Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
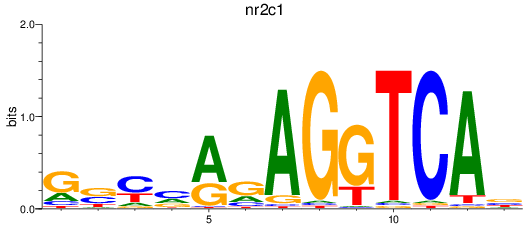
Results for nr2c1
Z-value: 0.75
Transcription factors associated with nr2c1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
nr2c1
|
ENSDARG00000045527 | nuclear receptor subfamily 2, group C, member 1 |
|
nr2c1
|
ENSDARG00000111803 | nuclear receptor subfamily 2, group C, member 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| nr2c1 | dr11_v1_chr4_-_25859305_25859305 | -0.14 | 1.6e-01 | Click! |
Activity profile of nr2c1 motif
Sorted Z-values of nr2c1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr3_+_30257582 | 9.68 |
ENSDART00000159497
ENSDART00000103457 ENSDART00000121883 |
mybpc2a
|
myosin binding protein C, fast type a |
| chr11_+_11201096 | 8.83 |
ENSDART00000171916
ENSDART00000171521 ENSDART00000087105 ENSDART00000159603 |
myom2a
|
myomesin 2a |
| chr22_-_7050 | 8.28 |
ENSDART00000127829
|
atad3
|
ATPase family, AAA domain containing 3 |
| chr2_+_38039857 | 7.96 |
ENSDART00000159951
|
casq1a
|
calsequestrin 1a |
| chr17_-_5583345 | 7.11 |
ENSDART00000035944
|
clic5a
|
chloride intracellular channel 5a |
| chr1_+_58353661 | 6.93 |
ENSDART00000140074
|
si:dkey-222h21.2
|
si:dkey-222h21.2 |
| chr2_+_42191592 | 6.61 |
ENSDART00000144716
|
cavin4a
|
caveolae associated protein 4a |
| chr18_+_1615 | 6.48 |
ENSDART00000082450
|
homer2
|
homer scaffolding protein 2 |
| chr12_-_26423439 | 5.96 |
ENSDART00000113978
|
synpo2lb
|
synaptopodin 2-like b |
| chr15_+_24644251 | 5.78 |
ENSDART00000181660
|
smtnl
|
smoothelin, like |
| chr15_+_24644016 | 5.75 |
ENSDART00000043292
|
smtnl
|
smoothelin, like |
| chr11_+_36243774 | 5.23 |
ENSDART00000023323
|
zgc:172270
|
zgc:172270 |
| chr1_+_58182400 | 5.07 |
ENSDART00000144784
|
si:ch211-15j1.1
|
si:ch211-15j1.1 |
| chr10_+_35417099 | 5.00 |
ENSDART00000063398
|
hhla2a.1
|
HERV-H LTR-associating 2a, tandem duplicate 1 |
| chr11_+_77526 | 4.87 |
ENSDART00000193521
|
CABZ01072242.1
|
|
| chr19_-_5369486 | 4.61 |
ENSDART00000105004
|
krt17
|
keratin 17 |
| chr23_-_39636195 | 4.52 |
ENSDART00000144439
|
vwa1
|
von Willebrand factor A domain containing 1 |
| chr8_+_50534948 | 4.47 |
ENSDART00000174435
|
PEBP4
|
phosphatidylethanolamine binding protein 4 |
| chr23_-_35069805 | 4.38 |
ENSDART00000087219
|
BX294434.1
|
|
| chr20_+_15015557 | 4.21 |
ENSDART00000039345
|
myoc
|
myocilin |
| chr1_-_58868306 | 3.90 |
ENSDART00000166615
|
dnm2b
|
dynamin 2b |
| chr6_+_27339962 | 3.88 |
ENSDART00000193726
|
klhl30
|
kelch-like family member 30 |
| chr19_-_10771558 | 3.60 |
ENSDART00000085165
|
slc9a3.2
|
solute carrier family 9, subfamily A (NHE3, cation proton antiporter 3), member 3, tandem duplicate 2 |
| chr25_+_3306620 | 3.55 |
ENSDART00000182085
ENSDART00000034704 |
slc25a3b
|
solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 3b |
| chr10_+_42374770 | 3.49 |
ENSDART00000020000
|
COX5B
|
zgc:86599 |
| chr3_-_34561624 | 3.13 |
ENSDART00000129313
|
sept9a
|
septin 9a |
| chr4_+_77993103 | 2.86 |
ENSDART00000174392
|
terfa
|
telomeric repeat binding factor a |
| chr22_-_15587360 | 2.67 |
ENSDART00000142717
ENSDART00000138978 |
tpm4a
|
tropomyosin 4a |
| chr10_+_42374957 | 2.67 |
ENSDART00000147926
|
COX5B
|
zgc:86599 |
| chr19_-_25519310 | 2.67 |
ENSDART00000089882
|
C1GALT1 (1 of many)
|
si:dkey-202e17.1 |
| chr6_+_60036767 | 2.62 |
ENSDART00000155009
|
tgm8
|
transglutaminase 8 |
| chr13_+_28819768 | 2.59 |
ENSDART00000191401
ENSDART00000188895 ENSDART00000101653 |
CU639469.1
|
|
| chr8_-_979735 | 2.59 |
ENSDART00000149612
|
znf366
|
zinc finger protein 366 |
| chr19_+_2631565 | 2.55 |
ENSDART00000171487
|
fam126a
|
family with sequence similarity 126, member A |
| chr19_-_25519612 | 2.51 |
ENSDART00000133150
|
C1GALT1 (1 of many)
|
si:dkey-202e17.1 |
| chr11_+_45343245 | 2.49 |
ENSDART00000160904
|
si:ch73-100l22.3
|
si:ch73-100l22.3 |
| chr17_+_53424415 | 2.37 |
ENSDART00000157022
|
slc9a1b
|
solute carrier family 9 member A1b |
| chr12_+_49125510 | 2.31 |
ENSDART00000185804
|
FO704607.1
|
|
| chr3_+_1179601 | 2.27 |
ENSDART00000173378
|
triobpb
|
TRIO and F-actin binding protein b |
| chr13_-_303137 | 2.22 |
ENSDART00000099131
|
chs1
|
chitin synthase 1 |
| chr11_+_24314148 | 2.19 |
ENSDART00000171491
|
rem1
|
RAS (RAD and GEM)-like GTP-binding 1 |
| chr1_-_51720633 | 2.15 |
ENSDART00000045894
|
rnaseh2a
|
ribonuclease H2, subunit A |
| chr13_-_7575216 | 2.13 |
ENSDART00000159443
|
pitx3
|
paired-like homeodomain 3 |
| chr12_+_19138452 | 2.07 |
ENSDART00000141346
ENSDART00000066397 |
phf5a
|
PHD finger protein 5A |
| chr10_+_20128267 | 2.04 |
ENSDART00000064615
|
dmtn
|
dematin actin binding protein |
| chr4_+_77993977 | 1.95 |
ENSDART00000174118
|
terfa
|
telomeric repeat binding factor a |
| chr14_+_989733 | 1.92 |
ENSDART00000161487
ENSDART00000127317 |
si:ch73-308l14.2
|
si:ch73-308l14.2 |
| chr16_+_30002605 | 1.90 |
ENSDART00000160555
|
sema6e
|
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6E |
| chr24_-_21674950 | 1.89 |
ENSDART00000123216
ENSDART00000046211 |
lnx2a
|
ligand of numb-protein X 2a |
| chr23_-_45504991 | 1.87 |
ENSDART00000148761
|
col24a1
|
collagen type XXIV alpha 1 |
| chr17_+_52524452 | 1.82 |
ENSDART00000016614
|
dlst
|
dihydrolipoamide S-succinyltransferase |
| chr1_+_36612660 | 1.81 |
ENSDART00000190784
|
ednraa
|
endothelin receptor type Aa |
| chr24_-_1021318 | 1.80 |
ENSDART00000181403
|
ralaa
|
v-ral simian leukemia viral oncogene homolog Aa (ras related) |
| chr11_+_24313931 | 1.79 |
ENSDART00000017599
ENSDART00000166045 |
rem1
|
RAS (RAD and GEM)-like GTP-binding 1 |
| chr4_-_965267 | 1.79 |
ENSDART00000093289
|
atp23
|
ATP23 metallopeptidase and ATP synthase assembly factor homolog |
| chr16_-_38518270 | 1.76 |
ENSDART00000131899
|
rspo2
|
R-spondin 2 |
| chr17_-_47305034 | 1.74 |
ENSDART00000150116
|
alk
|
ALK receptor tyrosine kinase |
| chr25_+_19008497 | 1.72 |
ENSDART00000104420
|
samm50
|
SAMM50 sorting and assembly machinery component |
| chr7_+_65586624 | 1.70 |
ENSDART00000184344
|
mical2a
|
microtubule associated monooxygenase, calponin and LIM domain containing 2a |
| chr16_+_25809235 | 1.70 |
ENSDART00000181196
|
irgq1
|
immunity-related GTPase family, q1 |
| chr4_-_2975461 | 1.67 |
ENSDART00000150794
|
plekha5
|
pleckstrin homology domain containing, family A member 5 |
| chr2_-_56649883 | 1.66 |
ENSDART00000191786
|
gpx4b
|
glutathione peroxidase 4b |
| chr9_-_14504834 | 1.65 |
ENSDART00000056103
|
nrp2b
|
neuropilin 2b |
| chr5_+_21211135 | 1.61 |
ENSDART00000088492
|
bmp10
|
bone morphogenetic protein 10 |
| chr7_-_26571994 | 1.55 |
ENSDART00000128801
|
si:dkey-62k3.6
|
si:dkey-62k3.6 |
| chr23_-_45705525 | 1.55 |
ENSDART00000148959
|
ednrab
|
endothelin receptor type Ab |
| chr4_-_4387012 | 1.53 |
ENSDART00000191836
|
CU468826.3
|
Danio rerio U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit-related protein 1-like (LOC100331497), mRNA. |
| chr1_-_44314 | 1.50 |
ENSDART00000160050
|
tmem39a
|
transmembrane protein 39A |
| chr17_-_49995366 | 1.50 |
ENSDART00000075223
|
cox7a2a
|
cytochrome c oxidase subunit VIIa polypeptide 2a (liver) |
| chr23_+_27912079 | 1.47 |
ENSDART00000171859
|
BX539336.1
|
|
| chr8_+_21159122 | 1.44 |
ENSDART00000033491
|
spryd4
|
SPRY domain containing 4 |
| chr13_+_713941 | 1.44 |
ENSDART00000109737
|
cox15
|
cytochrome c oxidase assembly homolog 15 (yeast) |
| chr4_-_16451375 | 1.43 |
ENSDART00000192700
ENSDART00000128835 |
wu:fc23c09
|
wu:fc23c09 |
| chr1_+_58303892 | 1.40 |
ENSDART00000147678
|
CR769768.1
|
|
| chr5_-_26330313 | 1.32 |
ENSDART00000148656
|
arvcfb
|
ARVCF, delta catenin family member b |
| chr11_+_25101220 | 1.32 |
ENSDART00000183700
|
ndrg3a
|
ndrg family member 3a |
| chr8_+_8893535 | 1.31 |
ENSDART00000139150
|
otud5a
|
OTU deubiquitinase 5a |
| chr1_+_58119117 | 1.31 |
ENSDART00000146788
|
si:ch211-15j1.5
|
si:ch211-15j1.5 |
| chr1_+_58067815 | 1.25 |
ENSDART00000156678
|
si:ch211-114l13.4
|
si:ch211-114l13.4 |
| chr12_+_23866368 | 1.24 |
ENSDART00000188652
ENSDART00000192478 |
svila
|
supervillin a |
| chr7_-_2090594 | 1.20 |
ENSDART00000183166
|
si:cabz01007802.1
|
si:cabz01007802.1 |
| chr3_-_36419641 | 1.19 |
ENSDART00000173545
|
cog1
|
component of oligomeric golgi complex 1 |
| chr23_+_39346774 | 1.19 |
ENSDART00000190985
|
src
|
v-src avian sarcoma (Schmidt-Ruppin A-2) viral oncogene homolog |
| chr10_-_38443312 | 1.19 |
ENSDART00000141260
|
gdpd5a
|
glycerophosphodiester phosphodiesterase domain containing 5a |
| chr18_+_35742838 | 1.18 |
ENSDART00000088504
ENSDART00000140386 |
rasgrp4
|
RAS guanyl releasing protein 4 |
| chr1_+_58260886 | 1.11 |
ENSDART00000110453
|
si:dkey-222h21.8
|
si:dkey-222h21.8 |
| chr1_+_58150000 | 1.07 |
ENSDART00000137836
|
si:ch211-15j1.3
|
si:ch211-15j1.3 |
| chr6_-_40583253 | 1.07 |
ENSDART00000032603
|
tspo
|
translocator protein |
| chr8_+_2530065 | 1.04 |
ENSDART00000063943
|
mrpl40
|
mitochondrial ribosomal protein L40 |
| chr3_+_27770110 | 1.03 |
ENSDART00000017962
|
eci1
|
enoyl-CoA delta isomerase 1 |
| chr18_-_43866526 | 1.03 |
ENSDART00000111309
|
treh
|
trehalase (brush-border membrane glycoprotein) |
| chr8_-_12468744 | 1.03 |
ENSDART00000135019
|
FIBCD1 (1 of many)
|
si:dkeyp-51b7.3 |
| chr3_+_32671146 | 0.97 |
ENSDART00000039466
|
rnf25
|
ring finger protein 25 |
| chr10_-_41400049 | 0.92 |
ENSDART00000009838
|
gpat4
|
glycerol-3-phosphate acyltransferase 4 |
| chr22_-_38543630 | 0.89 |
ENSDART00000172029
|
si:ch211-126j24.1
|
si:ch211-126j24.1 |
| chr6_+_8176486 | 0.88 |
ENSDART00000193308
|
nfil3-5
|
nuclear factor, interleukin 3 regulated, member 5 |
| chr3_+_34159192 | 0.80 |
ENSDART00000151119
|
carm1
|
coactivator-associated arginine methyltransferase 1 |
| chr15_+_19797918 | 0.77 |
ENSDART00000113314
|
si:ch211-229d2.5
|
si:ch211-229d2.5 |
| chr13_+_45980163 | 0.77 |
ENSDART00000074547
ENSDART00000005195 |
bsdc1
|
BSD domain containing 1 |
| chr21_+_38089036 | 0.77 |
ENSDART00000147219
|
klf8
|
Kruppel-like factor 8 |
| chr19_+_7595924 | 0.77 |
ENSDART00000140083
|
s100s
|
S100 calcium binding protein S |
| chr7_-_51793333 | 0.76 |
ENSDART00000180654
|
BX957362.5
|
|
| chr6_+_40952031 | 0.75 |
ENSDART00000189219
|
patz1
|
POZ (BTB) and AT hook containing zinc finger 1 |
| chr20_+_36806398 | 0.72 |
ENSDART00000153317
|
abracl
|
ABRA C-terminal like |
| chr20_-_26936887 | 0.72 |
ENSDART00000160827
|
ftr79
|
finTRIM family, member 79 |
| chr7_-_51032128 | 0.63 |
ENSDART00000182781
ENSDART00000121574 |
col4a6
|
collagen, type IV, alpha 6 |
| chr20_+_43379029 | 0.63 |
ENSDART00000142486
ENSDART00000186486 |
unc93a
|
unc-93 homolog A |
| chr20_-_29499363 | 0.63 |
ENSDART00000152889
ENSDART00000153252 ENSDART00000170972 ENSDART00000166420 ENSDART00000163079 |
rrm2
|
ribonucleotide reductase M2 polypeptide |
| chr23_+_22597624 | 0.60 |
ENSDART00000054337
|
gpr157
|
G protein-coupled receptor 157 |
| chr16_-_39131666 | 0.55 |
ENSDART00000075517
|
gdf6a
|
growth differentiation factor 6a |
| chr17_+_654759 | 0.54 |
ENSDART00000193703
|
CABZ01079748.1
|
|
| chr1_+_59328030 | 0.54 |
ENSDART00000172464
|
CABZ01052576.1
|
|
| chr16_-_1503023 | 0.53 |
ENSDART00000036348
|
sim1a
|
single-minded family bHLH transcription factor 1a |
| chr23_-_22598002 | 0.53 |
ENSDART00000146966
|
si:ch211-28e16.5
|
si:ch211-28e16.5 |
| chr25_+_27410352 | 0.51 |
ENSDART00000154362
|
pot1
|
protection of telomeres 1 homolog |
| chr25_+_12012 | 0.51 |
ENSDART00000170348
|
cntn1a
|
contactin 1a |
| chr15_-_1844048 | 0.49 |
ENSDART00000102410
|
taf15
|
TAF15 RNA polymerase II, TATA box binding protein (TBP)-associated factor |
| chr19_-_47571456 | 0.48 |
ENSDART00000158071
ENSDART00000165841 |
rrm2
|
ribonucleotide reductase M2 polypeptide |
| chr3_+_40289418 | 0.48 |
ENSDART00000017304
|
cpsf4
|
cleavage and polyadenylation specific factor 4 |
| chr10_-_322769 | 0.46 |
ENSDART00000165244
|
akt2l
|
v-akt murine thymoma viral oncogene homolog 2, like |
| chr1_+_58442694 | 0.46 |
ENSDART00000160897
|
zgc:194906
|
zgc:194906 |
| chr13_-_31878043 | 0.43 |
ENSDART00000187720
|
syt14a
|
synaptotagmin XIVa |
| chr1_+_44582369 | 0.43 |
ENSDART00000003022
ENSDART00000137980 |
med19b
|
mediator complex subunit 19b |
| chr3_+_62216727 | 0.43 |
ENSDART00000180069
|
BX470259.4
|
|
| chr16_-_25680666 | 0.42 |
ENSDART00000132693
ENSDART00000140539 ENSDART00000015302 |
tomm40
|
translocase of outer mitochondrial membrane 40 homolog (yeast) |
| chr22_+_2417105 | 0.42 |
ENSDART00000106415
|
zgc:113220
|
zgc:113220 |
| chr14_+_30413312 | 0.42 |
ENSDART00000186864
|
cnot7
|
CCR4-NOT transcription complex, subunit 7 |
| chr21_-_45728069 | 0.41 |
ENSDART00000185101
|
CU302316.1
|
|
| chr11_+_30729745 | 0.41 |
ENSDART00000103270
|
slc22a7a
|
solute carrier family 22 (organic anion transporter), member 7a |
| chr25_+_21054065 | 0.41 |
ENSDART00000148472
|
chrm2b
|
cholinergic receptor, muscarinic 2b |
| chr23_-_46126444 | 0.40 |
ENSDART00000030004
|
CABZ01071903.1
|
|
| chr23_+_39346930 | 0.39 |
ENSDART00000102843
|
src
|
v-src avian sarcoma (Schmidt-Ruppin A-2) viral oncogene homolog |
| chr15_+_1004680 | 0.37 |
ENSDART00000157310
|
si:dkey-77f5.8
|
si:dkey-77f5.8 |
| chr5_-_16472719 | 0.37 |
ENSDART00000162071
|
piwil2
|
piwi-like RNA-mediated gene silencing 2 |
| chr2_-_21618629 | 0.37 |
ENSDART00000027587
|
ralab
|
v-ral simian leukemia viral oncogene homolog Ab (ras related) |
| chr21_+_13245302 | 0.36 |
ENSDART00000189498
|
specc1lb
|
sperm antigen with calponin homology and coiled-coil domains 1-like b |
| chr5_-_66301142 | 0.36 |
ENSDART00000067541
|
prlrb
|
prolactin receptor b |
| chr12_-_44199316 | 0.35 |
ENSDART00000170378
|
si:ch73-329n5.1
|
si:ch73-329n5.1 |
| chr22_-_3595439 | 0.33 |
ENSDART00000083308
|
ptprsa
|
protein tyrosine phosphatase, receptor type, s, a |
| chr10_-_40939706 | 0.32 |
ENSDART00000059795
ENSDART00000190510 |
bmp1b
|
bone morphogenetic protein 1b |
| chr6_+_40523370 | 0.32 |
ENSDART00000033819
|
prkcda
|
protein kinase C, delta a |
| chr5_-_23280098 | 0.31 |
ENSDART00000126540
ENSDART00000051533 |
plp1b
|
proteolipid protein 1b |
| chr2_-_53925308 | 0.31 |
ENSDART00000190705
|
FO834855.1
|
|
| chr7_-_46744080 | 0.30 |
ENSDART00000193328
|
tshz3b
|
teashirt zinc finger homeobox 3b |
| chr1_+_24387659 | 0.30 |
ENSDART00000130356
|
qdprb2
|
quinoid dihydropteridine reductase b2 |
| chr7_+_7495034 | 0.29 |
ENSDART00000102629
ENSDART00000180475 |
nit1
|
nitrilase 1 |
| chr20_-_25877793 | 0.29 |
ENSDART00000177333
ENSDART00000184619 |
exoc1
|
exocyst complex component 1 |
| chr3_-_30609659 | 0.28 |
ENSDART00000182516
ENSDART00000187047 ENSDART00000110597 |
syt3
|
synaptotagmin III |
| chr7_-_20756013 | 0.28 |
ENSDART00000185259
|
chd3
|
chromodomain helicase DNA binding protein 3 |
| chr12_+_33433882 | 0.28 |
ENSDART00000185053
|
fasn
|
fatty acid synthase |
| chr5_-_68927728 | 0.28 |
ENSDART00000132838
|
ank1a
|
ankyrin 1, erythrocytic a |
| chr18_-_43866001 | 0.27 |
ENSDART00000150218
|
treh
|
trehalase (brush-border membrane glycoprotein) |
| chr24_+_39641991 | 0.26 |
ENSDART00000142182
|
luc7l
|
LUC7-like (S. cerevisiae) |
| chr13_+_42406883 | 0.26 |
ENSDART00000084354
|
cpeb3
|
cytoplasmic polyadenylation element binding protein 3 |
| chr4_-_77300601 | 0.25 |
ENSDART00000163015
ENSDART00000174015 |
slco1f2
|
solute carrier organic anion transporter family, member 1F2 |
| chr8_+_10920205 | 0.25 |
ENSDART00000179742
|
BX321886.1
|
|
| chr5_+_29851433 | 0.25 |
ENSDART00000143434
|
ubash3ba
|
ubiquitin associated and SH3 domain containing Ba |
| chr3_+_18062871 | 0.25 |
ENSDART00000112384
|
ankrd40
|
ankyrin repeat domain 40 |
| chr19_-_27564980 | 0.24 |
ENSDART00000171967
|
si:dkeyp-46h3.8
|
si:dkeyp-46h3.8 |
| chr9_+_40546677 | 0.22 |
ENSDART00000132911
|
vwc2l
|
von Willebrand factor C domain containing protein 2-like |
| chr12_+_48681601 | 0.22 |
ENSDART00000187831
|
uros
|
uroporphyrinogen III synthase |
| chr1_+_58221372 | 0.21 |
ENSDART00000138467
|
si:dkey-222h21.10
|
si:dkey-222h21.10 |
| chr12_+_27096835 | 0.21 |
ENSDART00000149475
|
ttll6
|
tubulin tyrosine ligase-like family, member 6 |
| chr8_+_32742650 | 0.20 |
ENSDART00000138117
|
hmcn2
|
hemicentin 2 |
| chr2_+_43755274 | 0.15 |
ENSDART00000138620
|
arhgap12a
|
Rho GTPase activating protein 12a |
| chr16_-_2558653 | 0.15 |
ENSDART00000110365
|
adcy3a
|
adenylate cyclase 3a |
| chr24_+_22022109 | 0.14 |
ENSDART00000133686
|
ropn1l
|
rhophilin associated tail protein 1-like |
| chr19_-_3193443 | 0.13 |
ENSDART00000179855
|
si:ch211-133n4.6
|
si:ch211-133n4.6 |
| chr7_+_73332486 | 0.13 |
ENSDART00000174119
ENSDART00000092051 ENSDART00000192388 |
CABZ01081780.1
|
|
| chr8_+_8699085 | 0.12 |
ENSDART00000021209
|
uxt
|
ubiquitously-expressed, prefoldin-like chaperone |
| chr8_+_37111643 | 0.11 |
ENSDART00000061336
|
ren
|
renin |
| chr2_-_38141997 | 0.11 |
ENSDART00000143469
|
trim110
|
tripartite motif containing 110 |
| chr23_-_2448234 | 0.11 |
ENSDART00000082097
|
LO018513.1
|
|
| chr10_+_29266392 | 0.09 |
ENSDART00000153577
|
crebzf
|
CREB/ATF bZIP transcription factor |
| chr2_+_32016516 | 0.09 |
ENSDART00000135040
|
mycb
|
MYC proto-oncogene, bHLH transcription factor b |
| chr5_+_19479200 | 0.08 |
ENSDART00000137703
|
aldh3b2
|
aldehyde dehydrogenase 3 family, member B2 |
| chr2_-_51388214 | 0.07 |
ENSDART00000163838
|
si:dkeyp-104b3.18
|
si:dkeyp-104b3.18 |
| chr19_-_27564458 | 0.06 |
ENSDART00000123155
|
si:dkeyp-46h3.6
|
si:dkeyp-46h3.6 |
| chr18_-_3456117 | 0.05 |
ENSDART00000158508
|
myo7aa
|
myosin VIIAa |
| chr15_+_21711671 | 0.05 |
ENSDART00000136151
|
NKAPD1
|
zgc:162339 |
| chr5_-_22615087 | 0.05 |
ENSDART00000146035
|
zgc:113208
|
zgc:113208 |
| chr21_+_15883546 | 0.05 |
ENSDART00000186325
|
fam169ab
|
family with sequence similarity 169, member Ab |
| chr21_-_43171249 | 0.04 |
ENSDART00000021037
ENSDART00000148630 |
hspa4a
|
heat shock protein 4a |
| chr23_+_39970425 | 0.03 |
ENSDART00000165376
|
fyco1a
|
FYVE and coiled-coil domain containing 1a |
| chr10_+_36354863 | 0.03 |
ENSDART00000159675
|
or104-1
|
odorant receptor, family C, subfamily 104, member 1 |
| chr16_-_560574 | 0.03 |
ENSDART00000148452
|
irx2a
|
iroquois homeobox 2a |
| chr10_-_8197049 | 0.02 |
ENSDART00000129467
|
dhx29
|
DEAH (Asp-Glu-Ala-His) box polypeptide 29 |
| chr17_+_1514711 | 0.01 |
ENSDART00000165641
|
akt1
|
v-akt murine thymoma viral oncogene homolog 1 |
| chr15_+_21672700 | 0.00 |
ENSDART00000187043
|
si:dkey-40g16.5
|
si:dkey-40g16.5 |
| chr3_+_54047342 | 0.00 |
ENSDART00000178486
|
olfm2a
|
olfactomedin 2a |
Network of associatons between targets according to the STRING database.
First level regulatory network of nr2c1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.7 | 8.0 | GO:0014809 | regulation of skeletal muscle contraction by calcium ion signaling(GO:0014722) regulation of skeletal muscle contraction by regulation of release of sequestered calcium ion(GO:0014809) |
| 1.2 | 4.8 | GO:0032207 | regulation of telomere maintenance via recombination(GO:0032207) negative regulation of telomere maintenance via recombination(GO:0032208) |
| 0.8 | 6.5 | GO:0021627 | olfactory nerve development(GO:0021553) olfactory nerve morphogenesis(GO:0021627) olfactory nerve formation(GO:0021628) |
| 0.8 | 4.0 | GO:1901841 | regulation of high voltage-gated calcium channel activity(GO:1901841) negative regulation of high voltage-gated calcium channel activity(GO:1901842) |
| 0.6 | 2.5 | GO:1903723 | negative regulation of centriole elongation(GO:1903723) |
| 0.6 | 1.8 | GO:0033512 | L-lysine catabolic process to acetyl-CoA via saccharopine(GO:0033512) |
| 0.6 | 1.8 | GO:0033615 | mitochondrial proton-transporting ATP synthase complex assembly(GO:0033615) |
| 0.6 | 1.8 | GO:0033335 | anal fin development(GO:0033335) |
| 0.6 | 1.7 | GO:0045040 | outer mitochondrial membrane organization(GO:0007008) protein import into mitochondrial outer membrane(GO:0045040) |
| 0.5 | 6.0 | GO:0097369 | sodium ion import(GO:0097369) sodium ion import across plasma membrane(GO:0098719) sodium ion import into cell(GO:1990118) |
| 0.5 | 1.6 | GO:0050847 | progesterone receptor signaling pathway(GO:0050847) |
| 0.5 | 5.2 | GO:0001765 | membrane raft assembly(GO:0001765) plasma membrane raft assembly(GO:0044854) plasma membrane raft organization(GO:0044857) caveola assembly(GO:0070836) |
| 0.5 | 5.0 | GO:0031294 | lymphocyte costimulation(GO:0031294) T cell costimulation(GO:0031295) |
| 0.4 | 2.2 | GO:1901073 | chitin biosynthetic process(GO:0006031) glucosamine-containing compound biosynthetic process(GO:1901073) |
| 0.4 | 1.3 | GO:0005991 | trehalose metabolic process(GO:0005991) |
| 0.4 | 2.5 | GO:0038063 | collagen-activated tyrosine kinase receptor signaling pathway(GO:0038063) collagen-activated signaling pathway(GO:0038065) |
| 0.4 | 2.2 | GO:0043137 | DNA replication, removal of RNA primer(GO:0043137) |
| 0.3 | 1.8 | GO:0035176 | social behavior(GO:0035176) intraspecies interaction between organisms(GO:0051703) |
| 0.3 | 10.2 | GO:0032231 | regulation of actin filament bundle assembly(GO:0032231) |
| 0.3 | 1.7 | GO:0010735 | positive regulation of transcription via serum response element binding(GO:0010735) |
| 0.3 | 2.3 | GO:1900026 | positive regulation of substrate adhesion-dependent cell spreading(GO:1900026) |
| 0.2 | 1.5 | GO:0097250 | mitochondrial respiratory chain supercomplex assembly(GO:0097250) |
| 0.2 | 1.5 | GO:0003319 | cardioblast migration to the midline involved in heart rudiment formation(GO:0003319) |
| 0.2 | 3.9 | GO:0098884 | postsynaptic neurotransmitter receptor internalization(GO:0098884) |
| 0.2 | 0.8 | GO:0045600 | positive regulation of fat cell differentiation(GO:0045600) |
| 0.2 | 1.4 | GO:1901097 | negative regulation of autophagosome maturation(GO:1901097) |
| 0.2 | 0.5 | GO:0072020 | proximal straight tubule development(GO:0072020) |
| 0.2 | 3.5 | GO:0035435 | phosphate ion transmembrane transport(GO:0035435) |
| 0.2 | 1.7 | GO:0071678 | olfactory bulb axon guidance(GO:0071678) |
| 0.1 | 1.2 | GO:0071684 | hatching(GO:0035188) organism emergence from protective structure(GO:0071684) |
| 0.1 | 0.6 | GO:0042706 | retinal cone cell fate determination(GO:0042671) eye photoreceptor cell fate commitment(GO:0042706) photoreceptor cell fate determination(GO:0043703) retinal cone cell fate commitment(GO:0046551) photoreceptor cell fate commitment(GO:0046552) camera-type eye photoreceptor cell fate commitment(GO:0060220) |
| 0.1 | 0.4 | GO:0097095 | frontonasal suture morphogenesis(GO:0097095) |
| 0.1 | 2.6 | GO:0018149 | peptide cross-linking(GO:0018149) |
| 0.1 | 0.5 | GO:0051972 | regulation of telomerase activity(GO:0051972) |
| 0.1 | 1.1 | GO:0009263 | deoxyribonucleotide biosynthetic process(GO:0009263) |
| 0.1 | 0.4 | GO:0030718 | germ-line stem cell population maintenance(GO:0030718) |
| 0.1 | 2.1 | GO:0021984 | adenohypophysis development(GO:0021984) |
| 0.1 | 0.4 | GO:0038161 | prolactin signaling pathway(GO:0038161) |
| 0.1 | 2.7 | GO:0055003 | cardiac myofibril assembly(GO:0055003) |
| 0.1 | 2.0 | GO:0030032 | lamellipodium assembly(GO:0030032) |
| 0.1 | 6.6 | GO:0035023 | regulation of Rho protein signal transduction(GO:0035023) |
| 0.1 | 0.8 | GO:0021702 | cerebellar Purkinje cell layer formation(GO:0021694) cerebellar Purkinje cell differentiation(GO:0021702) |
| 0.1 | 1.5 | GO:0017121 | phospholipid scrambling(GO:0017121) |
| 0.1 | 1.7 | GO:0006783 | heme biosynthetic process(GO:0006783) |
| 0.1 | 1.3 | GO:0071108 | protein K48-linked deubiquitination(GO:0071108) |
| 0.1 | 0.5 | GO:0098787 | mRNA cleavage involved in mRNA processing(GO:0098787) pre-mRNA cleavage required for polyadenylation(GO:0098789) |
| 0.1 | 7.1 | GO:0006821 | chloride transport(GO:0006821) |
| 0.0 | 8.7 | GO:0006936 | muscle contraction(GO:0006936) |
| 0.0 | 4.6 | GO:0031101 | fin regeneration(GO:0031101) |
| 0.0 | 1.2 | GO:0051014 | actin filament severing(GO:0051014) |
| 0.0 | 1.6 | GO:0060840 | artery development(GO:0060840) |
| 0.0 | 0.4 | GO:0043928 | exonucleolytic nuclear-transcribed mRNA catabolic process involved in deadenylation-dependent decay(GO:0043928) |
| 0.0 | 1.3 | GO:0016339 | calcium-dependent cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0016339) |
| 0.0 | 1.7 | GO:0000187 | activation of MAPK activity(GO:0000187) |
| 0.0 | 1.1 | GO:0060319 | primitive erythrocyte differentiation(GO:0060319) |
| 0.0 | 1.2 | GO:0006891 | intra-Golgi vesicle-mediated transport(GO:0006891) |
| 0.0 | 2.9 | GO:0042552 | myelination(GO:0042552) |
| 0.0 | 3.1 | GO:0061640 | cytoskeleton-dependent cytokinesis(GO:0061640) |
| 0.0 | 1.0 | GO:0051092 | positive regulation of NF-kappaB transcription factor activity(GO:0051092) |
| 0.0 | 1.9 | GO:0048843 | negative regulation of axon extension involved in axon guidance(GO:0048843) negative regulation of axon guidance(GO:1902668) |
| 0.0 | 0.3 | GO:0046146 | tetrahydrobiopterin biosynthetic process(GO:0006729) tetrahydrobiopterin metabolic process(GO:0046146) |
| 0.0 | 0.2 | GO:0043589 | skin morphogenesis(GO:0043589) mesenchymal cell migration(GO:0090497) |
| 0.0 | 0.3 | GO:0021772 | olfactory bulb development(GO:0021772) olfactory lobe development(GO:0021988) |
| 0.0 | 0.4 | GO:0030150 | protein import into mitochondrial matrix(GO:0030150) |
| 0.0 | 0.3 | GO:2000766 | negative regulation of cytoplasmic translation(GO:2000766) |
| 0.0 | 0.9 | GO:0032922 | circadian regulation of gene expression(GO:0032922) |
| 0.0 | 0.1 | GO:0048240 | sperm capacitation(GO:0048240) |
| 0.0 | 2.5 | GO:0017157 | regulation of exocytosis(GO:0017157) |
| 0.0 | 1.2 | GO:0001945 | lymph vessel development(GO:0001945) |
| 0.0 | 0.5 | GO:0006635 | fatty acid beta-oxidation(GO:0006635) |
| 0.0 | 1.7 | GO:0030178 | negative regulation of Wnt signaling pathway(GO:0030178) |
| 0.0 | 0.3 | GO:0043252 | sodium-independent organic anion transport(GO:0043252) |
| 0.0 | 3.2 | GO:0007005 | mitochondrion organization(GO:0007005) |
| 0.0 | 0.2 | GO:0018095 | protein polyglutamylation(GO:0018095) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 8.0 | GO:0033018 | sarcoplasmic reticulum lumen(GO:0033018) |
| 0.6 | 1.9 | GO:0005592 | collagen type XI trimer(GO:0005592) |
| 0.5 | 2.2 | GO:0032299 | ribonuclease H2 complex(GO:0032299) |
| 0.4 | 8.8 | GO:0031430 | M band(GO:0031430) |
| 0.4 | 3.6 | GO:0031526 | brush border membrane(GO:0031526) |
| 0.3 | 4.8 | GO:0070187 | telosome(GO:0070187) |
| 0.3 | 2.2 | GO:0030428 | cell septum(GO:0030428) |
| 0.3 | 11.8 | GO:0044853 | caveola(GO:0005901) plasma membrane raft(GO:0044853) |
| 0.2 | 1.7 | GO:0001401 | mitochondrial sorting and assembly machinery complex(GO:0001401) |
| 0.2 | 3.9 | GO:0098844 | postsynaptic endocytic zone membrane(GO:0098844) |
| 0.2 | 1.8 | GO:0031314 | extrinsic component of mitochondrial inner membrane(GO:0031314) |
| 0.2 | 1.8 | GO:0045252 | oxoglutarate dehydrogenase complex(GO:0045252) |
| 0.2 | 0.6 | GO:0098645 | collagen type IV trimer(GO:0005587) network-forming collagen trimer(GO:0098642) collagen network(GO:0098645) basement membrane collagen trimer(GO:0098651) |
| 0.1 | 4.2 | GO:0005791 | rough endoplasmic reticulum(GO:0005791) |
| 0.1 | 7.1 | GO:0034707 | chloride channel complex(GO:0034707) |
| 0.1 | 1.2 | GO:0017119 | Golgi transport complex(GO:0017119) |
| 0.1 | 2.1 | GO:0005686 | U2 snRNP(GO:0005686) U12-type spliceosomal complex(GO:0005689) |
| 0.1 | 3.1 | GO:0032156 | septin ring(GO:0005940) septin complex(GO:0031105) septin cytoskeleton(GO:0032156) |
| 0.1 | 0.4 | GO:1990923 | PET complex(GO:1990923) |
| 0.1 | 0.5 | GO:0000783 | telomere cap complex(GO:0000782) nuclear telomere cap complex(GO:0000783) nuclear chromosome, telomeric region(GO:0000784) |
| 0.1 | 6.0 | GO:0030018 | Z disc(GO:0030018) |
| 0.1 | 3.1 | GO:0031305 | integral component of mitochondrial inner membrane(GO:0031305) |
| 0.0 | 0.3 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.0 | 4.1 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 0.4 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.0 | 0.4 | GO:0030015 | CCR4-NOT core complex(GO:0030015) |
| 0.0 | 2.5 | GO:0005814 | centriole(GO:0005814) |
| 0.0 | 3.0 | GO:0005884 | actin filament(GO:0005884) |
| 0.0 | 1.0 | GO:0005762 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.0 | 0.3 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.0 | 1.5 | GO:0005746 | mitochondrial respiratory chain(GO:0005746) |
| 0.0 | 0.3 | GO:0005685 | U1 snRNP(GO:0005685) |
| 0.0 | 0.3 | GO:1990124 | messenger ribonucleoprotein complex(GO:1990124) |
| 0.0 | 1.6 | GO:0005924 | cell-substrate adherens junction(GO:0005924) focal adhesion(GO:0005925) |
| 0.0 | 4.2 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.0 | 0.1 | GO:0016272 | prefoldin complex(GO:0016272) |
| 0.0 | 1.1 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.0 | 4.1 | GO:0015629 | actin cytoskeleton(GO:0015629) |
| 0.0 | 0.3 | GO:0000145 | exocyst(GO:0000145) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 5.3 | GO:0098505 | G-rich strand telomeric DNA binding(GO:0098505) |
| 0.6 | 1.8 | GO:0016748 | succinyltransferase activity(GO:0016748) |
| 0.5 | 6.0 | GO:0015385 | monovalent cation:proton antiporter activity(GO:0005451) sodium:proton antiporter activity(GO:0015385) potassium:proton antiporter activity(GO:0015386) |
| 0.5 | 3.4 | GO:0004962 | endothelin receptor activity(GO:0004962) |
| 0.4 | 1.3 | GO:0004555 | alpha,alpha-trehalase activity(GO:0004555) trehalase activity(GO:0015927) |
| 0.3 | 1.7 | GO:0047066 | phospholipid-hydroperoxide glutathione peroxidase activity(GO:0047066) |
| 0.3 | 2.2 | GO:0004523 | RNA-DNA hybrid ribonuclease activity(GO:0004523) |
| 0.3 | 1.7 | GO:0043914 | NADPH:sulfur oxidoreductase activity(GO:0043914) |
| 0.3 | 2.2 | GO:0004100 | chitin synthase activity(GO:0004100) |
| 0.2 | 3.5 | GO:0005315 | inorganic phosphate transmembrane transporter activity(GO:0005315) |
| 0.2 | 1.7 | GO:0010314 | phosphatidylinositol-5-phosphate binding(GO:0010314) |
| 0.2 | 0.8 | GO:0008469 | histone-arginine N-methyltransferase activity(GO:0008469) |
| 0.2 | 1.1 | GO:0004748 | ribonucleoside-diphosphate reductase activity, thioredoxin disulfide as acceptor(GO:0004748) oxidoreductase activity, acting on CH or CH2 groups, disulfide as acceptor(GO:0016728) ribonucleoside-diphosphate reductase activity(GO:0061731) |
| 0.2 | 4.0 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.1 | 1.7 | GO:0005021 | vascular endothelial growth factor-activated receptor activity(GO:0005021) |
| 0.1 | 2.6 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 0.1 | 1.5 | GO:0017128 | phospholipid scramblase activity(GO:0017128) |
| 0.1 | 1.0 | GO:0004300 | enoyl-CoA hydratase activity(GO:0004300) |
| 0.1 | 1.4 | GO:0016653 | oxidoreductase activity, acting on NAD(P)H, heme protein as acceptor(GO:0016653) |
| 0.1 | 0.4 | GO:0004925 | prolactin receptor activity(GO:0004925) |
| 0.1 | 1.3 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.1 | 0.3 | GO:0004155 | 6,7-dihydropteridine reductase activity(GO:0004155) NADH binding(GO:0070404) |
| 0.1 | 7.1 | GO:0005254 | chloride channel activity(GO:0005254) |
| 0.1 | 2.2 | GO:0019003 | GDP binding(GO:0019003) |
| 0.1 | 1.5 | GO:0016676 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors(GO:0016675) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.0 | 0.4 | GO:0034584 | piRNA binding(GO:0034584) |
| 0.0 | 16.8 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.0 | 8.4 | GO:0060090 | binding, bridging(GO:0060090) |
| 0.0 | 13.2 | GO:0003779 | actin binding(GO:0003779) |
| 0.0 | 0.4 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 0.0 | 0.6 | GO:0030552 | cAMP binding(GO:0030552) |
| 0.0 | 0.8 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.0 | 1.9 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.0 | 0.4 | GO:0008320 | protein transmembrane transporter activity(GO:0008320) |
| 0.0 | 0.4 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.0 | 3.3 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 0.3 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.0 | 1.6 | GO:0070851 | growth factor receptor binding(GO:0070851) |
| 0.0 | 8.3 | GO:0016887 | ATPase activity(GO:0016887) |
| 0.0 | 0.9 | GO:0044325 | ion channel binding(GO:0044325) |
| 0.0 | 2.7 | GO:0003714 | transcription corepressor activity(GO:0003714) |
| 0.0 | 0.3 | GO:0000900 | translation repressor activity, nucleic acid binding(GO:0000900) |
| 0.0 | 0.2 | GO:0070739 | protein-glutamic acid ligase activity(GO:0070739) tubulin-glutamic acid ligase activity(GO:0070740) |
| 0.0 | 1.3 | GO:0045296 | cadherin binding(GO:0045296) |
| 0.0 | 3.4 | GO:0008017 | microtubule binding(GO:0008017) |
| 0.0 | 0.3 | GO:0015347 | sodium-independent organic anion transmembrane transporter activity(GO:0015347) |
| 0.0 | 0.3 | GO:0004697 | protein kinase C activity(GO:0004697) calcium-dependent protein kinase C activity(GO:0004698) |
| 0.0 | 1.5 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.6 | ST STAT3 PATHWAY | STAT3 Pathway |
| 0.1 | 2.5 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.0 | 3.6 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
| 0.0 | 1.2 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.0 | 6.5 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 0.8 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.0 | 0.8 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.0 | 1.2 | PID E2F PATHWAY | E2F transcription factor network |
| 0.0 | 0.8 | PID AR PATHWAY | Coregulation of Androgen receptor activity |
| 0.0 | 1.8 | NABA SECRETED FACTORS | Genes encoding secreted soluble factors |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.3 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 0.2 | 1.6 | REACTOME SIGNAL REGULATORY PROTEIN SIRP FAMILY INTERACTIONS | Genes involved in Signal regulatory protein (SIRP) family interactions |
| 0.1 | 1.8 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.1 | 6.2 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.1 | 1.0 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.1 | 1.1 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.1 | 2.5 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.1 | 2.1 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.0 | 0.5 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.0 | 0.4 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.0 | 0.3 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.0 | 2.1 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |
| 0.0 | 0.2 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.0 | 0.3 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |