Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
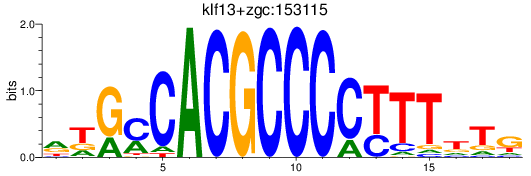
Results for klf13+zgc:153115
Z-value: 0.72
Transcription factors associated with klf13+zgc:153115
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
klf13
|
ENSDARG00000061368 | Kruppel-like factor 13 |
|
zgc
|
ENSDARG00000069342 | 153115 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| zgc:153115 | dr11_v1_chr2_-_7246338_7246338 | -0.18 | 8.1e-02 | Click! |
| klf13 | dr11_v1_chr25_-_34512102_34512102 | -0.07 | 5.2e-01 | Click! |
Activity profile of klf13+zgc:153115 motif
Sorted Z-values of klf13+zgc:153115 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr16_-_17197546 | 13.35 |
ENSDART00000139939
ENSDART00000135146 ENSDART00000063800 ENSDART00000163606 |
gapdh
|
glyceraldehyde-3-phosphate dehydrogenase |
| chr4_-_9852318 | 6.74 |
ENSDART00000080702
|
glt8d2
|
glycosyltransferase 8 domain containing 2 |
| chr19_+_9305964 | 4.61 |
ENSDART00000136241
|
si:ch73-15n24.1
|
si:ch73-15n24.1 |
| chr25_+_6122823 | 4.55 |
ENSDART00000191824
ENSDART00000067514 |
rbpms2a
|
RNA binding protein with multiple splicing 2a |
| chr9_+_30421489 | 3.88 |
ENSDART00000145025
ENSDART00000132058 |
zgc:113314
|
zgc:113314 |
| chr4_-_20177868 | 3.79 |
ENSDART00000003621
|
sinup
|
siaz-interacting nuclear protein |
| chr16_-_42523744 | 3.73 |
ENSDART00000017185
|
tbx20
|
T-box 20 |
| chr8_-_32491430 | 3.48 |
ENSDART00000163962
ENSDART00000168929 |
si:dkey-164f24.2
|
si:dkey-164f24.2 |
| chr8_+_25302172 | 3.08 |
ENSDART00000046182
ENSDART00000145316 |
gstm.3
|
glutathione S-transferase mu tandem duplicate 3 |
| chr6_-_25165693 | 2.81 |
ENSDART00000167259
|
znf326
|
zinc finger protein 326 |
| chr7_+_38349667 | 2.59 |
ENSDART00000010046
|
rhpn2
|
rhophilin, Rho GTPase binding protein 2 |
| chr12_-_3053873 | 2.47 |
ENSDART00000023796
ENSDART00000137148 |
dcxr
|
dicarbonyl/L-xylulose reductase |
| chr4_-_5597802 | 2.39 |
ENSDART00000136229
|
vegfab
|
vascular endothelial growth factor Ab |
| chr21_+_6197223 | 2.28 |
ENSDART00000147716
|
si:dkey-93m18.3
|
si:dkey-93m18.3 |
| chr19_+_47290287 | 2.26 |
ENSDART00000078382
|
tpmt.1
|
thiopurine S-methyltransferase, tandem duplicate 1 |
| chr1_-_12109216 | 2.25 |
ENSDART00000079930
|
mttp
|
microsomal triglyceride transfer protein |
| chr22_+_835728 | 2.23 |
ENSDART00000003325
|
dennd2db
|
DENN/MADD domain containing 2Db |
| chr12_+_30046320 | 2.23 |
ENSDART00000179904
ENSDART00000153394 |
ablim1b
|
actin binding LIM protein 1b |
| chr2_-_48153945 | 2.23 |
ENSDART00000146553
|
pfkpb
|
phosphofructokinase, platelet b |
| chr10_+_22034477 | 2.21 |
ENSDART00000133304
ENSDART00000134189 ENSDART00000021240 ENSDART00000100526 |
npm1a
|
nucleophosmin 1a |
| chr14_+_7048930 | 2.18 |
ENSDART00000109138
|
hbegfa
|
heparin-binding EGF-like growth factor a |
| chr23_-_31645760 | 2.18 |
ENSDART00000035031
|
sgk1
|
serum/glucocorticoid regulated kinase 1 |
| chr16_-_7228276 | 2.11 |
ENSDART00000149030
|
nt5c3a
|
5'-nucleotidase, cytosolic IIIA |
| chr10_-_44017642 | 2.08 |
ENSDART00000135240
ENSDART00000014669 |
acads
|
acyl-CoA dehydrogenase short chain |
| chr23_+_12160900 | 2.05 |
ENSDART00000136046
|
ppp1r3da
|
protein phosphatase 1, regulatory subunit 3Da |
| chr10_+_4875262 | 2.02 |
ENSDART00000165942
|
palm2
|
paralemmin 2 |
| chr19_+_46113828 | 2.02 |
ENSDART00000159331
ENSDART00000161826 |
rbm24a
|
RNA binding motif protein 24a |
| chr5_+_32924669 | 1.94 |
ENSDART00000085219
|
lmo4a
|
LIM domain only 4a |
| chr12_+_6391243 | 1.90 |
ENSDART00000152765
|
prkg1b
|
protein kinase, cGMP-dependent, type Ib |
| chr7_+_4474880 | 1.89 |
ENSDART00000143528
|
si:dkey-83f18.14
|
si:dkey-83f18.14 |
| chr1_-_34335752 | 1.89 |
ENSDART00000140157
|
si:dkey-24h22.5
|
si:dkey-24h22.5 |
| chr18_-_46258612 | 1.85 |
ENSDART00000153930
|
si:dkey-244a7.1
|
si:dkey-244a7.1 |
| chr12_-_3053699 | 1.70 |
ENSDART00000139721
|
dcxr
|
dicarbonyl/L-xylulose reductase |
| chr2_+_43894368 | 1.66 |
ENSDART00000143885
|
gbp3
|
guanylate binding protein 3 |
| chr10_+_41668483 | 1.55 |
ENSDART00000127073
|
lrrc75bb
|
leucine rich repeat containing 75Bb |
| chr21_+_15704556 | 1.53 |
ENSDART00000024858
ENSDART00000146909 |
chchd10
|
coiled-coil-helix-coiled-coil-helix domain containing 10 |
| chr18_+_33725576 | 1.52 |
ENSDART00000146816
|
si:dkey-145c18.5
|
si:dkey-145c18.5 |
| chr22_+_1006573 | 1.51 |
ENSDART00000123458
|
pparda
|
peroxisome proliferator-activated receptor delta a |
| chr19_+_47299212 | 1.44 |
ENSDART00000158262
|
tpmt.1
|
thiopurine S-methyltransferase, tandem duplicate 1 |
| chr10_+_1899595 | 1.40 |
ENSDART00000040271
|
CU861476.1
|
|
| chr3_+_19299309 | 1.38 |
ENSDART00000046297
ENSDART00000146955 |
ldlra
|
low density lipoprotein receptor a |
| chr5_+_68807170 | 1.36 |
ENSDART00000017849
|
her7
|
hairy and enhancer of split related-7 |
| chr6_+_23122789 | 1.26 |
ENSDART00000049226
ENSDART00000067560 |
acox1
|
acyl-CoA oxidase 1, palmitoyl |
| chr21_+_6114709 | 1.25 |
ENSDART00000065858
|
fpgs
|
folylpolyglutamate synthase |
| chr21_+_6114305 | 1.23 |
ENSDART00000141607
|
fpgs
|
folylpolyglutamate synthase |
| chr9_+_907459 | 1.22 |
ENSDART00000034850
ENSDART00000144114 |
dbi
|
diazepam binding inhibitor (GABA receptor modulator, acyl-CoA binding protein) |
| chr10_+_10728870 | 1.18 |
ENSDART00000109282
|
swi5
|
SWI5 homologous recombination repair protein |
| chr25_+_35913614 | 1.17 |
ENSDART00000022437
|
gpia
|
glucose-6-phosphate isomerase a |
| chr18_+_20566817 | 1.15 |
ENSDART00000100716
|
bida
|
BH3 interacting domain death agonist |
| chr4_-_5455506 | 1.14 |
ENSDART00000156593
ENSDART00000154676 |
si:dkey-14d8.22
|
si:dkey-14d8.22 |
| chr23_+_30006206 | 1.14 |
ENSDART00000150153
|
tnfrsf9a
|
tumor necrosis factor receptor superfamily, member 9a |
| chr1_-_47161996 | 1.11 |
ENSDART00000053153
|
mhc1zba
|
major histocompatibility complex class I ZBA |
| chr13_-_50200348 | 1.11 |
ENSDART00000038391
|
pkz
|
protein kinase containing Z-DNA binding domains |
| chr16_+_35395597 | 1.06 |
ENSDART00000158143
|
si:dkey-34d22.5
|
si:dkey-34d22.5 |
| chr8_-_4100365 | 1.05 |
ENSDART00000142846
|
cux2b
|
cut-like homeobox 2b |
| chr3_+_726000 | 1.04 |
ENSDART00000158510
|
dicp1.17
|
diverse immunoglobulin domain-containing protein 1.17 |
| chr7_-_73846995 | 1.04 |
ENSDART00000188079
|
FP236812.4
|
|
| chr20_+_23501535 | 1.02 |
ENSDART00000177922
ENSDART00000058532 |
palld
|
palladin, cytoskeletal associated protein |
| chr1_+_7189856 | 1.00 |
ENSDART00000092114
|
si:ch73-383l1.1
|
si:ch73-383l1.1 |
| chr16_-_9425451 | 0.96 |
ENSDART00000149163
|
ccr8.1
|
chemokine (C-C motif) receptor 8.1 |
| chr17_+_8323348 | 0.95 |
ENSDART00000169900
|
cdc42bpaa
|
CDC42 binding protein kinase alpha (DMPK-like) a |
| chr19_+_40832493 | 0.92 |
ENSDART00000145421
|
si:ch211-120e1.1
|
si:ch211-120e1.1 |
| chr18_-_3166726 | 0.91 |
ENSDART00000165002
|
aqp11
|
aquaporin 11 |
| chr4_+_9679840 | 0.90 |
ENSDART00000190889
|
BX901962.12
|
|
| chr19_-_2421793 | 0.87 |
ENSDART00000180238
|
TMEM196 (1 of many)
|
transmembrane protein 196 |
| chr13_-_50200042 | 0.87 |
ENSDART00000074230
|
pkz
|
protein kinase containing Z-DNA binding domains |
| chr22_-_3182965 | 0.87 |
ENSDART00000158009
|
lonp1
|
lon peptidase 1, mitochondrial |
| chr23_-_35483163 | 0.87 |
ENSDART00000138660
ENSDART00000113643 ENSDART00000189269 |
fbxo25
|
F-box protein 25 |
| chr23_+_44157682 | 0.86 |
ENSDART00000164474
ENSDART00000149928 |
si:ch73-106g13.1
|
si:ch73-106g13.1 |
| chr25_-_37284370 | 0.84 |
ENSDART00000103222
|
nudt7
|
nudix (nucleoside diphosphate linked moiety X)-type motif 7 |
| chr10_+_3299829 | 0.84 |
ENSDART00000183684
|
zgc:56235
|
zgc:56235 |
| chr16_-_13613475 | 0.83 |
ENSDART00000139102
|
dbpb
|
D site albumin promoter binding protein b |
| chr7_-_58729894 | 0.83 |
ENSDART00000149347
|
chchd7
|
coiled-coil-helix-coiled-coil-helix domain containing 7 |
| chr7_-_73815262 | 0.83 |
ENSDART00000185351
|
zgc:165555
|
zgc:165555 |
| chr17_-_45386823 | 0.83 |
ENSDART00000156002
|
tmem206
|
transmembrane protein 206 |
| chr1_-_52201266 | 0.83 |
ENSDART00000143805
ENSDART00000023757 |
rab3da
|
RAB3D, member RAS oncogene family, a |
| chr15_-_1590858 | 0.82 |
ENSDART00000081875
|
nnr
|
nanor |
| chr17_+_2162916 | 0.80 |
ENSDART00000103775
|
pak6a
|
p21 protein (Cdc42/Rac)-activated kinase 6a |
| chr21_+_37436907 | 0.79 |
ENSDART00000182611
ENSDART00000076328 |
pgrmc1
|
progesterone receptor membrane component 1 |
| chr9_+_7068832 | 0.78 |
ENSDART00000186685
|
BX629348.1
|
|
| chr10_-_34002185 | 0.78 |
ENSDART00000046599
|
zar1l
|
zygote arrest 1-like |
| chr10_-_4980150 | 0.77 |
ENSDART00000093228
|
mat2al
|
methionine adenosyltransferase II, alpha-like |
| chr10_+_35265083 | 0.77 |
ENSDART00000048831
|
tmem120a
|
transmembrane protein 120A |
| chr15_-_20468302 | 0.76 |
ENSDART00000018514
|
dlc
|
deltaC |
| chr10_-_10864331 | 0.75 |
ENSDART00000122657
|
nrarpa
|
NOTCH regulated ankyrin repeat protein a |
| chr1_-_34450622 | 0.75 |
ENSDART00000083736
|
lmo7b
|
LIM domain 7b |
| chr19_+_30495109 | 0.74 |
ENSDART00000192229
|
BX510301.1
|
|
| chr22_-_20924747 | 0.73 |
ENSDART00000185845
ENSDART00000179672 |
ell
|
elongation factor RNA polymerase II |
| chr25_+_3507368 | 0.73 |
ENSDART00000157777
|
zgc:153293
|
zgc:153293 |
| chr20_+_492529 | 0.72 |
ENSDART00000141709
|
FO904980.1
|
|
| chr22_+_508290 | 0.71 |
ENSDART00000135403
|
nuak2
|
NUAK family, SNF1-like kinase, 2 |
| chr21_+_45502621 | 0.71 |
ENSDART00000166719
|
si:dkey-223p19.2
|
si:dkey-223p19.2 |
| chr15_-_41762530 | 0.70 |
ENSDART00000187125
ENSDART00000154971 |
ftr91
|
finTRIM family, member 91 |
| chr19_+_46237665 | 0.68 |
ENSDART00000159391
|
vps28
|
vacuolar protein sorting 28 (yeast) |
| chr24_+_17005647 | 0.68 |
ENSDART00000149149
|
zfx
|
zinc finger protein, X-linked |
| chr21_+_45502773 | 0.68 |
ENSDART00000160059
ENSDART00000165704 |
si:dkey-223p19.2
|
si:dkey-223p19.2 |
| chr7_-_64152087 | 0.68 |
ENSDART00000165234
|
CR354398.1
|
|
| chr24_-_6671549 | 0.68 |
ENSDART00000114700
|
unm_sa821
|
un-named sa821 |
| chr18_+_41549764 | 0.66 |
ENSDART00000059135
|
bcl7bb
|
BCL tumor suppressor 7Bb |
| chr18_+_40584288 | 0.64 |
ENSDART00000087692
|
si:ch211-132b12.3
|
si:ch211-132b12.3 |
| chr22_-_6562618 | 0.64 |
ENSDART00000106100
|
zgc:171490
|
zgc:171490 |
| chr9_+_3035128 | 0.63 |
ENSDART00000140693
|
si:ch211-173m16.2
|
si:ch211-173m16.2 |
| chr13_+_21676235 | 0.63 |
ENSDART00000137804
ENSDART00000134950 ENSDART00000129653 |
mtg1
|
mitochondrial ribosome-associated GTPase 1 |
| chr7_+_29044888 | 0.63 |
ENSDART00000086871
|
gfod2
|
glucose-fructose oxidoreductase domain containing 2 |
| chr13_-_44808783 | 0.62 |
ENSDART00000099984
|
glo1
|
glyoxalase 1 |
| chr8_-_53490376 | 0.62 |
ENSDART00000158789
|
chdh
|
choline dehydrogenase |
| chr12_-_22670279 | 0.60 |
ENSDART00000164888
|
zcchc4
|
zinc finger, CCHC domain containing 4 |
| chr17_-_45387134 | 0.59 |
ENSDART00000010975
|
tmem206
|
transmembrane protein 206 |
| chr16_+_43077909 | 0.59 |
ENSDART00000014140
|
rundc3b
|
RUN domain containing 3b |
| chr12_-_34295298 | 0.59 |
ENSDART00000182789
|
CU855552.1
|
|
| chr3_+_5297493 | 0.57 |
ENSDART00000138596
|
si:ch211-150d5.3
|
si:ch211-150d5.3 |
| chr20_-_46554440 | 0.56 |
ENSDART00000043298
ENSDART00000060680 |
fosab
|
v-fos FBJ murine osteosarcoma viral oncogene homolog Ab |
| chr7_+_29153018 | 0.55 |
ENSDART00000138065
|
tmppe
|
transmembrane protein with metallophosphoesterase domain |
| chr16_-_26132122 | 0.55 |
ENSDART00000157787
|
lipeb
|
lipase, hormone-sensitive b |
| chr22_-_6801876 | 0.49 |
ENSDART00000135726
|
si:ch1073-188e1.1
|
si:ch1073-188e1.1 |
| chr25_+_25123385 | 0.47 |
ENSDART00000163892
|
ldha
|
lactate dehydrogenase A4 |
| chr7_+_20260172 | 0.47 |
ENSDART00000012450
|
dvl2
|
dishevelled segment polarity protein 2 |
| chr21_-_13231101 | 0.47 |
ENSDART00000190943
|
setb
|
SET nuclear proto-oncogene b |
| chr15_-_19443997 | 0.46 |
ENSDART00000114936
|
esamb
|
endothelial cell adhesion molecule b |
| chr10_-_5821584 | 0.45 |
ENSDART00000166388
|
il6st
|
interleukin 6 signal transducer |
| chr13_-_27916439 | 0.44 |
ENSDART00000139081
ENSDART00000087097 |
ogfrl1
|
opioid growth factor receptor-like 1 |
| chr7_+_52364748 | 0.44 |
ENSDART00000006795
|
gnrhr2
|
gonadotropin releasing hormone receptor 2 |
| chr4_+_14957360 | 0.43 |
ENSDART00000002770
ENSDART00000111882 ENSDART00000148292 |
tspan33a
|
tetraspanin 33a |
| chr19_+_29854223 | 0.43 |
ENSDART00000109729
ENSDART00000157419 |
si:ch73-130a3.4
|
si:ch73-130a3.4 |
| chr8_-_14179798 | 0.43 |
ENSDART00000040645
ENSDART00000146749 |
rhoaa
|
ras homolog gene family, member Aa |
| chr7_-_50517023 | 0.42 |
ENSDART00000073910
|
adamtsl5
|
ADAMTS like 5 |
| chr7_+_58730201 | 0.41 |
ENSDART00000073640
|
plag1
|
pleiomorphic adenoma gene 1 |
| chr23_-_43393083 | 0.41 |
ENSDART00000112758
|
zgc:174862
|
zgc:174862 |
| chr6_-_54111928 | 0.40 |
ENSDART00000083880
|
hyal2a
|
hyaluronoglucosaminidase 2a |
| chr22_-_6777976 | 0.39 |
ENSDART00000187946
|
CT583625.5
|
|
| chr13_-_42673978 | 0.39 |
ENSDART00000133848
ENSDART00000099738 ENSDART00000099729 ENSDART00000169083 |
lrrfip2
|
leucine rich repeat (in FLII) interacting protein 2 |
| chr4_-_9592402 | 0.39 |
ENSDART00000114060
|
cdnf
|
cerebral dopamine neurotrophic factor |
| chr14_+_80685 | 0.38 |
ENSDART00000188443
|
stag3
|
stromal antigen 3 |
| chr23_-_18707418 | 0.37 |
ENSDART00000144668
ENSDART00000141205 ENSDART00000016765 |
zgc:103759
|
zgc:103759 |
| chr9_-_19489264 | 0.37 |
ENSDART00000122894
|
wdr4
|
WD repeat domain 4 |
| chr7_-_70665965 | 0.36 |
ENSDART00000097710
|
ppargc1a
|
peroxisome proliferator-activated receptor gamma, coactivator 1 alpha |
| chr21_-_13230925 | 0.36 |
ENSDART00000023834
|
setb
|
SET nuclear proto-oncogene b |
| chr17_-_6765877 | 0.36 |
ENSDART00000193102
|
CR352340.1
|
|
| chr24_-_36238054 | 0.35 |
ENSDART00000155725
|
tmem241
|
transmembrane protein 241 |
| chr6_-_10034145 | 0.34 |
ENSDART00000185999
|
nudt15
|
nudix (nucleoside diphosphate linked moiety X)-type motif 15 |
| chr13_+_13033424 | 0.34 |
ENSDART00000159441
|
letm1
|
leucine zipper-EF-hand containing transmembrane protein 1 |
| chr3_+_634682 | 0.34 |
ENSDART00000164305
|
dicp1.1
|
diverse immunoglobulin domain-containing protein 1.1 |
| chr3_-_16039619 | 0.32 |
ENSDART00000143324
|
spsb3a
|
splA/ryanodine receptor domain and SOCS box containing 3a |
| chr23_-_5818992 | 0.32 |
ENSDART00000148730
|
csrp1a
|
cysteine and glycine-rich protein 1a |
| chr19_-_32944050 | 0.32 |
ENSDART00000137611
|
azin1b
|
antizyme inhibitor 1b |
| chr21_+_28535203 | 0.31 |
ENSDART00000184950
|
CR749775.2
|
|
| chr3_+_59117136 | 0.31 |
ENSDART00000182745
|
MGAT5B
|
mannosyl (alpha-1,6-)-glycoprotein beta-1,6-N-acetyl-glucosaminyltransferase, isozyme B |
| chr17_+_53250802 | 0.30 |
ENSDART00000143819
|
VASH1
|
vasohibin 1 |
| chr24_+_35387517 | 0.30 |
ENSDART00000058571
|
snai2
|
snail family zinc finger 2 |
| chr24_-_37688398 | 0.29 |
ENSDART00000141414
|
zgc:112185
|
zgc:112185 |
| chr10_-_1625080 | 0.29 |
ENSDART00000137285
|
nup155
|
nucleoporin 155 |
| chr17_-_18898115 | 0.29 |
ENSDART00000028044
|
galcb
|
galactosylceramidase b |
| chr9_-_1984604 | 0.29 |
ENSDART00000082339
|
hoxd12a
|
homeobox D12a |
| chr6_-_43233373 | 0.28 |
ENSDART00000188945
|
trnt1
|
tRNA nucleotidyl transferase, CCA-adding, 1 |
| chr3_+_24612191 | 0.27 |
ENSDART00000190998
|
zgc:171506
|
zgc:171506 |
| chr21_-_11657043 | 0.25 |
ENSDART00000141297
|
cast
|
calpastatin |
| chr24_-_39633224 | 0.25 |
ENSDART00000105646
|
fam234a
|
family with sequence similarity 234, member A |
| chr11_-_36341028 | 0.25 |
ENSDART00000146093
|
sort1a
|
sortilin 1a |
| chr18_+_7543347 | 0.24 |
ENSDART00000103467
|
arf5
|
ADP-ribosylation factor 5 |
| chr4_-_12718367 | 0.24 |
ENSDART00000035259
|
mgst1.1
|
microsomal glutathione S-transferase 1.1 |
| chr14_-_24101897 | 0.23 |
ENSDART00000143695
|
cpeb4a
|
cytoplasmic polyadenylation element binding protein 4a |
| chr7_-_32895668 | 0.22 |
ENSDART00000141828
|
ano5b
|
anoctamin 5b |
| chr22_+_19528851 | 0.21 |
ENSDART00000145079
|
si:dkey-78l4.13
|
si:dkey-78l4.13 |
| chr8_-_13046089 | 0.21 |
ENSDART00000137784
|
si:dkey-208b23.5
|
si:dkey-208b23.5 |
| chr11_+_13071645 | 0.20 |
ENSDART00000162259
|
zfyve9b
|
zinc finger, FYVE domain containing 9b |
| chr9_+_8380728 | 0.20 |
ENSDART00000133501
|
si:ch1073-75o15.4
|
si:ch1073-75o15.4 |
| chr25_+_1304173 | 0.19 |
ENSDART00000155229
|
rxfp3.3b
|
relaxin/insulin-like family peptide receptor 3.3b |
| chr8_-_40183197 | 0.18 |
ENSDART00000005118
|
gpx8
|
glutathione peroxidase 8 (putative) |
| chr20_-_25748407 | 0.18 |
ENSDART00000063152
|
chst14
|
carbohydrate (N-acetylgalactosamine 4-0) sulfotransferase 14 |
| chr20_+_46311242 | 0.18 |
ENSDART00000038696
|
flvcr2b
|
feline leukemia virus subgroup C cellular receptor family, member 2b |
| chr12_+_499881 | 0.17 |
ENSDART00000167527
|
mprip
|
myosin phosphatase Rho interacting protein |
| chr12_-_4540564 | 0.15 |
ENSDART00000106566
|
CABZ01030107.1
|
|
| chr10_+_21444654 | 0.14 |
ENSDART00000140113
ENSDART00000184386 ENSDART00000019252 |
fbxw11b
|
F-box and WD repeat domain containing 11b |
| chr2_+_13069168 | 0.14 |
ENSDART00000192832
|
prkag2b
|
protein kinase, AMP-activated, gamma 2 non-catalytic subunit b |
| chr14_+_6963312 | 0.14 |
ENSDART00000150050
|
hnrnpaba
|
heterogeneous nuclear ribonucleoprotein A/Ba |
| chr9_+_19489304 | 0.13 |
ENSDART00000151920
|
si:ch211-140m22.7
|
si:ch211-140m22.7 |
| chr10_-_41980797 | 0.13 |
ENSDART00000076575
|
rhof
|
ras homolog family member F |
| chr23_+_42338325 | 0.13 |
ENSDART00000169660
|
cyp2aa7
|
cytochrome P450, family 2, subfamily AA, polypeptide 7 |
| chr6_-_43233700 | 0.12 |
ENSDART00000113706
|
trnt1
|
tRNA nucleotidyl transferase, CCA-adding, 1 |
| chr7_-_3470451 | 0.11 |
ENSDART00000173387
|
si:ch211-285c6.6
|
si:ch211-285c6.6 |
| chr15_-_1001177 | 0.10 |
ENSDART00000160730
|
zgc:162936
|
zgc:162936 |
| chr4_+_76492691 | 0.09 |
ENSDART00000186337
|
CU104731.1
|
|
| chr19_+_22727940 | 0.09 |
ENSDART00000052509
|
trhrb
|
thyrotropin-releasing hormone receptor b |
| chr8_+_29963024 | 0.09 |
ENSDART00000149372
|
ptch1
|
patched 1 |
| chr10_+_41765616 | 0.09 |
ENSDART00000170682
|
rnf34b
|
ring finger protein 34b |
| chr25_+_3788443 | 0.08 |
ENSDART00000189747
|
chid1
|
chitinase domain containing 1 |
| chr2_-_56655769 | 0.08 |
ENSDART00000113589
|
gpx4b
|
glutathione peroxidase 4b |
| chr11_+_5499661 | 0.06 |
ENSDART00000027850
|
slc35e1
|
solute carrier family 35, member E1 |
| chr3_-_27915270 | 0.06 |
ENSDART00000115370
|
mettl22
|
methyltransferase like 22 |
| chr6_+_54576520 | 0.06 |
ENSDART00000093199
ENSDART00000127519 ENSDART00000157142 |
tead3b
|
TEA domain family member 3 b |
| chr15_+_17752928 | 0.06 |
ENSDART00000155314
|
si:ch211-213d14.2
|
si:ch211-213d14.2 |
| chr2_-_3614005 | 0.05 |
ENSDART00000110399
|
pter
|
phosphotriesterase related |
| chr7_+_9880811 | 0.05 |
ENSDART00000128376
|
lins1
|
lines homolog 1 |
| chr11_+_34921492 | 0.05 |
ENSDART00000128070
|
gnai2a
|
guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 2a |
| chr16_+_39271123 | 0.04 |
ENSDART00000043823
ENSDART00000141801 |
osbpl10b
|
oxysterol binding protein-like 10b |
| chr20_+_53541389 | 0.03 |
ENSDART00000100110
|
pak6b
|
p21 protein (Cdc42/Rac)-activated kinase 6b |
| chr3_+_1211242 | 0.02 |
ENSDART00000171287
ENSDART00000165769 |
poldip3
|
polymerase (DNA-directed), delta interacting protein 3 |
| chr21_-_41070182 | 0.01 |
ENSDART00000026064
|
larsb
|
leucyl-tRNA synthetase b |
| chr17_+_6765621 | 0.01 |
ENSDART00000156637
ENSDART00000007622 |
afg1la
|
AFG1 like ATPase a |
| chr23_-_10745288 | 0.01 |
ENSDART00000140745
ENSDART00000013768 |
eif4e3
|
eukaryotic translation initiation factor 4E family member 3 |
Network of associatons between targets according to the STRING database.
First level regulatory network of klf13+zgc:153115
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 4.2 | GO:0005997 | xylulose metabolic process(GO:0005997) |
| 0.9 | 3.7 | GO:0055021 | regulation of cardiac muscle tissue growth(GO:0055021) regulation of cardiac muscle cell proliferation(GO:0060043) |
| 0.9 | 4.5 | GO:0051145 | smooth muscle cell differentiation(GO:0051145) negative regulation of muscle cell differentiation(GO:0051148) |
| 0.7 | 2.0 | GO:2001014 | skeletal muscle fiber differentiation(GO:0098528) regulation of skeletal muscle cell differentiation(GO:2001014) |
| 0.6 | 2.5 | GO:0046900 | tetrahydrofolylpolyglutamate metabolic process(GO:0046900) |
| 0.5 | 2.1 | GO:0019626 | short-chain fatty acid catabolic process(GO:0019626) |
| 0.4 | 13.3 | GO:0050821 | protein stabilization(GO:0050821) |
| 0.3 | 1.3 | GO:0033540 | fatty acid beta-oxidation using acyl-CoA oxidase(GO:0033540) |
| 0.3 | 1.1 | GO:2001244 | positive regulation of intrinsic apoptotic signaling pathway(GO:2001244) |
| 0.3 | 2.2 | GO:0001579 | medium-chain fatty acid transport(GO:0001579) |
| 0.3 | 3.8 | GO:0001840 | neural plate development(GO:0001840) |
| 0.3 | 0.8 | GO:1903537 | meiotic cell cycle process involved in oocyte maturation(GO:1903537) regulation of meiotic cell cycle process involved in oocyte maturation(GO:1903538) |
| 0.3 | 0.8 | GO:0060842 | arterial endothelial cell differentiation(GO:0060842) |
| 0.2 | 2.2 | GO:0000056 | ribosomal small subunit export from nucleus(GO:0000056) |
| 0.2 | 2.2 | GO:0045741 | positive regulation of epidermal growth factor-activated receptor activity(GO:0045741) positive regulation of receptor activity(GO:2000273) |
| 0.2 | 1.4 | GO:0032367 | intracellular cholesterol transport(GO:0032367) |
| 0.2 | 0.7 | GO:0006178 | guanine salvage(GO:0006178) GMP salvage(GO:0032263) guanine biosynthetic process(GO:0046099) |
| 0.2 | 2.4 | GO:0060754 | mast cell chemotaxis(GO:0002551) regulation of mast cell chemotaxis(GO:0060753) positive regulation of mast cell chemotaxis(GO:0060754) mast cell migration(GO:0097531) |
| 0.2 | 0.6 | GO:0019695 | choline metabolic process(GO:0019695) |
| 0.2 | 2.2 | GO:0061620 | glucose catabolic process(GO:0006007) NADH regeneration(GO:0006735) glycolytic process through fructose-6-phosphate(GO:0061615) glycolytic process through glucose-6-phosphate(GO:0061620) canonical glycolysis(GO:0061621) glucose catabolic process to pyruvate(GO:0061718) |
| 0.2 | 1.5 | GO:0010887 | negative regulation of cholesterol storage(GO:0010887) |
| 0.1 | 0.7 | GO:0042795 | snRNA transcription from RNA polymerase II promoter(GO:0042795) |
| 0.1 | 0.4 | GO:0072020 | proximal straight tubule development(GO:0072020) distal convoluted tubule development(GO:0072025) late distal convoluted tubule development(GO:0072068) |
| 0.1 | 0.3 | GO:0034214 | protein hexamerization(GO:0034214) |
| 0.1 | 0.8 | GO:0006556 | S-adenosylmethionine biosynthetic process(GO:0006556) |
| 0.1 | 0.3 | GO:0090435 | protein targeting to nuclear inner membrane(GO:0036228) protein localization to nuclear envelope(GO:0090435) |
| 0.1 | 0.8 | GO:0009109 | coenzyme catabolic process(GO:0009109) |
| 0.1 | 2.1 | GO:0005979 | regulation of glycogen biosynthetic process(GO:0005979) regulation of glucan biosynthetic process(GO:0010962) |
| 0.1 | 0.4 | GO:0036265 | RNA (guanine-N7)-methylation(GO:0036265) |
| 0.1 | 2.2 | GO:0030032 | lamellipodium assembly(GO:0030032) |
| 0.1 | 0.8 | GO:0045444 | fat cell differentiation(GO:0045444) |
| 0.1 | 0.7 | GO:0043328 | protein targeting to vacuole involved in ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043328) |
| 0.1 | 3.1 | GO:0006749 | glutathione metabolic process(GO:0006749) |
| 0.1 | 0.2 | GO:1904158 | axonemal central apparatus assembly(GO:1904158) |
| 0.1 | 1.4 | GO:0071350 | interleukin-15-mediated signaling pathway(GO:0035723) response to interleukin-15(GO:0070672) cellular response to interleukin-15(GO:0071350) |
| 0.1 | 0.8 | GO:0002043 | blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:0002043) |
| 0.1 | 0.5 | GO:0090177 | establishment of planar polarity of embryonic epithelium(GO:0042249) establishment of planar polarity involved in neural tube closure(GO:0090177) regulation of establishment of planar polarity involved in neural tube closure(GO:0090178) planar cell polarity pathway involved in neural tube closure(GO:0090179) |
| 0.1 | 0.4 | GO:0048680 | positive regulation of axon regeneration(GO:0048680) positive regulation of neuron projection regeneration(GO:0070572) |
| 0.1 | 0.4 | GO:1902766 | skeletal muscle satellite cell migration(GO:1902766) |
| 0.1 | 2.7 | GO:0032784 | regulation of DNA-templated transcription, elongation(GO:0032784) |
| 0.1 | 0.9 | GO:0051131 | chaperone-mediated protein complex assembly(GO:0051131) |
| 0.1 | 0.6 | GO:0061026 | cardiac muscle tissue regeneration(GO:0061026) |
| 0.1 | 1.2 | GO:0051156 | glucose 6-phosphate metabolic process(GO:0051156) |
| 0.1 | 0.3 | GO:0015846 | polyamine transport(GO:0015846) polyamine transmembrane transport(GO:1902047) regulation of polyamine transmembrane transport(GO:1902267) positive regulation of polyamine transmembrane transport(GO:1902269) |
| 0.1 | 1.4 | GO:0001757 | somite specification(GO:0001757) |
| 0.0 | 1.5 | GO:0014866 | skeletal myofibril assembly(GO:0014866) |
| 0.0 | 0.3 | GO:0046070 | dGTP catabolic process(GO:0006203) purine nucleoside triphosphate catabolic process(GO:0009146) purine deoxyribonucleoside triphosphate catabolic process(GO:0009217) dGTP metabolic process(GO:0046070) |
| 0.0 | 0.6 | GO:0031167 | rRNA methylation(GO:0031167) |
| 0.0 | 0.4 | GO:0016074 | snoRNA metabolic process(GO:0016074) |
| 0.0 | 0.2 | GO:0002931 | response to ischemia(GO:0002931) |
| 0.0 | 0.3 | GO:0097341 | inhibition of cysteine-type endopeptidase activity(GO:0097340) zymogen inhibition(GO:0097341) |
| 0.0 | 0.3 | GO:0046477 | glycosylceramide catabolic process(GO:0046477) |
| 0.0 | 0.6 | GO:0019433 | triglyceride catabolic process(GO:0019433) |
| 0.0 | 2.0 | GO:0051607 | defense response to virus(GO:0051607) |
| 0.0 | 0.4 | GO:0030214 | hyaluronan catabolic process(GO:0030214) |
| 0.0 | 0.2 | GO:0006895 | Golgi to endosome transport(GO:0006895) |
| 0.0 | 0.2 | GO:0050655 | dermatan sulfate proteoglycan metabolic process(GO:0050655) |
| 0.0 | 0.8 | GO:0042149 | cellular response to glucose starvation(GO:0042149) |
| 0.0 | 0.1 | GO:0097037 | heme export(GO:0097037) |
| 0.0 | 0.8 | GO:0006904 | vesicle docking involved in exocytosis(GO:0006904) |
| 0.0 | 1.5 | GO:0031076 | embryonic camera-type eye development(GO:0031076) |
| 0.0 | 1.7 | GO:0006334 | nucleosome assembly(GO:0006334) |
| 0.0 | 0.4 | GO:0071542 | dopaminergic neuron differentiation(GO:0071542) |
| 0.0 | 0.4 | GO:0001966 | thigmotaxis(GO:0001966) |
| 0.0 | 0.9 | GO:0018107 | peptidyl-threonine phosphorylation(GO:0018107) |
| 0.0 | 3.5 | GO:0006954 | inflammatory response(GO:0006954) |
| 0.0 | 3.5 | GO:0032259 | methylation(GO:0032259) |
| 0.0 | 1.2 | GO:0019722 | calcium-mediated signaling(GO:0019722) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.8 | GO:0044609 | DBIRD complex(GO:0044609) |
| 0.2 | 2.2 | GO:0005945 | 6-phosphofructokinase complex(GO:0005945) |
| 0.1 | 1.2 | GO:0033061 | DNA recombinase mediator complex(GO:0033061) |
| 0.1 | 1.7 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.1 | 0.3 | GO:0044611 | nuclear pore inner ring(GO:0044611) |
| 0.1 | 0.4 | GO:0043527 | tRNA methyltransferase complex(GO:0043527) |
| 0.0 | 1.0 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.0 | 2.5 | GO:0042641 | actomyosin(GO:0042641) |
| 0.0 | 0.7 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 0.0 | 0.8 | GO:0046930 | pore complex(GO:0046930) |
| 0.0 | 1.9 | GO:0098857 | membrane raft(GO:0045121) membrane microdomain(GO:0098857) |
| 0.0 | 1.6 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.0 | 0.8 | GO:0005637 | nuclear inner membrane(GO:0005637) |
| 0.0 | 24.1 | GO:0005829 | cytosol(GO:0005829) |
| 0.0 | 0.2 | GO:1990124 | messenger ribonucleoprotein complex(GO:1990124) |
| 0.0 | 1.3 | GO:0005777 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.0 | 0.4 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 0.7 | GO:0008023 | transcription elongation factor complex(GO:0008023) |
| 0.0 | 3.6 | GO:0000785 | chromatin(GO:0000785) |
| 0.0 | 0.1 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.0 | 2.7 | GO:0009897 | external side of plasma membrane(GO:0009897) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.3 | 13.3 | GO:0004365 | glyceraldehyde-3-phosphate dehydrogenase (NAD+) (phosphorylating) activity(GO:0004365) glyceraldehyde-3-phosphate dehydrogenase (NAD(P)+) (phosphorylating) activity(GO:0043891) |
| 1.2 | 3.7 | GO:0008119 | thiopurine S-methyltransferase activity(GO:0008119) |
| 0.7 | 2.0 | GO:1990715 | mRNA CDS binding(GO:1990715) sequence-specific mRNA binding(GO:1990825) |
| 0.6 | 2.4 | GO:0043183 | vascular endothelial growth factor receptor 1 binding(GO:0043183) |
| 0.6 | 2.2 | GO:1904121 | phosphatidylethanolamine transporter activity(GO:1904121) |
| 0.5 | 2.0 | GO:0004694 | eukaryotic translation initiation factor 2alpha kinase activity(GO:0004694) |
| 0.3 | 1.4 | GO:0030228 | lipoprotein particle receptor activity(GO:0030228) |
| 0.3 | 1.9 | GO:0004692 | cGMP-dependent protein kinase activity(GO:0004692) |
| 0.3 | 1.3 | GO:0003997 | acyl-CoA oxidase activity(GO:0003997) |
| 0.2 | 0.7 | GO:0004422 | hypoxanthine phosphoribosyltransferase activity(GO:0004422) |
| 0.2 | 2.2 | GO:0003872 | 6-phosphofructokinase activity(GO:0003872) |
| 0.2 | 0.8 | GO:0004478 | methionine adenosyltransferase activity(GO:0004478) |
| 0.2 | 0.8 | GO:0015288 | porin activity(GO:0015288) |
| 0.1 | 2.1 | GO:0003995 | acyl-CoA dehydrogenase activity(GO:0003995) |
| 0.1 | 2.1 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.1 | 3.2 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.1 | 0.4 | GO:1990174 | phosphodiesterase decapping endonuclease activity(GO:1990174) |
| 0.1 | 0.4 | GO:0004968 | gonadotropin-releasing hormone receptor activity(GO:0004968) |
| 0.1 | 2.5 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.1 | 0.4 | GO:0033906 | hyaluronoglucuronidase activity(GO:0033906) |
| 0.1 | 2.1 | GO:0008253 | 5'-nucleotidase activity(GO:0008253) |
| 0.1 | 3.3 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.1 | 0.9 | GO:0004176 | ATP-dependent peptidase activity(GO:0004176) |
| 0.1 | 0.3 | GO:0051139 | calcium:proton antiporter activity(GO:0015369) metal ion:proton antiporter activity(GO:0051139) |
| 0.1 | 2.5 | GO:0016881 | acid-amino acid ligase activity(GO:0016881) |
| 0.1 | 1.1 | GO:0016861 | intramolecular oxidoreductase activity, interconverting aldoses and ketoses(GO:0016861) |
| 0.1 | 0.6 | GO:0016433 | rRNA (adenine) methyltransferase activity(GO:0016433) |
| 0.1 | 0.6 | GO:0033878 | hormone-sensitive lipase activity(GO:0033878) |
| 0.1 | 0.5 | GO:0004459 | L-lactate dehydrogenase activity(GO:0004459) |
| 0.1 | 1.4 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 0.1 | 1.5 | GO:0001103 | RNA polymerase II repressing transcription factor binding(GO:0001103) |
| 0.1 | 0.3 | GO:0047429 | nucleoside-triphosphate diphosphatase activity(GO:0047429) |
| 0.1 | 2.2 | GO:0015459 | potassium channel regulator activity(GO:0015459) |
| 0.1 | 1.2 | GO:0000062 | fatty-acyl-CoA binding(GO:0000062) |
| 0.1 | 0.3 | GO:0042978 | ornithine decarboxylase activator activity(GO:0042978) |
| 0.0 | 0.6 | GO:0005332 | gamma-aminobutyric acid:sodium symporter activity(GO:0005332) |
| 0.0 | 0.4 | GO:0004985 | opioid receptor activity(GO:0004985) |
| 0.0 | 0.8 | GO:0016289 | CoA hydrolase activity(GO:0016289) |
| 0.0 | 0.6 | GO:0016846 | carbon-sulfur lyase activity(GO:0016846) |
| 0.0 | 0.3 | GO:0010859 | calcium-dependent cysteine-type endopeptidase inhibitor activity(GO:0010859) |
| 0.0 | 4.5 | GO:0042803 | protein homodimerization activity(GO:0042803) |
| 0.0 | 4.4 | GO:0016614 | oxidoreductase activity, acting on CH-OH group of donors(GO:0016614) |
| 0.0 | 0.8 | GO:0005112 | Notch binding(GO:0005112) |
| 0.0 | 1.2 | GO:0019957 | C-C chemokine binding(GO:0019957) |
| 0.0 | 0.3 | GO:0004602 | glutathione peroxidase activity(GO:0004602) |
| 0.0 | 0.4 | GO:0016423 | tRNA (guanine) methyltransferase activity(GO:0016423) |
| 0.0 | 1.3 | GO:0019955 | cytokine binding(GO:0019955) |
| 0.0 | 0.7 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.0 | 0.2 | GO:0000900 | translation repressor activity, nucleic acid binding(GO:0000900) |
| 0.0 | 0.1 | GO:0004997 | thyrotropin-releasing hormone receptor activity(GO:0004997) |
| 0.0 | 0.1 | GO:0015232 | heme transporter activity(GO:0015232) |
| 0.0 | 0.3 | GO:0042805 | actinin binding(GO:0042805) |
| 0.0 | 0.5 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.4 | GO:0030374 | ligand-dependent nuclear receptor transcription coactivator activity(GO:0030374) |
| 0.0 | 0.3 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.0 | 4.6 | GO:0016757 | transferase activity, transferring glycosyl groups(GO:0016757) |
| 0.0 | 4.1 | GO:0003924 | GTPase activity(GO:0003924) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 13.8 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.1 | 1.7 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.1 | 2.2 | PID FOXO PATHWAY | FoxO family signaling |
| 0.0 | 0.5 | PID WNT CANONICAL PATHWAY | Canonical Wnt signaling pathway |
| 0.0 | 0.4 | PID IL27 PATHWAY | IL27-mediated signaling events |
| 0.0 | 0.4 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 13.3 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.2 | 2.2 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.1 | 2.1 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.1 | 1.3 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.1 | 0.7 | REACTOME MEMBRANE BINDING AND TARGETTING OF GAG PROTEINS | Genes involved in Membrane binding and targetting of GAG proteins |
| 0.0 | 0.4 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 0.0 | 2.3 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.0 | 0.5 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.0 | 0.5 | REACTOME PYRUVATE METABOLISM | Genes involved in Pyruvate metabolism |
| 0.0 | 0.6 | REACTOME ELONGATION ARREST AND RECOVERY | Genes involved in Elongation arrest and recovery |
| 0.0 | 0.4 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |