Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
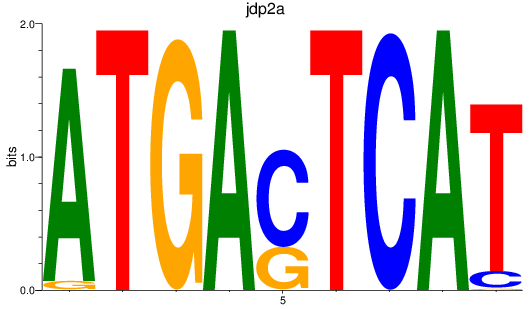
Results for jdp2a
Z-value: 1.06
Transcription factors associated with jdp2a
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
jdp2a
|
ENSDARG00000040137 | Jun dimerization protein 2a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| jdp2a | dr11_v1_chr17_-_50220228_50220228 | 0.69 | 8.1e-15 | Click! |
Activity profile of jdp2a motif
Sorted Z-values of jdp2a motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_-_21832441 | 20.17 |
ENSDART00000151272
ENSDART00000151442 ENSDART00000150168 ENSDART00000148797 ENSDART00000128196 ENSDART00000149259 ENSDART00000052556 ENSDART00000149658 ENSDART00000149639 ENSDART00000148424 |
mbpa
|
myelin basic protein a |
| chr7_-_13381129 | 12.42 |
ENSDART00000164326
|
si:ch73-119p20.1
|
si:ch73-119p20.1 |
| chr9_+_38372216 | 11.40 |
ENSDART00000141895
|
plcd4b
|
phospholipase C, delta 4b |
| chr11_-_3552067 | 11.30 |
ENSDART00000163656
|
CAMK2N1
|
si:dkey-33m11.6 |
| chr2_+_47581997 | 10.72 |
ENSDART00000112579
|
scg2b
|
secretogranin II (chromogranin C), b |
| chr1_+_45323142 | 10.48 |
ENSDART00000132210
|
emp1
|
epithelial membrane protein 1 |
| chr1_+_12766351 | 10.18 |
ENSDART00000165785
|
pcdh10a
|
protocadherin 10a |
| chr12_+_7497882 | 9.17 |
ENSDART00000134608
|
phyhiplb
|
phytanoyl-CoA 2-hydroxylase interacting protein-like b |
| chr1_+_31113951 | 8.66 |
ENSDART00000129362
|
eef1a1b
|
eukaryotic translation elongation factor 1 alpha 1b |
| chr2_+_47582681 | 8.63 |
ENSDART00000187579
|
scg2b
|
secretogranin II (chromogranin C), b |
| chr10_+_4499943 | 8.58 |
ENSDART00000125299
|
plk2a
|
polo-like kinase 2a (Drosophila) |
| chr1_-_45647846 | 8.55 |
ENSDART00000186881
|
BX511120.1
|
|
| chr25_+_30196039 | 8.35 |
ENSDART00000005299
|
hsd17b12a
|
hydroxysteroid (17-beta) dehydrogenase 12a |
| chr7_+_48460239 | 8.16 |
ENSDART00000052113
|
lingo1b
|
leucine rich repeat and Ig domain containing 1b |
| chr14_-_34044369 | 8.14 |
ENSDART00000149396
ENSDART00000123607 ENSDART00000190746 |
cyfip2
|
cytoplasmic FMR1 interacting protein 2 |
| chr6_+_40354424 | 7.98 |
ENSDART00000047416
|
slc4a8
|
solute carrier family 4, sodium bicarbonate cotransporter, member 8 |
| chr5_+_34549845 | 7.96 |
ENSDART00000139317
|
aif1l
|
allograft inflammatory factor 1-like |
| chr7_-_52963493 | 7.92 |
ENSDART00000052029
|
cart3
|
cocaine- and amphetamine-regulated transcript 3 |
| chr14_+_34486629 | 7.61 |
ENSDART00000131861
|
tmsb2
|
thymosin beta 2 |
| chr25_+_21324588 | 7.39 |
ENSDART00000151842
|
lrrn3a
|
leucine rich repeat neuronal 3a |
| chr18_+_21408794 | 7.16 |
ENSDART00000140161
|
necab2
|
N-terminal EF-hand calcium binding protein 2 |
| chr7_+_23577723 | 7.01 |
ENSDART00000049885
|
si:dkey-172j4.3
|
si:dkey-172j4.3 |
| chr23_+_4253957 | 6.96 |
ENSDART00000008761
|
arl6ip5b
|
ADP-ribosylation factor-like 6 interacting protein 5b |
| chr1_+_45323400 | 6.92 |
ENSDART00000148906
ENSDART00000132366 |
emp1
|
epithelial membrane protein 1 |
| chr2_+_30379650 | 6.89 |
ENSDART00000129542
|
crispld1b
|
cysteine-rich secretory protein LCCL domain containing 1b |
| chr24_-_24849091 | 6.83 |
ENSDART00000133649
ENSDART00000038290 |
crhb
|
corticotropin releasing hormone b |
| chr20_-_40319890 | 6.83 |
ENSDART00000075112
|
clvs2
|
clavesin 2 |
| chr4_-_69189894 | 6.77 |
ENSDART00000169596
|
si:ch211-209j12.1
|
si:ch211-209j12.1 |
| chr3_+_24986145 | 6.68 |
ENSDART00000055428
|
cbx7a
|
chromobox homolog 7a |
| chr19_-_103289 | 6.56 |
ENSDART00000143118
|
adgrb1b
|
adhesion G protein-coupled receptor B1b |
| chr17_-_7861219 | 6.33 |
ENSDART00000148604
|
syne1b
|
spectrin repeat containing, nuclear envelope 1b |
| chr13_-_21701323 | 6.14 |
ENSDART00000164112
|
si:dkey-191g9.7
|
si:dkey-191g9.7 |
| chr7_+_20587506 | 6.07 |
ENSDART00000172416
ENSDART00000170633 |
si:dkey-19b23.7
|
si:dkey-19b23.7 |
| chr14_+_24935131 | 6.05 |
ENSDART00000170871
|
ppargc1b
|
peroxisome proliferator-activated receptor gamma, coactivator 1 beta |
| chr8_-_14375890 | 5.89 |
ENSDART00000090306
|
xpr1a
|
xenotropic and polytropic retrovirus receptor 1a |
| chr5_-_14564878 | 5.81 |
ENSDART00000160511
|
sema4c
|
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4C |
| chr24_-_24797455 | 5.57 |
ENSDART00000138741
|
pde7a
|
phosphodiesterase 7A |
| chr8_-_22288004 | 5.52 |
ENSDART00000100042
|
si:ch211-147a11.3
|
si:ch211-147a11.3 |
| chr10_-_34772211 | 5.49 |
ENSDART00000145450
ENSDART00000134307 |
dclk1a
|
doublecortin-like kinase 1a |
| chr5_+_59534286 | 5.48 |
ENSDART00000189341
|
gtf2ird1
|
GTF2I repeat domain containing 1 |
| chr3_+_30921246 | 5.35 |
ENSDART00000076850
|
cldni
|
claudin i |
| chr10_+_45128375 | 5.21 |
ENSDART00000164805
|
camk2b2
|
calcium/calmodulin-dependent protein kinase (CaM kinase) II beta 2 |
| chr1_-_22861348 | 5.11 |
ENSDART00000139412
|
SMIM18
|
si:dkey-92j12.6 |
| chr8_+_24854600 | 5.07 |
ENSDART00000156570
|
slc6a17
|
solute carrier family 6 (neutral amino acid transporter), member 17 |
| chr12_+_20336070 | 4.98 |
ENSDART00000066385
|
zgc:163057
|
zgc:163057 |
| chr7_+_7019911 | 4.80 |
ENSDART00000172421
|
rbm14b
|
RNA binding motif protein 14b |
| chr19_-_5805923 | 4.78 |
ENSDART00000134340
|
si:ch211-264f5.8
|
si:ch211-264f5.8 |
| chr2_-_32738535 | 4.74 |
ENSDART00000135293
|
nrbp2a
|
nuclear receptor binding protein 2a |
| chr18_-_13360106 | 4.73 |
ENSDART00000091512
|
cmip
|
c-Maf inducing protein |
| chr17_+_33495194 | 4.72 |
ENSDART00000033691
|
pth2
|
parathyroid hormone 2 |
| chr7_+_23875269 | 4.68 |
ENSDART00000101406
|
rab39bb
|
RAB39B, member RAS oncogene family b |
| chr7_+_38770167 | 4.66 |
ENSDART00000190827
|
arhgap1
|
Rho GTPase activating protein 1 |
| chr3_+_31177972 | 4.54 |
ENSDART00000185954
|
clec19a
|
C-type lectin domain containing 19A |
| chr6_-_33685325 | 4.51 |
ENSDART00000181883
|
mast2
|
microtubule associated serine/threonine kinase 2 |
| chr8_+_44549290 | 4.42 |
ENSDART00000160801
|
si:dkey-220o5.5
|
si:dkey-220o5.5 |
| chr1_+_2190714 | 4.31 |
ENSDART00000132126
|
mbnl2
|
muscleblind-like splicing regulator 2 |
| chr7_+_528593 | 4.31 |
ENSDART00000091955
|
nrxn2b
|
neurexin 2b |
| chr8_-_22288258 | 4.28 |
ENSDART00000140978
ENSDART00000100046 |
si:ch211-147a11.3
|
si:ch211-147a11.3 |
| chr24_-_21404367 | 4.25 |
ENSDART00000152093
|
atp8a2
|
ATPase phospholipid transporting 8A2 |
| chr12_+_4971515 | 4.24 |
ENSDART00000161076
|
arhgap27
|
Rho GTPase activating protein 27 |
| chr24_+_41940299 | 4.22 |
ENSDART00000022349
|
epb41l3a
|
erythrocyte membrane protein band 4.1-like 3a |
| chr19_+_16595332 | 4.08 |
ENSDART00000151254
|
ptprua
|
protein tyrosine phosphatase, receptor type, U, a |
| chr14_+_20900989 | 4.06 |
ENSDART00000106209
|
zgc:162941
|
zgc:162941 |
| chr15_-_33834577 | 3.98 |
ENSDART00000163354
|
mmp13b
|
matrix metallopeptidase 13b |
| chr18_+_16750080 | 3.98 |
ENSDART00000136320
|
rnf141
|
ring finger protein 141 |
| chr4_+_68087932 | 3.96 |
ENSDART00000150494
|
znf1096
|
zinc finger protein 1096 |
| chr5_+_17624463 | 3.90 |
ENSDART00000183869
ENSDART00000081064 |
fbrsl1
|
fibrosin-like 1 |
| chr22_-_24297510 | 3.87 |
ENSDART00000163297
|
si:ch211-117l17.6
|
si:ch211-117l17.6 |
| chr15_-_47838391 | 3.86 |
ENSDART00000180337
|
kbtbd3
|
kelch repeat and BTB (POZ) domain containing 3 |
| chr15_+_24704798 | 3.82 |
ENSDART00000192470
|
LRRC75A
|
si:dkey-151p21.7 |
| chr22_+_19247255 | 3.82 |
ENSDART00000144053
|
si:dkey-21e2.10
|
si:dkey-21e2.10 |
| chr8_+_10561922 | 3.81 |
ENSDART00000133348
|
fam19a5l
|
family with sequence similarity 19 (chemokine (C-C motif)-like), member A5-like |
| chr3_+_20156956 | 3.79 |
ENSDART00000125281
|
ngfra
|
nerve growth factor receptor a (TNFR superfamily, member 16) |
| chr21_-_2248239 | 3.78 |
ENSDART00000169448
|
si:ch73-299h12.1
|
si:ch73-299h12.1 |
| chr12_+_29173523 | 3.64 |
ENSDART00000153212
|
adgra1a
|
adhesion G protein-coupled receptor A1a |
| chr3_+_7617353 | 3.60 |
ENSDART00000165551
|
zgc:109949
|
zgc:109949 |
| chr10_+_37145007 | 3.56 |
ENSDART00000131777
|
cuedc1a
|
CUE domain containing 1a |
| chr1_+_52518176 | 3.46 |
ENSDART00000003278
|
tacr3l
|
tachykinin receptor 3-like |
| chr17_+_12942021 | 3.38 |
ENSDART00000192514
|
nfkbiab
|
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha b |
| chr22_+_9862243 | 3.30 |
ENSDART00000105942
|
si:dkey-253d23.3
|
si:dkey-253d23.3 |
| chr4_-_38421217 | 3.27 |
ENSDART00000164517
|
znf1092
|
szinc finger protein 1092 |
| chr19_-_7495006 | 3.21 |
ENSDART00000148836
|
rfx5
|
regulatory factor X, 5 |
| chr6_+_55428924 | 3.19 |
ENSDART00000018270
|
ncoa5
|
nuclear receptor coactivator 5 |
| chr2_+_47582488 | 3.18 |
ENSDART00000149967
|
scg2b
|
secretogranin II (chromogranin C), b |
| chr12_+_28117365 | 3.07 |
ENSDART00000066290
|
UTS2R
|
urotensin 2 receptor |
| chr3_+_39922602 | 3.06 |
ENSDART00000193937
|
cacna1ha
|
calcium channel, voltage-dependent, T type, alpha 1H subunit a |
| chr2_+_9821757 | 3.02 |
ENSDART00000018408
ENSDART00000141227 ENSDART00000144681 ENSDART00000148227 |
anxa13l
|
annexin A13, like |
| chr14_-_6936731 | 3.01 |
ENSDART00000162196
|
clk4a
|
CDC-like kinase 4a |
| chr23_-_37514493 | 2.99 |
ENSDART00000083267
|
dnajc16
|
DnaJ (Hsp40) homolog, subfamily C, member 16 |
| chr3_-_45250924 | 2.97 |
ENSDART00000109017
|
usp31
|
ubiquitin specific peptidase 31 |
| chr15_+_25489406 | 2.93 |
ENSDART00000162482
|
zgc:152863
|
zgc:152863 |
| chr17_+_41302660 | 2.93 |
ENSDART00000059480
|
fosl2
|
fos-like antigen 2 |
| chr21_+_25765734 | 2.92 |
ENSDART00000021664
|
cldnb
|
claudin b |
| chr12_-_35386910 | 2.89 |
ENSDART00000153453
|
camk2g1
|
calcium/calmodulin-dependent protein kinase (CaM kinase) II gamma 1 |
| chr11_+_13630107 | 2.85 |
ENSDART00000172220
|
si:ch211-1a19.3
|
si:ch211-1a19.3 |
| chr9_-_30346279 | 2.84 |
ENSDART00000089488
|
sytl5
|
synaptotagmin-like 5 |
| chr6_-_8295657 | 2.83 |
ENSDART00000185135
|
slc44a2
|
solute carrier family 44 (choline transporter), member 2 |
| chr24_-_21973163 | 2.81 |
ENSDART00000131406
|
acot9.1
|
acyl-CoA thioesterase 9, tandem duplicate 1 |
| chr9_+_33154841 | 2.80 |
ENSDART00000132465
|
dopey2
|
dopey family member 2 |
| chr24_-_21973365 | 2.79 |
ENSDART00000081204
ENSDART00000030592 |
acot9.1
|
acyl-CoA thioesterase 9, tandem duplicate 1 |
| chr25_-_173165 | 2.77 |
ENSDART00000193594
|
CABZ01114053.1
|
|
| chr11_+_13629528 | 2.76 |
ENSDART00000186509
|
si:ch211-1a19.3
|
si:ch211-1a19.3 |
| chr4_+_54899568 | 2.76 |
ENSDART00000162786
|
si:dkey-56m15.8
|
si:dkey-56m15.8 |
| chr18_-_19095141 | 2.71 |
ENSDART00000090962
ENSDART00000090922 |
dennd4a
|
DENN/MADD domain containing 4A |
| chr24_+_17140938 | 2.71 |
ENSDART00000149134
|
mllt10
|
MLLT10, histone lysine methyltransferase DOT1L cofactor |
| chr8_+_43053519 | 2.71 |
ENSDART00000147178
|
prnpa
|
prion protein a |
| chr23_+_29570386 | 2.66 |
ENSDART00000193192
|
lzic
|
leucine zipper and CTNNBIP1 domain containing |
| chr22_+_16320076 | 2.65 |
ENSDART00000164161
|
osbpl1a
|
oxysterol binding protein-like 1A |
| chr3_+_33440615 | 2.62 |
ENSDART00000146005
|
gtpbp1
|
GTP binding protein 1 |
| chr13_+_30421472 | 2.61 |
ENSDART00000143569
|
zmiz1a
|
zinc finger, MIZ-type containing 1a |
| chr12_-_17147473 | 2.59 |
ENSDART00000106035
|
stambpl1
|
STAM binding protein-like 1 |
| chr22_+_17399124 | 2.58 |
ENSDART00000145769
|
rabgap1l
|
RAB GTPase activating protein 1-like |
| chr19_-_21716593 | 2.53 |
ENSDART00000155126
|
znf516
|
zinc finger protein 516 |
| chr11_-_25324534 | 2.52 |
ENSDART00000158598
|
si:ch211-232b12.5
|
si:ch211-232b12.5 |
| chr21_-_18824434 | 2.51 |
ENSDART00000156333
|
si:dkey-112m2.1
|
si:dkey-112m2.1 |
| chr24_-_32665283 | 2.48 |
ENSDART00000038364
|
ca2
|
carbonic anhydrase II |
| chr6_+_11250316 | 2.46 |
ENSDART00000137122
|
atg9a
|
ATG9 autophagy related 9 homolog A (S. cerevisiae) |
| chr19_-_36675023 | 2.46 |
ENSDART00000132471
|
csmd2
|
CUB and Sushi multiple domains 2 |
| chr16_+_42829735 | 2.45 |
ENSDART00000014956
|
polr3glb
|
polymerase (RNA) III (DNA directed) polypeptide G like b |
| chr2_-_23391266 | 2.44 |
ENSDART00000159048
|
ivns1abpb
|
influenza virus NS1A binding protein b |
| chr23_-_1017428 | 2.43 |
ENSDART00000110588
ENSDART00000183158 |
cdh26.1
|
cadherin 26, tandem duplicate 1 |
| chr20_+_9683994 | 2.42 |
ENSDART00000053831
|
ptger2b
|
prostaglandin E receptor 2b (subtype EP2) |
| chr2_+_9822319 | 2.41 |
ENSDART00000144078
ENSDART00000144371 |
anxa13l
|
annexin A13, like |
| chr4_-_73299225 | 2.41 |
ENSDART00000174178
ENSDART00000174198 |
si:cabz01021430.2
|
si:cabz01021430.2 |
| chr8_+_25900049 | 2.39 |
ENSDART00000124300
ENSDART00000127618 ENSDART00000024009 |
rhoab
|
ras homolog gene family, member Ab |
| chr7_+_22688781 | 2.37 |
ENSDART00000173509
|
ugt5g1
|
UDP glucuronosyltransferase 5 family, polypeptide G1 |
| chr23_-_1017605 | 2.36 |
ENSDART00000138290
|
cdh26.1
|
cadherin 26, tandem duplicate 1 |
| chr1_+_51312752 | 2.36 |
ENSDART00000063938
|
mast1a
|
microtubule associated serine/threonine kinase 1a |
| chr9_+_32358514 | 2.32 |
ENSDART00000144608
|
plcl1
|
phospholipase C like 1 |
| chr14_+_24934736 | 2.31 |
ENSDART00000191821
|
ppargc1b
|
peroxisome proliferator-activated receptor gamma, coactivator 1 beta |
| chr7_-_19369002 | 2.31 |
ENSDART00000165680
|
ntn4
|
netrin 4 |
| chr25_-_18330503 | 2.30 |
ENSDART00000104496
|
dusp6
|
dual specificity phosphatase 6 |
| chr14_+_20156477 | 2.30 |
ENSDART00000123434
|
fmr1
|
fragile X mental retardation 1 |
| chr16_+_13855039 | 2.27 |
ENSDART00000113764
ENSDART00000143983 |
zgc:174888
|
zgc:174888 |
| chr11_-_22371105 | 2.21 |
ENSDART00000146873
|
tmem183a
|
transmembrane protein 183A |
| chr20_+_23173710 | 2.21 |
ENSDART00000074172
|
sgcb
|
sarcoglycan, beta (dystrophin-associated glycoprotein) |
| chr6_+_49901465 | 2.21 |
ENSDART00000023515
|
chmp4ba
|
charged multivesicular body protein 4Ba |
| chr3_+_24197934 | 2.13 |
ENSDART00000055609
|
atf4b
|
activating transcription factor 4b |
| chr22_-_5839686 | 2.13 |
ENSDART00000145406
|
si:rp71-36a1.5
|
si:rp71-36a1.5 |
| chr2_-_48356787 | 2.11 |
ENSDART00000098004
|
per2
|
period circadian clock 2 |
| chr3_-_27646070 | 2.04 |
ENSDART00000122031
ENSDART00000151027 |
si:ch211-157c3.4
|
si:ch211-157c3.4 |
| chr19_-_38539670 | 2.04 |
ENSDART00000136775
|
col16a1
|
collagen, type XVI, alpha 1 |
| chr2_-_37837472 | 2.04 |
ENSDART00000165347
|
mettl17
|
methyltransferase like 17 |
| chr21_+_3093419 | 2.03 |
ENSDART00000162520
|
SHC3
|
SHC adaptor protein 3 |
| chr14_+_6287871 | 2.01 |
ENSDART00000105344
|
tbc1d2
|
TBC1 domain family, member 2 |
| chr10_+_35422802 | 2.00 |
ENSDART00000147303
|
hhla2a.1
|
HERV-H LTR-associating 2a, tandem duplicate 1 |
| chr4_+_45274792 | 1.95 |
ENSDART00000150295
|
znf1138
|
zinc finger protein 1138 |
| chr7_-_30174882 | 1.92 |
ENSDART00000110409
|
frmd5
|
FERM domain containing 5 |
| chr21_-_2172924 | 1.92 |
ENSDART00000169262
|
zgc:163077
|
zgc:163077 |
| chr3_-_48259289 | 1.91 |
ENSDART00000160717
|
znf750
|
zinc finger protein 750 |
| chr1_-_35694978 | 1.91 |
ENSDART00000136157
|
si:dkey-27h10.2
|
si:dkey-27h10.2 |
| chr5_+_20035284 | 1.90 |
ENSDART00000191808
|
sgsm1a
|
small G protein signaling modulator 1a |
| chr3_-_61185746 | 1.89 |
ENSDART00000028219
|
pvalb4
|
parvalbumin 4 |
| chr2_-_15362923 | 1.88 |
ENSDART00000166926
|
olfm3b
|
olfactomedin 3b |
| chr9_+_2499627 | 1.88 |
ENSDART00000160782
|
wipf1a
|
WAS/WASL interacting protein family, member 1a |
| chr24_-_5691956 | 1.87 |
ENSDART00000189112
|
dia1b
|
deleted in autism 1b |
| chr1_-_54765262 | 1.87 |
ENSDART00000150362
|
si:ch211-197k17.3
|
si:ch211-197k17.3 |
| chr14_+_6615564 | 1.86 |
ENSDART00000139292
|
si:dkeyp-44a8.2
|
si:dkeyp-44a8.2 |
| chr7_-_60351876 | 1.83 |
ENSDART00000098563
|
plcb3
|
phospholipase C, beta 3 (phosphatidylinositol-specific) |
| chr16_-_12984631 | 1.79 |
ENSDART00000184863
|
cacng7b
|
calcium channel, voltage-dependent, gamma subunit 7b |
| chr12_-_7639120 | 1.78 |
ENSDART00000126712
ENSDART00000126219 |
ccdc6b
|
coiled-coil domain containing 6b |
| chr14_+_49382180 | 1.77 |
ENSDART00000158329
|
tnip1
|
TNFAIP3 interacting protein 1 |
| chr4_+_58057168 | 1.73 |
ENSDART00000161097
|
si:dkey-57k17.1
|
si:dkey-57k17.1 |
| chr1_+_46598764 | 1.72 |
ENSDART00000053240
|
cab39l
|
calcium binding protein 39-like |
| chr22_+_19405517 | 1.70 |
ENSDART00000138245
ENSDART00000155144 |
si:dkey-78l4.2
|
si:dkey-78l4.2 |
| chr20_-_35512932 | 1.68 |
ENSDART00000137690
|
adgrf3b
|
adhesion G protein-coupled receptor F3b |
| chr21_-_2173300 | 1.67 |
ENSDART00000172276
|
zgc:163077
|
zgc:163077 |
| chr22_+_9862466 | 1.66 |
ENSDART00000146864
|
si:dkey-253d23.3
|
si:dkey-253d23.3 |
| chr16_+_23403602 | 1.61 |
ENSDART00000159848
|
s100w
|
S100 calcium binding protein W |
| chr5_+_22393501 | 1.61 |
ENSDART00000185276
|
si:dkey-27p18.7
|
si:dkey-27p18.7 |
| chr20_+_32478151 | 1.57 |
ENSDART00000145269
|
ostm1
|
osteopetrosis associated transmembrane protein 1 |
| chr16_+_37876779 | 1.48 |
ENSDART00000140148
|
si:ch211-198c19.1
|
si:ch211-198c19.1 |
| chr3_+_45401472 | 1.48 |
ENSDART00000156693
|
baiap3
|
BAI1-associated protein 3 |
| chr4_-_56157199 | 1.47 |
ENSDART00000169806
|
si:ch211-207e19.15
|
si:ch211-207e19.15 |
| chr19_-_31035155 | 1.47 |
ENSDART00000161882
|
bzw2
|
basic leucine zipper and W2 domains 2 |
| chr12_+_47280839 | 1.46 |
ENSDART00000187574
|
RYR2
|
ryanodine receptor 2 |
| chr22_-_26595027 | 1.44 |
ENSDART00000184162
|
CABZ01072309.1
|
|
| chr19_+_33701366 | 1.43 |
ENSDART00000162517
|
runx1t1
|
runt-related transcription factor 1; translocated to, 1 (cyclin D-related) |
| chr2_-_44746723 | 1.42 |
ENSDART00000041806
|
acsm3
|
acyl-CoA synthetase medium chain family member 3 |
| chr11_+_2416064 | 1.41 |
ENSDART00000067117
|
ube2v1
|
ubiquitin-conjugating enzyme E2 variant 1 |
| chr8_+_20037368 | 1.40 |
ENSDART00000029175
ENSDART00000170792 |
fcho1
|
FCH domain only 1 |
| chr4_-_77267787 | 1.37 |
ENSDART00000190346
|
zgc:174310
|
zgc:174310 |
| chrM_+_8894 | 1.32 |
ENSDART00000093611
|
mt-atp8
|
ATP synthase 8, mitochondrial |
| chr5_-_69523816 | 1.26 |
ENSDART00000112692
|
si:ch211-157p22.10
|
si:ch211-157p22.10 |
| chr12_+_23371542 | 1.26 |
ENSDART00000078896
|
waca
|
WW domain containing adaptor with coiled-coil a |
| chr2_+_21229909 | 1.24 |
ENSDART00000046127
|
vipr1a
|
vasoactive intestinal peptide receptor 1a |
| chr22_+_19220459 | 1.24 |
ENSDART00000163070
|
si:dkey-21e2.7
|
si:dkey-21e2.7 |
| chr1_-_11689525 | 1.20 |
ENSDART00000090689
|
brat1
|
BRCA1-associated ATM activator 1 |
| chr1_+_52137528 | 1.16 |
ENSDART00000007079
ENSDART00000074265 |
asna1
|
arsA arsenite transporter, ATP-binding, homolog 1 (bacterial) |
| chr15_-_13254480 | 1.15 |
ENSDART00000190499
|
zgc:172282
|
zgc:172282 |
| chr3_+_35498119 | 1.12 |
ENSDART00000178963
|
tnrc6a
|
trinucleotide repeat containing 6a |
| chr10_-_32877348 | 1.10 |
ENSDART00000018977
ENSDART00000133421 |
rabgef1
|
RAB guanine nucleotide exchange factor (GEF) 1 |
| chr15_+_847077 | 1.10 |
ENSDART00000188586
|
si:dkey-7i4.5
|
si:dkey-7i4.5 |
| chr4_-_69550757 | 1.10 |
ENSDART00000168344
|
znf1101
|
zinc finger protein 1101 |
| chr13_+_22964868 | 1.07 |
ENSDART00000142129
|
tacr2
|
tachykinin receptor 2 |
| chr7_+_5529967 | 1.04 |
ENSDART00000142709
|
si:dkeyp-67a8.4
|
si:dkeyp-67a8.4 |
| chr22_+_16497670 | 1.04 |
ENSDART00000014330
|
ier5
|
immediate early response 5 |
| chr3_-_39648021 | 1.03 |
ENSDART00000055171
|
grapa
|
GRB2-related adaptor protein a |
| chr4_+_7822773 | 1.02 |
ENSDART00000171391
|
si:ch1073-67j19.2
|
si:ch1073-67j19.2 |
| chr11_-_24458786 | 1.01 |
ENSDART00000089713
|
mxra8a
|
matrix-remodelling associated 8a |
Network of associatons between targets according to the STRING database.
First level regulatory network of jdp2a
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.3 | 6.8 | GO:0035902 | response to immobilization stress(GO:0035902) |
| 1.7 | 5.1 | GO:0015824 | proline transport(GO:0015824) |
| 1.6 | 6.6 | GO:0090387 | phagosome maturation involved in apoptotic cell clearance(GO:0090386) phagolysosome assembly involved in apoptotic cell clearance(GO:0090387) |
| 1.4 | 4.1 | GO:0016998 | cell wall macromolecule catabolic process(GO:0016998) |
| 1.0 | 8.3 | GO:0006703 | estrogen biosynthetic process(GO:0006703) |
| 1.0 | 7.2 | GO:0042984 | amyloid precursor protein biosynthetic process(GO:0042983) regulation of amyloid precursor protein biosynthetic process(GO:0042984) |
| 1.0 | 7.9 | GO:0008343 | adult feeding behavior(GO:0008343) |
| 0.9 | 4.5 | GO:0032655 | interleukin-12 production(GO:0032615) regulation of interleukin-12 production(GO:0032655) |
| 0.8 | 7.6 | GO:0042989 | sequestering of actin monomers(GO:0042989) |
| 0.7 | 2.8 | GO:0015871 | choline transport(GO:0015871) |
| 0.6 | 4.5 | GO:1902093 | positive regulation of sperm motility(GO:1902093) |
| 0.6 | 2.5 | GO:0015670 | carbon dioxide transport(GO:0015670) |
| 0.6 | 10.2 | GO:0097324 | melanocyte migration(GO:0097324) |
| 0.5 | 22.5 | GO:1903670 | regulation of sprouting angiogenesis(GO:1903670) |
| 0.5 | 8.1 | GO:0001964 | startle response(GO:0001964) |
| 0.5 | 2.4 | GO:0034695 | response to prostaglandin E(GO:0034695) cellular response to prostaglandin E stimulus(GO:0071380) |
| 0.5 | 4.2 | GO:1904861 | excitatory synapse assembly(GO:1904861) |
| 0.4 | 8.0 | GO:0097178 | ruffle assembly(GO:0097178) |
| 0.4 | 2.3 | GO:0048755 | branching morphogenesis of a nerve(GO:0048755) |
| 0.4 | 2.9 | GO:1905207 | regulation of cardiocyte differentiation(GO:1905207) regulation of cardiac muscle cell differentiation(GO:2000725) |
| 0.4 | 2.1 | GO:0042753 | positive regulation of circadian rhythm(GO:0042753) |
| 0.3 | 8.0 | GO:0015701 | bicarbonate transport(GO:0015701) |
| 0.3 | 2.4 | GO:0044319 | wound healing, spreading of cells(GO:0044319) epiboly involved in wound healing(GO:0090505) skeletal muscle satellite cell migration(GO:1902766) |
| 0.3 | 2.3 | GO:0060420 | regulation of heart growth(GO:0060420) |
| 0.3 | 10.3 | GO:0007340 | acrosome reaction(GO:0007340) |
| 0.3 | 7.0 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.3 | 2.3 | GO:0032228 | regulation of synaptic transmission, GABAergic(GO:0032228) |
| 0.3 | 3.1 | GO:0045956 | positive regulation of calcium ion-dependent exocytosis(GO:0045956) positive regulation of regulated secretory pathway(GO:1903307) |
| 0.3 | 6.3 | GO:0007097 | nuclear migration(GO:0007097) establishment of nucleus localization(GO:0040023) |
| 0.2 | 2.5 | GO:0044805 | late nucleophagy(GO:0044805) |
| 0.2 | 5.6 | GO:0019933 | cAMP-mediated signaling(GO:0019933) |
| 0.2 | 5.0 | GO:0015669 | gas transport(GO:0015669) oxygen transport(GO:0015671) |
| 0.2 | 7.4 | GO:0032482 | Rab protein signal transduction(GO:0032482) |
| 0.2 | 4.0 | GO:0051865 | protein autoubiquitination(GO:0051865) |
| 0.2 | 6.8 | GO:0007040 | lysosome organization(GO:0007040) lytic vacuole organization(GO:0080171) |
| 0.2 | 1.3 | GO:0044783 | G1 DNA damage checkpoint(GO:0044783) |
| 0.2 | 5.9 | GO:0006817 | phosphate ion transport(GO:0006817) |
| 0.2 | 0.4 | GO:0051228 | mitotic spindle disassembly(GO:0051228) spindle disassembly(GO:0051230) |
| 0.2 | 2.0 | GO:0031294 | lymphocyte costimulation(GO:0031294) T cell costimulation(GO:0031295) |
| 0.2 | 2.5 | GO:0090279 | regulation of calcium ion import(GO:0090279) |
| 0.2 | 8.6 | GO:0032465 | regulation of cytokinesis(GO:0032465) |
| 0.2 | 5.2 | GO:0045332 | phospholipid translocation(GO:0045332) |
| 0.1 | 5.2 | GO:0035118 | embryonic pectoral fin morphogenesis(GO:0035118) |
| 0.1 | 7.5 | GO:0051091 | positive regulation of sequence-specific DNA binding transcription factor activity(GO:0051091) |
| 0.1 | 0.7 | GO:0042102 | positive regulation of T cell proliferation(GO:0042102) |
| 0.1 | 2.2 | GO:0031468 | nuclear envelope reassembly(GO:0031468) |
| 0.1 | 4.0 | GO:0044243 | collagen catabolic process(GO:0030574) multicellular organism catabolic process(GO:0044243) |
| 0.1 | 2.2 | GO:0030277 | maintenance of gastrointestinal epithelium(GO:0030277) intestinal epithelial structure maintenance(GO:0060729) |
| 0.1 | 8.2 | GO:0021549 | cerebellum development(GO:0021549) |
| 0.1 | 1.8 | GO:0051968 | positive regulation of synaptic transmission, glutamatergic(GO:0051968) |
| 0.1 | 0.7 | GO:0001672 | regulation of chromatin assembly or disassembly(GO:0001672) |
| 0.1 | 5.4 | GO:0048843 | negative regulation of axon extension involved in axon guidance(GO:0048843) negative regulation of axon guidance(GO:1902668) |
| 0.1 | 0.3 | GO:0097095 | frontonasal suture morphogenesis(GO:0097095) |
| 0.1 | 2.6 | GO:0070536 | protein K63-linked deubiquitination(GO:0070536) |
| 0.1 | 1.8 | GO:0043124 | negative regulation of I-kappaB kinase/NF-kappaB signaling(GO:0043124) |
| 0.1 | 8.2 | GO:0042552 | myelination(GO:0042552) |
| 0.1 | 4.2 | GO:0043297 | apical junction assembly(GO:0043297) bicellular tight junction assembly(GO:0070830) |
| 0.1 | 0.3 | GO:0010939 | regulation of necrotic cell death(GO:0010939) regulation of necroptotic process(GO:0060544) negative regulation of necroptotic process(GO:0060546) negative regulation of necrotic cell death(GO:0060547) |
| 0.1 | 2.4 | GO:0006383 | transcription from RNA polymerase III promoter(GO:0006383) |
| 0.1 | 2.3 | GO:0070831 | basement membrane assembly(GO:0070831) |
| 0.1 | 5.1 | GO:0035249 | synaptic transmission, glutamatergic(GO:0035249) |
| 0.1 | 2.9 | GO:0072114 | pronephros morphogenesis(GO:0072114) |
| 0.1 | 1.1 | GO:0035278 | miRNA mediated inhibition of translation(GO:0035278) negative regulation of translation, ncRNA-mediated(GO:0040033) regulation of translation, ncRNA-mediated(GO:0045974) |
| 0.1 | 1.8 | GO:0015986 | energy coupled proton transport, down electrochemical gradient(GO:0015985) ATP synthesis coupled proton transport(GO:0015986) |
| 0.1 | 6.7 | GO:0051051 | negative regulation of transport(GO:0051051) |
| 0.1 | 1.4 | GO:0006301 | postreplication repair(GO:0006301) |
| 0.1 | 7.0 | GO:0006637 | acyl-CoA metabolic process(GO:0006637) thioester metabolic process(GO:0035383) |
| 0.1 | 4.2 | GO:0006414 | translational elongation(GO:0006414) |
| 0.1 | 0.2 | GO:0070221 | sulfide oxidation(GO:0019418) sulfide oxidation, using sulfide:quinone oxidoreductase(GO:0070221) |
| 0.1 | 0.9 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.1 | 0.5 | GO:0071376 | neutrophil homeostasis(GO:0001780) response to corticotropin-releasing hormone(GO:0043435) cellular response to corticotropin-releasing hormone stimulus(GO:0071376) |
| 0.1 | 5.4 | GO:0007218 | neuropeptide signaling pathway(GO:0007218) |
| 0.1 | 6.5 | GO:0090630 | activation of GTPase activity(GO:0090630) |
| 0.0 | 4.3 | GO:0000381 | regulation of alternative mRNA splicing, via spliceosome(GO:0000381) |
| 0.0 | 1.4 | GO:0048268 | clathrin coat assembly(GO:0048268) |
| 0.0 | 4.7 | GO:0006888 | ER to Golgi vesicle-mediated transport(GO:0006888) |
| 0.0 | 0.4 | GO:0016199 | axon midline choice point recognition(GO:0016199) |
| 0.0 | 1.0 | GO:0051904 | melanosome transport(GO:0032402) pigment granule transport(GO:0051904) |
| 0.0 | 1.2 | GO:0010212 | response to ionizing radiation(GO:0010212) |
| 0.0 | 1.6 | GO:0050852 | T cell receptor signaling pathway(GO:0050852) |
| 0.0 | 0.5 | GO:0010824 | regulation of centrosome duplication(GO:0010824) |
| 0.0 | 2.3 | GO:0030048 | actin filament-based movement(GO:0030048) |
| 0.0 | 4.8 | GO:0071772 | BMP signaling pathway(GO:0030509) response to BMP(GO:0071772) cellular response to BMP stimulus(GO:0071773) |
| 0.0 | 4.4 | GO:0043087 | regulation of GTPase activity(GO:0043087) |
| 0.0 | 0.3 | GO:0033505 | floor plate formation(GO:0021508) floor plate morphogenesis(GO:0033505) |
| 0.0 | 2.9 | GO:0007266 | Rho protein signal transduction(GO:0007266) |
| 0.0 | 0.7 | GO:0018230 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 0.0 | 4.3 | GO:0006887 | exocytosis(GO:0006887) |
| 0.0 | 0.6 | GO:0032011 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.0 | 0.1 | GO:0030878 | thyroid gland development(GO:0030878) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 20.2 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.6 | 2.3 | GO:0044326 | dendritic spine neck(GO:0044326) dendritic filopodium(GO:1902737) |
| 0.6 | 4.5 | GO:0097225 | sperm midpiece(GO:0097225) |
| 0.5 | 8.1 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.4 | 5.0 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 0.4 | 6.3 | GO:0005640 | nuclear outer membrane(GO:0005640) |
| 0.3 | 22.5 | GO:0030141 | secretory granule(GO:0030141) |
| 0.2 | 2.9 | GO:1990589 | ATF4-CREB1 transcription factor complex(GO:1990589) |
| 0.2 | 1.4 | GO:0031371 | ubiquitin conjugating enzyme complex(GO:0031371) |
| 0.2 | 8.0 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.2 | 2.2 | GO:0000815 | ESCRT III complex(GO:0000815) |
| 0.1 | 3.1 | GO:0034706 | voltage-gated sodium channel complex(GO:0001518) sodium channel complex(GO:0034706) |
| 0.1 | 2.4 | GO:0005666 | DNA-directed RNA polymerase III complex(GO:0005666) |
| 0.1 | 2.3 | GO:0043256 | basal lamina(GO:0005605) laminin complex(GO:0043256) |
| 0.1 | 0.4 | GO:0060171 | stereocilium membrane(GO:0060171) |
| 0.1 | 6.8 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.1 | 0.3 | GO:0030289 | protein phosphatase 4 complex(GO:0030289) |
| 0.1 | 1.8 | GO:0045263 | proton-transporting ATP synthase complex, coupling factor F(o)(GO:0045263) |
| 0.1 | 2.2 | GO:0090665 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.1 | 8.6 | GO:0000922 | spindle pole(GO:0000922) |
| 0.1 | 0.4 | GO:0034098 | VCP-NPL4-UFD1 AAA ATPase complex(GO:0034098) |
| 0.1 | 2.5 | GO:0000407 | pre-autophagosomal structure(GO:0000407) |
| 0.1 | 3.6 | GO:0098978 | glutamatergic synapse(GO:0098978) |
| 0.1 | 4.8 | GO:0016342 | catenin complex(GO:0016342) |
| 0.1 | 0.9 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.1 | 2.4 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 1.8 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.0 | 8.1 | GO:0070382 | exocytic vesicle(GO:0070382) |
| 0.0 | 1.4 | GO:0005905 | clathrin-coated pit(GO:0005905) |
| 0.0 | 0.8 | GO:0030119 | AP-type membrane coat adaptor complex(GO:0030119) |
| 0.0 | 10.3 | GO:0031012 | extracellular matrix(GO:0031012) |
| 0.0 | 3.4 | GO:0070160 | bicellular tight junction(GO:0005923) occluding junction(GO:0070160) |
| 0.0 | 4.8 | GO:0005681 | spliceosomal complex(GO:0005681) |
| 0.0 | 4.3 | GO:0045211 | postsynaptic membrane(GO:0045211) |
| 0.0 | 0.5 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.0 | 2.0 | GO:0005765 | lysosomal membrane(GO:0005765) lytic vacuole membrane(GO:0098852) |
| 0.0 | 1.1 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.0 | 6.7 | GO:0009986 | cell surface(GO:0009986) |
| 0.0 | 0.6 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.0 | 0.6 | GO:0030139 | endocytic vesicle(GO:0030139) |
| 0.0 | 0.3 | GO:0070822 | Sin3-type complex(GO:0070822) |
| 0.0 | 0.3 | GO:0005921 | gap junction(GO:0005921) |
| 0.0 | 3.5 | GO:0005768 | endosome(GO:0005768) |
| 0.0 | 1.6 | GO:0048471 | perinuclear region of cytoplasm(GO:0048471) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.0 | 5.9 | GO:0000822 | inositol hexakisphosphate binding(GO:0000822) |
| 1.4 | 8.4 | GO:0030331 | estrogen receptor binding(GO:0030331) |
| 1.3 | 20.2 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 1.0 | 8.3 | GO:0004303 | estradiol 17-beta-dehydrogenase activity(GO:0004303) |
| 1.0 | 4.1 | GO:0003796 | lysozyme activity(GO:0003796) |
| 0.7 | 4.4 | GO:0042169 | SH2 domain binding(GO:0042169) |
| 0.7 | 2.8 | GO:0015220 | choline transmembrane transporter activity(GO:0015220) |
| 0.6 | 4.5 | GO:0004995 | tachykinin receptor activity(GO:0004995) |
| 0.6 | 8.0 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.5 | 3.8 | GO:0005035 | death receptor activity(GO:0005035) |
| 0.5 | 3.2 | GO:1990380 | Lys48-specific deubiquitinase activity(GO:1990380) |
| 0.4 | 3.1 | GO:0008332 | low voltage-gated calcium channel activity(GO:0008332) |
| 0.4 | 15.1 | GO:0004629 | phosphatidylinositol phospholipase C activity(GO:0004435) phospholipase C activity(GO:0004629) |
| 0.4 | 5.0 | GO:0031720 | haptoglobin binding(GO:0031720) |
| 0.4 | 7.6 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.4 | 5.7 | GO:0048018 | receptor activator activity(GO:0030546) receptor agonist activity(GO:0048018) |
| 0.3 | 8.2 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.3 | 2.7 | GO:0004784 | superoxide dismutase activity(GO:0004784) oxidoreductase activity, acting on superoxide radicals as acceptor(GO:0016721) |
| 0.3 | 6.8 | GO:0080025 | phosphatidylinositol-3,5-bisphosphate binding(GO:0080025) |
| 0.3 | 7.0 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.3 | 4.6 | GO:0000340 | RNA 7-methylguanosine cap binding(GO:0000340) |
| 0.3 | 2.1 | GO:0001222 | transcription corepressor binding(GO:0001222) |
| 0.2 | 2.4 | GO:0004957 | prostaglandin E receptor activity(GO:0004957) |
| 0.2 | 5.6 | GO:0047617 | acyl-CoA hydrolase activity(GO:0047617) |
| 0.2 | 5.6 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.2 | 1.2 | GO:0004999 | vasoactive intestinal polypeptide receptor activity(GO:0004999) |
| 0.2 | 7.7 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.2 | 8.1 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.2 | 2.3 | GO:0008330 | protein tyrosine/threonine phosphatase activity(GO:0008330) |
| 0.2 | 2.6 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.1 | 4.3 | GO:0099529 | neurotransmitter receptor activity involved in regulation of postsynaptic membrane potential(GO:0099529) transmitter-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1904315) |
| 0.1 | 0.9 | GO:0001641 | group II metabotropic glutamate receptor activity(GO:0001641) |
| 0.1 | 2.8 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.1 | 1.4 | GO:0016408 | C-acyltransferase activity(GO:0016408) |
| 0.1 | 2.6 | GO:0015248 | sterol transporter activity(GO:0015248) |
| 0.1 | 4.2 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.1 | 5.4 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.1 | 2.6 | GO:0030374 | ligand-dependent nuclear receptor transcription coactivator activity(GO:0030374) |
| 0.1 | 1.3 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.1 | 0.2 | GO:0070224 | sulfide:quinone oxidoreductase activity(GO:0070224) |
| 0.1 | 1.6 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.1 | 1.4 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.1 | 9.9 | GO:0000287 | magnesium ion binding(GO:0000287) |
| 0.1 | 2.7 | GO:0031491 | nucleosome binding(GO:0031491) |
| 0.1 | 2.3 | GO:0045182 | translation regulator activity(GO:0045182) |
| 0.1 | 2.7 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.0 | 0.9 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.0 | 0.2 | GO:0047192 | 1-alkylglycerophosphocholine O-acetyltransferase activity(GO:0047192) |
| 0.0 | 1.5 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.0 | 0.5 | GO:0035258 | steroid hormone receptor binding(GO:0035258) |
| 0.0 | 4.7 | GO:0005179 | hormone activity(GO:0005179) |
| 0.0 | 2.4 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.0 | 12.0 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.0 | 0.3 | GO:0043028 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) cysteine-type endopeptidase regulator activity involved in apoptotic process(GO:0043028) |
| 0.0 | 0.2 | GO:0038036 | sphingosine-1-phosphate receptor activity(GO:0038036) |
| 0.0 | 8.9 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.0 | 21.0 | GO:0004674 | protein serine/threonine kinase activity(GO:0004674) |
| 0.0 | 1.8 | GO:0016247 | channel regulator activity(GO:0016247) |
| 0.0 | 3.1 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 0.8 | GO:0031593 | polyubiquitin binding(GO:0031593) |
| 0.0 | 0.7 | GO:0019706 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.0 | 0.4 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.0 | 2.0 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 1.8 | GO:0015078 | hydrogen ion transmembrane transporter activity(GO:0015078) |
| 0.0 | 0.4 | GO:0008484 | sulfuric ester hydrolase activity(GO:0008484) |
| 0.0 | 2.1 | GO:0001228 | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001228) |
| 0.0 | 11.6 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.0 | 3.2 | GO:0005085 | guanyl-nucleotide exchange factor activity(GO:0005085) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 7.6 | PID ECADHERIN NASCENT AJ PATHWAY | E-cadherin signaling in the nascent adherens junction |
| 0.1 | 2.5 | PID IL8 CXCR1 PATHWAY | IL8- and CXCR1-mediated signaling events |
| 0.1 | 4.7 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.1 | 10.4 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.1 | 2.1 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.1 | 2.9 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.1 | 2.3 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.1 | 2.3 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
| 0.1 | 1.1 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.1 | 2.0 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
| 0.1 | 2.0 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.0 | 1.4 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.0 | 0.5 | ST PHOSPHOINOSITIDE 3 KINASE PATHWAY | PI3K Pathway |
| 0.0 | 0.6 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.0 | 0.4 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.0 | 0.3 | PID FCER1 PATHWAY | Fc-epsilon receptor I signaling in mast cells |
| 0.0 | 0.3 | PID NFKAPPAB CANONICAL PATHWAY | Canonical NF-kappaB pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 2.5 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.2 | 2.3 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.2 | 2.8 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.2 | 2.0 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.1 | 1.5 | REACTOME REGULATION OF AMPK ACTIVITY VIA LKB1 | Genes involved in Regulation of AMPK activity via LKB1 |
| 0.1 | 11.7 | REACTOME G ALPHA S SIGNALLING EVENTS | Genes involved in G alpha (s) signalling events |
| 0.1 | 0.9 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.1 | 8.0 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.1 | 4.3 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.1 | 2.1 | REACTOME BMAL1 CLOCK NPAS2 ACTIVATES CIRCADIAN EXPRESSION | Genes involved in BMAL1:CLOCK/NPAS2 Activates Circadian Expression |
| 0.1 | 2.3 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.1 | 0.6 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.1 | 3.5 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.1 | 1.1 | REACTOME REGULATORY RNA PATHWAYS | Genes involved in Regulatory RNA pathways |
| 0.0 | 2.0 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 2.6 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.0 | 3.8 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 0.5 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 0.2 | REACTOME ACYL CHAIN REMODELLING OF PC | Genes involved in Acyl chain remodelling of PC |
| 0.0 | 0.6 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.0 | 0.3 | REACTOME APOPTOTIC CLEAVAGE OF CELLULAR PROTEINS | Genes involved in Apoptotic cleavage of cellular proteins |
| 0.0 | 0.5 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |