Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
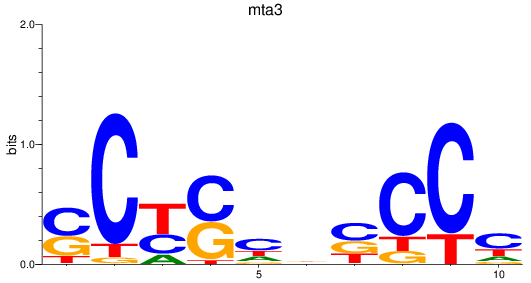
Results for mta3
Z-value: 0.26
Transcription factors associated with mta3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
mta3
|
ENSDARG00000054903 | metastasis associated 1 family, member 3 |
|
mta3
|
ENSDARG00000113436 | metastasis associated 1 family, member 3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| mta3 | dr11_v1_chr12_+_25223843_25223873 | -0.02 | 8.5e-01 | Click! |
Activity profile of mta3 motif
Sorted Z-values of mta3 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr17_-_2578026 | 1.46 |
ENSDART00000065821
|
zp3.2
|
zona pellucida glycoprotein 3, tandem duplicate 2 |
| chr7_+_1467863 | 1.25 |
ENSDART00000173433
|
emc4
|
ER membrane protein complex subunit 4 |
| chr5_-_54712159 | 1.23 |
ENSDART00000149207
|
ccnb1
|
cyclin B1 |
| chr18_-_45736 | 1.08 |
ENSDART00000148373
ENSDART00000148950 |
gatm
|
glycine amidinotransferase (L-arginine:glycine amidinotransferase) |
| chr18_-_46010 | 0.99 |
ENSDART00000052641
|
gatm
|
glycine amidinotransferase (L-arginine:glycine amidinotransferase) |
| chr22_-_193234 | 0.87 |
ENSDART00000131067
|
fbxo42
|
F-box protein 42 |
| chr12_-_290413 | 0.66 |
ENSDART00000152496
|
adprm
|
ADP-ribose/CDP-alcohol diphosphatase, manganese-dependent |
| chr2_-_22535 | 0.65 |
ENSDART00000157877
|
CABZ01092282.1
|
|
| chr1_+_15062 | 0.60 |
ENSDART00000169005
|
cep97
|
centrosomal protein 97 |
| chr5_-_23800376 | 0.47 |
ENSDART00000134184
|
gbgt1l4
|
globoside alpha-1,3-N-acetylgalactosaminyltransferase 1, like 4 |
| chr3_+_16566361 | 0.46 |
ENSDART00000132869
ENSDART00000080771 |
slc6a16a
|
solute carrier family 6, member 16a |
| chr21_+_44581654 | 0.44 |
ENSDART00000187201
ENSDART00000180360 ENSDART00000191543 ENSDART00000180039 ENSDART00000186308 |
TMEM164
|
transmembrane protein 164 |
| chr14_-_237130 | 0.42 |
ENSDART00000164988
|
bod1l1
|
biorientation of chromosomes in cell division 1-like 1 |
| chr21_+_45712247 | 0.40 |
ENSDART00000163982
|
camlg
|
calcium modulating ligand |
| chr22_+_26703026 | 0.39 |
ENSDART00000158756
|
crebbpa
|
CREB binding protein a |
| chr13_+_1100197 | 0.36 |
ENSDART00000139560
|
ppp3r1a
|
protein phosphatase 3, regulatory subunit B, alpha a |
| chr5_+_20453874 | 0.36 |
ENSDART00000124545
ENSDART00000008402 |
sart3
|
squamous cell carcinoma antigen recognized by T cells 3 |
| chr22_+_344763 | 0.35 |
ENSDART00000181934
|
CU914780.1
|
|
| chr12_+_13348918 | 0.35 |
ENSDART00000181373
|
rnasen
|
ribonuclease type III, nuclear |
| chr16_-_17207754 | 0.34 |
ENSDART00000063804
|
wu:fj39g12
|
wu:fj39g12 |
| chr11_-_20956309 | 0.34 |
ENSDART00000188659
|
CABZ01008739.1
|
|
| chr7_+_1579236 | 0.33 |
ENSDART00000172830
|
supt16h
|
SPT16 homolog, facilitates chromatin remodeling subunit |
| chr20_-_14665002 | 0.29 |
ENSDART00000152816
|
scrn2
|
secernin 2 |
| chr5_+_45139196 | 0.28 |
ENSDART00000113738
|
smarca2
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 |
| chr6_-_12270226 | 0.28 |
ENSDART00000180473
|
pkp4
|
plakophilin 4 |
| chr14_-_572742 | 0.21 |
ENSDART00000168951
|
fgf2
|
fibroblast growth factor 2 |
| chr23_-_24825863 | 0.20 |
ENSDART00000112493
|
syt6a
|
synaptotagmin VIa |
| chr14_-_2369849 | 0.20 |
ENSDART00000180422
ENSDART00000189731 ENSDART00000111748 |
pcdhb
|
protocadherin b |
| chr7_+_1579510 | 0.20 |
ENSDART00000190525
|
supt16h
|
SPT16 homolog, facilitates chromatin remodeling subunit |
| chr11_+_44579865 | 0.19 |
ENSDART00000173425
|
nid1b
|
nidogen 1b |
| chr20_-_54435287 | 0.18 |
ENSDART00000148632
|
yy1b
|
YY1 transcription factor b |
| chr7_-_6592142 | 0.17 |
ENSDART00000160137
|
kcnj10a
|
potassium inwardly-rectifying channel, subfamily J, member 10a |
| chr16_-_280835 | 0.14 |
ENSDART00000190541
|
LO017917.1
|
|
| chr9_-_1434484 | 0.14 |
ENSDART00000093412
|
osbpl6
|
oxysterol binding protein-like 6 |
| chr5_+_66433287 | 0.14 |
ENSDART00000170757
|
kntc1
|
kinetochore associated 1 |
| chr12_-_14551077 | 0.13 |
ENSDART00000188717
|
FO704673.1
|
|
| chr13_+_31545812 | 0.12 |
ENSDART00000076527
|
ppm1aa
|
protein phosphatase, Mg2+/Mn2+ dependent, 1Aa |
| chr25_-_35143360 | 0.11 |
ENSDART00000188033
|
zgc:165555
|
zgc:165555 |
| chr11_-_32723851 | 0.11 |
ENSDART00000155592
|
pcdh17
|
protocadherin 17 |
| chr17_-_25630822 | 0.11 |
ENSDART00000126201
ENSDART00000105503 ENSDART00000151878 |
rab3gap2
|
RAB3 GTPase activating protein subunit 2 (non-catalytic) |
| chr4_-_13255700 | 0.11 |
ENSDART00000162277
ENSDART00000026593 |
grip1
|
glutamate receptor interacting protein 1 |
| chr11_-_44543082 | 0.09 |
ENSDART00000099568
|
gpr137bb
|
G protein-coupled receptor 137Bb |
| chr7_+_17560808 | 0.08 |
ENSDART00000159699
ENSDART00000165724 ENSDART00000129017 ENSDART00000101955 |
nitr1b
|
novel immune-type receptor 1b |
| chr23_+_2825940 | 0.07 |
ENSDART00000135781
|
plcg1
|
phospholipase C, gamma 1 |
| chr22_-_7809904 | 0.07 |
ENSDART00000182269
ENSDART00000097201 ENSDART00000138522 |
si:ch73-44m9.3
si:ch73-44m9.1
|
si:ch73-44m9.3 si:ch73-44m9.1 |
| chr5_-_25123807 | 0.07 |
ENSDART00000183171
|
abca2
|
ATP-binding cassette, sub-family A (ABC1), member 2 |
| chr4_+_29909870 | 0.06 |
ENSDART00000150324
|
znf1111
|
zinc finger protein 1111 |
| chr3_-_5228137 | 0.06 |
ENSDART00000137105
|
myh9b
|
myosin, heavy chain 9b, non-muscle |
| chr4_-_34799395 | 0.06 |
ENSDART00000170744
|
si:dkey-146m20.13
|
si:dkey-146m20.13 |
| chr8_+_43056153 | 0.05 |
ENSDART00000186377
|
prnpa
|
prion protein a |
| chr22_-_367569 | 0.02 |
ENSDART00000041895
|
ssu72
|
SSU72 homolog, RNA polymerase II CTD phosphatase |
Network of associatons between targets according to the STRING database.
First level regulatory network of mta3
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.1 | GO:0006601 | creatine metabolic process(GO:0006600) creatine biosynthetic process(GO:0006601) |
| 0.2 | 1.3 | GO:0045050 | protein insertion into ER membrane by stop-transfer membrane-anchor sequence(GO:0045050) |
| 0.1 | 1.2 | GO:0007080 | mitotic metaphase plate congression(GO:0007080) |
| 0.1 | 0.4 | GO:1903003 | positive regulation of protein deubiquitination(GO:1903003) |
| 0.1 | 1.5 | GO:2000344 | binding of sperm to zona pellucida(GO:0007339) egg coat formation(GO:0035803) regulation of acrosome reaction(GO:0060046) positive regulation of acrosome reaction(GO:2000344) |
| 0.1 | 0.4 | GO:0032515 | negative regulation of phosphoprotein phosphatase activity(GO:0032515) |
| 0.0 | 0.2 | GO:0031507 | heterochromatin assembly(GO:0031507) |
| 0.0 | 0.5 | GO:0034724 | DNA replication-independent nucleosome organization(GO:0034724) |
| 0.0 | 0.1 | GO:0045671 | negative regulation of osteoclast differentiation(GO:0045671) |
| 0.0 | 0.5 | GO:0030259 | lipid glycosylation(GO:0030259) |
| 0.0 | 0.1 | GO:0035477 | regulation of angioblast cell migration involved in selective angioblast sprouting(GO:0035477) |
| 0.0 | 0.2 | GO:0036268 | swimming(GO:0036268) |
| 0.0 | 0.3 | GO:0032467 | positive regulation of cytokinesis(GO:0032467) |
| 0.0 | 0.3 | GO:0007168 | receptor guanylyl cyclase signaling pathway(GO:0007168) |
| 0.0 | 0.2 | GO:0030323 | respiratory tube development(GO:0030323) lung development(GO:0030324) |
| 0.0 | 0.2 | GO:0055090 | acylglycerol homeostasis(GO:0055090) triglyceride homeostasis(GO:0070328) |
| 0.0 | 0.2 | GO:0007340 | acrosome reaction(GO:0007340) |
| 0.0 | 0.1 | GO:0070650 | actin filament bundle distribution(GO:0070650) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.2 | GO:0097125 | cyclin B1-CDK1 complex(GO:0097125) |
| 0.1 | 1.3 | GO:0072546 | ER membrane protein complex(GO:0072546) |
| 0.0 | 0.5 | GO:0035101 | FACT complex(GO:0035101) |
| 0.0 | 0.4 | GO:0000940 | condensed chromosome outer kinetochore(GO:0000940) |
| 0.0 | 0.4 | GO:0015030 | Cajal body(GO:0015030) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.3 | GO:0032977 | membrane insertase activity(GO:0032977) |
| 0.1 | 1.5 | GO:0035804 | structural constituent of egg coat(GO:0035804) |
| 0.1 | 2.1 | GO:0016769 | transferase activity, transferring nitrogenous groups(GO:0016769) |
| 0.1 | 0.3 | GO:0070004 | cysteine-type exopeptidase activity(GO:0070004) |
| 0.0 | 0.7 | GO:0047631 | ADP-ribose diphosphatase activity(GO:0047631) |
| 0.0 | 0.4 | GO:0017070 | U6 snRNA binding(GO:0017070) |
| 0.0 | 0.4 | GO:0051721 | protein phosphatase 2A binding(GO:0051721) |
| 0.0 | 1.2 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.0 | 0.1 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.2 | PID FOXO PATHWAY | FoxO family signaling |
| 0.0 | 0.2 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.2 | REACTOME RECRUITMENT OF NUMA TO MITOTIC CENTROSOMES | Genes involved in Recruitment of NuMA to mitotic centrosomes |
| 0.0 | 0.2 | REACTOME SIGNALING BY ACTIVATED POINT MUTANTS OF FGFR1 | Genes involved in Signaling by activated point mutants of FGFR1 |
| 0.0 | 0.5 | REACTOME ELONGATION ARREST AND RECOVERY | Genes involved in Elongation arrest and recovery |
| 0.0 | 0.1 | REACTOME ROLE OF SECOND MESSENGERS IN NETRIN1 SIGNALING | Genes involved in Role of second messengers in netrin-1 signaling |
| 0.0 | 2.1 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |