Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
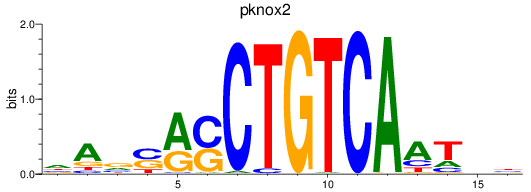
Results for pknox2
Z-value: 0.76
Transcription factors associated with pknox2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
pknox2
|
ENSDARG00000055349 | pbx/knotted 1 homeobox 2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| pknox2 | dr11_v1_chr10_-_31104983_31104983 | 0.65 | 9.1e-13 | Click! |
Activity profile of pknox2 motif
Sorted Z-values of pknox2 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr3_-_36260102 | 9.56 |
ENSDART00000126588
|
rac3a
|
Rac family small GTPase 3a |
| chr6_+_14949950 | 9.18 |
ENSDART00000149202
ENSDART00000149949 |
pou3f3b
|
POU class 3 homeobox 3b |
| chr11_-_32723851 | 8.55 |
ENSDART00000155592
|
pcdh17
|
protocadherin 17 |
| chr22_-_23253481 | 8.47 |
ENSDART00000054807
|
lhx9
|
LIM homeobox 9 |
| chr1_-_14234076 | 7.43 |
ENSDART00000040049
|
camk2d2
|
calcium/calmodulin-dependent protein kinase (CaM kinase) II delta 2 |
| chr6_+_38381957 | 7.34 |
ENSDART00000087300
|
gabrb3
|
gamma-aminobutyric acid (GABA) A receptor, beta 3 |
| chr19_-_9712530 | 6.89 |
ENSDART00000134816
|
slc2a3a
|
solute carrier family 2 (facilitated glucose transporter), member 3a |
| chr22_-_23253252 | 6.63 |
ENSDART00000175556
|
lhx9
|
LIM homeobox 9 |
| chr1_-_14233815 | 6.46 |
ENSDART00000044896
|
camk2d2
|
calcium/calmodulin-dependent protein kinase (CaM kinase) II delta 2 |
| chr9_+_6009077 | 6.43 |
ENSDART00000057484
|
st6gal2a
|
ST6 beta-galactosamide alpha-2,6-sialyltranferase 2a |
| chr8_-_33114202 | 6.36 |
ENSDART00000098840
|
ralgps1
|
Ral GEF with PH domain and SH3 binding motif 1 |
| chr20_+_28434196 | 5.90 |
ENSDART00000034245
|
dpf3
|
D4, zinc and double PHD fingers, family 3 |
| chr16_+_11151699 | 5.83 |
ENSDART00000140674
|
cicb
|
capicua transcriptional repressor b |
| chr12_+_17504559 | 5.38 |
ENSDART00000020628
|
cyth3a
|
cytohesin 3a |
| chr5_-_46273938 | 5.28 |
ENSDART00000080033
|
si:ch211-130m23.3
|
si:ch211-130m23.3 |
| chr11_+_28218141 | 5.26 |
ENSDART00000043756
|
ephb2b
|
eph receptor B2b |
| chr13_+_32446169 | 5.23 |
ENSDART00000143325
|
nt5c1ba
|
5'-nucleotidase, cytosolic IB a |
| chr5_-_18911114 | 5.22 |
ENSDART00000014434
|
bri3bp
|
bri3 binding protein |
| chr5_+_15667874 | 5.11 |
ENSDART00000127015
|
srrm4
|
serine/arginine repetitive matrix 4 |
| chr11_+_40649412 | 5.00 |
ENSDART00000043016
ENSDART00000134560 |
slc45a1
|
solute carrier family 45, member 1 |
| chr10_+_37500234 | 4.86 |
ENSDART00000132096
ENSDART00000099473 |
msi2a
|
musashi RNA-binding protein 2a |
| chr4_-_17629444 | 4.76 |
ENSDART00000108814
|
nrip2
|
nuclear receptor interacting protein 2 |
| chr21_+_38312549 | 4.72 |
ENSDART00000065159
|
zgc:158291
|
zgc:158291 |
| chr21_-_34658266 | 4.69 |
ENSDART00000023038
|
dacha
|
dachshund a |
| chr4_-_20081621 | 4.65 |
ENSDART00000024647
|
dennd6b
|
DENN/MADD domain containing 6B |
| chr20_+_26095530 | 4.59 |
ENSDART00000139350
|
syne1a
|
spectrin repeat containing, nuclear envelope 1a |
| chr7_-_28565230 | 4.40 |
ENSDART00000028887
|
tmem9b
|
TMEM9 domain family, member B |
| chr5_+_57743815 | 4.38 |
ENSDART00000005090
|
alg9
|
ALG9, alpha-1,2-mannosyltransferase |
| chr21_+_39336285 | 4.30 |
ENSDART00000139677
|
si:ch211-274p24.4
|
si:ch211-274p24.4 |
| chr19_+_42693855 | 4.30 |
ENSDART00000136873
|
clasp2
|
cytoplasmic linker associated protein 2 |
| chr15_+_16897554 | 4.29 |
ENSDART00000154679
|
ypel2b
|
yippee-like 2b |
| chr10_+_25982212 | 4.28 |
ENSDART00000128292
ENSDART00000108808 |
frem2a
|
Fras1 related extracellular matrix protein 2a |
| chr3_-_16039619 | 4.22 |
ENSDART00000143324
|
spsb3a
|
splA/ryanodine receptor domain and SOCS box containing 3a |
| chr18_+_22793743 | 4.15 |
ENSDART00000150106
|
gnao1a
|
guanine nucleotide binding protein (G protein), alpha activating activity polypeptide O, a |
| chr20_-_28433990 | 4.07 |
ENSDART00000182824
ENSDART00000193381 |
wdr21
|
WD repeat domain 21 |
| chr23_-_29667544 | 4.00 |
ENSDART00000059339
|
clstn1
|
calsyntenin 1 |
| chr20_-_39188476 | 3.84 |
ENSDART00000152858
|
rcan2
|
regulator of calcineurin 2 |
| chr16_-_563235 | 3.82 |
ENSDART00000016303
|
irx2a
|
iroquois homeobox 2a |
| chr14_+_49152341 | 3.78 |
ENSDART00000084114
|
nsd1a
|
nuclear receptor binding SET domain protein 1a |
| chr5_+_19320554 | 3.74 |
ENSDART00000165119
|
rusc2
|
RUN and SH3 domain containing 2 |
| chr3_-_40836081 | 3.72 |
ENSDART00000143135
|
wipi2
|
WD repeat domain, phosphoinositide interacting 2 |
| chr16_+_13855039 | 3.67 |
ENSDART00000113764
ENSDART00000143983 |
zgc:174888
|
zgc:174888 |
| chr20_-_47731768 | 3.66 |
ENSDART00000031167
|
tfap2d
|
transcription factor AP-2 delta (activating enhancer binding protein 2 delta) |
| chr5_-_13766651 | 3.55 |
ENSDART00000134064
|
mxd1
|
MAX dimerization protein 1 |
| chr3_+_28576173 | 3.52 |
ENSDART00000151189
|
sept12
|
septin 12 |
| chr8_+_25959940 | 3.47 |
ENSDART00000143011
ENSDART00000140626 |
si:dkey-72l14.4
|
si:dkey-72l14.4 |
| chr3_-_15227666 | 3.41 |
ENSDART00000065807
|
kctd13
|
potassium channel tetramerization domain containing 13 |
| chr5_-_60159116 | 3.40 |
ENSDART00000147675
|
si:dkey-280e8.1
|
si:dkey-280e8.1 |
| chr5_-_32489796 | 3.28 |
ENSDART00000168870
|
gpr107
|
G protein-coupled receptor 107 |
| chr7_-_19369002 | 3.26 |
ENSDART00000165680
|
ntn4
|
netrin 4 |
| chr4_+_9279784 | 3.22 |
ENSDART00000014897
|
srgap1b
|
SLIT-ROBO Rho GTPase activating protein 1b |
| chr22_-_21046654 | 3.22 |
ENSDART00000064902
|
ssbp4
|
single stranded DNA binding protein 4 |
| chr2_-_32558795 | 3.18 |
ENSDART00000140026
|
smarcd3a
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 3a |
| chr10_-_43029001 | 3.14 |
ENSDART00000171494
|
ssbp2
|
single-stranded DNA binding protein 2 |
| chr5_-_12219572 | 3.14 |
ENSDART00000167834
|
nos1
|
nitric oxide synthase 1 (neuronal) |
| chr2_+_38742338 | 3.11 |
ENSDART00000177290
|
carmil3
|
capping protein regulator and myosin 1 linker 3 |
| chr17_+_23964132 | 3.05 |
ENSDART00000154823
|
xpo1b
|
exportin 1 (CRM1 homolog, yeast) b |
| chr6_+_43903209 | 3.01 |
ENSDART00000006435
|
gpr27
|
G protein-coupled receptor 27 |
| chr20_-_27857676 | 2.98 |
ENSDART00000192934
|
syndig1l
|
synapse differentiation inducing 1-like |
| chr5_+_27137473 | 2.89 |
ENSDART00000181833
|
unc5db
|
unc-5 netrin receptor Db |
| chr6_-_39160422 | 2.87 |
ENSDART00000148661
|
stat2
|
signal transducer and activator of transcription 2 |
| chr5_-_23675222 | 2.70 |
ENSDART00000135153
|
TBC1D8B
|
si:dkey-110k5.6 |
| chr7_-_26262978 | 2.69 |
ENSDART00000137769
|
ap1s1
|
adaptor-related protein complex 1, sigma 1 subunit |
| chr19_+_41075488 | 2.60 |
ENSDART00000138687
|
ppp1r9a
|
protein phosphatase 1, regulatory subunit 9A |
| chr20_-_28433616 | 2.58 |
ENSDART00000169289
|
wdr21
|
WD repeat domain 21 |
| chr4_+_4267451 | 2.52 |
ENSDART00000192069
|
ano2
|
anoctamin 2 |
| chr11_-_843811 | 2.49 |
ENSDART00000173331
|
atg7
|
ATG7 autophagy related 7 homolog (S. cerevisiae) |
| chr10_+_42521943 | 2.49 |
ENSDART00000010420
ENSDART00000075303 |
actr1
|
ARP1 actin related protein 1, centractin |
| chr17_-_14780578 | 2.48 |
ENSDART00000154690
|
si:ch211-266o15.1
|
si:ch211-266o15.1 |
| chr7_-_68373495 | 2.47 |
ENSDART00000167440
|
zfhx3
|
zinc finger homeobox 3 |
| chr3_-_23596532 | 2.42 |
ENSDART00000124921
|
ube2z
|
ubiquitin-conjugating enzyme E2Z |
| chr23_-_4019928 | 2.40 |
ENSDART00000021062
|
slc9a8
|
solute carrier family 9, subfamily A (NHE8, cation proton antiporter 8), member 8 |
| chr15_-_46718759 | 2.36 |
ENSDART00000085926
ENSDART00000154577 |
zgc:162872
|
zgc:162872 |
| chr19_-_25427255 | 2.30 |
ENSDART00000036854
|
glcci1a
|
glucocorticoid induced 1a |
| chr10_+_10738880 | 2.30 |
ENSDART00000004181
|
slc27a4
|
solute carrier family 27 (fatty acid transporter), member 4 |
| chr7_-_26263183 | 2.28 |
ENSDART00000079357
ENSDART00000190369 ENSDART00000193154 ENSDART00000101109 |
ap1s1
|
adaptor-related protein complex 1, sigma 1 subunit |
| chr5_-_17601759 | 2.28 |
ENSDART00000138387
|
si:ch211-130h14.6
|
si:ch211-130h14.6 |
| chr13_+_22712406 | 2.28 |
ENSDART00000132847
|
si:ch211-134m17.9
|
si:ch211-134m17.9 |
| chr18_-_26126258 | 2.21 |
ENSDART00000145875
|
ANKRD34C
|
ankyrin repeat domain 34C |
| chr23_+_25893020 | 2.19 |
ENSDART00000144769
|
pkig
|
protein kinase (cAMP-dependent, catalytic) inhibitor gamma |
| chr3_-_16055432 | 2.16 |
ENSDART00000123621
ENSDART00000023859 |
atp6v0ca
|
ATPase H+ transporting V0 subunit ca |
| chr8_+_44623540 | 2.15 |
ENSDART00000141513
|
grk5l
|
G protein-coupled receptor kinase 5 like |
| chr4_+_26056548 | 2.10 |
ENSDART00000171204
|
SCYL2
|
si:ch211-244b2.1 |
| chr11_+_38280454 | 2.03 |
ENSDART00000171496
|
CDK18
|
si:dkey-166c18.1 |
| chr3_-_23596809 | 2.00 |
ENSDART00000156897
|
ube2z
|
ubiquitin-conjugating enzyme E2Z |
| chr12_+_28854963 | 1.96 |
ENSDART00000153227
|
nfe2l1b
|
nuclear factor, erythroid 2-like 1b |
| chr6_+_41452979 | 1.91 |
ENSDART00000007353
|
wdr82
|
WD repeat domain 82 |
| chr18_+_17786548 | 1.88 |
ENSDART00000189493
ENSDART00000146133 |
ZNF423
|
si:ch211-216l23.1 |
| chr4_+_9279515 | 1.87 |
ENSDART00000048707
|
srgap1b
|
SLIT-ROBO Rho GTPase activating protein 1b |
| chr22_-_17574511 | 1.86 |
ENSDART00000181496
|
pip5k1ca
|
phosphatidylinositol-4-phosphate 5-kinase, type I, gamma a |
| chr24_+_33392698 | 1.76 |
ENSDART00000122579
|
si:ch73-173p19.1
|
si:ch73-173p19.1 |
| chr10_+_36695597 | 1.75 |
ENSDART00000169015
ENSDART00000171392 |
rab6a
|
RAB6A, member RAS oncogene family |
| chr2_-_11027258 | 1.69 |
ENSDART00000081072
ENSDART00000193824 ENSDART00000187036 ENSDART00000097741 |
ssbp3a
|
single stranded DNA binding protein 3a |
| chr11_+_44503774 | 1.67 |
ENSDART00000169295
|
ero1b
|
endoplasmic reticulum oxidoreductase beta |
| chr2_-_41723165 | 1.60 |
ENSDART00000155577
|
zgc:110158
|
zgc:110158 |
| chr23_-_4019699 | 1.54 |
ENSDART00000159780
|
slc9a8
|
solute carrier family 9, subfamily A (NHE8, cation proton antiporter 8), member 8 |
| chr19_-_6134802 | 1.52 |
ENSDART00000140051
|
cica
|
capicua transcriptional repressor a |
| chr20_+_32478151 | 1.50 |
ENSDART00000145269
|
ostm1
|
osteopetrosis associated transmembrane protein 1 |
| chr3_+_40315410 | 1.49 |
ENSDART00000083241
ENSDART00000132827 |
slc29a4
|
solute carrier family 29 (equilibrative nucleoside transporter), member 4 |
| chr22_+_2417105 | 1.46 |
ENSDART00000106415
|
zgc:113220
|
zgc:113220 |
| chr12_+_36971952 | 1.43 |
ENSDART00000125900
|
hs3st3b1b
|
heparan sulfate (glucosamine) 3-O-sulfotransferase 3B1b |
| chr5_-_41124241 | 1.41 |
ENSDART00000083561
|
mtmr12
|
myotubularin related protein 12 |
| chr17_+_33999630 | 1.38 |
ENSDART00000167085
ENSDART00000155030 ENSDART00000168522 ENSDART00000191799 ENSDART00000189684 ENSDART00000153942 ENSDART00000187272 ENSDART00000127692 |
gphna
|
gephyrin a |
| chr12_-_36740781 | 1.36 |
ENSDART00000105484
|
si:ch211-216b21.2
|
si:ch211-216b21.2 |
| chr24_-_34034931 | 1.33 |
ENSDART00000169227
|
asic1c
|
acid-sensing (proton-gated) ion channel 1c |
| chr2_-_38117538 | 1.33 |
ENSDART00000013676
|
chd8
|
chromodomain helicase DNA binding protein 8 |
| chr5_-_69437422 | 1.32 |
ENSDART00000073676
|
isca1
|
iron-sulfur cluster assembly 1 |
| chr16_-_563732 | 1.31 |
ENSDART00000183394
|
irx2a
|
iroquois homeobox 2a |
| chr7_-_29723761 | 1.29 |
ENSDART00000173560
|
vps13c
|
vacuolar protein sorting 13 homolog C (S. cerevisiae) |
| chr4_+_25693463 | 1.29 |
ENSDART00000132864
|
acot18
|
acyl-CoA thioesterase 18 |
| chr11_+_2506516 | 1.28 |
ENSDART00000130886
ENSDART00000189767 |
NABP2
|
si:ch73-190f16.2 |
| chr12_-_33659328 | 1.28 |
ENSDART00000153457
|
tmem94
|
transmembrane protein 94 |
| chr1_+_19849168 | 1.24 |
ENSDART00000111454
|
si:dkey-82j4.2
|
si:dkey-82j4.2 |
| chr2_+_3696038 | 1.20 |
ENSDART00000150499
ENSDART00000136906 ENSDART00000164152 ENSDART00000128514 ENSDART00000108964 |
egfra
|
epidermal growth factor receptor a (erythroblastic leukemia viral (v-erb-b) oncogene homolog, avian) |
| chr18_+_17786710 | 1.17 |
ENSDART00000190203
ENSDART00000187095 ENSDART00000083296 |
ZNF423
|
si:ch211-216l23.1 |
| chr23_-_21758253 | 1.16 |
ENSDART00000046613
|
vps13d
|
vacuolar protein sorting 13 homolog D (S. cerevisiae) |
| chr2_-_41723487 | 1.15 |
ENSDART00000170171
|
zgc:110158
|
zgc:110158 |
| chr24_-_21511737 | 1.14 |
ENSDART00000109587
|
rnf6
|
ring finger protein (C3H2C3 type) 6 |
| chr7_+_73332486 | 1.13 |
ENSDART00000174119
ENSDART00000092051 ENSDART00000192388 |
CABZ01081780.1
|
|
| chr3_-_20793655 | 1.09 |
ENSDART00000163473
ENSDART00000159457 |
spop
|
speckle-type POZ protein |
| chr15_+_23528010 | 1.08 |
ENSDART00000152786
|
si:dkey-182i3.8
|
si:dkey-182i3.8 |
| chr7_-_56766973 | 1.07 |
ENSDART00000020967
|
csnk2a2a
|
casein kinase 2, alpha prime polypeptide a |
| chr15_+_23528310 | 1.07 |
ENSDART00000152523
|
si:dkey-182i3.8
|
si:dkey-182i3.8 |
| chr12_+_48204891 | 1.06 |
ENSDART00000190534
ENSDART00000164427 |
ndr2
|
nodal-related 2 |
| chr20_-_20270191 | 1.03 |
ENSDART00000009356
|
ppp2r5ea
|
protein phosphatase 2, regulatory subunit B', epsilon isoform a |
| chr16_+_48714048 | 1.00 |
ENSDART00000148709
ENSDART00000150121 |
brd2b
|
bromodomain containing 2b |
| chr3_-_15889508 | 0.97 |
ENSDART00000148363
|
cramp1
|
cramped chromatin regulator homolog 1 |
| chr11_+_5880562 | 0.92 |
ENSDART00000129663
ENSDART00000130768 ENSDART00000160909 |
dazap1
|
DAZ associated protein 1 |
| chr2_+_30489846 | 0.92 |
ENSDART00000145732
|
march6
|
membrane-associated ring finger (C3HC4) 6 |
| chr14_-_32893785 | 0.91 |
ENSDART00000169380
|
slc25a43
|
solute carrier family 25, member 43 |
| chr6_+_58915889 | 0.89 |
ENSDART00000083628
|
ddit3
|
DNA-damage-inducible transcript 3 |
| chr8_+_30425700 | 0.83 |
ENSDART00000012132
|
cbwd
|
COBW domain containing |
| chr22_-_21046843 | 0.83 |
ENSDART00000133982
|
ssbp4
|
single stranded DNA binding protein 4 |
| chr17_-_11418513 | 0.82 |
ENSDART00000064412
ENSDART00000151847 |
arid4a
|
AT rich interactive domain 4A (RBP1-like) |
| chr19_-_30800004 | 0.81 |
ENSDART00000128560
ENSDART00000045504 ENSDART00000125893 |
trit1
|
tRNA isopentenyltransferase 1 |
| chr5_-_13645995 | 0.80 |
ENSDART00000099665
ENSDART00000166957 |
purba
|
purine-rich element binding protein Ba |
| chr10_+_11355841 | 0.77 |
ENSDART00000193067
ENSDART00000064215 |
cops4
|
COP9 constitutive photomorphogenic homolog subunit 4 (Arabidopsis) |
| chr16_+_40575742 | 0.73 |
ENSDART00000161503
|
ccne2
|
cyclin E2 |
| chr7_+_30725473 | 0.71 |
ENSDART00000085716
|
mtmr10
|
myotubularin related protein 10 |
| chr22_-_22231720 | 0.68 |
ENSDART00000160165
|
ap3d1
|
adaptor-related protein complex 3, delta 1 subunit |
| chr24_+_10413484 | 0.65 |
ENSDART00000111014
|
myca
|
MYC proto-oncogene, bHLH transcription factor a |
| chr16_+_44768361 | 0.56 |
ENSDART00000036302
|
upk1a
|
uroplakin 1a |
| chr7_-_56766100 | 0.54 |
ENSDART00000189934
|
csnk2a2a
|
casein kinase 2, alpha prime polypeptide a |
| chr16_-_13595027 | 0.53 |
ENSDART00000060004
|
ntd5
|
ntl-dependent gene 5 |
| chr6_+_23931236 | 0.52 |
ENSDART00000166079
|
gadd45ab
|
growth arrest and DNA-damage-inducible, alpha, b |
| chr9_-_12885201 | 0.52 |
ENSDART00000124957
|
ankzf1
|
ankyrin repeat and zinc finger domain containing 1 |
| chr12_+_18899396 | 0.50 |
ENSDART00000105858
|
xrcc6
|
X-ray repair complementing defective repair in Chinese hamster cells 6 |
| chr7_-_37555208 | 0.49 |
ENSDART00000148905
ENSDART00000150229 |
cylda
|
cylindromatosis (turban tumor syndrome), a |
| chr13_-_18011168 | 0.49 |
ENSDART00000144813
|
march8
|
membrane-associated ring finger (C3HC4) 8 |
| chr19_-_81477 | 0.47 |
ENSDART00000159815
|
hnrnpr
|
heterogeneous nuclear ribonucleoprotein R |
| chr14_-_689841 | 0.45 |
ENSDART00000125969
|
ube2ka
|
ubiquitin-conjugating enzyme E2Ka (UBC1 homolog, yeast) |
| chr20_+_26690036 | 0.38 |
ENSDART00000103232
|
foxf2b
|
forkhead box F2b |
| chr12_+_3871452 | 0.37 |
ENSDART00000066546
|
nif3l1
|
NIF3 NGG1 interacting factor 3-like 1 (S. cerevisiae) |
| chr19_+_25465025 | 0.27 |
ENSDART00000018553
|
rpa3
|
replication protein A3 |
| chr22_-_20720427 | 0.25 |
ENSDART00000105532
|
oaz1a
|
ornithine decarboxylase antizyme 1a |
| chr25_+_14870043 | 0.25 |
ENSDART00000035714
ENSDART00000171835 |
dnajc24
|
DnaJ (Hsp40) homolog, subfamily C, member 24 |
| chr17_+_27158706 | 0.22 |
ENSDART00000151829
|
rps6ka1
|
ribosomal protein S6 kinase a, polypeptide 1 |
| chr7_+_52761841 | 0.16 |
ENSDART00000111444
|
ppip5k1a
|
diphosphoinositol pentakisphosphate kinase 1a |
| chr23_+_24705424 | 0.14 |
ENSDART00000104029
|
c1qtnf12
|
C1q and TNF related 12 |
| chr15_-_20528494 | 0.14 |
ENSDART00000048423
|
timm50
|
translocase of inner mitochondrial membrane 50 homolog (S. cerevisiae) |
| chr3_+_13190012 | 0.12 |
ENSDART00000179747
ENSDART00000109876 ENSDART00000124824 ENSDART00000130261 |
sun1
|
Sad1 and UNC84 domain containing 1 |
| chr17_-_23673864 | 0.11 |
ENSDART00000104738
ENSDART00000128958 |
ptena
|
phosphatase and tensin homolog A |
| chr23_+_33963619 | 0.10 |
ENSDART00000140666
ENSDART00000084792 |
plpbp
|
pyridoxal phosphate binding protein |
| chr3_-_25646149 | 0.01 |
ENSDART00000122735
|
usp43b
|
ubiquitin specific peptidase 43b |
Network of associatons between targets according to the STRING database.
First level regulatory network of pknox2
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.0 | 15.1 | GO:0097378 | interneuron axon guidance(GO:0097376) spinal cord interneuron axon guidance(GO:0097377) dorsal spinal cord interneuron axon guidance(GO:0097378) |
| 1.0 | 3.1 | GO:0098923 | retrograde trans-synaptic signaling by soluble gas(GO:0098923) retrograde trans-synaptic signaling by nitric oxide(GO:0098924) |
| 1.0 | 4.0 | GO:0030951 | establishment or maintenance of microtubule cytoskeleton polarity(GO:0030951) |
| 0.6 | 1.9 | GO:0080182 | histone H3-K4 trimethylation(GO:0080182) |
| 0.5 | 1.4 | GO:0042040 | establishment of synaptic specificity at neuromuscular junction(GO:0007529) molybdenum incorporation into molybdenum-molybdopterin complex(GO:0018315) metal incorporation into metallo-molybdopterin complex(GO:0042040) glycine receptor clustering(GO:0072579) |
| 0.4 | 5.2 | GO:0046085 | adenosine metabolic process(GO:0046085) |
| 0.4 | 5.1 | GO:0021754 | facial nucleus development(GO:0021754) |
| 0.4 | 2.2 | GO:2000480 | negative regulation of cAMP-dependent protein kinase activity(GO:2000480) |
| 0.4 | 2.5 | GO:0006501 | C-terminal protein lipidation(GO:0006501) |
| 0.4 | 3.2 | GO:0032986 | nucleosome disassembly(GO:0006337) chromatin disassembly(GO:0031498) protein-DNA complex disassembly(GO:0032986) |
| 0.4 | 2.5 | GO:0045053 | protein retention in Golgi apparatus(GO:0045053) |
| 0.3 | 3.1 | GO:0000056 | ribosomal small subunit export from nucleus(GO:0000056) |
| 0.3 | 8.7 | GO:0021952 | central nervous system projection neuron axonogenesis(GO:0021952) |
| 0.3 | 2.3 | GO:0042760 | very long-chain fatty acid catabolic process(GO:0042760) |
| 0.3 | 3.7 | GO:0034497 | protein localization to pre-autophagosomal structure(GO:0034497) |
| 0.3 | 5.1 | GO:0035778 | pronephric nephron tubule epithelial cell differentiation(GO:0035778) cell differentiation involved in pronephros development(GO:0039014) nephron tubule epithelial cell differentiation(GO:0072160) |
| 0.3 | 1.7 | GO:0034975 | protein folding in endoplasmic reticulum(GO:0034975) |
| 0.3 | 2.9 | GO:0038007 | netrin-activated signaling pathway(GO:0038007) |
| 0.3 | 2.9 | GO:0071357 | response to type I interferon(GO:0034340) type I interferon signaling pathway(GO:0060337) cellular response to type I interferon(GO:0071357) |
| 0.2 | 2.2 | GO:0060036 | notochord cell vacuolation(GO:0060036) |
| 0.2 | 0.9 | GO:2001244 | positive regulation of intrinsic apoptotic signaling pathway(GO:2001244) |
| 0.2 | 1.1 | GO:0001709 | cell fate determination(GO:0001709) |
| 0.2 | 3.5 | GO:0032418 | lysosome localization(GO:0032418) |
| 0.2 | 1.3 | GO:0070207 | protein homotrimerization(GO:0070207) |
| 0.2 | 1.3 | GO:2000178 | negative regulation of neural precursor cell proliferation(GO:2000178) |
| 0.2 | 1.3 | GO:0097428 | protein maturation by iron-sulfur cluster transfer(GO:0097428) |
| 0.2 | 4.3 | GO:0007020 | microtubule nucleation(GO:0007020) |
| 0.2 | 5.9 | GO:0055008 | cardiac muscle tissue morphogenesis(GO:0055008) |
| 0.2 | 6.3 | GO:0060219 | camera-type eye photoreceptor cell differentiation(GO:0060219) |
| 0.2 | 5.8 | GO:0060319 | primitive erythrocyte differentiation(GO:0060319) |
| 0.1 | 2.3 | GO:0010569 | regulation of double-strand break repair via homologous recombination(GO:0010569) |
| 0.1 | 0.5 | GO:0002495 | antigen processing and presentation of peptide antigen via MHC class II(GO:0002495) |
| 0.1 | 5.4 | GO:0032011 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.1 | 3.3 | GO:0070831 | basement membrane assembly(GO:0070831) |
| 0.1 | 0.7 | GO:0048490 | anterograde synaptic vesicle transport(GO:0048490) synaptic vesicle cytoskeletal transport(GO:0099514) synaptic vesicle transport along microtubule(GO:0099517) |
| 0.1 | 2.1 | GO:1902017 | regulation of cilium assembly(GO:1902017) |
| 0.1 | 0.5 | GO:0071480 | response to gamma radiation(GO:0010332) cellular response to gamma radiation(GO:0071480) |
| 0.1 | 1.5 | GO:0015858 | nucleoside transport(GO:0015858) nucleoside transmembrane transport(GO:1901642) |
| 0.1 | 4.2 | GO:0007212 | dopamine receptor signaling pathway(GO:0007212) |
| 0.1 | 7.3 | GO:1902476 | chloride transmembrane transport(GO:1902476) |
| 0.1 | 1.2 | GO:0072554 | endothelial tube morphogenesis(GO:0061154) blood vessel lumenization(GO:0072554) |
| 0.1 | 9.6 | GO:0008360 | regulation of cell shape(GO:0008360) |
| 0.1 | 0.5 | GO:1990108 | protein linear deubiquitination(GO:1990108) |
| 0.1 | 5.1 | GO:0030336 | negative regulation of cell migration(GO:0030336) |
| 0.1 | 0.4 | GO:0051145 | smooth muscle cell differentiation(GO:0051145) |
| 0.1 | 1.7 | GO:0048936 | peripheral nervous system neuron axonogenesis(GO:0048936) |
| 0.1 | 2.3 | GO:0072015 | glomerular visceral epithelial cell development(GO:0072015) glomerular epithelial cell development(GO:0072310) |
| 0.1 | 3.0 | GO:0033500 | carbohydrate homeostasis(GO:0033500) glucose homeostasis(GO:0042593) |
| 0.1 | 6.4 | GO:0019722 | calcium-mediated signaling(GO:0019722) |
| 0.1 | 0.5 | GO:0000185 | activation of MAPKKK activity(GO:0000185) |
| 0.1 | 0.9 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
| 0.1 | 1.0 | GO:0002574 | thrombocyte differentiation(GO:0002574) |
| 0.0 | 1.0 | GO:0031952 | regulation of protein autophosphorylation(GO:0031952) |
| 0.0 | 2.8 | GO:0051453 | regulation of intracellular pH(GO:0051453) |
| 0.0 | 0.1 | GO:0046324 | regulation of glucose import(GO:0046324) |
| 0.0 | 4.4 | GO:0006487 | protein N-linked glycosylation(GO:0006487) |
| 0.0 | 3.5 | GO:0033334 | fin morphogenesis(GO:0033334) |
| 0.0 | 2.0 | GO:0034599 | cellular response to oxidative stress(GO:0034599) |
| 0.0 | 6.4 | GO:0006486 | protein glycosylation(GO:0006486) macromolecule glycosylation(GO:0043413) |
| 0.0 | 13.9 | GO:0045944 | positive regulation of transcription from RNA polymerase II promoter(GO:0045944) |
| 0.0 | 4.7 | GO:0048839 | ear development(GO:0043583) inner ear development(GO:0048839) |
| 0.0 | 1.3 | GO:0046856 | phosphatidylinositol dephosphorylation(GO:0046856) |
| 0.0 | 1.6 | GO:0018107 | peptidyl-threonine phosphorylation(GO:0018107) peptidyl-threonine modification(GO:0018210) |
| 0.0 | 0.2 | GO:0032781 | positive regulation of ATPase activity(GO:0032781) |
| 0.0 | 13.1 | GO:0007420 | brain development(GO:0007420) |
| 0.0 | 2.2 | GO:0072583 | clathrin-mediated endocytosis(GO:0072583) |
| 0.0 | 5.0 | GO:0030198 | extracellular matrix organization(GO:0030198) extracellular structure organization(GO:0043062) |
| 0.0 | 1.7 | GO:0061640 | cytoskeleton-dependent cytokinesis(GO:0061640) |
| 0.0 | 1.4 | GO:0030433 | ER-associated ubiquitin-dependent protein catabolic process(GO:0030433) |
| 0.0 | 0.8 | GO:0006400 | tRNA modification(GO:0006400) |
| 0.0 | 0.3 | GO:0006596 | polyamine biosynthetic process(GO:0006596) |
| 0.0 | 0.7 | GO:1904029 | regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0000079) regulation of cyclin-dependent protein kinase activity(GO:1904029) |
| 0.0 | 4.3 | GO:0042127 | regulation of cell proliferation(GO:0042127) |
| 0.0 | 3.6 | GO:0032259 | methylation(GO:0032259) |
| 0.0 | 0.2 | GO:0006020 | inositol metabolic process(GO:0006020) |
| 0.0 | 2.3 | GO:0000122 | negative regulation of transcription from RNA polymerase II promoter(GO:0000122) |
| 0.0 | 0.3 | GO:0006298 | mismatch repair(GO:0006298) |
| 0.0 | 4.4 | GO:0043066 | negative regulation of apoptotic process(GO:0043066) |
| 0.0 | 4.0 | GO:0007264 | small GTPase mediated signal transduction(GO:0007264) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 3.7 | GO:0034045 | pre-autophagosomal structure membrane(GO:0034045) |
| 0.4 | 9.1 | GO:0071565 | nBAF complex(GO:0071565) |
| 0.3 | 4.6 | GO:0005640 | nuclear outer membrane(GO:0005640) |
| 0.2 | 7.3 | GO:1902710 | GABA receptor complex(GO:1902710) GABA-A receptor complex(GO:1902711) |
| 0.2 | 0.7 | GO:0097134 | cyclin E1-CDK2 complex(GO:0097134) |
| 0.2 | 4.3 | GO:0045180 | basal cortex(GO:0045180) |
| 0.2 | 4.9 | GO:0005844 | polysome(GO:0005844) |
| 0.1 | 2.2 | GO:0033179 | proton-transporting V-type ATPase, V0 domain(GO:0033179) |
| 0.1 | 4.5 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.1 | 2.5 | GO:0000407 | pre-autophagosomal structure(GO:0000407) |
| 0.1 | 1.6 | GO:0005956 | protein kinase CK2 complex(GO:0005956) |
| 0.1 | 2.4 | GO:0005605 | basal lamina(GO:0005605) laminin complex(GO:0043256) |
| 0.1 | 6.4 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.1 | 1.9 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.1 | 3.5 | GO:0005940 | septin ring(GO:0005940) septin complex(GO:0031105) septin cytoskeleton(GO:0032156) |
| 0.1 | 2.6 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.1 | 5.1 | GO:0005684 | U2-type spliceosomal complex(GO:0005684) |
| 0.1 | 0.8 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.1 | 1.3 | GO:0044665 | MLL1/2 complex(GO:0044665) MLL1 complex(GO:0071339) |
| 0.1 | 4.2 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.1 | 1.2 | GO:0045178 | basal part of cell(GO:0045178) |
| 0.1 | 5.7 | GO:0030117 | membrane coat(GO:0030117) coated membrane(GO:0048475) |
| 0.0 | 3.9 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.0 | 2.3 | GO:0005902 | microvillus(GO:0005902) |
| 0.0 | 2.2 | GO:0030136 | clathrin-coated vesicle(GO:0030136) |
| 0.0 | 4.0 | GO:0045211 | postsynaptic membrane(GO:0045211) |
| 0.0 | 3.1 | GO:0042383 | sarcolemma(GO:0042383) |
| 0.0 | 1.0 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.0 | 22.6 | GO:0043005 | neuron projection(GO:0043005) |
| 0.0 | 5.6 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.0 | 0.1 | GO:0034992 | microtubule organizing center attachment site(GO:0034992) LINC complex(GO:0034993) |
| 0.0 | 0.1 | GO:0005744 | mitochondrial inner membrane presequence translocase complex(GO:0005744) |
| 0.0 | 3.6 | GO:0000785 | chromatin(GO:0000785) |
| 0.0 | 1.1 | GO:0055037 | recycling endosome(GO:0055037) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 4.4 | GO:0000026 | alpha-1,2-mannosyltransferase activity(GO:0000026) |
| 0.8 | 4.2 | GO:0051430 | mu-type opioid receptor binding(GO:0031852) corticotropin-releasing hormone receptor 1 binding(GO:0051430) |
| 0.8 | 6.4 | GO:0003835 | beta-galactoside alpha-2,6-sialyltransferase activity(GO:0003835) |
| 0.6 | 3.8 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 0.6 | 5.0 | GO:0008515 | sugar:proton symporter activity(GO:0005351) cation:sugar symporter activity(GO:0005402) sucrose:proton symporter activity(GO:0008506) sucrose transmembrane transporter activity(GO:0008515) disaccharide transmembrane transporter activity(GO:0015154) oligosaccharide transmembrane transporter activity(GO:0015157) |
| 0.5 | 2.5 | GO:0019778 | Atg12 activating enzyme activity(GO:0019778) Atg8 activating enzyme activity(GO:0019779) |
| 0.5 | 1.4 | GO:0061598 | molybdopterin adenylyltransferase activity(GO:0061598) molybdopterin molybdotransferase activity(GO:0061599) |
| 0.4 | 3.1 | GO:0004517 | nitric-oxide synthase activity(GO:0004517) |
| 0.4 | 1.2 | GO:0005006 | epidermal growth factor-activated receptor activity(GO:0005006) |
| 0.4 | 2.2 | GO:0004862 | cAMP-dependent protein kinase inhibitor activity(GO:0004862) |
| 0.4 | 3.9 | GO:0015386 | monovalent cation:proton antiporter activity(GO:0005451) sodium:proton antiporter activity(GO:0015385) potassium:proton antiporter activity(GO:0015386) |
| 0.3 | 2.3 | GO:0031957 | very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 0.3 | 5.3 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.3 | 3.1 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.3 | 13.9 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.3 | 2.9 | GO:0005042 | netrin receptor activity(GO:0005042) |
| 0.3 | 1.3 | GO:0044736 | acid-sensing ion channel activity(GO:0044736) |
| 0.3 | 4.8 | GO:0070001 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.2 | 3.8 | GO:0046975 | histone methyltransferase activity (H3-K36 specific)(GO:0046975) |
| 0.2 | 3.3 | GO:0032050 | clathrin heavy chain binding(GO:0032050) |
| 0.2 | 7.3 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 0.2 | 1.5 | GO:0008504 | monoamine transmembrane transporter activity(GO:0008504) |
| 0.2 | 0.9 | GO:0034046 | poly(G) binding(GO:0034046) |
| 0.2 | 1.2 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 0.2 | 3.6 | GO:0008253 | 5'-nucleotidase activity(GO:0008253) |
| 0.2 | 3.7 | GO:0080025 | phosphatidylinositol-3,5-bisphosphate binding(GO:0080025) |
| 0.2 | 15.9 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.2 | 1.1 | GO:0070698 | type I activin receptor binding(GO:0070698) |
| 0.1 | 2.1 | GO:0004703 | G-protein coupled receptor kinase activity(GO:0004703) |
| 0.1 | 1.7 | GO:0016671 | protein disulfide isomerase activity(GO:0003756) oxidoreductase activity, acting on a sulfur group of donors, disulfide as acceptor(GO:0016671) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
| 0.1 | 4.3 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.1 | 0.8 | GO:0032422 | purine-rich negative regulatory element binding(GO:0032422) |
| 0.1 | 4.4 | GO:0004869 | cysteine-type endopeptidase inhibitor activity(GO:0004869) |
| 0.1 | 1.9 | GO:0042800 | histone methyltransferase activity (H3-K4 specific)(GO:0042800) |
| 0.1 | 0.7 | GO:0008467 | [heparan sulfate]-glucosamine 3-sulfotransferase 1 activity(GO:0008467) |
| 0.1 | 1.3 | GO:0002039 | p53 binding(GO:0002039) |
| 0.1 | 0.3 | GO:0008073 | ornithine decarboxylase inhibitor activity(GO:0008073) |
| 0.1 | 1.0 | GO:0019211 | phosphatase activator activity(GO:0019211) protein phosphatase activator activity(GO:0072542) |
| 0.1 | 1.3 | GO:0047617 | acyl-CoA hydrolase activity(GO:0047617) |
| 0.1 | 1.9 | GO:0016307 | phosphatidylinositol phosphate kinase activity(GO:0016307) |
| 0.0 | 1.3 | GO:0004438 | phosphatidylinositol-3-phosphatase activity(GO:0004438) |
| 0.0 | 0.2 | GO:0000829 | inositol heptakisphosphate kinase activity(GO:0000829) diphosphoinositol-pentakisphosphate kinase activity(GO:0033857) |
| 0.0 | 1.3 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 0.0 | 0.5 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.0 | 5.7 | GO:0042393 | histone binding(GO:0042393) |
| 0.0 | 11.4 | GO:0003924 | GTPase activity(GO:0003924) |
| 0.0 | 0.5 | GO:0042162 | telomeric DNA binding(GO:0042162) |
| 0.0 | 4.1 | GO:0001228 | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001228) |
| 0.0 | 0.2 | GO:0004711 | ribosomal protein S6 kinase activity(GO:0004711) |
| 0.0 | 0.9 | GO:0005347 | ATP transmembrane transporter activity(GO:0005347) |
| 0.0 | 5.7 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.0 | 0.5 | GO:0042287 | MHC protein binding(GO:0042287) |
| 0.0 | 32.9 | GO:0000981 | RNA polymerase II transcription factor activity, sequence-specific DNA binding(GO:0000981) |
| 0.0 | 5.3 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.0 | 0.2 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.0 | 4.2 | GO:0005085 | guanyl-nucleotide exchange factor activity(GO:0005085) |
| 0.0 | 0.7 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.0 | 4.0 | GO:0044822 | mRNA binding(GO:0003729) poly(A) RNA binding(GO:0044822) |
| 0.0 | 0.8 | GO:0016765 | transferase activity, transferring alkyl or aryl (other than methyl) groups(GO:0016765) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 2.9 | ST TYPE I INTERFERON PATHWAY | Type I Interferon (alpha/beta IFN) Pathway |
| 0.3 | 3.8 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.0 | 3.5 | PID HDAC CLASSI PATHWAY | Signaling events mediated by HDAC Class I |
| 0.0 | 2.4 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.0 | 0.7 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.0 | 2.5 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.0 | 0.5 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.0 | 1.1 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.0 | 0.9 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.0 | 1.3 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 7.3 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.5 | 5.0 | REACTOME NEF MEDIATED DOWNREGULATION OF MHC CLASS I COMPLEX CELL SURFACE EXPRESSION | Genes involved in Nef mediated downregulation of MHC class I complex cell surface expression |
| 0.3 | 2.9 | REACTOME REGULATION OF IFNA SIGNALING | Genes involved in Regulation of IFNA signaling |
| 0.2 | 3.1 | REACTOME NITRIC OXIDE STIMULATES GUANYLATE CYCLASE | Genes involved in Nitric oxide stimulates guanylate cyclase |
| 0.2 | 3.8 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.1 | 4.4 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.1 | 0.9 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.1 | 1.7 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.1 | 3.3 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.1 | 0.5 | REACTOME INTEGRATION OF PROVIRUS | Genes involved in Integration of provirus |
| 0.1 | 0.7 | REACTOME SCFSKP2 MEDIATED DEGRADATION OF P27 P21 | Genes involved in SCF(Skp2)-mediated degradation of p27/p21 |
| 0.1 | 2.7 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.0 | 8.0 | REACTOME ANTIGEN PROCESSING UBIQUITINATION PROTEASOME DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |
| 0.0 | 2.8 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.0 | 0.3 | REACTOME REMOVAL OF THE FLAP INTERMEDIATE FROM THE C STRAND | Genes involved in Removal of the Flap Intermediate from the C-strand |
| 0.0 | 0.2 | REACTOME CREB PHOSPHORYLATION THROUGH THE ACTIVATION OF RAS | Genes involved in CREB phosphorylation through the activation of Ras |