Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
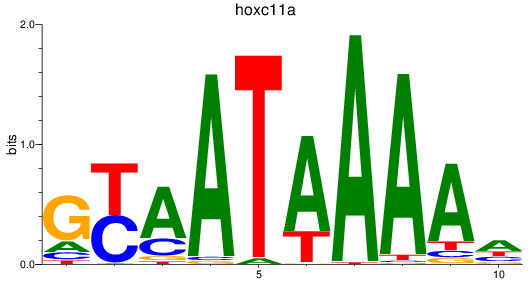
Results for hoxc11a
Z-value: 0.97
Transcription factors associated with hoxc11a
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
hoxc11a
|
ENSDARG00000070351 | homeobox C11a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| hoxc11a | dr11_v1_chr23_+_36074798_36074880 | 0.34 | 9.6e-04 | Click! |
Activity profile of hoxc11a motif
Sorted Z-values of hoxc11a motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr7_+_39386982 | 13.01 |
ENSDART00000146702
|
tnni2b.2
|
troponin I type 2b (skeletal, fast), tandem duplicate 2 |
| chr12_-_25916530 | 9.73 |
ENSDART00000186386
|
sncgb
|
synuclein, gamma b (breast cancer-specific protein 1) |
| chr6_-_60147517 | 9.23 |
ENSDART00000083453
|
slc32a1
|
solute carrier family 32 (GABA vesicular transporter), member 1 |
| chr25_+_29161609 | 8.57 |
ENSDART00000180752
|
pkmb
|
pyruvate kinase M1/2b |
| chr17_-_12389259 | 8.10 |
ENSDART00000185724
|
snap25b
|
synaptosomal-associated protein, 25b |
| chr5_-_40510397 | 6.89 |
ENSDART00000146237
ENSDART00000051065 |
fsta
|
follistatin a |
| chr20_-_25709247 | 6.73 |
ENSDART00000146711
|
si:dkeyp-117h8.2
|
si:dkeyp-117h8.2 |
| chr21_+_5129513 | 6.46 |
ENSDART00000102572
|
thbs4b
|
thrombospondin 4b |
| chr10_+_36650222 | 6.44 |
ENSDART00000126963
|
ucp3
|
uncoupling protein 3 |
| chr15_-_33896159 | 6.18 |
ENSDART00000159791
|
mag
|
myelin associated glycoprotein |
| chr8_-_34052019 | 5.89 |
ENSDART00000040126
ENSDART00000159208 ENSDART00000048994 ENSDART00000098822 |
pbx3b
|
pre-B-cell leukemia homeobox 3b |
| chr24_-_6158933 | 5.81 |
ENSDART00000021609
|
gad2
|
glutamate decarboxylase 2 |
| chr9_+_44431174 | 5.66 |
ENSDART00000149726
|
ppp1r1c
|
protein phosphatase 1, regulatory (inhibitor) subunit 1C |
| chr6_-_40697585 | 5.54 |
ENSDART00000113196
|
si:ch211-157b11.14
|
si:ch211-157b11.14 |
| chr3_+_29714775 | 5.50 |
ENSDART00000041388
|
cacng2a
|
calcium channel, voltage-dependent, gamma subunit 2a |
| chr24_-_8732519 | 5.28 |
ENSDART00000082351
|
tfap2a
|
transcription factor AP-2 alpha |
| chr7_+_25059845 | 5.14 |
ENSDART00000077215
|
ppp2r5b
|
protein phosphatase 2, regulatory subunit B', beta |
| chr23_-_27571667 | 5.11 |
ENSDART00000008174
|
pfkma
|
phosphofructokinase, muscle a |
| chr17_+_52822831 | 5.07 |
ENSDART00000193368
|
meis2a
|
Meis homeobox 2a |
| chr17_-_26721007 | 5.04 |
ENSDART00000034580
|
calm1a
|
calmodulin 1a |
| chr18_-_14941840 | 4.89 |
ENSDART00000091729
|
mlc1
|
megalencephalic leukoencephalopathy with subcortical cysts 1 |
| chr14_-_12837052 | 4.83 |
ENSDART00000165004
ENSDART00000043180 |
gria3b
|
glutamate receptor, ionotropic, AMPA 3b |
| chr16_-_26074529 | 4.73 |
ENSDART00000148653
ENSDART00000148923 |
tmem145
|
transmembrane protein 145 |
| chr9_+_44430974 | 4.61 |
ENSDART00000056846
|
ppp1r1c
|
protein phosphatase 1, regulatory (inhibitor) subunit 1C |
| chr6_+_12853655 | 4.36 |
ENSDART00000156341
|
fam117ba
|
family with sequence similarity 117, member Ba |
| chr9_+_34425736 | 4.17 |
ENSDART00000135147
|
si:ch211-218d20.15
|
si:ch211-218d20.15 |
| chr17_-_30975978 | 4.16 |
ENSDART00000051697
|
evla
|
Enah/Vasp-like a |
| chr9_-_41784799 | 4.15 |
ENSDART00000144573
ENSDART00000112542 ENSDART00000190486 |
obsl1b
|
obscurin-like 1b |
| chr2_+_21128391 | 4.07 |
ENSDART00000136814
|
pip4k2ab
|
phosphatidylinositol-5-phosphate 4-kinase, type II, alpha b |
| chr14_-_12837432 | 4.05 |
ENSDART00000178444
|
gria3b
|
glutamate receptor, ionotropic, AMPA 3b |
| chr2_+_2223837 | 3.96 |
ENSDART00000101038
ENSDART00000129354 |
tmie
|
transmembrane inner ear |
| chr23_-_12014931 | 3.92 |
ENSDART00000134652
|
si:dkey-178k16.1
|
si:dkey-178k16.1 |
| chr6_+_52804267 | 3.89 |
ENSDART00000065681
|
matn4
|
matrilin 4 |
| chr11_-_18253111 | 3.87 |
ENSDART00000125984
|
mustn1b
|
musculoskeletal, embryonic nuclear protein 1b |
| chr11_-_6188413 | 3.79 |
ENSDART00000109972
|
ccl44
|
chemokine (C-C motif) ligand 44 |
| chr7_+_36041509 | 3.79 |
ENSDART00000162850
|
irx3a
|
iroquois homeobox 3a |
| chr14_+_34966598 | 3.72 |
ENSDART00000004550
|
rnf145a
|
ring finger protein 145a |
| chr5_+_36611128 | 3.71 |
ENSDART00000097684
|
nova1
|
neuro-oncological ventral antigen 1 |
| chr16_+_32995882 | 3.67 |
ENSDART00000170157
|
prss35
|
protease, serine, 35 |
| chr17_-_30975707 | 3.64 |
ENSDART00000138346
|
evla
|
Enah/Vasp-like a |
| chr14_+_49135264 | 3.64 |
ENSDART00000084119
|
si:ch1073-44g3.1
|
si:ch1073-44g3.1 |
| chr1_-_22512063 | 3.63 |
ENSDART00000031546
ENSDART00000190987 |
chrna6
|
cholinergic receptor, nicotinic, alpha 6 |
| chr25_+_20089986 | 3.62 |
ENSDART00000143441
ENSDART00000184073 |
tnni4b.2
|
troponin I4b, tandem duplicate 2 |
| chr19_-_10243148 | 3.55 |
ENSDART00000148073
|
shisa7b
|
shisa family member 7 |
| chr3_+_24134418 | 3.54 |
ENSDART00000156204
|
si:ch211-246i5.5
|
si:ch211-246i5.5 |
| chr24_+_26134029 | 3.42 |
ENSDART00000185134
|
tmtopsb
|
teleost multiple tissue opsin b |
| chr24_+_39105051 | 3.37 |
ENSDART00000115297
|
mss51
|
MSS51 mitochondrial translational activator |
| chr1_+_46194333 | 3.28 |
ENSDART00000010894
|
sox1b
|
SRY (sex determining region Y)-box 1b |
| chr17_+_52822422 | 3.19 |
ENSDART00000158273
ENSDART00000161414 |
meis2a
|
Meis homeobox 2a |
| chr8_+_11687254 | 3.16 |
ENSDART00000042040
|
atp2a2a
|
ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 2a |
| chr24_+_5840258 | 3.11 |
ENSDART00000087034
|
trpc1
|
transient receptor potential cation channel, subfamily C, member 1 |
| chr4_+_5741733 | 3.07 |
ENSDART00000110243
|
pou3f2a
|
POU class 3 homeobox 2a |
| chr3_+_16265924 | 2.97 |
ENSDART00000122519
|
st8sia6
|
ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 6 |
| chr11_+_25257022 | 2.96 |
ENSDART00000156052
|
tp53inp2
|
tumor protein p53 inducible nuclear protein 2 |
| chr25_+_37126921 | 2.96 |
ENSDART00000124331
|
si:ch1073-174d20.1
|
si:ch1073-174d20.1 |
| chr20_+_30490682 | 2.92 |
ENSDART00000184871
|
myt1la
|
myelin transcription factor 1-like, a |
| chr10_+_18877362 | 2.91 |
ENSDART00000138334
|
ppp2r2ab
|
protein phosphatase 2, regulatory subunit B, alpha b |
| chr22_-_16377960 | 2.83 |
ENSDART00000168170
|
ttc39c
|
tetratricopeptide repeat domain 39C |
| chr8_+_39619087 | 2.83 |
ENSDART00000134822
|
msi1
|
musashi RNA-binding protein 1 |
| chr8_-_52413032 | 2.82 |
ENSDART00000183039
|
CABZ01070469.1
|
|
| chr23_+_28648864 | 2.81 |
ENSDART00000189096
|
l1cama
|
L1 cell adhesion molecule, paralog a |
| chr6_-_6448519 | 2.81 |
ENSDART00000180157
ENSDART00000191112 |
si:ch211-194e18.2
|
si:ch211-194e18.2 |
| chr21_-_20328375 | 2.77 |
ENSDART00000079593
|
slc26a1
|
solute carrier family 26 (anion exchanger), member 1 |
| chr11_-_12998400 | 2.77 |
ENSDART00000018614
|
chrna4b
|
cholinergic receptor, nicotinic, alpha 4b |
| chr19_-_28789404 | 2.75 |
ENSDART00000191453
ENSDART00000026992 |
sox4a
|
SRY (sex determining region Y)-box 4a |
| chr7_+_30875273 | 2.75 |
ENSDART00000173693
|
apba2b
|
amyloid beta (A4) precursor protein-binding, family A, member 2b |
| chr14_+_15597049 | 2.74 |
ENSDART00000159732
|
si:dkey-203a12.8
|
si:dkey-203a12.8 |
| chr14_+_15495088 | 2.71 |
ENSDART00000165765
ENSDART00000188577 |
si:dkey-203a12.6
|
si:dkey-203a12.6 |
| chr2_+_31948352 | 2.70 |
ENSDART00000192611
|
ankhb
|
ANKH inorganic pyrophosphate transport regulator b |
| chr6_-_26559921 | 2.67 |
ENSDART00000104532
|
sox14
|
SRY (sex determining region Y)-box 14 |
| chr11_+_30244356 | 2.64 |
ENSDART00000036050
ENSDART00000150080 |
rs1a
|
retinoschisin 1a |
| chr14_-_33613794 | 2.63 |
ENSDART00000010022
|
xpnpep2
|
X-prolyl aminopeptidase (aminopeptidase P) 2, membrane-bound |
| chr5_+_31946973 | 2.61 |
ENSDART00000189876
ENSDART00000163366 |
ungb
|
uracil DNA glycosylase b |
| chr12_-_46959990 | 2.58 |
ENSDART00000084557
|
lhpp
|
phospholysine phosphohistidine inorganic pyrophosphate phosphatase |
| chr22_-_34551568 | 2.57 |
ENSDART00000148147
|
rnf123
|
ring finger protein 123 |
| chr10_-_34889053 | 2.55 |
ENSDART00000136966
|
ccdc169
|
coiled-coil domain containing 169 |
| chr24_+_5237753 | 2.55 |
ENSDART00000106488
ENSDART00000005901 |
plod2
|
procollagen-lysine, 2-oxoglutarate 5-dioxygenase 2 |
| chr6_-_37468971 | 2.51 |
ENSDART00000126379
|
plcxd1
|
phosphatidylinositol-specific phospholipase C, X domain containing 1 |
| chr23_-_30431333 | 2.51 |
ENSDART00000146633
|
camta1a
|
calmodulin binding transcription activator 1a |
| chr18_-_15373620 | 2.46 |
ENSDART00000031752
|
rfx4
|
regulatory factor X, 4 |
| chr5_-_37881345 | 2.43 |
ENSDART00000084819
|
arhgap35b
|
Rho GTPase activating protein 35b |
| chr10_-_34772211 | 2.37 |
ENSDART00000145450
ENSDART00000134307 |
dclk1a
|
doublecortin-like kinase 1a |
| chr25_+_16080181 | 2.34 |
ENSDART00000061753
|
far1
|
fatty acyl CoA reductase 1 |
| chr5_-_42272517 | 2.27 |
ENSDART00000137692
ENSDART00000164363 |
si:ch211-207c6.2
|
si:ch211-207c6.2 |
| chr16_-_35329803 | 2.26 |
ENSDART00000161729
ENSDART00000157700 ENSDART00000184584 ENSDART00000174713 ENSDART00000162518 |
ptprub
|
protein tyrosine phosphatase, receptor type, U, b |
| chr12_+_27117609 | 2.22 |
ENSDART00000076154
|
hoxb8b
|
homeobox B8b |
| chr23_+_13124085 | 2.21 |
ENSDART00000139475
|
samd10b
|
sterile alpha motif domain containing 10b |
| chr21_+_19070921 | 2.20 |
ENSDART00000029874
|
nkx6.1
|
NK6 homeobox 1 |
| chr6_-_41138854 | 2.17 |
ENSDART00000128723
ENSDART00000151055 ENSDART00000132484 |
slc6a22.1
|
solute carrier family 6 member 22, tandem duplicate 1 |
| chr9_-_16853462 | 2.15 |
ENSDART00000160273
|
CT573248.2
|
|
| chr14_+_45306073 | 2.15 |
ENSDART00000034606
ENSDART00000173301 |
sfxn5b
|
sideroflexin 5b |
| chr6_-_12296170 | 2.12 |
ENSDART00000155685
|
pkp4
|
plakophilin 4 |
| chr24_-_29822913 | 2.11 |
ENSDART00000160929
|
aglb
|
amylo-alpha-1, 6-glucosidase, 4-alpha-glucanotransferase b |
| chr17_-_19019635 | 2.07 |
ENSDART00000126666
|
flrt2
|
fibronectin leucine rich transmembrane protein 2 |
| chr1_+_961607 | 2.06 |
ENSDART00000184660
|
n6amt1
|
N-6 adenine-specific DNA methyltransferase 1 |
| chr17_-_18888959 | 2.05 |
ENSDART00000080029
|
ak7b
|
adenylate kinase 7b |
| chr5_+_20147830 | 2.03 |
ENSDART00000098727
|
svopa
|
SV2 related protein a |
| chr20_-_47731768 | 1.99 |
ENSDART00000031167
|
tfap2d
|
transcription factor AP-2 delta (activating enhancer binding protein 2 delta) |
| chr11_-_36009924 | 1.96 |
ENSDART00000189959
ENSDART00000167472 ENSDART00000191211 ENSDART00000191662 ENSDART00000191780 ENSDART00000192622 ENSDART00000179911 |
itpr1b
|
inositol 1,4,5-trisphosphate receptor, type 1b |
| chr6_+_40629066 | 1.96 |
ENSDART00000103757
|
slc6a11a
|
solute carrier family 6 (neurotransmitter transporter), member 11a |
| chr18_-_25905574 | 1.91 |
ENSDART00000143899
ENSDART00000163369 |
sema4ba
|
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4Ba |
| chr2_-_24962820 | 1.89 |
ENSDART00000182767
|
hltf
|
helicase-like transcription factor |
| chr11_+_2198831 | 1.89 |
ENSDART00000160515
|
hoxc6b
|
homeobox C6b |
| chr15_-_13254480 | 1.88 |
ENSDART00000190499
|
zgc:172282
|
zgc:172282 |
| chr23_-_33038423 | 1.88 |
ENSDART00000180539
|
plxna2
|
plexin A2 |
| chr10_-_29903165 | 1.84 |
ENSDART00000078800
|
lim2.1
|
lens intrinsic membrane protein 2.1 |
| chr1_+_45217425 | 1.80 |
ENSDART00000179983
ENSDART00000074683 |
EVI5L
|
si:ch211-239f4.1 |
| chr1_-_19215336 | 1.77 |
ENSDART00000162949
ENSDART00000170680 |
ptprdb
|
protein tyrosine phosphatase, receptor type, D, b |
| chr9_-_42696408 | 1.75 |
ENSDART00000144744
|
col5a2a
|
collagen, type V, alpha 2a |
| chr22_+_31821815 | 1.74 |
ENSDART00000159825
|
dock3
|
dedicator of cytokinesis 3 |
| chr20_+_41906960 | 1.72 |
ENSDART00000193460
|
cep85l
|
centrosomal protein 85, like |
| chr18_+_16330025 | 1.72 |
ENSDART00000142353
|
nts
|
neurotensin |
| chr23_+_11669109 | 1.71 |
ENSDART00000091416
|
cntn3a.1
|
contactin 3a, tandem duplicate 1 |
| chr23_-_7799184 | 1.67 |
ENSDART00000190946
ENSDART00000165427 |
myt1b
|
myelin transcription factor 1b |
| chr2_+_45068366 | 1.65 |
ENSDART00000142175
|
si:dkey-76d14.2
|
si:dkey-76d14.2 |
| chr23_+_11669337 | 1.62 |
ENSDART00000131355
|
cntn3a.1
|
contactin 3a, tandem duplicate 1 |
| chr19_+_21362553 | 1.60 |
ENSDART00000122002
|
tshz1
|
teashirt zinc finger homeobox 1 |
| chr23_-_18030399 | 1.58 |
ENSDART00000136967
|
pm20d1.1
|
peptidase M20 domain containing 1, tandem duplicate 1 |
| chr7_+_61480296 | 1.57 |
ENSDART00000083255
|
adam19a
|
ADAM metallopeptidase domain 19a |
| chr25_-_6389713 | 1.55 |
ENSDART00000083539
|
sin3aa
|
SIN3 transcription regulator family member Aa |
| chr20_+_20726231 | 1.53 |
ENSDART00000147112
|
zgc:193541
|
zgc:193541 |
| chr19_+_42469058 | 1.53 |
ENSDART00000076915
|
si:dkey-166k12.1
|
si:dkey-166k12.1 |
| chr14_+_23184517 | 1.52 |
ENSDART00000181410
|
enox2
|
ecto-NOX disulfide-thiol exchanger 2 |
| chr6_+_23931236 | 1.45 |
ENSDART00000166079
|
gadd45ab
|
growth arrest and DNA-damage-inducible, alpha, b |
| chr12_+_19356623 | 1.44 |
ENSDART00000078284
|
dmc1
|
DNA meiotic recombinase 1 |
| chr6_-_46768040 | 1.43 |
ENSDART00000154071
|
igfn1.2
|
immunoglobulin-like and fibronectin type III domain containing 1, tandem duplicate 2 |
| chr24_+_9412450 | 1.42 |
ENSDART00000132724
|
si:ch211-285f17.1
|
si:ch211-285f17.1 |
| chr9_+_917060 | 1.40 |
ENSDART00000082390
|
tmem37
|
transmembrane protein 37 |
| chr14_+_4276394 | 1.38 |
ENSDART00000038301
|
gnpda2
|
glucosamine-6-phosphate deaminase 2 |
| chr22_+_5176255 | 1.37 |
ENSDART00000092647
|
cers1
|
ceramide synthase 1 |
| chr2_-_32738535 | 1.36 |
ENSDART00000135293
|
nrbp2a
|
nuclear receptor binding protein 2a |
| chr7_-_30367650 | 1.34 |
ENSDART00000075519
|
aldh1a2
|
aldehyde dehydrogenase 1 family, member A2 |
| chr12_-_1406480 | 1.33 |
ENSDART00000152308
|
MED9
|
si:ch73-105m5.1 |
| chr4_+_20954929 | 1.33 |
ENSDART00000143674
|
nav3
|
neuron navigator 3 |
| chr12_+_27022517 | 1.33 |
ENSDART00000152975
|
msl1b
|
male-specific lethal 1 homolog b (Drosophila) |
| chr5_+_33519943 | 1.32 |
ENSDART00000131316
|
ms4a17c.1
|
membrane-spanning 4-domains, subfamily A, member 17C.1 |
| chr4_-_16836006 | 1.31 |
ENSDART00000010777
|
ldhba
|
lactate dehydrogenase Ba |
| chr17_+_52823015 | 1.30 |
ENSDART00000160507
ENSDART00000186979 |
meis2a
|
Meis homeobox 2a |
| chr18_-_42333428 | 1.30 |
ENSDART00000034225
|
cntn5
|
contactin 5 |
| chr11_+_41243325 | 1.25 |
ENSDART00000170657
|
pax7a
|
paired box 7a |
| chr7_-_26457208 | 1.21 |
ENSDART00000173519
|
zgc:172079
|
zgc:172079 |
| chr23_+_10146542 | 1.20 |
ENSDART00000048073
|
zgc:171775
|
zgc:171775 |
| chr3_+_52545014 | 1.20 |
ENSDART00000018908
|
slc27a1a
|
solute carrier family 27 (fatty acid transporter), member 1a |
| chr11_+_25044082 | 1.17 |
ENSDART00000123263
|
phf20a
|
PHD finger protein 20, a |
| chr22_-_4644484 | 1.14 |
ENSDART00000167748
|
fbn2b
|
fibrillin 2b |
| chr9_-_48370645 | 1.14 |
ENSDART00000140185
|
col28a2a
|
collagen, type XXVIII, alpha 2a |
| chr19_+_5418006 | 1.12 |
ENSDART00000132874
|
eif1b
|
eukaryotic translation initiation factor 1B |
| chr20_-_45060241 | 1.09 |
ENSDART00000185227
|
klhl29
|
kelch-like family member 29 |
| chr23_+_25354856 | 1.05 |
ENSDART00000109023
ENSDART00000147440 |
fmnl3
|
formin-like 3 |
| chr4_-_5652030 | 1.05 |
ENSDART00000010903
|
rsph9
|
radial spoke head 9 homolog |
| chr9_-_1949915 | 1.05 |
ENSDART00000190712
|
hoxd3a
|
homeobox D3a |
| chr6_+_35362225 | 1.04 |
ENSDART00000133783
ENSDART00000102483 |
rgs4
|
regulator of G protein signaling 4 |
| chr1_+_5402476 | 1.02 |
ENSDART00000040204
|
tuba8l2
|
tubulin, alpha 8 like 2 |
| chr13_-_8892514 | 1.02 |
ENSDART00000144553
|
plekhh2
|
pleckstrin homology domain containing, family H (with MyTH4 domain) member 2 |
| chr1_+_45217619 | 1.00 |
ENSDART00000125037
|
EVI5L
|
si:ch211-239f4.1 |
| chr7_+_29080684 | 0.98 |
ENSDART00000173709
ENSDART00000173576 |
acd
|
ACD, shelterin complex subunit and telomerase recruitment factor |
| chr1_+_36437585 | 0.98 |
ENSDART00000189182
|
pou4f2
|
POU class 4 homeobox 2 |
| chr5_+_37785152 | 0.98 |
ENSDART00000053511
ENSDART00000189812 |
myo1ca
|
myosin Ic, paralog a |
| chr6_-_50203682 | 0.97 |
ENSDART00000083999
ENSDART00000143050 |
raly
|
RALY heterogeneous nuclear ribonucleoprotein |
| chr23_+_36052944 | 0.96 |
ENSDART00000103149
|
hoxc13a
|
homeobox C13a |
| chr2_+_19785540 | 0.93 |
ENSDART00000149789
|
ptbp2b
|
polypyrimidine tract binding protein 2b |
| chr2_+_35733335 | 0.93 |
ENSDART00000113489
|
rasal2
|
RAS protein activator like 2 |
| chr20_-_4883673 | 0.92 |
ENSDART00000145540
ENSDART00000053877 |
zdhhc14
|
zinc finger, DHHC-type containing 14 |
| chr8_-_45867358 | 0.92 |
ENSDART00000132810
|
adam9
|
ADAM metallopeptidase domain 9 |
| chr15_-_19772372 | 0.90 |
ENSDART00000152729
|
picalmb
|
phosphatidylinositol binding clathrin assembly protein b |
| chr13_-_50614639 | 0.87 |
ENSDART00000170527
|
vent
|
ventral expressed homeobox |
| chr22_-_17052381 | 0.85 |
ENSDART00000138382
|
nfia
|
nuclear factor I/A |
| chr11_-_21586157 | 0.82 |
ENSDART00000190095
|
srgap2
|
SLIT-ROBO Rho GTPase activating protein 2 |
| chr19_+_19747430 | 0.82 |
ENSDART00000166129
|
hoxa9a
|
homeobox A9a |
| chr16_+_10918252 | 0.81 |
ENSDART00000172949
|
pou2f2a
|
POU class 2 homeobox 2a |
| chr13_+_29519115 | 0.81 |
ENSDART00000086711
|
chst3a
|
carbohydrate (chondroitin 6) sulfotransferase 3a |
| chr11_+_6116503 | 0.80 |
ENSDART00000176170
|
nr2f6b
|
nuclear receptor subfamily 2, group F, member 6b |
| chr17_-_17764801 | 0.80 |
ENSDART00000155261
|
slirp
|
SRA stem-loop interacting RNA binding protein |
| chr1_+_27153859 | 0.79 |
ENSDART00000180184
|
bnc2
|
basonuclin 2 |
| chr19_+_19652439 | 0.79 |
ENSDART00000165934
|
hibadha
|
3-hydroxyisobutyrate dehydrogenase a |
| chr8_+_22977304 | 0.76 |
ENSDART00000136784
|
samd10a
|
sterile alpha motif domain containing 10a |
| chr8_+_23731483 | 0.74 |
ENSDART00000099751
|
fance
|
Fanconi anemia, complementation group E |
| chr5_-_63218919 | 0.73 |
ENSDART00000149979
|
tecta
|
tectorin alpha |
| chr20_+_27428505 | 0.72 |
ENSDART00000002641
|
kif26aa
|
kinesin family member 26Aa |
| chr19_-_1002959 | 0.71 |
ENSDART00000168138
|
ehmt2
|
euchromatic histone-lysine N-methyltransferase 2 |
| chr10_-_25591194 | 0.69 |
ENSDART00000131640
|
tiam1a
|
T cell lymphoma invasion and metastasis 1a |
| chr8_-_11324143 | 0.65 |
ENSDART00000008215
|
pip5k1bb
|
phosphatidylinositol-4-phosphate 5-kinase, type I, beta b |
| chr1_+_2321384 | 0.64 |
ENSDART00000157662
|
farp1
|
FERM, RhoGEF (ARHGEF) and pleckstrin domain protein 1 (chondrocyte-derived) |
| chr11_+_37049347 | 0.64 |
ENSDART00000109235
|
bicd2
|
bicaudal D homolog 2 (Drosophila) |
| chr20_-_27733683 | 0.63 |
ENSDART00000103317
ENSDART00000138139 |
zgc:153157
|
zgc:153157 |
| chr3_+_24190207 | 0.61 |
ENSDART00000034762
|
prr15la
|
proline rich 15-like a |
| chr17_-_26610814 | 0.61 |
ENSDART00000133402
ENSDART00000016608 |
mrpl57
|
mitochondrial ribosomal protein L57 |
| chr19_+_7810028 | 0.60 |
ENSDART00000081592
ENSDART00000140719 |
aqp10b
|
aquaporin 10b |
| chr12_-_11560794 | 0.59 |
ENSDART00000149098
ENSDART00000169975 |
plekha1b
|
pleckstrin homology domain containing, family A (phosphoinositide binding specific) member 1b |
| chr3_-_19200571 | 0.58 |
ENSDART00000131503
ENSDART00000012335 |
rfx1a
|
regulatory factor X, 1a (influences HLA class II expression) |
| chr3_+_52545400 | 0.58 |
ENSDART00000184183
|
slc27a1a
|
solute carrier family 27 (fatty acid transporter), member 1a |
| chr16_+_38940758 | 0.57 |
ENSDART00000102482
ENSDART00000136215 |
eny2
|
enhancer of yellow 2 homolog (Drosophila) |
| chr16_-_21903083 | 0.57 |
ENSDART00000165849
|
setdb1b
|
SET domain, bifurcated 1b |
| chr10_-_10864331 | 0.57 |
ENSDART00000122657
|
nrarpa
|
NOTCH regulated ankyrin repeat protein a |
| chr2_-_4925025 | 0.51 |
ENSDART00000160257
|
dlg1l
|
discs, large (Drosophila) homolog 1, like |
| chr6_-_28222592 | 0.50 |
ENSDART00000131126
|
bcl6a
|
B-cell CLL/lymphoma 6a (zinc finger protein 51) |
| chr8_+_20825987 | 0.49 |
ENSDART00000133309
|
si:ch211-133l5.4
|
si:ch211-133l5.4 |
| chr11_+_37049105 | 0.48 |
ENSDART00000155408
|
bicd2
|
bicaudal D homolog 2 (Drosophila) |
| chr8_+_12951155 | 0.48 |
ENSDART00000081601
|
cept1a
|
choline/ethanolamine phosphotransferase 1a |
| chr7_-_26844064 | 0.47 |
ENSDART00000162241
|
si:ch211-107p11.3
|
si:ch211-107p11.3 |
Network of associatons between targets according to the STRING database.
First level regulatory network of hoxc11a
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.9 | 5.8 | GO:0009449 | gamma-aminobutyric acid biosynthetic process(GO:0009449) |
| 1.2 | 9.2 | GO:0098700 | neurotransmitter loading into synaptic vesicle(GO:0098700) |
| 1.1 | 6.5 | GO:0035989 | tendon development(GO:0035989) |
| 1.0 | 8.0 | GO:1990845 | adaptive thermogenesis(GO:1990845) |
| 0.8 | 3.9 | GO:0035988 | chondrocyte proliferation(GO:0035988) |
| 0.7 | 2.8 | GO:0019532 | oxalate transport(GO:0019532) |
| 0.6 | 2.5 | GO:0017185 | peptidyl-lysine hydroxylation(GO:0017185) |
| 0.6 | 1.9 | GO:0003403 | optic vesicle formation(GO:0003403) |
| 0.6 | 6.9 | GO:0032926 | negative regulation of activin receptor signaling pathway(GO:0032926) |
| 0.6 | 8.1 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 0.5 | 2.6 | GO:0097510 | base-excision repair, AP site formation via deaminated base removal(GO:0097510) |
| 0.5 | 5.1 | GO:0006735 | glucose catabolic process(GO:0006007) NADH regeneration(GO:0006735) glycolytic process through fructose-6-phosphate(GO:0061615) glycolytic process through glucose-6-phosphate(GO:0061620) canonical glycolysis(GO:0061621) glucose catabolic process to pyruvate(GO:0061718) |
| 0.5 | 1.4 | GO:0006043 | glucosamine metabolic process(GO:0006041) glucosamine catabolic process(GO:0006043) |
| 0.4 | 4.9 | GO:0034394 | protein localization to cell surface(GO:0034394) |
| 0.4 | 1.7 | GO:0003322 | pancreatic A cell development(GO:0003322) |
| 0.4 | 3.5 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.4 | 9.7 | GO:0071542 | dopaminergic neuron differentiation(GO:0071542) |
| 0.4 | 1.1 | GO:2000793 | cell proliferation involved in heart valve morphogenesis(GO:0003249) regulation of cell proliferation involved in heart valve morphogenesis(GO:0003250) endocardial cell development(GO:0060958) cell proliferation involved in heart valve development(GO:2000793) |
| 0.4 | 3.0 | GO:0021628 | olfactory nerve development(GO:0021553) olfactory nerve morphogenesis(GO:0021627) olfactory nerve formation(GO:0021628) |
| 0.4 | 1.4 | GO:0042148 | strand invasion(GO:0042148) |
| 0.4 | 1.8 | GO:0071072 | heat generation(GO:0031649) regulation of heat generation(GO:0031650) positive regulation of heat generation(GO:0031652) regulation of phospholipid biosynthetic process(GO:0071071) negative regulation of phospholipid biosynthetic process(GO:0071072) |
| 0.4 | 1.1 | GO:1904158 | axonemal central apparatus assembly(GO:1904158) |
| 0.3 | 1.3 | GO:0061113 | pancreas morphogenesis(GO:0061113) |
| 0.3 | 1.0 | GO:0051973 | positive regulation of telomerase activity(GO:0051973) |
| 0.3 | 5.5 | GO:0098943 | neurotransmitter receptor transport, postsynaptic endosome to lysosome(GO:0098943) |
| 0.3 | 16.6 | GO:0003009 | skeletal muscle contraction(GO:0003009) |
| 0.3 | 4.4 | GO:0071698 | olfactory placode development(GO:0071698) |
| 0.3 | 2.5 | GO:0021910 | smoothened signaling pathway involved in ventral spinal cord patterning(GO:0021910) |
| 0.3 | 5.5 | GO:0014823 | response to activity(GO:0014823) |
| 0.3 | 1.5 | GO:0007624 | ultradian rhythm(GO:0007624) |
| 0.3 | 2.9 | GO:0070262 | peptidyl-serine dephosphorylation(GO:0070262) |
| 0.3 | 7.8 | GO:0051289 | protein homotetramerization(GO:0051289) |
| 0.2 | 1.2 | GO:0050938 | regulation of xanthophore differentiation(GO:0050938) |
| 0.2 | 5.1 | GO:0031952 | regulation of protein autophosphorylation(GO:0031952) |
| 0.2 | 2.8 | GO:0035094 | response to nicotine(GO:0035094) |
| 0.2 | 2.1 | GO:0015746 | tricarboxylic acid transport(GO:0006842) citrate transport(GO:0015746) |
| 0.2 | 4.0 | GO:0060088 | auditory receptor cell stereocilium organization(GO:0060088) |
| 0.2 | 5.0 | GO:0021555 | midbrain-hindbrain boundary morphogenesis(GO:0021555) |
| 0.2 | 2.2 | GO:0003310 | pancreatic A cell differentiation(GO:0003310) |
| 0.2 | 0.8 | GO:2001223 | negative regulation of neuron migration(GO:2001223) |
| 0.2 | 1.4 | GO:0000185 | activation of MAPKKK activity(GO:0000185) |
| 0.2 | 0.8 | GO:1902369 | negative regulation of RNA catabolic process(GO:1902369) |
| 0.2 | 8.6 | GO:0006096 | glycolytic process(GO:0006096) |
| 0.2 | 2.1 | GO:0005980 | polysaccharide catabolic process(GO:0000272) glycogen catabolic process(GO:0005980) glucan catabolic process(GO:0009251) cellular polysaccharide catabolic process(GO:0044247) |
| 0.1 | 0.4 | GO:0060898 | eye field cell fate commitment involved in camera-type eye formation(GO:0060898) |
| 0.1 | 1.3 | GO:0043984 | histone H4-K16 acetylation(GO:0043984) |
| 0.1 | 0.8 | GO:0006574 | valine catabolic process(GO:0006574) |
| 0.1 | 0.7 | GO:0035889 | otolith tethering(GO:0035889) |
| 0.1 | 2.8 | GO:0045773 | positive regulation of axon extension(GO:0045773) |
| 0.1 | 0.7 | GO:0051570 | regulation of histone H3-K9 methylation(GO:0051570) |
| 0.1 | 2.1 | GO:0032467 | positive regulation of cytokinesis(GO:0032467) |
| 0.1 | 2.7 | GO:0035435 | phosphate ion transmembrane transport(GO:0035435) |
| 0.1 | 3.1 | GO:0006828 | manganese ion transport(GO:0006828) |
| 0.1 | 3.3 | GO:0070593 | dendrite self-avoidance(GO:0070593) |
| 0.1 | 2.8 | GO:0032474 | otolith morphogenesis(GO:0032474) |
| 0.1 | 1.1 | GO:0072393 | microtubule anchoring at microtubule organizing center(GO:0072393) |
| 0.1 | 1.3 | GO:0072576 | liver morphogenesis(GO:0072576) |
| 0.1 | 3.0 | GO:0010508 | positive regulation of autophagy(GO:0010508) |
| 0.1 | 2.9 | GO:0045879 | negative regulation of smoothened signaling pathway(GO:0045879) |
| 0.1 | 3.5 | GO:0045746 | negative regulation of Notch signaling pathway(GO:0045746) |
| 0.1 | 9.9 | GO:0009880 | embryonic pattern specification(GO:0009880) |
| 0.1 | 0.8 | GO:0009954 | proximal/distal pattern formation(GO:0009954) |
| 0.1 | 3.8 | GO:0048247 | lymphocyte chemotaxis(GO:0048247) |
| 0.1 | 5.5 | GO:0030866 | cortical actin cytoskeleton organization(GO:0030866) |
| 0.1 | 0.6 | GO:0060017 | parathyroid gland development(GO:0060017) |
| 0.1 | 2.1 | GO:0006165 | nucleoside diphosphate phosphorylation(GO:0006165) |
| 0.1 | 2.0 | GO:0051209 | release of sequestered calcium ion into cytosol(GO:0051209) |
| 0.1 | 2.7 | GO:0021761 | limbic system development(GO:0021761) hypothalamus development(GO:0021854) |
| 0.1 | 0.9 | GO:0042661 | regulation of mesodermal cell fate specification(GO:0042661) |
| 0.1 | 0.6 | GO:0016578 | histone deubiquitination(GO:0016578) |
| 0.1 | 0.6 | GO:0002043 | blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:0002043) |
| 0.1 | 3.5 | GO:0018298 | protein-chromophore linkage(GO:0018298) |
| 0.1 | 4.2 | GO:0010842 | retina layer formation(GO:0010842) |
| 0.1 | 0.5 | GO:0006646 | phosphatidylethanolamine biosynthetic process(GO:0006646) |
| 0.1 | 0.8 | GO:0006044 | N-acetylglucosamine metabolic process(GO:0006044) |
| 0.1 | 0.6 | GO:0061647 | histone H3-K9 methylation(GO:0051567) histone H3-K9 modification(GO:0061647) |
| 0.1 | 0.4 | GO:0006501 | C-terminal protein lipidation(GO:0006501) |
| 0.0 | 0.2 | GO:0019859 | thymine catabolic process(GO:0006210) uracil catabolic process(GO:0006212) thymine metabolic process(GO:0019859) uracil metabolic process(GO:0019860) |
| 0.0 | 3.4 | GO:0031018 | endocrine pancreas development(GO:0031018) |
| 0.0 | 3.7 | GO:0000381 | regulation of alternative mRNA splicing, via spliceosome(GO:0000381) |
| 0.0 | 0.4 | GO:0050909 | sensory perception of taste(GO:0050909) |
| 0.0 | 0.3 | GO:0006833 | water transport(GO:0006833) |
| 0.0 | 1.4 | GO:0035336 | long-chain fatty-acyl-CoA metabolic process(GO:0035336) |
| 0.0 | 0.8 | GO:0039020 | pronephric nephron tubule development(GO:0039020) |
| 0.0 | 3.2 | GO:0008016 | regulation of heart contraction(GO:0008016) |
| 0.0 | 1.9 | GO:1902668 | negative regulation of axon extension involved in axon guidance(GO:0048843) negative regulation of axon guidance(GO:1902668) |
| 0.0 | 0.9 | GO:0070142 | synaptic vesicle budding from presynaptic endocytic zone membrane(GO:0016185) synaptic vesicle budding(GO:0070142) |
| 0.0 | 1.0 | GO:0018230 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 0.0 | 0.3 | GO:0016560 | protein import into peroxisome matrix, docking(GO:0016560) protein to membrane docking(GO:0022615) |
| 0.0 | 0.8 | GO:0003094 | glomerular filtration(GO:0003094) |
| 0.0 | 1.6 | GO:0016575 | histone deacetylation(GO:0016575) |
| 0.0 | 4.1 | GO:0035725 | sodium ion transmembrane transport(GO:0035725) |
| 0.0 | 4.7 | GO:0046488 | phosphatidylinositol metabolic process(GO:0046488) |
| 0.0 | 0.4 | GO:0060997 | dendritic spine morphogenesis(GO:0060997) |
| 0.0 | 1.4 | GO:0046513 | ceramide biosynthetic process(GO:0046513) |
| 0.0 | 1.0 | GO:0030050 | vesicle transport along actin filament(GO:0030050) actin filament-based transport(GO:0099515) |
| 0.0 | 3.6 | GO:0042391 | regulation of membrane potential(GO:0042391) |
| 0.0 | 0.7 | GO:0050772 | positive regulation of axonogenesis(GO:0050772) |
| 0.0 | 0.4 | GO:0034315 | regulation of Arp2/3 complex-mediated actin nucleation(GO:0034315) |
| 0.0 | 9.3 | GO:0007420 | brain development(GO:0007420) |
| 0.0 | 0.3 | GO:1900153 | regulation of nuclear-transcribed mRNA poly(A) tail shortening(GO:0060211) positive regulation of nuclear-transcribed mRNA poly(A) tail shortening(GO:0060213) regulation of mRNA catabolic process(GO:0061013) positive regulation of mRNA catabolic process(GO:0061014) regulation of nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay(GO:1900151) positive regulation of nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay(GO:1900153) |
| 0.0 | 0.5 | GO:0036297 | interstrand cross-link repair(GO:0036297) |
| 0.0 | 0.5 | GO:0050768 | negative regulation of neurogenesis(GO:0050768) |
| 0.0 | 1.2 | GO:0016573 | histone acetylation(GO:0016573) |
| 0.0 | 0.1 | GO:0016540 | protein autoprocessing(GO:0016540) |
| 0.0 | 1.4 | GO:0006888 | ER to Golgi vesicle-mediated transport(GO:0006888) |
| 0.0 | 1.0 | GO:0031101 | fin regeneration(GO:0031101) |
| 0.0 | 0.9 | GO:0007229 | integrin-mediated signaling pathway(GO:0007229) |
| 0.0 | 2.9 | GO:0030198 | extracellular matrix organization(GO:0030198) extracellular structure organization(GO:0043062) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 8.1 | GO:0070032 | synaptobrevin 2-SNAP-25-syntaxin-1a-complexin I complex(GO:0070032) |
| 0.6 | 9.2 | GO:0030285 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.5 | 5.1 | GO:0005945 | 6-phosphofructokinase complex(GO:0005945) |
| 0.4 | 1.7 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.4 | 17.9 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.4 | 16.6 | GO:0005861 | troponin complex(GO:0005861) |
| 0.4 | 1.1 | GO:0001534 | radial spoke(GO:0001534) |
| 0.3 | 8.1 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.2 | 1.0 | GO:0060171 | stereocilium membrane(GO:0060171) |
| 0.2 | 9.7 | GO:0044306 | axon terminus(GO:0043679) neuron projection terminus(GO:0044306) |
| 0.2 | 11.6 | GO:0016528 | sarcoplasm(GO:0016528) sarcoplasmic reticulum(GO:0016529) |
| 0.2 | 2.8 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.2 | 2.8 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.2 | 1.3 | GO:0072487 | MSL complex(GO:0072487) |
| 0.1 | 1.6 | GO:0016580 | Sin3 complex(GO:0016580) |
| 0.1 | 0.6 | GO:0070390 | transcription export complex 2(GO:0070390) |
| 0.1 | 10.0 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.1 | 1.9 | GO:0002116 | semaphorin receptor complex(GO:0002116) |
| 0.1 | 0.9 | GO:0098833 | presynaptic endocytic zone(GO:0098833) presynaptic endocytic zone membrane(GO:0098835) extrinsic component of presynaptic membrane(GO:0098888) extrinsic component of presynaptic endocytic zone(GO:0098894) |
| 0.1 | 1.2 | GO:0044545 | NSL complex(GO:0044545) |
| 0.1 | 3.0 | GO:0016605 | PML body(GO:0016605) |
| 0.1 | 3.5 | GO:0001750 | photoreceptor outer segment(GO:0001750) |
| 0.1 | 0.7 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.1 | 1.0 | GO:0070187 | telosome(GO:0070187) |
| 0.1 | 2.6 | GO:0005778 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.1 | 1.1 | GO:0016282 | eukaryotic 43S preinitiation complex(GO:0016282) |
| 0.0 | 0.4 | GO:0034045 | pre-autophagosomal structure membrane(GO:0034045) |
| 0.0 | 3.7 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.0 | 2.1 | GO:0031305 | integral component of mitochondrial inner membrane(GO:0031305) |
| 0.0 | 2.7 | GO:0044309 | dendritic spine(GO:0043197) neuron spine(GO:0044309) |
| 0.0 | 1.4 | GO:0000794 | condensed nuclear chromosome(GO:0000794) |
| 0.0 | 0.1 | GO:0010369 | chromocenter(GO:0010369) |
| 0.0 | 0.5 | GO:0031594 | neuromuscular junction(GO:0031594) |
| 0.0 | 11.9 | GO:0043005 | neuron projection(GO:0043005) |
| 0.0 | 4.5 | GO:0005911 | cell-cell junction(GO:0005911) |
| 0.0 | 3.1 | GO:0034703 | cation channel complex(GO:0034703) |
| 0.0 | 6.5 | GO:0031966 | mitochondrial membrane(GO:0031966) |
| 0.0 | 6.0 | GO:0045202 | synapse(GO:0045202) |
| 0.0 | 1.7 | GO:0030133 | transport vesicle(GO:0030133) |
| 0.0 | 2.1 | GO:0070161 | adherens junction(GO:0005912) anchoring junction(GO:0070161) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.2 | 9.7 | GO:1903136 | cuprous ion binding(GO:1903136) |
| 1.9 | 5.8 | GO:0004351 | glutamate decarboxylase activity(GO:0004351) |
| 1.6 | 6.4 | GO:0017077 | oxidative phosphorylation uncoupler activity(GO:0017077) |
| 1.4 | 8.6 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 1.0 | 13.4 | GO:0015185 | gamma-aminobutyric acid transmembrane transporter activity(GO:0015185) |
| 0.8 | 2.5 | GO:0070815 | peptidyl-lysine 5-dioxygenase activity(GO:0070815) |
| 0.8 | 7.8 | GO:0005522 | profilin binding(GO:0005522) |
| 0.6 | 2.6 | GO:0101006 | inorganic diphosphatase activity(GO:0004427) protein histidine phosphatase activity(GO:0101006) |
| 0.6 | 8.9 | GO:0004971 | AMPA glutamate receptor activity(GO:0004971) |
| 0.6 | 2.8 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.5 | 2.7 | GO:0030504 | inorganic diphosphate transmembrane transporter activity(GO:0030504) |
| 0.5 | 2.1 | GO:0004134 | glycogen debranching enzyme activity(GO:0004133) 4-alpha-glucanotransferase activity(GO:0004134) amylo-alpha-1,6-glucosidase activity(GO:0004135) |
| 0.5 | 1.4 | GO:0000150 | recombinase activity(GO:0000150) |
| 0.5 | 5.1 | GO:0003872 | 6-phosphofructokinase activity(GO:0003872) |
| 0.5 | 1.4 | GO:0004342 | glucosamine-6-phosphate deaminase activity(GO:0004342) |
| 0.4 | 2.1 | GO:0004127 | cytidylate kinase activity(GO:0004127) |
| 0.4 | 3.0 | GO:0003828 | alpha-N-acetylneuraminate alpha-2,8-sialyltransferase activity(GO:0003828) |
| 0.4 | 2.1 | GO:0015137 | citrate transmembrane transporter activity(GO:0015137) tricarboxylic acid transmembrane transporter activity(GO:0015142) |
| 0.3 | 6.9 | GO:0048185 | activin binding(GO:0048185) |
| 0.3 | 6.2 | GO:0033691 | sialic acid binding(GO:0033691) |
| 0.3 | 2.3 | GO:0080019 | fatty-acyl-CoA reductase (alcohol-forming) activity(GO:0080019) |
| 0.3 | 10.3 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.3 | 5.1 | GO:0019211 | phosphatase activator activity(GO:0019211) protein phosphatase activator activity(GO:0072542) |
| 0.3 | 6.4 | GO:0022848 | acetylcholine-gated cation channel activity(GO:0022848) |
| 0.3 | 1.8 | GO:0031957 | very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 0.2 | 2.6 | GO:0004844 | uracil DNA N-glycosylase activity(GO:0004844) deaminated base DNA N-glycosylase activity(GO:0097506) |
| 0.2 | 0.8 | GO:0008459 | chondroitin 6-sulfotransferase activity(GO:0008459) |
| 0.2 | 8.1 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.2 | 0.8 | GO:0008442 | 3-hydroxyisobutyrate dehydrogenase activity(GO:0008442) |
| 0.2 | 1.3 | GO:0004459 | L-lactate dehydrogenase activity(GO:0004459) |
| 0.2 | 3.2 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.2 | 2.0 | GO:0005220 | inositol 1,4,5-trisphosphate-sensitive calcium-release channel activity(GO:0005220) |
| 0.1 | 2.8 | GO:0019531 | oxalate transmembrane transporter activity(GO:0019531) |
| 0.1 | 1.6 | GO:0004046 | aminoacylase activity(GO:0004046) |
| 0.1 | 1.1 | GO:0034452 | dynactin binding(GO:0034452) |
| 0.1 | 3.1 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.1 | 4.7 | GO:0016307 | phosphatidylinositol phosphate kinase activity(GO:0016307) |
| 0.1 | 2.7 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.1 | 3.3 | GO:0098632 | protein binding involved in cell-cell adhesion(GO:0098632) |
| 0.1 | 0.5 | GO:0004142 | diacylglycerol cholinephosphotransferase activity(GO:0004142) ethanolaminephosphotransferase activity(GO:0004307) |
| 0.1 | 1.3 | GO:0004029 | aldehyde dehydrogenase (NAD) activity(GO:0004029) |
| 0.1 | 1.4 | GO:0050291 | sphingosine N-acyltransferase activity(GO:0050291) |
| 0.1 | 3.8 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.1 | 0.2 | GO:0004031 | aldehyde oxidase activity(GO:0004031) oxidoreductase activity, acting on the aldehyde or oxo group of donors, oxygen as acceptor(GO:0016623) |
| 0.1 | 2.6 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
| 0.1 | 3.5 | GO:0008020 | G-protein coupled photoreceptor activity(GO:0008020) |
| 0.1 | 0.9 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.1 | 10.9 | GO:0001228 | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001228) |
| 0.1 | 5.5 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.1 | 0.6 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) |
| 0.1 | 1.9 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.1 | 0.4 | GO:0015168 | glycerol transmembrane transporter activity(GO:0015168) |
| 0.1 | 2.9 | GO:0019888 | protein phosphatase regulator activity(GO:0019888) |
| 0.0 | 0.2 | GO:0002061 | uracil binding(GO:0002058) pyrimidine nucleobase binding(GO:0002061) dihydropyrimidine dehydrogenase (NADP+) activity(GO:0017113) |
| 0.0 | 0.4 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 0.0 | 1.7 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.0 | 0.7 | GO:0002039 | p53 binding(GO:0002039) |
| 0.0 | 0.8 | GO:0031267 | small GTPase binding(GO:0031267) |
| 0.0 | 6.5 | GO:0008083 | growth factor activity(GO:0008083) |
| 0.0 | 4.8 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 0.3 | GO:0005052 | peroxisome matrix targeting signal-1 binding(GO:0005052) |
| 0.0 | 5.2 | GO:0003714 | transcription corepressor activity(GO:0003714) |
| 0.0 | 1.2 | GO:0004707 | MAP kinase activity(GO:0004707) |
| 0.0 | 1.9 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.0 | 1.0 | GO:0042162 | telomeric DNA binding(GO:0042162) |
| 0.0 | 2.0 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.0 | 2.5 | GO:0045296 | cadherin binding(GO:0045296) |
| 0.0 | 0.1 | GO:0008798 | beta-aspartyl-peptidase activity(GO:0008798) |
| 0.0 | 1.4 | GO:0005262 | calcium channel activity(GO:0005262) |
| 0.0 | 1.0 | GO:0019707 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.0 | 0.4 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.0 | 0.6 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.0 | 1.0 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.0 | 3.0 | GO:0000287 | magnesium ion binding(GO:0000287) |
| 0.0 | 1.0 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.0 | 3.5 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 0.6 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.0 | 0.5 | GO:0019902 | phosphatase binding(GO:0019902) |
| 0.0 | 1.1 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.0 | 0.2 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.0 | 3.5 | GO:0044822 | mRNA binding(GO:0003729) poly(A) RNA binding(GO:0044822) |
| 0.0 | 1.6 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 5.3 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.1 | 6.2 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.0 | 1.7 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.0 | 0.9 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 4.6 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 0.7 | PID BARD1 PATHWAY | BARD1 signaling events |
| 0.0 | 1.9 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 1.0 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.0 | 0.7 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.0 | 0.8 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.0 | 0.6 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.0 | 0.2 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 15.0 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 0.4 | 5.1 | REACTOME PLATELET SENSITIZATION BY LDL | Genes involved in Platelet sensitization by LDL |
| 0.4 | 3.6 | REACTOME PRESYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Presynaptic nicotinic acetylcholine receptors |
| 0.3 | 6.2 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |
| 0.2 | 1.9 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.1 | 3.0 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.1 | 2.3 | REACTOME PEROXISOMAL LIPID METABOLISM | Genes involved in Peroxisomal lipid metabolism |
| 0.1 | 6.4 | REACTOME RESPIRATORY ELECTRON TRANSPORT ATP SYNTHESIS BY CHEMIOSMOTIC COUPLING AND HEAT PRODUCTION BY UNCOUPLING PROTEINS | Genes involved in Respiratory electron transport, ATP synthesis by chemiosmotic coupling, and heat production by uncoupling proteins. |
| 0.1 | 1.0 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.1 | 2.5 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 0.7 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.0 | 2.8 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.0 | 1.4 | REACTOME MEIOTIC RECOMBINATION | Genes involved in Meiotic Recombination |
| 0.0 | 0.7 | REACTOME RNA POL I TRANSCRIPTION INITIATION | Genes involved in RNA Polymerase I Transcription Initiation |
| 0.0 | 1.0 | REACTOME G ALPHA Z SIGNALLING EVENTS | Genes involved in G alpha (z) signalling events |
| 0.0 | 1.7 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.0 | 0.8 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 0.0 | 1.3 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.0 | 0.4 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |
| 0.0 | 2.1 | REACTOME ANTIGEN PROCESSING UBIQUITINATION PROTEASOME DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |