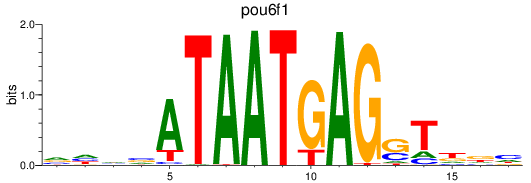
Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
Results for pou6f1
Z-value: 0.95
Transcription factors associated with pou6f1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
pou6f1
|
ENSDARG00000011570 | POU class 6 homeobox 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| pou6f1 | dr11_v1_chr23_-_33709964_33710029 | 0.65 | 1.8e-12 | Click! |
Activity profile of pou6f1 motif
Sorted Z-values of pou6f1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr14_+_50770537 | 21.21 |
ENSDART00000158723
|
sncb
|
synuclein, beta |
| chr15_-_16098531 | 20.75 |
ENSDART00000080377
|
aldoca
|
aldolase C, fructose-bisphosphate, a |
| chr4_+_12615836 | 17.14 |
ENSDART00000003583
|
lmo3
|
LIM domain only 3 |
| chr4_+_22365061 | 14.90 |
ENSDART00000039277
|
lhfpl3
|
LHFPL tetraspan subfamily member 3 |
| chr18_-_1185772 | 14.09 |
ENSDART00000143245
|
nptnb
|
neuroplastin b |
| chr7_+_38717624 | 13.77 |
ENSDART00000132522
|
syt13
|
synaptotagmin XIII |
| chr5_+_64732270 | 13.15 |
ENSDART00000134241
|
olfm1a
|
olfactomedin 1a |
| chr19_+_30662529 | 12.84 |
ENSDART00000175662
|
fam49al
|
family with sequence similarity 49, member A-like |
| chr3_-_28250722 | 12.42 |
ENSDART00000165936
|
rbfox1
|
RNA binding fox-1 homolog 1 |
| chr11_+_39672874 | 10.90 |
ENSDART00000046663
ENSDART00000157659 |
camta1b
|
calmodulin binding transcription activator 1b |
| chr19_-_5103313 | 10.15 |
ENSDART00000037007
|
tpi1a
|
triosephosphate isomerase 1a |
| chr11_-_7320211 | 9.61 |
ENSDART00000091664
|
apc2
|
adenomatosis polyposis coli 2 |
| chr21_-_26918901 | 9.03 |
ENSDART00000100685
|
lrfn4a
|
leucine rich repeat and fibronectin type III domain containing 4a |
| chr17_+_12698532 | 7.43 |
ENSDART00000064509
ENSDART00000136830 |
stmn4l
|
stathmin-like 4, like |
| chr24_-_32408404 | 7.38 |
ENSDART00000144157
|
si:ch211-56a11.2
|
si:ch211-56a11.2 |
| chr22_-_11829436 | 7.25 |
ENSDART00000126784
|
ptpn4b
|
protein tyrosine phosphatase, non-receptor type 4b |
| chr4_-_19028861 | 7.24 |
ENSDART00000166374
|
si:dkey-31f5.11
|
si:dkey-31f5.11 |
| chr12_+_18681477 | 7.07 |
ENSDART00000127981
ENSDART00000143979 |
rgs9b
|
regulator of G protein signaling 9b |
| chr24_+_41931585 | 6.81 |
ENSDART00000130310
|
epb41l3a
|
erythrocyte membrane protein band 4.1-like 3a |
| chr6_+_3004972 | 6.45 |
ENSDART00000186750
ENSDART00000183862 ENSDART00000191485 ENSDART00000171014 |
ptprfa
|
protein tyrosine phosphatase, receptor type, f, a |
| chr22_-_15385442 | 6.40 |
ENSDART00000090975
|
tmem264
|
transmembrane protein 264 |
| chr9_+_11034314 | 6.07 |
ENSDART00000032695
|
asic4a
|
acid-sensing (proton-gated) ion channel family member 4a |
| chr14_-_31465905 | 6.07 |
ENSDART00000173108
|
gpc3
|
glypican 3 |
| chr22_+_16535575 | 5.74 |
ENSDART00000083063
|
tal1
|
T-cell acute lymphocytic leukemia 1 |
| chr13_-_21701323 | 5.68 |
ENSDART00000164112
|
si:dkey-191g9.7
|
si:dkey-191g9.7 |
| chr9_+_41612642 | 5.65 |
ENSDART00000138473
|
spegb
|
SPEG complex locus b |
| chr2_-_12243213 | 5.60 |
ENSDART00000113081
|
gpr158b
|
G protein-coupled receptor 158b |
| chr3_+_33341640 | 5.48 |
ENSDART00000186352
|
pyya
|
peptide YYa |
| chr22_+_2830703 | 5.42 |
ENSDART00000145463
ENSDART00000144785 |
si:dkey-20i20.8
|
si:dkey-20i20.8 |
| chr2_+_29249204 | 5.40 |
ENSDART00000168957
|
cdh18a
|
cadherin 18, type 2a |
| chr12_+_32729470 | 5.32 |
ENSDART00000175712
|
rbfox3a
|
RNA binding fox-1 homolog 3a |
| chr1_+_25801648 | 5.30 |
ENSDART00000129471
|
gucy1b1
|
guanylate cyclase 1 soluble subunit beta 1 |
| chr21_+_25231160 | 5.27 |
ENSDART00000063089
ENSDART00000139127 |
gng8
|
guanine nucleotide binding protein (G protein), gamma 8 |
| chr19_-_31402429 | 5.10 |
ENSDART00000137292
|
tmem106bb
|
transmembrane protein 106Bb |
| chr22_+_2793382 | 5.10 |
ENSDART00000144980
|
si:dkey-20i20.8
|
si:dkey-20i20.8 |
| chr24_-_21404367 | 4.92 |
ENSDART00000152093
|
atp8a2
|
ATPase phospholipid transporting 8A2 |
| chr19_-_9503473 | 4.91 |
ENSDART00000091615
|
iffo1a
|
intermediate filament family orphan 1a |
| chr16_-_5154024 | 4.64 |
ENSDART00000132069
ENSDART00000060635 |
dctn3
|
dynactin 3 (p22) |
| chr13_+_22119798 | 4.45 |
ENSDART00000173206
ENSDART00000078652 ENSDART00000165842 |
camk2g2
|
calcium/calmodulin-dependent protein kinase (CaM kinase) II gamma 2 |
| chr17_+_22072337 | 4.27 |
ENSDART00000063716
ENSDART00000141684 |
nphp1
|
nephronophthisis 1 |
| chr13_+_43400443 | 4.21 |
ENSDART00000084321
|
dact2
|
dishevelled-binding antagonist of beta-catenin 2 |
| chr22_-_10019768 | 4.17 |
ENSDART00000162077
|
BX324216.2
|
|
| chr5_-_35953472 | 4.10 |
ENSDART00000143448
|
rxfp2l
|
relaxin/insulin-like family peptide receptor 2, like |
| chr11_+_29770966 | 4.06 |
ENSDART00000088624
ENSDART00000124471 |
rpgrb
|
retinitis pigmentosa GTPase regulator b |
| chr2_+_9552456 | 4.05 |
ENSDART00000056896
|
dnajb4
|
DnaJ (Hsp40) homolog, subfamily B, member 4 |
| chr3_-_21288202 | 4.05 |
ENSDART00000191766
ENSDART00000187319 |
fam171a2a
|
family with sequence similarity 171, member A2a |
| chr1_-_57280585 | 3.90 |
ENSDART00000152220
|
si:dkey-27j5.5
|
si:dkey-27j5.5 |
| chr6_-_17849786 | 3.81 |
ENSDART00000172709
|
rptor
|
regulatory associated protein of MTOR, complex 1 |
| chr4_+_14826299 | 3.76 |
ENSDART00000067035
|
shisal1a
|
shisa like 1a |
| chr24_+_14595680 | 3.72 |
ENSDART00000137337
ENSDART00000091784 ENSDART00000136026 |
thtpa
|
thiamine triphosphatase |
| chr2_-_48966431 | 3.69 |
ENSDART00000147948
|
kcnj9
|
potassium inwardly-rectifying channel, subfamily J, member 9 |
| chr17_-_45150763 | 3.63 |
ENSDART00000155043
ENSDART00000156786 ENSDART00000191147 |
tmed8
|
transmembrane p24 trafficking protein 8 |
| chr19_-_8600383 | 3.52 |
ENSDART00000081546
|
trim46b
|
tripartite motif containing 46b |
| chr15_+_24704798 | 3.48 |
ENSDART00000192470
|
LRRC75A
|
si:dkey-151p21.7 |
| chr4_-_18202291 | 3.39 |
ENSDART00000169704
|
anks1b
|
ankyrin repeat and sterile alpha motif domain containing 1B |
| chr25_-_20666328 | 3.15 |
ENSDART00000098076
|
csk
|
C-terminal Src kinase |
| chr8_-_10961991 | 2.84 |
ENSDART00000139603
|
trim33
|
tripartite motif containing 33 |
| chr14_+_46003973 | 2.70 |
ENSDART00000145241
ENSDART00000141545 |
slu7
|
SLU7 homolog, splicing factor |
| chr9_-_6624663 | 2.64 |
ENSDART00000092537
|
GPR45
|
si:dkeyp-118h3.5 |
| chr19_+_46222918 | 2.55 |
ENSDART00000158703
|
vps28
|
vacuolar protein sorting 28 (yeast) |
| chr16_-_24044664 | 2.53 |
ENSDART00000136982
|
bcam
|
basal cell adhesion molecule (Lutheran blood group) |
| chr9_-_14504834 | 2.40 |
ENSDART00000056103
|
nrp2b
|
neuropilin 2b |
| chr19_+_46222428 | 2.26 |
ENSDART00000183984
|
vps28
|
vacuolar protein sorting 28 (yeast) |
| chr4_+_25431990 | 2.19 |
ENSDART00000176982
|
prkcq
|
protein kinase C, theta |
| chr13_+_3938526 | 2.17 |
ENSDART00000012759
|
yipf3
|
Yip1 domain family, member 3 |
| chr11_+_21587950 | 2.15 |
ENSDART00000162251
|
FAM72B
|
zgc:101564 |
| chr2_-_50372467 | 2.13 |
ENSDART00000108900
|
cntnap2b
|
contactin associated protein like 2b |
| chr9_-_38036984 | 2.10 |
ENSDART00000134574
|
hacd2
|
3-hydroxyacyl-CoA dehydratase 2 |
| chr25_-_20666754 | 2.09 |
ENSDART00000158418
|
csk
|
C-terminal Src kinase |
| chr7_-_19364813 | 2.08 |
ENSDART00000173977
|
ntn4
|
netrin 4 |
| chr22_+_1462997 | 2.05 |
ENSDART00000157476
|
si:dkeyp-53d3.5
|
si:dkeyp-53d3.5 |
| chr9_-_32672129 | 1.90 |
ENSDART00000140581
|
gzm3.4
|
granzyme 3, tandem duplicate 4 |
| chr19_+_42469058 | 1.77 |
ENSDART00000076915
|
si:dkey-166k12.1
|
si:dkey-166k12.1 |
| chr17_-_49438873 | 1.76 |
ENSDART00000004424
|
znf292a
|
zinc finger protein 292a |
| chr7_+_44715224 | 1.75 |
ENSDART00000184630
|
si:dkey-56m19.5
|
si:dkey-56m19.5 |
| chr5_-_39736383 | 1.73 |
ENSDART00000127123
|
rasgef1ba
|
RasGEF domain family, member 1Ba |
| chr22_+_2504524 | 1.71 |
ENSDART00000136686
|
znf1170
|
zinc finger protein 1170 |
| chr7_+_53498152 | 1.55 |
ENSDART00000184497
|
znf609b
|
zinc finger protein 609b |
| chr18_+_34181655 | 1.54 |
ENSDART00000130831
ENSDART00000109535 |
gmps
|
guanine monophosphate synthase |
| chr13_-_24825691 | 1.49 |
ENSDART00000142745
|
slka
|
STE20-like kinase a |
| chr7_+_48297842 | 1.48 |
ENSDART00000052123
|
slc25a44b
|
solute carrier family 25, member 44 b |
| chr15_-_931815 | 1.47 |
ENSDART00000106627
ENSDART00000102239 ENSDART00000184754 ENSDART00000186733 |
zgc:162936
|
zgc:162936 |
| chr16_-_25368048 | 1.41 |
ENSDART00000132445
|
rbfa
|
ribosome binding factor A |
| chr20_+_22067337 | 1.38 |
ENSDART00000152636
|
clocka
|
clock circadian regulator a |
| chr3_-_33935200 | 1.29 |
ENSDART00000151211
|
gtf2f1
|
general transcription factor IIF, polypeptide 1 |
| chr7_-_9803154 | 1.29 |
ENSDART00000055593
|
aldh1a3
|
aldehyde dehydrogenase 1 family, member A3 |
| chr15_-_33640890 | 1.28 |
ENSDART00000167705
|
stard13b
|
StAR-related lipid transfer (START) domain containing 13b |
| chr2_-_27619954 | 1.27 |
ENSDART00000144826
|
tgs1
|
trimethylguanosine synthase 1 |
| chr18_+_3243292 | 1.14 |
ENSDART00000166580
|
pak1
|
p21 protein (Cdc42/Rac)-activated kinase 1 |
| chr8_-_7474997 | 1.14 |
ENSDART00000146555
|
gata1b
|
GATA binding protein 1b |
| chr1_+_19649545 | 1.12 |
ENSDART00000054575
|
tmem192
|
transmembrane protein 192 |
| chr23_+_41912151 | 1.11 |
ENSDART00000191115
|
podn
|
podocan |
| chr25_+_37480285 | 1.03 |
ENSDART00000166187
|
CABZ01095001.1
|
|
| chr17_-_49443399 | 1.02 |
ENSDART00000155293
|
cga
|
glycoprotein hormones, alpha polypeptide |
| chr15_-_33172246 | 0.97 |
ENSDART00000158666
|
nbeab
|
neurobeachin b |
| chr1_+_33634186 | 0.91 |
ENSDART00000075626
|
htr1fb
|
5-hydroxytryptamine (serotonin) receptor 1Fb |
| chr3_+_57038033 | 0.88 |
ENSDART00000162930
|
bahcc1a
|
BAH domain and coiled-coil containing 1a |
| chr14_+_44804326 | 0.88 |
ENSDART00000079866
ENSDART00000172974 |
slc30a9
|
solute carrier family 30 (zinc transporter), member 9 |
| chr6_-_29105727 | 0.81 |
ENSDART00000184355
|
fam69ab
|
family with sequence similarity 69, member Ab |
| chr11_-_14813029 | 0.81 |
ENSDART00000004920
ENSDART00000122645 |
sgta
|
small glutamine-rich tetratricopeptide repeat (TPR)-containing, alpha |
| chr2_-_57941037 | 0.72 |
ENSDART00000131420
|
si:dkeyp-68b7.5
|
si:dkeyp-68b7.5 |
| chr23_+_45860676 | 0.67 |
ENSDART00000012234
|
syap1
|
synapse associated protein 1 |
| chr15_-_5720583 | 0.66 |
ENSDART00000158034
ENSDART00000190332 ENSDART00000109599 |
urb1
|
URB1 ribosome biogenesis 1 homolog (S. cerevisiae) |
| chr12_-_22509069 | 0.52 |
ENSDART00000179755
ENSDART00000109707 |
neurl4
|
neuralized E3 ubiquitin protein ligase 4 |
| chr11_-_26180714 | 0.52 |
ENSDART00000173597
|
kaznb
|
kazrin, periplakin interacting protein b |
| chr4_-_72609118 | 0.46 |
ENSDART00000174077
|
si:cabz01054396.2
|
si:cabz01054396.2 |
| chr12_+_34854562 | 0.46 |
ENSDART00000130366
|
si:dkey-21c1.4
|
si:dkey-21c1.4 |
| chr22_-_11520405 | 0.44 |
ENSDART00000063157
|
slc26a11
|
solute carrier family 26 (anion exchanger), member 11 |
| chr13_-_5965423 | 0.42 |
ENSDART00000144931
|
actr2b
|
ARP2 actin related protein 2b homolog |
| chr4_-_72609735 | 0.39 |
ENSDART00000174299
ENSDART00000159227 |
si:cabz01054394.6
|
si:cabz01054394.6 |
| chr21_-_22547496 | 0.28 |
ENSDART00000166835
ENSDART00000089030 |
myo5b
|
myosin VB |
| chr25_-_8625601 | 0.22 |
ENSDART00000155280
|
GDPGP1
|
zgc:153343 |
| chr21_-_40938382 | 0.22 |
ENSDART00000008593
|
yipf5
|
Yip1 domain family, member 5 |
| chr3_-_33934788 | 0.19 |
ENSDART00000151411
|
gtf2f1
|
general transcription factor IIF, polypeptide 1 |
| chr7_-_22941472 | 0.18 |
ENSDART00000190334
|
tnfsf10l
|
TNF superfamily member 10, like |
| chr1_-_51710225 | 0.16 |
ENSDART00000057601
ENSDART00000152745 |
snrpb2
|
small nuclear ribonucleoprotein polypeptide B2 |
| chr11_+_11303458 | 0.13 |
ENSDART00000162486
ENSDART00000160703 |
si:dkey-23f9.4
|
si:dkey-23f9.4 |
| chr24_-_16979728 | 0.11 |
ENSDART00000005331
|
klhl15
|
kelch-like family member 15 |
| chr24_+_22485710 | 0.02 |
ENSDART00000146058
|
si:dkey-40h20.1
|
si:dkey-40h20.1 |
| chr25_-_1333676 | 0.02 |
ENSDART00000183182
ENSDART00000188907 |
cln6b
|
CLN6, transmembrane ER protein b |
Network of associatons between targets according to the STRING database.
First level regulatory network of pou6f1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.5 | 10.1 | GO:0046166 | methylglyoxal biosynthetic process(GO:0019242) glyceraldehyde-3-phosphate biosynthetic process(GO:0046166) |
| 1.9 | 5.7 | GO:0045601 | regulation of endothelial cell differentiation(GO:0045601) |
| 1.5 | 13.1 | GO:0061075 | positive regulation of neural retina development(GO:0061075) positive regulation of retina development in camera-type eye(GO:1902868) |
| 1.3 | 5.3 | GO:0099548 | trans-synaptic signaling by soluble gas(GO:0099543) trans-synaptic signaling by nitric oxide(GO:0099548) |
| 1.0 | 5.2 | GO:0060368 | regulation of Fc receptor mediated stimulatory signaling pathway(GO:0060368) |
| 0.9 | 20.7 | GO:0030388 | fructose 1,6-bisphosphate metabolic process(GO:0030388) |
| 0.8 | 21.2 | GO:0071542 | dopaminergic neuron differentiation(GO:0071542) |
| 0.8 | 9.6 | GO:0031111 | negative regulation of microtubule depolymerization(GO:0007026) negative regulation of microtubule polymerization or depolymerization(GO:0031111) |
| 0.8 | 6.1 | GO:0060976 | coronary vasculature development(GO:0060976) |
| 0.8 | 6.8 | GO:1904861 | excitatory synapse assembly(GO:1904861) |
| 0.7 | 6.5 | GO:0097374 | sensory neuron axon guidance(GO:0097374) |
| 0.7 | 4.2 | GO:1900108 | negative regulation of nodal signaling pathway(GO:1900108) |
| 0.6 | 3.7 | GO:0006772 | thiamine metabolic process(GO:0006772) thiamine-containing compound metabolic process(GO:0042723) |
| 0.5 | 4.8 | GO:0043328 | protein targeting to vacuole involved in ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043328) |
| 0.5 | 14.1 | GO:0070593 | dendrite self-avoidance(GO:0070593) |
| 0.4 | 2.1 | GO:0071205 | protein localization to juxtaparanode region of axon(GO:0071205) protein localization to axon(GO:0099612) |
| 0.4 | 2.4 | GO:0071678 | olfactory bulb axon guidance(GO:0071678) |
| 0.4 | 4.1 | GO:0046548 | retinal rod cell development(GO:0046548) |
| 0.4 | 2.8 | GO:0048569 | post-embryonic organ development(GO:0048569) |
| 0.3 | 1.0 | GO:0010893 | positive regulation of steroid biosynthetic process(GO:0010893) |
| 0.3 | 4.1 | GO:0030816 | activation of adenylate cyclase activity(GO:0007190) positive regulation of cAMP metabolic process(GO:0030816) positive regulation of cAMP biosynthetic process(GO:0030819) positive regulation of adenylate cyclase activity(GO:0045762) |
| 0.3 | 5.1 | GO:0032418 | lysosome localization(GO:0032418) |
| 0.3 | 13.8 | GO:0014046 | dopamine secretion(GO:0014046) regulation of dopamine secretion(GO:0014059) |
| 0.3 | 12.4 | GO:0021761 | limbic system development(GO:0021761) hypothalamus development(GO:0021854) |
| 0.3 | 0.8 | GO:1903332 | regulation of protein folding(GO:1903332) positive regulation of protein folding(GO:1903334) regulation of chaperone-mediated protein folding(GO:1903644) positive regulation of chaperone-mediated protein folding(GO:1903646) |
| 0.2 | 0.7 | GO:0090069 | regulation of ribosome biogenesis(GO:0090069) |
| 0.2 | 7.4 | GO:0007019 | microtubule depolymerization(GO:0007019) |
| 0.2 | 5.5 | GO:0007631 | feeding behavior(GO:0007631) |
| 0.2 | 1.1 | GO:0003232 | bulbus arteriosus development(GO:0003232) |
| 0.2 | 7.4 | GO:0010921 | regulation of phosphatase activity(GO:0010921) |
| 0.2 | 1.5 | GO:0046037 | GMP biosynthetic process(GO:0006177) GMP metabolic process(GO:0046037) |
| 0.1 | 4.9 | GO:0045332 | phospholipid translocation(GO:0045332) |
| 0.1 | 4.3 | GO:0001736 | establishment of planar polarity(GO:0001736) establishment of tissue polarity(GO:0007164) |
| 0.1 | 13.3 | GO:0007605 | sensory perception of sound(GO:0007605) |
| 0.1 | 4.5 | GO:0016339 | calcium-dependent cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0016339) |
| 0.1 | 4.1 | GO:0051085 | chaperone mediated protein folding requiring cofactor(GO:0051085) |
| 0.1 | 3.4 | GO:0048013 | ephrin receptor signaling pathway(GO:0048013) |
| 0.1 | 3.8 | GO:0030307 | positive regulation of cell growth(GO:0030307) TORC1 signaling(GO:0038202) cellular response to amino acid stimulus(GO:0071230) |
| 0.1 | 0.5 | GO:0042776 | mitochondrial ATP synthesis coupled proton transport(GO:0042776) |
| 0.1 | 2.1 | GO:0030497 | fatty acid elongation(GO:0030497) |
| 0.1 | 2.1 | GO:0070831 | basement membrane assembly(GO:0070831) |
| 0.1 | 1.5 | GO:0032968 | positive regulation of transcription elongation from RNA polymerase II promoter(GO:0032968) |
| 0.1 | 0.3 | GO:0003241 | growth involved in heart morphogenesis(GO:0003241) cardiac chamber ballooning(GO:0003242) cardiac muscle tissue growth involved in heart morphogenesis(GO:0003245) |
| 0.1 | 3.7 | GO:0010107 | potassium ion import(GO:0010107) potassium ion import across plasma membrane(GO:1990573) |
| 0.1 | 5.2 | GO:0000381 | regulation of alternative mRNA splicing, via spliceosome(GO:0000381) |
| 0.1 | 0.4 | GO:2001032 | positive regulation of double-strand break repair via homologous recombination(GO:1905168) regulation of double-strand break repair via nonhomologous end joining(GO:2001032) |
| 0.0 | 4.5 | GO:0001666 | response to hypoxia(GO:0001666) response to decreased oxygen levels(GO:0036293) |
| 0.0 | 1.4 | GO:0009648 | photoperiodism(GO:0009648) |
| 0.0 | 0.9 | GO:0007198 | adenylate cyclase-inhibiting serotonin receptor signaling pathway(GO:0007198) serotonin receptor signaling pathway(GO:0007210) |
| 0.0 | 0.9 | GO:0006882 | cellular zinc ion homeostasis(GO:0006882) zinc ion homeostasis(GO:0055069) |
| 0.0 | 7.3 | GO:0006470 | protein dephosphorylation(GO:0006470) |
| 0.0 | 3.5 | GO:0035725 | sodium ion transmembrane transport(GO:0035725) |
| 0.0 | 0.4 | GO:0008272 | sulfate transport(GO:0008272) |
| 0.0 | 0.2 | GO:2001238 | positive regulation of extrinsic apoptotic signaling pathway(GO:2001238) |
| 0.0 | 2.9 | GO:0000377 | RNA splicing, via transesterification reactions(GO:0000375) RNA splicing, via transesterification reactions with bulged adenosine as nucleophile(GO:0000377) mRNA splicing, via spliceosome(GO:0000398) |
| 0.0 | 2.2 | GO:0018105 | peptidyl-serine phosphorylation(GO:0018105) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 9.6 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.5 | 21.2 | GO:0043679 | axon terminus(GO:0043679) |
| 0.5 | 3.8 | GO:0031931 | TORC1 complex(GO:0031931) |
| 0.3 | 4.6 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.3 | 4.8 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 0.3 | 1.4 | GO:1990513 | CLOCK-BMAL transcription complex(GO:1990513) |
| 0.3 | 2.2 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.3 | 5.6 | GO:0031430 | M band(GO:0031430) |
| 0.3 | 0.8 | GO:0072380 | TRC complex(GO:0072380) |
| 0.3 | 2.1 | GO:0033010 | paranodal junction(GO:0033010) |
| 0.3 | 6.1 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.2 | 1.5 | GO:0005674 | transcription factor TFIIF complex(GO:0005674) |
| 0.2 | 3.5 | GO:0031680 | G-protein beta/gamma-subunit complex(GO:0031680) |
| 0.1 | 27.9 | GO:0030424 | axon(GO:0030424) |
| 0.1 | 2.1 | GO:0005605 | basal lamina(GO:0005605) laminin complex(GO:0043256) |
| 0.1 | 4.5 | GO:0016342 | catenin complex(GO:0016342) |
| 0.1 | 5.1 | GO:0031902 | late endosome membrane(GO:0031902) |
| 0.1 | 7.3 | GO:0009898 | cytoplasmic side of plasma membrane(GO:0009898) |
| 0.0 | 3.4 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.0 | 4.9 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 5.7 | GO:0045177 | apical part of cell(GO:0045177) |
| 0.0 | 13.1 | GO:0045202 | synapse(GO:0045202) |
| 0.0 | 0.5 | GO:0045263 | proton-transporting ATP synthase complex, coupling factor F(o)(GO:0045263) |
| 0.0 | 2.7 | GO:0016607 | nuclear speck(GO:0016607) |
| 0.0 | 12.8 | GO:0043005 | neuron projection(GO:0043005) |
| 0.0 | 3.1 | GO:0005911 | cell-cell junction(GO:0005911) |
| 0.0 | 2.1 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.0 | 0.4 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.0 | 1.1 | GO:0005765 | lysosomal membrane(GO:0005765) lytic vacuole membrane(GO:0098852) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.1 | 21.2 | GO:1903136 | cuprous ion binding(GO:1903136) |
| 2.6 | 20.7 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 2.5 | 10.1 | GO:0004807 | triose-phosphate isomerase activity(GO:0004807) methylglyoxal synthase activity(GO:0008929) |
| 1.1 | 5.5 | GO:0031841 | neuropeptide Y receptor binding(GO:0031841) type 2 neuropeptide Y receptor binding(GO:0031843) |
| 0.9 | 3.7 | GO:0050333 | thiamin-triphosphatase activity(GO:0050333) |
| 0.8 | 7.4 | GO:0004865 | protein serine/threonine phosphatase inhibitor activity(GO:0004865) |
| 0.5 | 2.1 | GO:0102345 | 3-hydroxyacyl-CoA dehydratase activity(GO:0018812) 3-hydroxy-behenoyl-CoA dehydratase activity(GO:0102344) 3-hydroxy-lignoceroyl-CoA dehydratase activity(GO:0102345) |
| 0.5 | 6.1 | GO:0015280 | ligand-gated sodium channel activity(GO:0015280) |
| 0.4 | 14.1 | GO:0098632 | protein binding involved in cell-cell adhesion(GO:0098632) |
| 0.4 | 3.7 | GO:0015467 | G-protein activated inward rectifier potassium channel activity(GO:0015467) |
| 0.4 | 1.5 | GO:0001096 | TFIIF-class transcription factor binding(GO:0001096) |
| 0.3 | 5.3 | GO:0004383 | guanylate cyclase activity(GO:0004383) |
| 0.3 | 13.8 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.3 | 2.7 | GO:0030628 | pre-mRNA 3'-splice site binding(GO:0030628) |
| 0.2 | 2.4 | GO:0005021 | vascular endothelial growth factor-activated receptor activity(GO:0005021) |
| 0.2 | 1.4 | GO:0070888 | E-box binding(GO:0070888) |
| 0.2 | 9.6 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.2 | 3.5 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.2 | 3.4 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.1 | 1.5 | GO:0016884 | carbon-nitrogen ligase activity, with glutamine as amido-N-donor(GO:0016884) |
| 0.1 | 4.1 | GO:0051087 | chaperone binding(GO:0051087) |
| 0.1 | 4.5 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.1 | 1.3 | GO:0004029 | aldehyde dehydrogenase (NAD) activity(GO:0004029) |
| 0.1 | 2.2 | GO:0010857 | protein kinase C activity(GO:0004697) calcium-dependent protein kinase C activity(GO:0004698) calcium-dependent protein serine/threonine kinase activity(GO:0009931) calcium-dependent protein kinase activity(GO:0010857) |
| 0.1 | 13.7 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.1 | 5.2 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.1 | 3.8 | GO:0030674 | protein binding, bridging(GO:0030674) |
| 0.1 | 17.7 | GO:0003729 | mRNA binding(GO:0003729) poly(A) RNA binding(GO:0044822) |
| 0.0 | 4.5 | GO:0045296 | cadherin binding(GO:0045296) |
| 0.0 | 0.5 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.0 | 0.9 | GO:0051378 | amine binding(GO:0043176) serotonin binding(GO:0051378) |
| 0.0 | 0.9 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 4.9 | GO:0000287 | magnesium ion binding(GO:0000287) |
| 0.0 | 1.5 | GO:0005347 | ATP transmembrane transporter activity(GO:0005347) |
| 0.0 | 0.2 | GO:0030619 | U1 snRNA binding(GO:0030619) |
| 0.0 | 10.9 | GO:0000989 | transcription factor activity, transcription factor binding(GO:0000989) transcription cofactor activity(GO:0003712) |
| 0.0 | 0.4 | GO:0008271 | secondary active sulfate transmembrane transporter activity(GO:0008271) |
| 0.0 | 7.4 | GO:0015631 | tubulin binding(GO:0015631) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 3.7 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
| 0.1 | 5.2 | SIG BCR SIGNALING PATHWAY | Members of the BCR signaling pathway |
| 0.1 | 1.1 | PID EPHA2 FWD PATHWAY | EPHA2 forward signaling |
| 0.1 | 2.2 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.1 | 3.8 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.1 | 6.1 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.1 | 4.2 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.0 | 2.1 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.0 | 1.1 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 1.0 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 5.2 | REACTOME PD1 SIGNALING | Genes involved in PD-1 signaling |
| 0.4 | 6.1 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.4 | 4.8 | REACTOME MEMBRANE BINDING AND TARGETTING OF GAG PROTEINS | Genes involved in Membrane binding and targetting of GAG proteins |
| 0.4 | 3.8 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.4 | 7.2 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.2 | 1.1 | REACTOME CD28 DEPENDENT VAV1 PATHWAY | Genes involved in CD28 dependent Vav1 pathway |
| 0.1 | 2.8 | REACTOME DOWNREGULATION OF SMAD2 3 SMAD4 TRANSCRIPTIONAL ACTIVITY | Genes involved in Downregulation of SMAD2/3:SMAD4 transcriptional activity |
| 0.1 | 1.0 | REACTOME GLYCOPROTEIN HORMONES | Genes involved in Glycoprotein hormones |
| 0.1 | 4.3 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.1 | 1.5 | REACTOME PURINE RIBONUCLEOSIDE MONOPHOSPHATE BIOSYNTHESIS | Genes involved in Purine ribonucleoside monophosphate biosynthesis |
| 0.1 | 1.5 | REACTOME VIRAL MESSENGER RNA SYNTHESIS | Genes involved in Viral Messenger RNA Synthesis |
| 0.1 | 4.6 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.1 | 4.9 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.1 | 1.3 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.1 | 3.7 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.0 | 0.3 | REACTOME REGULATION OF WATER BALANCE BY RENAL AQUAPORINS | Genes involved in Regulation of Water Balance by Renal Aquaporins |