Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
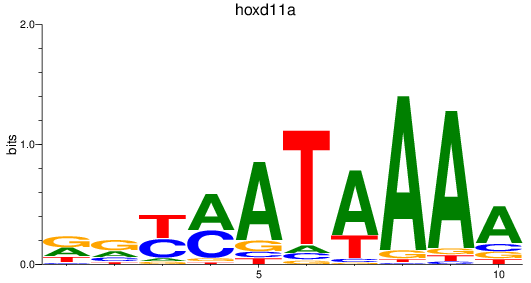
Results for hoxd11a
Z-value: 0.73
Transcription factors associated with hoxd11a
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
hoxd11a
|
ENSDARG00000059267 | homeobox D11a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| hoxd11a | dr11_v1_chr9_-_1978090_1978090 | 0.18 | 8.0e-02 | Click! |
Activity profile of hoxd11a motif
Sorted Z-values of hoxd11a motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr16_-_45917322 | 10.07 |
ENSDART00000060822
|
afp4
|
antifreeze protein type IV |
| chr23_+_39606108 | 10.01 |
ENSDART00000109464
|
g0s2
|
G0/G1 switch 2 |
| chr12_-_20373058 | 9.83 |
ENSDART00000066382
|
aqp8a.1
|
aquaporin 8a, tandem duplicate 1 |
| chr13_-_36034582 | 9.28 |
ENSDART00000133565
|
si:dkey-157l19.2
|
si:dkey-157l19.2 |
| chr23_+_7379728 | 8.12 |
ENSDART00000012194
|
gata5
|
GATA binding protein 5 |
| chr15_+_20403903 | 8.10 |
ENSDART00000134182
|
cox7a1
|
cytochrome c oxidase subunit VIIa polypeptide 1 (muscle) |
| chr1_+_17900306 | 7.52 |
ENSDART00000089480
|
cyp4v8
|
cytochrome P450, family 4, subfamily V, polypeptide 8 |
| chr4_+_7391110 | 6.88 |
ENSDART00000160708
ENSDART00000187823 |
tnni4a
|
troponin I4a |
| chr22_+_16308806 | 6.87 |
ENSDART00000162685
|
lrrc39
|
leucine rich repeat containing 39 |
| chr7_-_16562200 | 6.22 |
ENSDART00000169093
ENSDART00000173491 |
csrp3
|
cysteine and glycine-rich protein 3 (cardiac LIM protein) |
| chr15_+_14856307 | 5.68 |
ENSDART00000167213
|
diabloa
|
diablo, IAP-binding mitochondrial protein a |
| chr8_-_2616326 | 5.65 |
ENSDART00000027214
|
slc25a25a
|
solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 25a |
| chr11_-_25257045 | 5.60 |
ENSDART00000130477
|
snai1a
|
snail family zinc finger 1a |
| chr4_-_8043839 | 5.21 |
ENSDART00000190047
ENSDART00000057567 |
si:ch211-240l19.5
|
si:ch211-240l19.5 |
| chr13_-_22843562 | 5.17 |
ENSDART00000142738
|
pbld
|
phenazine biosynthesis like protein domain containing |
| chr17_-_15498275 | 5.10 |
ENSDART00000156905
ENSDART00000080661 |
si:ch211-266g18.10
|
si:ch211-266g18.10 |
| chr14_-_15171435 | 4.95 |
ENSDART00000159148
ENSDART00000166622 |
si:dkey-77g12.1
|
si:dkey-77g12.1 |
| chr25_+_29161609 | 4.86 |
ENSDART00000180752
|
pkmb
|
pyruvate kinase M1/2b |
| chr13_-_25719628 | 4.82 |
ENSDART00000135383
|
si:dkey-192p21.6
|
si:dkey-192p21.6 |
| chr8_+_30709685 | 4.77 |
ENSDART00000133989
|
upb1
|
ureidopropionase, beta |
| chr7_-_26306546 | 4.75 |
ENSDART00000140817
|
zgc:77439
|
zgc:77439 |
| chr1_-_9249943 | 4.74 |
ENSDART00000055011
|
zgc:136472
|
zgc:136472 |
| chr14_+_15495088 | 4.71 |
ENSDART00000165765
ENSDART00000188577 |
si:dkey-203a12.6
|
si:dkey-203a12.6 |
| chr3_+_19299309 | 4.61 |
ENSDART00000046297
ENSDART00000146955 |
ldlra
|
low density lipoprotein receptor a |
| chr8_+_40477264 | 4.54 |
ENSDART00000085559
|
gck
|
glucokinase (hexokinase 4) |
| chr20_+_26880668 | 4.43 |
ENSDART00000077769
|
serpinb1
|
serpin peptidase inhibitor, clade B (ovalbumin), member 1 |
| chr13_-_319996 | 4.32 |
ENSDART00000148675
|
ddx21
|
DEAD (Asp-Glu-Ala-Asp) box helicase 21 |
| chr14_+_32918172 | 4.22 |
ENSDART00000182867
|
lnx2b
|
ligand of numb-protein X 2b |
| chr1_-_14332283 | 4.20 |
ENSDART00000090025
|
wfs1a
|
Wolfram syndrome 1a (wolframin) |
| chr10_+_2587234 | 4.14 |
ENSDART00000126937
|
wu:fb59d01
|
wu:fb59d01 |
| chr16_+_13993285 | 4.11 |
ENSDART00000139130
ENSDART00000130353 |
si:dkey-85k15.7
fdps
|
si:dkey-85k15.7 farnesyl diphosphate synthase (farnesyl pyrophosphate synthetase, dimethylallyltranstransferase, geranyltranstransferase) |
| chr14_+_15597049 | 4.02 |
ENSDART00000159732
|
si:dkey-203a12.8
|
si:dkey-203a12.8 |
| chr11_-_18253111 | 4.01 |
ENSDART00000125984
|
mustn1b
|
musculoskeletal, embryonic nuclear protein 1b |
| chr21_-_19918286 | 3.88 |
ENSDART00000180816
|
ppp1r3b
|
protein phosphatase 1, regulatory subunit 3B |
| chr21_+_11969603 | 3.84 |
ENSDART00000142247
ENSDART00000140652 |
mlnl
|
motilin-like |
| chr6_+_14301214 | 3.84 |
ENSDART00000129491
|
tmem182b
|
transmembrane protein 182b |
| chr23_+_18722715 | 3.77 |
ENSDART00000137438
|
myh7bb
|
myosin, heavy chain 7B, cardiac muscle, beta b |
| chr20_+_52546186 | 3.74 |
ENSDART00000110777
ENSDART00000153377 ENSDART00000153013 ENSDART00000042704 |
eef1db
|
eukaryotic translation elongation factor 1 delta b (guanine nucleotide exchange protein) |
| chr11_-_25257595 | 3.65 |
ENSDART00000123567
|
snai1a
|
snail family zinc finger 1a |
| chr22_-_24818066 | 3.59 |
ENSDART00000143443
|
vtg6
|
vitellogenin 6 |
| chr8_+_28467893 | 3.59 |
ENSDART00000189724
|
slc52a3
|
solute carrier family 52 (riboflavin transporter), member 3 |
| chr23_+_44349252 | 3.56 |
ENSDART00000097644
|
ephb4b
|
eph receptor B4b |
| chr11_+_24313931 | 3.41 |
ENSDART00000017599
ENSDART00000166045 |
rem1
|
RAS (RAD and GEM)-like GTP-binding 1 |
| chr2_+_20406399 | 3.33 |
ENSDART00000006817
ENSDART00000137848 |
palmda
|
palmdelphin a |
| chr21_-_30648106 | 3.30 |
ENSDART00000160800
ENSDART00000177022 |
phka1b
|
phosphorylase kinase, alpha 1b (muscle) |
| chr11_+_42556395 | 3.29 |
ENSDART00000039206
|
rps23
|
ribosomal protein S23 |
| chr18_-_15559817 | 3.26 |
ENSDART00000061681
|
si:ch211-245j22.3
|
si:ch211-245j22.3 |
| chr20_-_33961697 | 3.23 |
ENSDART00000061765
|
selp
|
selectin P |
| chr3_+_49097775 | 3.16 |
ENSDART00000169185
|
zgc:123284
|
zgc:123284 |
| chr14_-_33613794 | 3.13 |
ENSDART00000010022
|
xpnpep2
|
X-prolyl aminopeptidase (aminopeptidase P) 2, membrane-bound |
| chr24_-_32025637 | 3.09 |
ENSDART00000180448
ENSDART00000159034 |
rsu1
|
Ras suppressor protein 1 |
| chr10_-_13116337 | 3.09 |
ENSDART00000164568
|
musk
|
muscle, skeletal, receptor tyrosine kinase |
| chr9_-_41784799 | 3.04 |
ENSDART00000144573
ENSDART00000112542 ENSDART00000190486 |
obsl1b
|
obscurin-like 1b |
| chr17_-_2690083 | 2.98 |
ENSDART00000135374
|
ptpn21
|
protein tyrosine phosphatase, non-receptor type 21 |
| chr22_-_4644484 | 2.92 |
ENSDART00000167748
|
fbn2b
|
fibrillin 2b |
| chr24_+_7782313 | 2.92 |
ENSDART00000111090
|
ptprh
|
protein tyrosine phosphatase, receptor type, h |
| chr24_+_39105051 | 2.90 |
ENSDART00000115297
|
mss51
|
MSS51 mitochondrial translational activator |
| chr2_+_52847049 | 2.90 |
ENSDART00000121980
|
creb3l3b
|
cAMP responsive element binding protein 3-like 3b |
| chr7_-_60096318 | 2.83 |
ENSDART00000189125
|
BX511067.1
|
|
| chr23_+_27068225 | 2.76 |
ENSDART00000054238
|
mipa
|
major intrinsic protein of lens fiber a |
| chr21_-_39024754 | 2.75 |
ENSDART00000056878
|
traf4b
|
tnf receptor-associated factor 4b |
| chr8_-_38159805 | 2.75 |
ENSDART00000112331
ENSDART00000180006 |
adgra2
|
adhesion G protein-coupled receptor A2 |
| chr21_-_33032715 | 2.74 |
ENSDART00000065349
ENSDART00000191502 ENSDART00000149386 |
slc34a1b
|
solute carrier family 34 (type II sodium/phosphate cotransporter), member 1b |
| chr23_+_2666944 | 2.73 |
ENSDART00000192861
|
CABZ01057928.1
|
|
| chr7_+_18075504 | 2.61 |
ENSDART00000173689
|
si:ch73-40a2.1
|
si:ch73-40a2.1 |
| chr14_+_34490445 | 2.60 |
ENSDART00000132193
ENSDART00000148044 |
wnt8a
|
wingless-type MMTV integration site family, member 8a |
| chr17_+_8799661 | 2.54 |
ENSDART00000105326
|
tonsl
|
tonsoku-like, DNA repair protein |
| chr23_+_8797143 | 2.53 |
ENSDART00000132992
|
sox18
|
SRY (sex determining region Y)-box 18 |
| chr13_-_37619159 | 2.44 |
ENSDART00000186348
|
zgc:152791
|
zgc:152791 |
| chr5_-_29531948 | 2.43 |
ENSDART00000098360
|
arrdc1a
|
arrestin domain containing 1a |
| chr9_+_54290896 | 2.40 |
ENSDART00000149175
|
pou4f3
|
POU class 4 homeobox 3 |
| chr15_+_29024895 | 2.39 |
ENSDART00000141164
ENSDART00000144126 |
si:ch211-137a8.2
|
si:ch211-137a8.2 |
| chr4_+_14899720 | 2.36 |
ENSDART00000187456
|
akr1b1
|
aldo-keto reductase family 1, member B1 (aldose reductase) |
| chr3_-_24456451 | 2.35 |
ENSDART00000024480
ENSDART00000156814 |
baiap2l2a
|
BAI1-associated protein 2-like 2a |
| chr5_-_23715861 | 2.34 |
ENSDART00000019992
|
gbgt1l1
|
globoside alpha-1,3-N-acetylgalactosaminyltransferase 1, like 1 |
| chr17_+_23554932 | 2.32 |
ENSDART00000135814
|
pank1a
|
pantothenate kinase 1a |
| chr11_-_15090118 | 2.24 |
ENSDART00000171118
|
slc1a8a
|
solute carrier family 1 (glutamate transporter), member 8a |
| chr9_-_23990416 | 2.19 |
ENSDART00000113176
|
col6a3
|
collagen, type VI, alpha 3 |
| chr8_+_41037541 | 2.15 |
ENSDART00000129344
|
gpat2
|
glycerol-3-phosphate acyltransferase 2, mitochondrial |
| chr16_-_10547781 | 2.14 |
ENSDART00000166495
|
g6fl
|
g6f-like |
| chr7_-_24112484 | 2.13 |
ENSDART00000111923
|
ajuba
|
ajuba LIM protein |
| chr10_+_35526528 | 2.13 |
ENSDART00000184110
|
phldb2a
|
pleckstrin homology-like domain, family B, member 2a |
| chr24_+_5237753 | 2.13 |
ENSDART00000106488
ENSDART00000005901 |
plod2
|
procollagen-lysine, 2-oxoglutarate 5-dioxygenase 2 |
| chr2_-_43196595 | 2.12 |
ENSDART00000141087
|
crema
|
cAMP responsive element modulator a |
| chr9_+_41080029 | 2.01 |
ENSDART00000141179
ENSDART00000019289 |
zgc:136439
|
zgc:136439 |
| chr24_+_29912509 | 2.01 |
ENSDART00000168422
|
frrs1b
|
ferric-chelate reductase 1b |
| chr9_+_30275139 | 1.98 |
ENSDART00000131781
|
rpgra
|
retinitis pigmentosa GTPase regulator a |
| chr15_-_44052927 | 1.96 |
ENSDART00000166209
|
wu:fb44b02
|
wu:fb44b02 |
| chr22_-_14115292 | 1.95 |
ENSDART00000105717
ENSDART00000165670 |
aox5
|
aldehyde oxidase 5 |
| chr24_+_1042594 | 1.95 |
ENSDART00000109117
|
si:dkey-192l18.9
|
si:dkey-192l18.9 |
| chr11_-_1392468 | 1.92 |
ENSDART00000004423
|
iars
|
isoleucyl-tRNA synthetase |
| chr22_-_32360464 | 1.90 |
ENSDART00000148886
|
pcbp4
|
poly(rC) binding protein 4 |
| chr7_+_66822229 | 1.86 |
ENSDART00000112109
|
lyve1a
|
lymphatic vessel endothelial hyaluronic receptor 1a |
| chr10_+_40660772 | 1.84 |
ENSDART00000148007
|
taar19l
|
trace amine associated receptor 19l |
| chr7_-_7692723 | 1.83 |
ENSDART00000183352
|
aadat
|
aminoadipate aminotransferase |
| chr22_-_30999000 | 1.83 |
ENSDART00000109841
|
ssuh2.3
|
ssu-2 homolog, tandem duplicate 3 |
| chr23_+_36052944 | 1.83 |
ENSDART00000103149
|
hoxc13a
|
homeobox C13a |
| chr2_-_30734098 | 1.83 |
ENSDART00000133769
|
rp1
|
retinitis pigmentosa 1 (autosomal dominant) |
| chr11_-_15090564 | 1.80 |
ENSDART00000162079
|
slc1a8a
|
solute carrier family 1 (glutamate transporter), member 8a |
| chr17_+_15788100 | 1.80 |
ENSDART00000027667
|
rragd
|
ras-related GTP binding D |
| chr10_+_26990095 | 1.79 |
ENSDART00000064111
|
faub
|
Finkel-Biskis-Reilly murine sarcoma virus (FBR-MuSV) ubiquitously expressed b |
| chr21_+_16980141 | 1.77 |
ENSDART00000101241
|
aqp3b
|
aquaporin 3b |
| chr15_+_31344472 | 1.75 |
ENSDART00000146695
ENSDART00000159182 ENSDART00000060125 |
or107-1
|
odorant receptor, family D, subfamily 107, member 1 |
| chr16_+_54829574 | 1.74 |
ENSDART00000148392
|
pabpc1a
|
poly(A) binding protein, cytoplasmic 1a |
| chr21_+_20396858 | 1.73 |
ENSDART00000003299
ENSDART00000146615 |
zgc:103482
|
zgc:103482 |
| chr7_+_2467057 | 1.70 |
ENSDART00000154517
|
si:dkey-125e8.3
|
si:dkey-125e8.3 |
| chr3_-_34089310 | 1.69 |
ENSDART00000151774
|
ighv5-5
|
immunoglobulin heavy variable 5-5 |
| chr19_-_14191592 | 1.69 |
ENSDART00000164594
|
tbxta
|
T-box transcription factor Ta |
| chr25_+_16356083 | 1.68 |
ENSDART00000125925
ENSDART00000125444 |
tead1a
|
TEA domain family member 1a |
| chr23_+_36087219 | 1.68 |
ENSDART00000154825
|
hoxc3a
|
homeobox C3a |
| chr7_+_24522308 | 1.67 |
ENSDART00000173542
|
btr09
|
bloodthirsty-related gene family, member 9 |
| chr20_+_46202188 | 1.67 |
ENSDART00000100523
|
taar13c
|
trace amine associated receptor 13c |
| chr7_-_7692992 | 1.63 |
ENSDART00000192619
|
aadat
|
aminoadipate aminotransferase |
| chr19_+_19761966 | 1.59 |
ENSDART00000163697
|
hoxa3a
|
homeobox A3a |
| chr24_+_25258904 | 1.59 |
ENSDART00000155714
|
gabrr3b
|
gamma-aminobutyric acid (GABA) A receptor, rho 3b |
| chr25_+_3549584 | 1.57 |
ENSDART00000165913
|
ccdc77
|
coiled-coil domain containing 77 |
| chr21_+_39197628 | 1.57 |
ENSDART00000113607
|
cpdb
|
carboxypeptidase D, b |
| chr14_-_4120636 | 1.57 |
ENSDART00000059230
|
irf2
|
interferon regulatory factor 2 |
| chr16_-_28658341 | 1.57 |
ENSDART00000148456
|
abcb4
|
ATP-binding cassette, sub-family B (MDR/TAP), member 4 |
| chr21_-_17296010 | 1.57 |
ENSDART00000114877
|
gfi1b
|
growth factor independent 1B transcription repressor |
| chr19_-_617246 | 1.50 |
ENSDART00000062551
|
cyp51
|
cytochrome P450, family 51 |
| chr10_+_41199660 | 1.49 |
ENSDART00000125314
|
adrb3b
|
adrenoceptor beta 3b |
| chr3_+_23710839 | 1.49 |
ENSDART00000151584
|
hoxb4a
|
homeobox B4a |
| chr10_+_36178713 | 1.49 |
ENSDART00000140816
|
or108-3
|
odorant receptor, family D, subfamily 108, member 3 |
| chr11_-_38899816 | 1.47 |
ENSDART00000141515
|
pimr127
|
Pim proto-oncogene, serine/threonine kinase, related 127 |
| chr3_+_49074008 | 1.46 |
ENSDART00000168864
|
zgc:112146
|
zgc:112146 |
| chr10_+_26612321 | 1.41 |
ENSDART00000134322
|
fhl1b
|
four and a half LIM domains 1b |
| chr12_-_11258404 | 1.40 |
ENSDART00000149229
|
si:ch73-30l9.1
|
si:ch73-30l9.1 |
| chr9_-_34986827 | 1.38 |
ENSDART00000137862
|
si:ch211-160b11.4
|
si:ch211-160b11.4 |
| chr22_-_9736050 | 1.38 |
ENSDART00000152919
|
si:dkey-286j17.4
|
si:dkey-286j17.4 |
| chr8_-_25338709 | 1.37 |
ENSDART00000131616
|
atp5pb
|
ATP synthase peripheral stalk-membrane subunit b |
| chr19_+_7810028 | 1.37 |
ENSDART00000081592
ENSDART00000140719 |
aqp10b
|
aquaporin 10b |
| chr1_+_44173245 | 1.35 |
ENSDART00000159450
ENSDART00000106048 ENSDART00000157763 |
ctnnd1
|
catenin (cadherin-associated protein), delta 1 |
| chr3_-_55121125 | 1.34 |
ENSDART00000125092
|
hbae1
|
hemoglobin, alpha embryonic 1 |
| chr16_+_54829293 | 1.32 |
ENSDART00000024729
|
pabpc1a
|
poly(A) binding protein, cytoplasmic 1a |
| chr9_-_23152092 | 1.32 |
ENSDART00000180155
ENSDART00000186935 |
lypd6b
|
LY6/PLAUR domain containing 6B |
| chr15_-_563877 | 1.32 |
ENSDART00000128032
|
cbln18
|
cerebellin 18 |
| chr12_-_18872717 | 1.30 |
ENSDART00000126300
|
shisa8b
|
shisa family member 8b |
| chr14_+_36497250 | 1.28 |
ENSDART00000184727
|
TENM3
|
si:dkey-237h12.3 |
| chr17_+_25519089 | 1.28 |
ENSDART00000041721
|
aim1a
|
crystallin beta-gamma domain containing 1a |
| chr7_+_18989241 | 1.26 |
ENSDART00000053213
|
zgc:194252
|
zgc:194252 |
| chr24_+_30392834 | 1.26 |
ENSDART00000162555
|
dpyda.1
|
dihydropyrimidine dehydrogenase a, tandem duplicate 1 |
| chr12_+_19356623 | 1.26 |
ENSDART00000078284
|
dmc1
|
DNA meiotic recombinase 1 |
| chr23_-_43718067 | 1.25 |
ENSDART00000015777
|
abce1
|
ATP-binding cassette, sub-family E (OABP), member 1 |
| chr2_-_32637592 | 1.25 |
ENSDART00000136353
|
si:dkeyp-73d8.8
|
si:dkeyp-73d8.8 |
| chr3_+_52545400 | 1.25 |
ENSDART00000184183
|
slc27a1a
|
solute carrier family 27 (fatty acid transporter), member 1a |
| chr20_-_49889111 | 1.24 |
ENSDART00000058858
|
kif13bb
|
kinesin family member 13Bb |
| chr3_+_31851004 | 1.23 |
ENSDART00000181422
|
BX324157.1
|
|
| chr24_-_27452488 | 1.22 |
ENSDART00000136433
|
ccl34b.8
|
chemokine (C-C motif) ligand 34b, duplicate 8 |
| chr20_-_34028967 | 1.21 |
ENSDART00000153408
ENSDART00000033817 |
scyl3
|
SCY1-like, kinase-like 3 |
| chr10_+_506538 | 1.20 |
ENSDART00000141713
|
si:ch211-242f23.3
|
si:ch211-242f23.3 |
| chr20_+_25879826 | 1.18 |
ENSDART00000018519
|
zgc:153896
|
zgc:153896 |
| chr6_+_13083146 | 1.17 |
ENSDART00000172158
|
l3hypdh
|
trans-L-3-hydroxyproline dehydratase |
| chr14_+_80685 | 1.16 |
ENSDART00000188443
|
stag3
|
stromal antigen 3 |
| chr25_+_15933411 | 1.16 |
ENSDART00000191581
|
ppfibp2b
|
PTPRF interacting protein, binding protein 2b (liprin beta 2) |
| chr12_-_18872927 | 1.15 |
ENSDART00000187717
|
shisa8b
|
shisa family member 8b |
| chr25_-_13614702 | 1.15 |
ENSDART00000165510
ENSDART00000190959 |
fa2h
|
fatty acid 2-hydroxylase |
| chr17_+_17764979 | 1.14 |
ENSDART00000105013
|
alkbh1
|
alkB homolog 1, histone H2A dioxygenase |
| chr15_+_5343186 | 1.13 |
ENSDART00000170086
|
CABZ01018874.1
|
|
| chr2_-_40196547 | 1.11 |
ENSDART00000168098
|
ccl34a.3
|
chemokine (C-C motif) ligand 34a, duplicate 3 |
| chr5_+_44805269 | 1.09 |
ENSDART00000136965
|
ctsla
|
cathepsin La |
| chr3_+_13482542 | 1.06 |
ENSDART00000161287
|
CABZ01079251.1
|
|
| chr20_+_43379029 | 1.06 |
ENSDART00000142486
ENSDART00000186486 |
unc93a
|
unc-93 homolog A |
| chr17_-_50355512 | 1.04 |
ENSDART00000041685
|
otofb
|
otoferlin b |
| chr3_-_31115601 | 1.02 |
ENSDART00000139090
|
arl6ip1
|
ADP-ribosylation factor-like 6 interacting protein 1 |
| chr3_+_52545014 | 1.00 |
ENSDART00000018908
|
slc27a1a
|
solute carrier family 27 (fatty acid transporter), member 1a |
| chr13_+_30169681 | 0.99 |
ENSDART00000138326
|
pald1b
|
phosphatase domain containing, paladin 1b |
| chr2_-_28420415 | 0.99 |
ENSDART00000183857
|
CABZ01056051.1
|
|
| chr14_+_3071110 | 0.99 |
ENSDART00000162445
|
hmgxb3
|
HMG box domain containing 3 |
| chr10_-_42882215 | 0.96 |
ENSDART00000180580
ENSDART00000187374 |
FP236542.1
|
|
| chr21_-_11970199 | 0.96 |
ENSDART00000114524
|
nop56
|
NOP56 ribonucleoprotein homolog |
| chr21_+_3775551 | 0.95 |
ENSDART00000111699
|
tor1
|
torsin family 1 |
| chr18_+_30508729 | 0.94 |
ENSDART00000185140
|
cox4i1
|
cytochrome c oxidase subunit IV isoform 1 |
| chr8_-_38317914 | 0.93 |
ENSDART00000125920
|
pdlim2
|
PDZ and LIM domain 2 (mystique) |
| chr15_-_16946124 | 0.92 |
ENSDART00000154923
|
hip1
|
huntingtin interacting protein 1 |
| chr10_-_8060573 | 0.92 |
ENSDART00000147104
ENSDART00000099030 |
si:ch211-251f6.6
|
si:ch211-251f6.6 |
| chr2_+_19578079 | 0.91 |
ENSDART00000144413
|
pimr50
|
Pim proto-oncogene, serine/threonine kinase, related 50 |
| chr22_-_10158038 | 0.91 |
ENSDART00000047444
|
rbck1
|
RanBP-type and C3HC4-type zinc finger containing 1 |
| chr6_+_60004871 | 0.88 |
ENSDART00000179832
|
CABZ01079267.1
|
|
| chr15_-_31406093 | 0.87 |
ENSDART00000123444
|
or111-8
|
odorant receptor, family D, subfamily 111, member 8 |
| chr16_+_20934353 | 0.87 |
ENSDART00000052660
|
skap2
|
src kinase associated phosphoprotein 2 |
| chr16_-_28268201 | 0.86 |
ENSDART00000121671
ENSDART00000141911 |
si:dkey-12j5.1
|
si:dkey-12j5.1 |
| chr6_+_52869892 | 0.85 |
ENSDART00000146143
|
si:dkeyp-3f10.16
|
si:dkeyp-3f10.16 |
| chr20_-_39367895 | 0.84 |
ENSDART00000136476
ENSDART00000021788 ENSDART00000180784 |
pbk
|
PDZ binding kinase |
| chr9_+_3282369 | 0.84 |
ENSDART00000044128
|
hat1
|
histone acetyltransferase 1 |
| chr15_+_29727799 | 0.83 |
ENSDART00000182006
|
zgc:153372
|
zgc:153372 |
| chr11_+_25257022 | 0.82 |
ENSDART00000156052
|
tp53inp2
|
tumor protein p53 inducible nuclear protein 2 |
| chr21_+_27278120 | 0.82 |
ENSDART00000193882
|
si:dkey-175m17.7
|
si:dkey-175m17.7 |
| chr12_+_27117609 | 0.82 |
ENSDART00000076154
|
hoxb8b
|
homeobox B8b |
| chr10_-_25328814 | 0.80 |
ENSDART00000123820
|
tmem135
|
transmembrane protein 135 |
| chr8_-_25033681 | 0.79 |
ENSDART00000003493
|
nfyal
|
nuclear transcription factor Y, alpha, like |
| chr25_+_33063762 | 0.78 |
ENSDART00000189974
|
tln2b
|
talin 2b |
| chr7_-_23873173 | 0.75 |
ENSDART00000101408
|
zgc:162171
|
zgc:162171 |
| chr20_-_2713120 | 0.74 |
ENSDART00000138753
ENSDART00000104606 |
rars2
|
arginyl-tRNA synthetase 2, mitochondrial (putative) |
| chr3_-_26183699 | 0.74 |
ENSDART00000147517
ENSDART00000140731 |
si:ch211-11k18.4
|
si:ch211-11k18.4 |
| chr15_+_29276508 | 0.74 |
ENSDART00000170537
ENSDART00000126559 |
rap1gap2a
|
RAP1 GTPase activating protein 2a |
| chr1_+_44826593 | 0.73 |
ENSDART00000162200
|
STX3
|
zgc:165520 |
| chr21_+_5580948 | 0.72 |
ENSDART00000160373
|
ly6m7
|
lymphocyte antigen 6 family member M7 |
| chr10_-_3332362 | 0.71 |
ENSDART00000007577
ENSDART00000055140 |
tor4aa
|
torsin family 4, member Aa |
| chr5_+_11290851 | 0.67 |
ENSDART00000180408
|
CABZ01077124.1
|
|
Network of associatons between targets according to the STRING database.
First level regulatory network of hoxd11a
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.5 | 7.5 | GO:0010430 | fatty acid omega-oxidation(GO:0010430) |
| 2.0 | 8.1 | GO:0048618 | post-embryonic foregut morphogenesis(GO:0048618) |
| 1.6 | 6.2 | GO:0035994 | response to muscle stretch(GO:0035994) |
| 1.3 | 8.1 | GO:0097250 | mitochondrial respiratory chain supercomplex assembly(GO:0097250) |
| 1.3 | 3.9 | GO:0005981 | regulation of glycogen catabolic process(GO:0005981) |
| 1.1 | 5.7 | GO:0008635 | activation of cysteine-type endopeptidase activity involved in apoptotic process by cytochrome c(GO:0008635) |
| 1.0 | 2.9 | GO:0003249 | cell proliferation involved in heart valve morphogenesis(GO:0003249) regulation of cell proliferation involved in heart valve morphogenesis(GO:0003250) endocardial cell development(GO:0060958) cell proliferation involved in heart valve development(GO:2000793) |
| 1.0 | 4.8 | GO:0019483 | beta-alanine metabolic process(GO:0019482) beta-alanine biosynthetic process(GO:0019483) |
| 0.9 | 2.7 | GO:0022009 | central nervous system vasculogenesis(GO:0022009) |
| 0.9 | 2.6 | GO:0039015 | spinal cord anterior/posterior patterning(GO:0021512) cell proliferation involved in pronephros development(GO:0039015) cell proliferation involved in kidney development(GO:0072111) |
| 0.8 | 4.0 | GO:0035988 | chondrocyte proliferation(GO:0035988) |
| 0.8 | 4.6 | GO:0032367 | intracellular cholesterol transport(GO:0032367) |
| 0.7 | 2.1 | GO:0060148 | negative regulation of hippo signaling(GO:0035331) positive regulation of posttranscriptional gene silencing(GO:0060148) positive regulation of gene silencing by miRNA(GO:2000637) |
| 0.7 | 2.8 | GO:0007250 | activation of NF-kappaB-inducing kinase activity(GO:0007250) |
| 0.7 | 4.1 | GO:0045337 | farnesyl diphosphate biosynthetic process(GO:0045337) |
| 0.7 | 3.4 | GO:1901841 | regulation of high voltage-gated calcium channel activity(GO:1901841) negative regulation of high voltage-gated calcium channel activity(GO:1901842) |
| 0.6 | 1.3 | GO:0006212 | uracil catabolic process(GO:0006212) uracil metabolic process(GO:0019860) |
| 0.6 | 3.6 | GO:0032218 | riboflavin transport(GO:0032218) |
| 0.5 | 3.2 | GO:0097241 | hematopoietic stem cell migration to bone marrow(GO:0097241) |
| 0.5 | 2.1 | GO:0017185 | peptidyl-lysine hydroxylation(GO:0017185) |
| 0.5 | 4.1 | GO:0006833 | water transport(GO:0006833) |
| 0.5 | 10.6 | GO:0008078 | mesodermal cell migration(GO:0008078) |
| 0.5 | 2.7 | GO:0044341 | sodium-dependent phosphate transport(GO:0044341) |
| 0.5 | 4.5 | GO:0032024 | positive regulation of insulin secretion(GO:0032024) |
| 0.4 | 2.2 | GO:0071071 | heat generation(GO:0031649) regulation of heat generation(GO:0031650) positive regulation of heat generation(GO:0031652) regulation of phospholipid biosynthetic process(GO:0071071) negative regulation of phospholipid biosynthetic process(GO:0071072) |
| 0.4 | 1.7 | GO:0072045 | convergent extension involved in nephron morphogenesis(GO:0072045) |
| 0.4 | 3.1 | GO:0071340 | skeletal muscle acetylcholine-gated channel clustering(GO:0071340) |
| 0.4 | 1.9 | GO:0033119 | negative regulation of RNA splicing(GO:0033119) negative regulation of mRNA splicing, via spliceosome(GO:0048025) |
| 0.3 | 10.1 | GO:0060030 | dorsal convergence(GO:0060030) |
| 0.3 | 1.3 | GO:0042148 | strand invasion(GO:0042148) |
| 0.3 | 2.2 | GO:1990511 | piRNA biosynthetic process(GO:1990511) |
| 0.3 | 1.5 | GO:0002025 | vasodilation by norepinephrine-epinephrine involved in regulation of systemic arterial blood pressure(GO:0002025) |
| 0.3 | 0.8 | GO:0018872 | arsonoacetate metabolic process(GO:0018872) |
| 0.3 | 2.4 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.3 | 2.3 | GO:2000251 | positive regulation of actin cytoskeleton reorganization(GO:2000251) |
| 0.2 | 6.9 | GO:0055008 | cardiac muscle tissue morphogenesis(GO:0055008) |
| 0.2 | 1.7 | GO:0050911 | detection of chemical stimulus involved in sensory perception of smell(GO:0050911) |
| 0.2 | 1.1 | GO:0035513 | oxidative RNA demethylation(GO:0035513) |
| 0.2 | 0.9 | GO:0071763 | nuclear membrane organization(GO:0071763) |
| 0.2 | 0.5 | GO:0050968 | chemosensory behavior(GO:0007635) detection of chemical stimulus involved in sensory perception of pain(GO:0050968) |
| 0.2 | 1.3 | GO:0060017 | parathyroid gland development(GO:0060017) |
| 0.2 | 0.9 | GO:0097039 | protein linear polyubiquitination(GO:0097039) |
| 0.2 | 1.7 | GO:0045948 | positive regulation of translational initiation(GO:0045948) |
| 0.2 | 2.3 | GO:0015937 | coenzyme A biosynthetic process(GO:0015937) |
| 0.2 | 0.8 | GO:0043508 | negative regulation of JUN kinase activity(GO:0043508) |
| 0.2 | 7.1 | GO:0030968 | endoplasmic reticulum unfolded protein response(GO:0030968) |
| 0.2 | 0.5 | GO:0071314 | response to cocaine(GO:0042220) cellular response to alkaloid(GO:0071312) cellular response to cocaine(GO:0071314) |
| 0.1 | 0.6 | GO:0038093 | Fc receptor signaling pathway(GO:0038093) |
| 0.1 | 2.3 | GO:0030259 | lipid glycosylation(GO:0030259) |
| 0.1 | 6.9 | GO:0003009 | skeletal muscle contraction(GO:0003009) |
| 0.1 | 2.5 | GO:0031297 | replication fork processing(GO:0031297) |
| 0.1 | 4.2 | GO:0070936 | protein K48-linked ubiquitination(GO:0070936) |
| 0.1 | 1.2 | GO:0072386 | plus-end-directed vesicle transport along microtubule(GO:0072383) plus-end-directed organelle transport along microtubule(GO:0072386) |
| 0.1 | 2.1 | GO:0035723 | interleukin-15-mediated signaling pathway(GO:0035723) response to interleukin-15(GO:0070672) cellular response to interleukin-15(GO:0071350) |
| 0.1 | 0.4 | GO:0010032 | meiotic chromosome condensation(GO:0010032) |
| 0.1 | 0.8 | GO:0006348 | chromatin silencing at telomere(GO:0006348) |
| 0.1 | 3.6 | GO:0032355 | response to estradiol(GO:0032355) |
| 0.1 | 1.0 | GO:0010996 | response to auditory stimulus(GO:0010996) |
| 0.1 | 4.9 | GO:0006096 | glycolytic process(GO:0006096) |
| 0.1 | 9.3 | GO:0002088 | lens development in camera-type eye(GO:0002088) |
| 0.1 | 0.9 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.1 | 0.9 | GO:1901970 | positive regulation of mitotic metaphase/anaphase transition(GO:0045842) positive regulation of mitotic sister chromatid separation(GO:1901970) positive regulation of metaphase/anaphase transition of cell cycle(GO:1902101) |
| 0.1 | 1.8 | GO:0032008 | positive regulation of TOR signaling(GO:0032008) |
| 0.1 | 1.4 | GO:1990798 | pancreas regeneration(GO:1990798) |
| 0.1 | 4.4 | GO:0001946 | lymphangiogenesis(GO:0001946) |
| 0.1 | 1.3 | GO:0033753 | ribosomal subunit export from nucleus(GO:0000054) ribosome localization(GO:0033750) establishment of ribosome localization(GO:0033753) rRNA-containing ribonucleoprotein complex export from nucleus(GO:0071428) |
| 0.1 | 1.7 | GO:0035329 | hippo signaling(GO:0035329) |
| 0.1 | 3.0 | GO:0001837 | epithelial to mesenchymal transition(GO:0001837) |
| 0.1 | 1.0 | GO:0006613 | cotranslational protein targeting to membrane(GO:0006613) |
| 0.1 | 3.7 | GO:0006414 | translational elongation(GO:0006414) |
| 0.1 | 4.4 | GO:0010951 | negative regulation of endopeptidase activity(GO:0010951) |
| 0.1 | 0.4 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
| 0.1 | 1.5 | GO:1902653 | cholesterol biosynthetic process(GO:0006695) secondary alcohol biosynthetic process(GO:1902653) |
| 0.1 | 0.5 | GO:0006388 | tRNA splicing, via endonucleolytic cleavage and ligation(GO:0006388) |
| 0.1 | 5.3 | GO:0071466 | cellular response to xenobiotic stimulus(GO:0071466) |
| 0.1 | 0.5 | GO:0006857 | oligopeptide transport(GO:0006857) |
| 0.0 | 3.3 | GO:0006073 | glycogen metabolic process(GO:0005977) cellular glucan metabolic process(GO:0006073) glucan metabolic process(GO:0044042) |
| 0.0 | 1.1 | GO:0015985 | energy coupled proton transport, down electrochemical gradient(GO:0015985) ATP synthesis coupled proton transport(GO:0015986) |
| 0.0 | 3.6 | GO:0070121 | Kupffer's vesicle development(GO:0070121) |
| 0.0 | 0.5 | GO:0043951 | negative regulation of cAMP-mediated signaling(GO:0043951) |
| 0.0 | 3.8 | GO:0019722 | calcium-mediated signaling(GO:0019722) |
| 0.0 | 0.7 | GO:0032438 | melanosome organization(GO:0032438) |
| 0.0 | 0.8 | GO:0010508 | positive regulation of autophagy(GO:0010508) |
| 0.0 | 0.3 | GO:0032986 | nucleosome disassembly(GO:0006337) chromatin disassembly(GO:0031498) protein-DNA complex disassembly(GO:0032986) |
| 0.0 | 2.2 | GO:0008045 | motor neuron axon guidance(GO:0008045) |
| 0.0 | 3.0 | GO:0007416 | synapse assembly(GO:0007416) |
| 0.0 | 0.5 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.0 | 0.7 | GO:0006418 | tRNA aminoacylation for protein translation(GO:0006418) |
| 0.0 | 1.8 | GO:0035082 | axoneme assembly(GO:0035082) |
| 0.0 | 1.7 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.0 | 2.1 | GO:0006909 | phagocytosis(GO:0006909) |
| 0.0 | 0.3 | GO:0050909 | sensory perception of taste(GO:0050909) |
| 0.0 | 2.7 | GO:0034976 | response to endoplasmic reticulum stress(GO:0034976) |
| 0.0 | 0.2 | GO:1905097 | regulation of guanyl-nucleotide exchange factor activity(GO:1905097) regulation of Rho guanyl-nucleotide exchange factor activity(GO:2001106) |
| 0.0 | 3.0 | GO:0008360 | regulation of cell shape(GO:0008360) |
| 0.0 | 0.9 | GO:0042113 | B cell activation(GO:0042113) |
| 0.0 | 0.5 | GO:0071456 | cellular response to hypoxia(GO:0071456) |
| 0.0 | 1.6 | GO:0060216 | definitive hemopoiesis(GO:0060216) |
| 0.0 | 1.2 | GO:0007528 | neuromuscular junction development(GO:0007528) |
| 0.0 | 3.1 | GO:0006417 | regulation of translation(GO:0006417) |
| 0.0 | 1.5 | GO:1902476 | chloride transmembrane transport(GO:1902476) |
| 0.0 | 5.1 | GO:0006412 | translation(GO:0006412) |
| 0.0 | 1.8 | GO:0031101 | fin regeneration(GO:0031101) |
| 0.0 | 1.6 | GO:0006518 | peptide metabolic process(GO:0006518) |
| 0.0 | 0.1 | GO:0034312 | diol biosynthetic process(GO:0034312) sphingosine biosynthetic process(GO:0046512) |
| 0.0 | 0.5 | GO:0046466 | sphingolipid catabolic process(GO:0030149) membrane lipid catabolic process(GO:0046466) |
| 0.0 | 1.2 | GO:0009615 | response to virus(GO:0009615) |
| 0.0 | 0.5 | GO:0051262 | protein tetramerization(GO:0051262) |
| 0.0 | 0.3 | GO:0030574 | collagen catabolic process(GO:0030574) multicellular organism catabolic process(GO:0044243) |
| 0.0 | 0.1 | GO:0051754 | meiotic sister chromatid cohesion, centromeric(GO:0051754) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 2.6 | GO:0097189 | apoptotic body(GO:0097189) |
| 0.7 | 2.7 | GO:1990909 | Wnt signalosome(GO:1990909) |
| 0.5 | 3.7 | GO:0005853 | eukaryotic translation elongation factor 1 complex(GO:0005853) |
| 0.4 | 2.5 | GO:0005662 | DNA replication factor A complex(GO:0005662) |
| 0.4 | 1.8 | GO:0034448 | EGO complex(GO:0034448) Gtr1-Gtr2 GTPase complex(GO:1990131) |
| 0.3 | 6.9 | GO:0031430 | M band(GO:0031430) |
| 0.3 | 3.3 | GO:0005964 | phosphorylase kinase complex(GO:0005964) |
| 0.2 | 3.9 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.2 | 5.1 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.2 | 1.0 | GO:0005784 | Sec61 translocon complex(GO:0005784) |
| 0.2 | 2.1 | GO:1990589 | ATF4-CREB1 transcription factor complex(GO:1990589) |
| 0.2 | 6.9 | GO:0005861 | troponin complex(GO:0005861) |
| 0.2 | 1.0 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.2 | 2.4 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.1 | 2.7 | GO:0005903 | brush border(GO:0005903) |
| 0.1 | 0.4 | GO:0031251 | PAN complex(GO:0031251) |
| 0.1 | 2.4 | GO:0034992 | microtubule organizing center attachment site(GO:0034992) LINC complex(GO:0034993) |
| 0.1 | 9.0 | GO:0005746 | mitochondrial respiratory chain(GO:0005746) |
| 0.1 | 5.1 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.1 | 0.9 | GO:0071797 | LUBAC complex(GO:0071797) |
| 0.1 | 0.3 | GO:0031213 | RSF complex(GO:0031213) |
| 0.1 | 2.1 | GO:0045180 | basal cortex(GO:0045180) |
| 0.1 | 3.1 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.1 | 1.4 | GO:0000276 | mitochondrial proton-transporting ATP synthase complex, coupling factor F(o)(GO:0000276) |
| 0.1 | 0.5 | GO:0072669 | tRNA-splicing ligase complex(GO:0072669) |
| 0.1 | 7.1 | GO:0030018 | Z disc(GO:0030018) |
| 0.1 | 0.1 | GO:0035253 | ciliary rootlet(GO:0035253) |
| 0.1 | 0.5 | GO:0097225 | sperm midpiece(GO:0097225) |
| 0.1 | 1.3 | GO:0005852 | eukaryotic translation initiation factor 3 complex(GO:0005852) |
| 0.0 | 1.5 | GO:1902711 | GABA receptor complex(GO:1902710) GABA-A receptor complex(GO:1902711) |
| 0.0 | 1.2 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.0 | 1.0 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
| 0.0 | 3.9 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.0 | 1.9 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.0 | 1.7 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.0 | 4.2 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.0 | 2.1 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.0 | 1.7 | GO:0005776 | autophagosome(GO:0005776) |
| 0.0 | 0.4 | GO:0034751 | aryl hydrocarbon receptor complex(GO:0034751) |
| 0.0 | 6.3 | GO:0005743 | mitochondrial inner membrane(GO:0005743) |
| 0.0 | 1.3 | GO:0000794 | condensed nuclear chromosome(GO:0000794) |
| 0.0 | 0.5 | GO:1990907 | beta-catenin-TCF complex(GO:1990907) |
| 0.0 | 1.8 | GO:0005930 | axoneme(GO:0005930) |
| 0.0 | 0.7 | GO:0048770 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.0 | 0.2 | GO:0016602 | CCAAT-binding factor complex(GO:0016602) |
| 0.0 | 1.2 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.0 | 5.0 | GO:0005730 | nucleolus(GO:0005730) |
| 0.0 | 0.3 | GO:0044666 | MLL3/4 complex(GO:0044666) |
| 0.0 | 0.4 | GO:0008278 | cohesin complex(GO:0008278) |
| 0.0 | 1.2 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.0 | 1.7 | GO:0016459 | myosin complex(GO:0016459) |
| 0.0 | 0.8 | GO:0001726 | ruffle(GO:0001726) |
| 0.0 | 9.6 | GO:0005739 | mitochondrion(GO:0005739) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.3 | 9.3 | GO:0051059 | NF-kappaB binding(GO:0051059) |
| 1.4 | 12.6 | GO:0015250 | water channel activity(GO:0015250) |
| 1.4 | 4.1 | GO:0004337 | dimethylallyltranstransferase activity(GO:0004161) geranyltranstransferase activity(GO:0004337) |
| 1.2 | 4.6 | GO:0030228 | lipoprotein particle receptor activity(GO:0030228) |
| 0.8 | 4.9 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 0.8 | 4.5 | GO:0008865 | glucokinase activity(GO:0004340) hexokinase activity(GO:0004396) glucose binding(GO:0005536) fructokinase activity(GO:0008865) mannokinase activity(GO:0019158) |
| 0.7 | 2.1 | GO:0070815 | peptidyl-lysine 5-dioxygenase activity(GO:0070815) |
| 0.6 | 1.9 | GO:0004031 | aldehyde oxidase activity(GO:0004031) oxidoreductase activity, acting on the aldehyde or oxo group of donors, oxygen as acceptor(GO:0016623) |
| 0.6 | 3.6 | GO:0032217 | riboflavin transporter activity(GO:0032217) |
| 0.5 | 4.8 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 0.5 | 3.1 | GO:0005372 | water transmembrane transporter activity(GO:0005372) |
| 0.5 | 1.6 | GO:0015462 | protein-transmembrane transporting ATPase activity(GO:0015462) efflux transmembrane transporter activity(GO:0015562) |
| 0.5 | 3.6 | GO:0045735 | nutrient reservoir activity(GO:0045735) |
| 0.5 | 3.5 | GO:0016212 | kynurenine-oxoglutarate transaminase activity(GO:0016212) kynurenine aminotransferase activity(GO:0036137) |
| 0.5 | 2.4 | GO:1990756 | protein binding, bridging involved in substrate recognition for ubiquitination(GO:1990756) |
| 0.5 | 2.7 | GO:0005436 | sodium:phosphate symporter activity(GO:0005436) |
| 0.4 | 1.3 | GO:0000150 | recombinase activity(GO:0000150) |
| 0.4 | 2.3 | GO:0004594 | pantothenate kinase activity(GO:0004594) |
| 0.4 | 1.5 | GO:0051380 | norepinephrine binding(GO:0051380) |
| 0.3 | 3.1 | GO:0038064 | protein tyrosine kinase collagen receptor activity(GO:0038062) collagen receptor activity(GO:0038064) |
| 0.3 | 9.0 | GO:0016676 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors(GO:0016675) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.3 | 2.2 | GO:0031957 | very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 0.3 | 1.1 | GO:0035516 | oxidative DNA demethylase activity(GO:0035516) |
| 0.3 | 0.8 | GO:0030792 | arsenite methyltransferase activity(GO:0030791) methylarsonite methyltransferase activity(GO:0030792) |
| 0.3 | 2.2 | GO:0004366 | glycerol-3-phosphate O-acyltransferase activity(GO:0004366) |
| 0.3 | 7.1 | GO:0042805 | actinin binding(GO:0042805) |
| 0.3 | 3.9 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.3 | 1.3 | GO:0017113 | uracil binding(GO:0002058) pyrimidine nucleobase binding(GO:0002061) dihydropyrimidine dehydrogenase (NADP+) activity(GO:0017113) |
| 0.2 | 1.7 | GO:0004984 | olfactory receptor activity(GO:0004984) |
| 0.2 | 1.3 | GO:0099602 | acetylcholine receptor regulator activity(GO:0030548) neurotransmitter receptor regulator activity(GO:0099602) |
| 0.2 | 3.1 | GO:0008266 | poly(U) RNA binding(GO:0008266) |
| 0.2 | 2.9 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.2 | 0.5 | GO:0016496 | substance P receptor activity(GO:0016496) |
| 0.1 | 0.4 | GO:0080132 | fatty acid alpha-hydroxylase activity(GO:0080132) |
| 0.1 | 3.4 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.1 | 0.5 | GO:0008117 | sphinganine-1-phosphate aldolase activity(GO:0008117) |
| 0.1 | 5.7 | GO:0005347 | ATP transmembrane transporter activity(GO:0005347) |
| 0.1 | 1.3 | GO:0043024 | ribosomal small subunit binding(GO:0043024) |
| 0.1 | 0.5 | GO:0060182 | apelin receptor activity(GO:0060182) |
| 0.1 | 3.1 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
| 0.1 | 2.1 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 0.1 | 10.3 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.1 | 1.0 | GO:0035612 | AP-2 adaptor complex binding(GO:0035612) |
| 0.1 | 0.8 | GO:0010485 | H4 histone acetyltransferase activity(GO:0010485) |
| 0.1 | 7.7 | GO:0004722 | protein serine/threonine phosphatase activity(GO:0004722) |
| 0.1 | 3.7 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.1 | 1.2 | GO:0031729 | CCR1 chemokine receptor binding(GO:0031726) CCR4 chemokine receptor binding(GO:0031729) |
| 0.1 | 2.6 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.1 | 1.6 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.1 | 0.2 | GO:0034416 | bisphosphoglycerate phosphatase activity(GO:0034416) |
| 0.1 | 4.8 | GO:0016811 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amides(GO:0016811) |
| 0.1 | 4.3 | GO:0003724 | RNA helicase activity(GO:0003724) |
| 0.1 | 1.9 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.1 | 9.2 | GO:0004497 | monooxygenase activity(GO:0004497) |
| 0.0 | 1.0 | GO:0030515 | snoRNA binding(GO:0030515) |
| 0.0 | 1.5 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 0.0 | 0.9 | GO:0019894 | kinesin binding(GO:0019894) |
| 0.0 | 0.9 | GO:0005003 | ephrin receptor activity(GO:0005003) |
| 0.0 | 0.8 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.0 | 6.9 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.0 | 1.7 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.0 | 0.6 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.0 | 2.6 | GO:0004714 | transmembrane receptor protein tyrosine kinase activity(GO:0004714) |
| 0.0 | 0.7 | GO:0004812 | aminoacyl-tRNA ligase activity(GO:0004812) ligase activity, forming carbon-oxygen bonds(GO:0016875) ligase activity, forming aminoacyl-tRNA and related compounds(GO:0016876) |
| 0.0 | 0.9 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.0 | 0.5 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.0 | 1.8 | GO:0001594 | trace-amine receptor activity(GO:0001594) |
| 0.0 | 5.1 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 2.9 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 0.8 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.0 | 0.1 | GO:0016454 | serine C-palmitoyltransferase activity(GO:0004758) C-palmitoyltransferase activity(GO:0016454) |
| 0.0 | 0.2 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.0 | 3.3 | GO:0005516 | calmodulin binding(GO:0005516) |
| 0.0 | 0.8 | GO:0004993 | G-protein coupled serotonin receptor activity(GO:0004993) |
| 0.0 | 3.0 | GO:0016853 | isomerase activity(GO:0016853) |
| 0.0 | 0.5 | GO:0008381 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 0.0 | 2.0 | GO:0016779 | nucleotidyltransferase activity(GO:0016779) |
| 0.0 | 1.2 | GO:0016836 | hydro-lyase activity(GO:0016836) |
| 0.0 | 0.3 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.0 | 0.8 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 3.2 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 0.0 | 0.3 | GO:0042800 | histone methyltransferase activity (H3-K4 specific)(GO:0042800) |
| 0.0 | 2.9 | GO:0003774 | motor activity(GO:0003774) |
| 0.0 | 0.3 | GO:0047617 | acyl-CoA hydrolase activity(GO:0047617) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 4.5 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 3.2 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.1 | 3.0 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.1 | 2.5 | ST GA13 PATHWAY | G alpha 13 Pathway |
| 0.1 | 1.9 | ST INTERLEUKIN 4 PATHWAY | Interleukin 4 (IL-4) Pathway |
| 0.1 | 1.4 | PID ECADHERIN KERATINOCYTE PATHWAY | E-cadherin signaling in keratinocytes |
| 0.1 | 2.2 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.0 | 6.6 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 0.6 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.0 | 1.8 | PID MTOR 4PATHWAY | mTOR signaling pathway |
| 0.0 | 2.6 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.0 | 0.5 | PID S1P META PATHWAY | Sphingosine 1-phosphate (S1P) pathway |
| 0.0 | 1.9 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
| 0.0 | 0.4 | PID AURORA B PATHWAY | Aurora B signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 4.8 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.3 | 3.5 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.2 | 4.5 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.2 | 3.1 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.2 | 4.1 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.1 | 10.4 | REACTOME PPARA ACTIVATES GENE EXPRESSION | Genes involved in PPARA Activates Gene Expression |
| 0.1 | 0.9 | REACTOME SIGNAL REGULATORY PROTEIN SIRP FAMILY INTERACTIONS | Genes involved in Signal regulatory protein (SIRP) family interactions |
| 0.1 | 2.2 | REACTOME SYNTHESIS OF PA | Genes involved in Synthesis of PA |
| 0.1 | 4.3 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.1 | 1.6 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION | Genes involved in TRAF6 mediated IRF7 activation |
| 0.1 | 2.4 | REACTOME METABOLISM OF STEROID HORMONES AND VITAMINS A AND D | Genes involved in Metabolism of steroid hormones and vitamins A and D |
| 0.1 | 6.7 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.1 | 1.9 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 0.1 | 3.3 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.1 | 1.4 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.1 | 3.5 | REACTOME CELL SURFACE INTERACTIONS AT THE VASCULAR WALL | Genes involved in Cell surface interactions at the vascular wall |
| 0.0 | 0.7 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.0 | 0.7 | REACTOME MITOCHONDRIAL TRNA AMINOACYLATION | Genes involved in Mitochondrial tRNA aminoacylation |
| 0.0 | 2.4 | REACTOME MEIOSIS | Genes involved in Meiosis |
| 0.0 | 1.0 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.0 | 0.6 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 0.4 | REACTOME CONVERSION FROM APC C CDC20 TO APC C CDH1 IN LATE ANAPHASE | Genes involved in Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase |
| 0.0 | 0.9 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |