 0.9 | GO:0005795 | Golgi stack(GO:0005795) |
| 0.0 | 0.8 | GO:0030426 | growth cone(GO:0030426) |
| 0.0 | 0.1 | GO:0070552 | BRISC complex(GO:0070552) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 3.7 | GO:0070738 | protein-glycine ligase activity(GO:0070735) tubulin-glycine ligase activity(GO:0070738) |
| 0.8 | 4.1 | GO:0005219 | ryanodine-sensitive calcium-release channel activity(GO:0005219) calcium-induced calcium release activity(GO:0048763) |
| 0.7 | 6.0 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.5 | 3.8 | GO:0001642 | group III metabotropic glutamate receptor activity(GO:0001642) |
| 0.4 | 1.7 | GO:0004134 | glycogen debranching enzyme activity(GO:0004133) 4-alpha-glucanotransferase activity(GO:0004134) amylo-alpha-1,6-glucosidase activity(GO:0004135) |
| 0.4 | 1.6 | GO:0031005 | filamin binding(GO:0031005) |
| 0.4 | 2.9 | GO:0003835 | beta-galactoside alpha-2,6-sialyltransferase activity(GO:0003835) |
| 0.3 | 0.9 | GO:0031704 | apelin receptor binding(GO:0031704) |
| 0.2 | 4.5 | GO:0004033 | aldo-keto reductase (NADP) activity(GO:0004033) |
| 0.2 | 4.4 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.2 | 4.9 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.1 | 1.2 | GO:0008494 | translation activator activity(GO:0008494) |
| 0.1 | 0.6 | GO:0004385 | guanylate kinase activity(GO:0004385) |
| 0.1 | 4.4 | GO:0019003 | GDP binding(GO:0019003) |
| 0.1 | 4.7 | GO:0044325 | ion channel binding(GO:0044325) |
| 0.1 | 0.3 | GO:0015126 | canalicular bile acid transmembrane transporter activity(GO:0015126) bile acid-exporting ATPase activity(GO:0015432) |
| 0.1 | 0.9 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 0.1 | 0.7 | GO:0005243 | gap junction channel activity(GO:0005243) |
| 0.0 | 4.7 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.0 | 2.1 | GO:0005251 | delayed rectifier potassium channel activity(GO:0005251) |
| 0.0 | 0.5 | GO:0004017 | adenylate kinase activity(GO:0004017) |
| 0.0 | 2.9 | GO:0045296 | cadherin binding(GO:0045296) |
| 0.0 | 1.7 | GO:0003725 | double-stranded RNA binding(GO:0003725) |
| 0.0 | 0.9 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.0 | 2.5 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 4.0 | GO:0051015 | actin filament binding(GO:0051015) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 4.4 | ST INTEGRIN SIGNALING PATHWAY | Integrin Signaling Pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.1 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |