Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
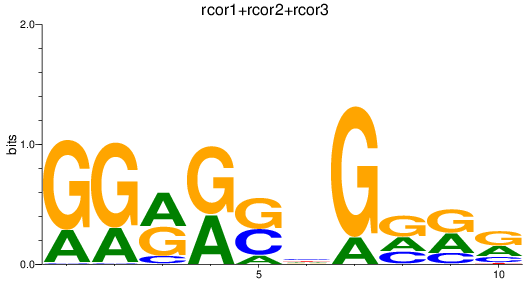
Results for rcor1+rcor2+rcor3
Z-value: 0.80
Transcription factors associated with rcor1+rcor2+rcor3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
rcor3
|
ENSDARG00000004502 | REST corepressor 3 |
|
rcor2
|
ENSDARG00000008278 | REST corepressor 2 |
|
rcor1
|
ENSDARG00000031434 | REST corepressor 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| rcor3 | dr11_v1_chr22_+_38276024_38276024 | 0.50 | 2.1e-07 | Click! |
| rcor2 | dr11_v1_chr7_-_24995631_24995640 | 0.43 | 1.2e-05 | Click! |
| rcor1 | dr11_v1_chr17_-_29271359_29271359 | -0.11 | 3.0e-01 | Click! |
Activity profile of rcor1+rcor2+rcor3 motif
Sorted Z-values of rcor1+rcor2+rcor3 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_-_22535 | 12.61 |
ENSDART00000157877
|
CABZ01092282.1
|
|
| chr16_-_27640995 | 9.17 |
ENSDART00000019658
|
nacad
|
NAC alpha domain containing |
| chr5_+_32228538 | 8.90 |
ENSDART00000077471
|
myhc4
|
myosin heavy chain 4 |
| chr11_-_44543082 | 7.85 |
ENSDART00000099568
|
gpr137bb
|
G protein-coupled receptor 137Bb |
| chr2_-_44282796 | 6.94 |
ENSDART00000163040
ENSDART00000166923 ENSDART00000056372 ENSDART00000109251 ENSDART00000132682 |
mpz
|
myelin protein zero |
| chr16_-_17207754 | 6.47 |
ENSDART00000063804
|
wu:fj39g12
|
wu:fj39g12 |
| chr23_+_44732863 | 5.36 |
ENSDART00000160044
ENSDART00000172268 |
atp1b2a
|
ATPase Na+/K+ transporting subunit beta 2a |
| chr3_-_32818607 | 5.36 |
ENSDART00000075465
|
mylpfa
|
myosin light chain, phosphorylatable, fast skeletal muscle a |
| chr6_+_58543336 | 5.16 |
ENSDART00000157018
|
stmn3
|
stathmin-like 3 |
| chr25_+_21324588 | 4.93 |
ENSDART00000151842
|
lrrn3a
|
leucine rich repeat neuronal 3a |
| chr7_+_58686860 | 4.86 |
ENSDART00000052332
|
penkb
|
proenkephalin b |
| chr11_+_14676236 | 4.84 |
ENSDART00000109308
|
si:ch73-60h1.1
|
si:ch73-60h1.1 |
| chr24_+_32411753 | 4.75 |
ENSDART00000058530
|
neurod6a
|
neuronal differentiation 6a |
| chr8_+_26868105 | 4.69 |
ENSDART00000005337
|
rimkla
|
ribosomal modification protein rimK-like family member A |
| chr3_+_59851537 | 4.63 |
ENSDART00000180997
|
CU693479.1
|
|
| chr4_+_21206283 | 4.52 |
ENSDART00000041861
|
syt1a
|
synaptotagmin Ia |
| chr7_-_33829824 | 4.35 |
ENSDART00000074729
|
uacab
|
uveal autoantigen with coiled-coil domains and ankyrin repeats b |
| chr6_-_58828113 | 3.81 |
ENSDART00000180934
|
kif5ab
|
kinesin family member 5A, b |
| chr8_+_16004154 | 3.71 |
ENSDART00000134787
ENSDART00000172510 ENSDART00000141173 |
elavl4
|
ELAV like neuron-specific RNA binding protein 4 |
| chr13_+_1100197 | 3.61 |
ENSDART00000139560
|
ppp3r1a
|
protein phosphatase 3, regulatory subunit B, alpha a |
| chr22_+_396840 | 3.56 |
ENSDART00000163198
|
capzb
|
capping protein (actin filament) muscle Z-line, beta |
| chr10_-_35542071 | 3.39 |
ENSDART00000162139
|
si:ch211-244c8.4
|
si:ch211-244c8.4 |
| chr16_-_25285469 | 3.35 |
ENSDART00000183943
ENSDART00000191103 ENSDART00000154543 |
prelid3a
|
PRELI domain containing 3A |
| chr22_+_12366516 | 3.34 |
ENSDART00000157802
|
r3hdm1
|
R3H domain containing 1 |
| chr8_+_24861264 | 3.28 |
ENSDART00000099607
|
slc6a17
|
solute carrier family 6 (neutral amino acid transporter), member 17 |
| chr6_-_52156427 | 3.27 |
ENSDART00000082821
|
rims4
|
regulating synaptic membrane exocytosis 4 |
| chr6_-_13408680 | 3.20 |
ENSDART00000151566
|
fmnl2b
|
formin-like 2b |
| chr22_+_10215558 | 3.18 |
ENSDART00000063274
|
kctd6a
|
potassium channel tetramerization domain containing 6a |
| chr16_+_13822137 | 3.13 |
ENSDART00000163251
|
flcn
|
folliculin |
| chr21_+_25643880 | 3.11 |
ENSDART00000192392
ENSDART00000145091 |
tmem151a
|
transmembrane protein 151A |
| chr11_-_44030962 | 3.07 |
ENSDART00000171910
|
FP016005.1
|
|
| chr20_+_52554352 | 3.07 |
ENSDART00000153217
ENSDART00000145230 |
eef1db
|
eukaryotic translation elongation factor 1 delta b (guanine nucleotide exchange protein) |
| chr1_-_25966068 | 3.06 |
ENSDART00000137869
ENSDART00000134192 |
synpo2b
|
synaptopodin 2b |
| chr5_+_13385837 | 2.99 |
ENSDART00000191190
|
ccl19a.1
|
chemokine (C-C motif) ligand 19a, tandem duplicate 1 |
| chr19_-_27966780 | 2.96 |
ENSDART00000110016
|
ube2ql1
|
ubiquitin-conjugating enzyme E2Q family-like 1 |
| chr19_+_6938289 | 2.92 |
ENSDART00000139122
ENSDART00000178832 |
flot1b
|
flotillin 1b |
| chr1_-_19502322 | 2.89 |
ENSDART00000181888
ENSDART00000044030 |
kitb
|
v-kit Hardy-Zuckerman 4 feline sarcoma viral oncogene homolog b |
| chr7_+_528593 | 2.88 |
ENSDART00000091955
|
nrxn2b
|
neurexin 2b |
| chr5_-_32445835 | 2.83 |
ENSDART00000170919
|
ncs1a
|
neuronal calcium sensor 1a |
| chr6_+_22597362 | 2.83 |
ENSDART00000131242
|
cygb2
|
cytoglobin 2 |
| chr11_-_6206520 | 2.79 |
ENSDART00000150199
ENSDART00000148246 ENSDART00000019440 |
pole4
|
polymerase (DNA-directed), epsilon 4, accessory subunit |
| chr15_-_12113045 | 2.69 |
ENSDART00000159879
|
dscaml1
|
Down syndrome cell adhesion molecule like 1 |
| chr3_+_33340939 | 2.67 |
ENSDART00000128786
|
pyya
|
peptide YYa |
| chr24_-_41320037 | 2.65 |
ENSDART00000129058
|
rheb
|
Ras homolog, mTORC1 binding |
| chr6_+_41255485 | 2.64 |
ENSDART00000042683
ENSDART00000186013 |
cadpsb
|
Ca2+-dependent activator protein for secretion b |
| chr25_+_20116407 | 2.63 |
ENSDART00000183615
ENSDART00000193243 |
bpgm
|
2,3-bisphosphoglycerate mutase |
| chr5_+_51102010 | 2.59 |
ENSDART00000110377
|
zgc:194398
|
zgc:194398 |
| chr11_-_32723851 | 2.55 |
ENSDART00000155592
|
pcdh17
|
protocadherin 17 |
| chr4_+_6572364 | 2.53 |
ENSDART00000122574
|
ppp1r3aa
|
protein phosphatase 1, regulatory subunit 3Aa |
| chr6_-_58828398 | 2.46 |
ENSDART00000090634
|
kif5ab
|
kinesin family member 5A, b |
| chr4_-_64703 | 2.46 |
ENSDART00000167851
|
CU856344.1
|
|
| chr25_+_2993855 | 2.46 |
ENSDART00000163009
|
neo1b
|
neogenin 1b |
| chr23_-_14990865 | 2.43 |
ENSDART00000147799
|
ndrg3b
|
ndrg family member 3b |
| chr21_-_26918901 | 2.42 |
ENSDART00000100685
|
lrfn4a
|
leucine rich repeat and fibronectin type III domain containing 4a |
| chr11_+_36231248 | 2.38 |
ENSDART00000131104
|
GPR62 (1 of many)
|
si:ch211-213o11.11 |
| chr1_+_45351890 | 2.38 |
ENSDART00000145486
|
si:ch211-243a20.3
|
si:ch211-243a20.3 |
| chr19_-_27196090 | 2.38 |
ENSDART00000045616
|
gabbr1b
|
gamma-aminobutyric acid (GABA) B receptor, 1b |
| chr4_+_9394426 | 2.35 |
ENSDART00000092013
|
tmtc1
|
transmembrane and tetratricopeptide repeat containing 1 |
| chr1_-_39943596 | 2.34 |
ENSDART00000149730
|
stox2a
|
storkhead box 2a |
| chr5_+_32009956 | 2.32 |
ENSDART00000188482
|
scai
|
suppressor of cancer cell invasion |
| chr1_-_25966411 | 2.30 |
ENSDART00000193375
|
synpo2b
|
synaptopodin 2b |
| chr4_-_9728730 | 2.26 |
ENSDART00000150265
|
mical3b
|
microtubule associated monooxygenase, calponin and LIM domain containing 3b |
| chr11_-_25734417 | 2.19 |
ENSDART00000103570
|
brpf3a
|
bromodomain and PHD finger containing, 3a |
| chr14_-_1949277 | 2.18 |
ENSDART00000159435
|
pcdh2g5
|
protocadherin 2 gamma 5 |
| chr10_-_26179805 | 2.18 |
ENSDART00000174797
|
trim3b
|
tripartite motif containing 3b |
| chr2_+_24867534 | 2.16 |
ENSDART00000158050
|
rab3aa
|
RAB3A, member RAS oncogene family, a |
| chr16_-_24518027 | 2.13 |
ENSDART00000134120
ENSDART00000143761 |
cadm4
|
cell adhesion molecule 4 |
| chr18_+_11970987 | 2.11 |
ENSDART00000144111
|
si:dkeyp-2c8.3
|
si:dkeyp-2c8.3 |
| chr22_-_12160283 | 2.11 |
ENSDART00000146785
ENSDART00000128176 |
tmem163b
|
transmembrane protein 163b |
| chr6_-_42418225 | 2.11 |
ENSDART00000002501
|
ip6k2a
|
inositol hexakisphosphate kinase 2a |
| chr20_-_53321499 | 2.10 |
ENSDART00000179894
ENSDART00000127427 ENSDART00000084952 |
wasf1
|
WAS protein family, member 1 |
| chr20_-_47348116 | 2.08 |
ENSDART00000162087
ENSDART00000160769 ENSDART00000164484 |
dtnba
|
dystrobrevin, beta a |
| chr12_+_48220584 | 2.08 |
ENSDART00000164392
|
lrrc20
|
leucine rich repeat containing 20 |
| chr3_+_60957512 | 1.98 |
ENSDART00000044096
|
helz
|
helicase with zinc finger |
| chr22_-_11724563 | 1.95 |
ENSDART00000190796
ENSDART00000184744 |
krt222
|
keratin 222 |
| chr17_+_41992054 | 1.93 |
ENSDART00000182878
ENSDART00000111537 |
kiz
|
kizuna centrosomal protein |
| chr16_-_12914288 | 1.90 |
ENSDART00000184221
|
cacng8b
|
calcium channel, voltage-dependent, gamma subunit 8b |
| chr22_+_10201826 | 1.90 |
ENSDART00000006513
ENSDART00000132641 |
pdhb
|
pyruvate dehydrogenase E1 beta subunit |
| chr1_+_19535144 | 1.87 |
ENSDART00000103089
|
si:dkey-245p14.4
|
si:dkey-245p14.4 |
| chr7_+_1467863 | 1.87 |
ENSDART00000173433
|
emc4
|
ER membrane protein complex subunit 4 |
| chr7_+_56735195 | 1.87 |
ENSDART00000082830
|
KIAA0895L
|
KIAA0895 like |
| chr22_-_7050 | 1.85 |
ENSDART00000127829
|
atad3
|
ATPase family, AAA domain containing 3 |
| chr2_+_50094873 | 1.84 |
ENSDART00000132307
|
zcchc2
|
zinc finger, CCHC domain containing 2 |
| chr2_+_45374163 | 1.83 |
ENSDART00000193650
|
camsap2b
|
calmodulin regulated spectrin-associated protein family, member 2b |
| chr24_-_26518972 | 1.80 |
ENSDART00000097792
|
tnikb
|
TRAF2 and NCK interacting kinase b |
| chr4_+_11311315 | 1.80 |
ENSDART00000051792
|
sema3aa
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3Aa |
| chr8_+_21629941 | 1.78 |
ENSDART00000140145
|
ajap1
|
adherens junctions associated protein 1 |
| chr8_+_28065803 | 1.77 |
ENSDART00000178481
|
kcnd3
|
potassium voltage-gated channel, Shal-related subfamily, member 3 |
| chr12_+_32159272 | 1.76 |
ENSDART00000153167
|
hlfb
|
hepatic leukemia factor b |
| chr16_+_17389116 | 1.72 |
ENSDART00000103750
ENSDART00000173448 |
fam131bb
|
family with sequence similarity 131, member Bb |
| chr3_+_9379877 | 1.71 |
ENSDART00000182080
|
LO018550.1
|
|
| chr9_-_27649406 | 1.70 |
ENSDART00000181270
ENSDART00000187112 |
stxbp5l
|
syntaxin binding protein 5-like |
| chr17_+_135590 | 1.70 |
ENSDART00000166339
|
klhdc2
|
kelch domain containing 2 |
| chr10_-_1718395 | 1.69 |
ENSDART00000137620
|
si:ch73-46j18.5
|
si:ch73-46j18.5 |
| chr8_+_29749017 | 1.68 |
ENSDART00000185144
|
mapk4
|
mitogen-activated protein kinase 4 |
| chr7_-_38477235 | 1.66 |
ENSDART00000084355
|
zgc:165481
|
zgc:165481 |
| chr20_-_2355357 | 1.65 |
ENSDART00000085281
|
EPB41L2
|
si:ch73-18b11.1 |
| chr3_+_37827373 | 1.65 |
ENSDART00000039517
|
asic2
|
acid-sensing (proton-gated) ion channel 2 |
| chr9_+_21535885 | 1.64 |
ENSDART00000141408
|
arhgef7a
|
Rho guanine nucleotide exchange factor (GEF) 7a |
| chr11_+_11271959 | 1.62 |
ENSDART00000190678
|
ptp4a1
|
protein tyrosine phosphatase type IVA, member 1 |
| chr19_+_7559901 | 1.62 |
ENSDART00000141189
|
pbxip1a
|
pre-B-cell leukemia homeobox interacting protein 1a |
| chr9_+_2393764 | 1.61 |
ENSDART00000172624
|
chn1
|
chimerin 1 |
| chr10_+_21722892 | 1.61 |
ENSDART00000162855
|
pcdh1g13
|
protocadherin 1 gamma 13 |
| chr16_+_29043813 | 1.59 |
ENSDART00000122681
|
nes
|
nestin |
| chr7_-_35379899 | 1.58 |
ENSDART00000183484
|
slc6a2
|
solute carrier family 6 (neurotransmitter transporter), member 2 |
| chr14_-_2369849 | 1.57 |
ENSDART00000180422
ENSDART00000189731 ENSDART00000111748 |
pcdhb
|
protocadherin b |
| chr14_+_34547554 | 1.53 |
ENSDART00000074819
|
gabrp
|
gamma-aminobutyric acid (GABA) A receptor, pi |
| chr19_-_2029777 | 1.52 |
ENSDART00000128639
|
CABZ01071939.1
|
|
| chr20_+_13969414 | 1.51 |
ENSDART00000049864
|
rd3
|
retinal degeneration 3 |
| chr3_-_56933578 | 1.48 |
ENSDART00000192185
|
hid1a
|
HID1 domain containing a |
| chr22_+_110158 | 1.48 |
ENSDART00000143698
|
prkar2ab
|
protein kinase, cAMP-dependent, regulatory, type II, alpha, B |
| chr20_-_26588736 | 1.46 |
ENSDART00000134337
|
exoc2
|
exocyst complex component 2 |
| chr12_+_32729470 | 1.45 |
ENSDART00000175712
|
rbfox3a
|
RNA binding fox-1 homolog 3a |
| chr11_-_19694334 | 1.44 |
ENSDART00000054735
|
SYNPR
|
si:dkey-30j16.3 |
| chr21_+_43561650 | 1.43 |
ENSDART00000085071
|
gpr185a
|
G protein-coupled receptor 185 a |
| chr11_+_45299447 | 1.42 |
ENSDART00000172999
|
mafgb
|
v-maf avian musculoaponeurotic fibrosarcoma oncogene homolog Gb |
| chr23_+_2825940 | 1.41 |
ENSDART00000135781
|
plcg1
|
phospholipase C, gamma 1 |
| chr14_-_2196267 | 1.40 |
ENSDART00000161674
ENSDART00000125674 |
pcdh2ab8
pcdh2ab9
|
protocadherin 2 alpha b 8 protocadherin 2 alpha b 9 |
| chr23_-_45504991 | 1.38 |
ENSDART00000148761
|
col24a1
|
collagen type XXIV alpha 1 |
| chr7_+_39624728 | 1.38 |
ENSDART00000173847
ENSDART00000173845 |
ptpn5
|
protein tyrosine phosphatase, non-receptor type 5 |
| chr25_-_37435060 | 1.37 |
ENSDART00000102855
ENSDART00000148566 |
pdhx
|
pyruvate dehydrogenase complex, component X |
| chr1_-_7894255 | 1.36 |
ENSDART00000167126
ENSDART00000145460 |
radil
|
Ras association and DIL domains |
| chr18_-_44611252 | 1.36 |
ENSDART00000173095
|
spred3
|
sprouty-related, EVH1 domain containing 3 |
| chr15_+_1397811 | 1.36 |
ENSDART00000102125
|
schip1
|
schwannomin interacting protein 1 |
| chr19_+_9533008 | 1.34 |
ENSDART00000104607
|
fam131ba
|
family with sequence similarity 131, member Ba |
| chr17_-_22048233 | 1.34 |
ENSDART00000155203
|
ttbk1b
|
tau tubulin kinase 1b |
| chr7_+_31051213 | 1.34 |
ENSDART00000148347
|
tjp1a
|
tight junction protein 1a |
| chr2_+_59015878 | 1.34 |
ENSDART00000148816
ENSDART00000122795 |
si:ch1073-391i24.1
|
si:ch1073-391i24.1 |
| chr15_-_33807758 | 1.34 |
ENSDART00000158445
|
pds5b
|
PDS5 cohesin associated factor B |
| chr15_+_47418565 | 1.33 |
ENSDART00000155709
|
clpb
|
ClpB homolog, mitochondrial AAA ATPase chaperonin |
| chr14_+_52408619 | 1.32 |
ENSDART00000163856
|
noa1
|
nitric oxide associated 1 |
| chr14_-_237130 | 1.31 |
ENSDART00000164988
|
bod1l1
|
biorientation of chromosomes in cell division 1-like 1 |
| chr9_-_10532591 | 1.30 |
ENSDART00000175269
|
thsd7ba
|
thrombospondin, type I, domain containing 7Ba |
| chr15_+_9327252 | 1.30 |
ENSDART00000144381
|
sgcg
|
sarcoglycan, gamma |
| chr13_-_51922290 | 1.30 |
ENSDART00000168648
|
srfb
|
serum response factor b |
| chr6_+_31684 | 1.29 |
ENSDART00000188853
ENSDART00000184553 |
CZQB01141835.2
|
|
| chr2_-_11258547 | 1.26 |
ENSDART00000165803
ENSDART00000193817 |
slc44a5a
|
solute carrier family 44, member 5a |
| chr22_-_38508112 | 1.24 |
ENSDART00000192004
|
klc4
|
kinesin light chain 4 |
| chr5_+_22098591 | 1.23 |
ENSDART00000143676
|
zc3h12b
|
zinc finger CCCH-type containing 12B |
| chr12_+_31494830 | 1.23 |
ENSDART00000153390
|
gpam
|
glycerol-3-phosphate acyltransferase, mitochondrial |
| chr9_-_54344405 | 1.23 |
ENSDART00000182939
|
CT998556.1
|
|
| chr3_+_22578369 | 1.21 |
ENSDART00000187695
ENSDART00000182678 ENSDART00000112270 |
tanc2a
|
tetratricopeptide repeat, ankyrin repeat and coiled-coil containing 2a |
| chr1_+_36552 | 1.21 |
ENSDART00000169685
|
hikeshi
|
Hikeshi, heat shock protein nuclear import factor |
| chr9_-_23824290 | 1.19 |
ENSDART00000059209
|
wdfy2
|
WD repeat and FYVE domain containing 2 |
| chr12_+_35203091 | 1.18 |
ENSDART00000153022
|
ndst2b
|
N-deacetylase/N-sulfotransferase (heparan glucosaminyl) 2b |
| chr12_-_49168398 | 1.18 |
ENSDART00000186608
|
FO704624.1
|
|
| chr12_-_11768889 | 1.17 |
ENSDART00000057866
|
atoh1c
|
atonal bHLH transcription factor 1c |
| chr22_+_30009926 | 1.15 |
ENSDART00000142529
|
si:dkey-286j15.1
|
si:dkey-286j15.1 |
| chr19_+_9050852 | 1.14 |
ENSDART00000151031
|
ash1l
|
ash1 (absent, small, or homeotic)-like (Drosophila) |
| chr6_-_7726849 | 1.14 |
ENSDART00000151511
|
slc25a38b
|
solute carrier family 25, member 38b |
| chr14_-_1990290 | 1.13 |
ENSDART00000183382
|
pcdh2g5
|
protocadherin 2 gamma 5 |
| chr25_-_32449235 | 1.13 |
ENSDART00000115343
|
atp8b4
|
ATPase phospholipid transporting 8B4 |
| chr3_-_23596532 | 1.13 |
ENSDART00000124921
|
ube2z
|
ubiquitin-conjugating enzyme E2Z |
| chr6_-_60031693 | 1.12 |
ENSDART00000160275
|
CABZ01079262.1
|
|
| chr20_-_46694573 | 1.12 |
ENSDART00000182557
|
AL929435.2
|
|
| chr14_+_18785727 | 1.12 |
ENSDART00000184452
|
si:ch211-111e20.1
|
si:ch211-111e20.1 |
| chr6_-_21873266 | 1.12 |
ENSDART00000151658
ENSDART00000151152 ENSDART00000151179 |
si:dkey-19e4.5
|
si:dkey-19e4.5 |
| chr10_-_5847904 | 1.09 |
ENSDART00000161096
|
ankrd55
|
ankyrin repeat domain 55 |
| chr21_-_43398457 | 1.09 |
ENSDART00000166530
|
ccni2
|
cyclin I family, member 2 |
| chr8_+_11687254 | 1.09 |
ENSDART00000042040
|
atp2a2a
|
ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 2a |
| chr10_-_7347311 | 1.08 |
ENSDART00000168885
|
nrg1
|
neuregulin 1 |
| chr17_-_23412705 | 1.08 |
ENSDART00000126995
|
si:ch211-149k12.3
|
si:ch211-149k12.3 |
| chr6_-_19023468 | 1.07 |
ENSDART00000184729
|
sept9b
|
septin 9b |
| chr3_-_24205339 | 1.06 |
ENSDART00000157135
|
si:dkey-110g7.8
|
si:dkey-110g7.8 |
| chr19_+_48018802 | 1.06 |
ENSDART00000161339
ENSDART00000166978 |
UBE2M
|
si:ch1073-205c8.3 |
| chr19_-_82504 | 1.04 |
ENSDART00000027864
ENSDART00000160560 |
hnrnpr
|
heterogeneous nuclear ribonucleoprotein R |
| chr1_-_59232267 | 1.04 |
ENSDART00000169658
ENSDART00000163257 |
akap8l
|
A kinase (PRKA) anchor protein 8-like |
| chr8_-_13177536 | 1.04 |
ENSDART00000100992
|
zgc:194990
|
zgc:194990 |
| chr14_+_24283915 | 1.03 |
ENSDART00000172868
|
klhl3
|
kelch-like family member 3 |
| chr14_-_2217285 | 1.00 |
ENSDART00000157949
ENSDART00000166150 ENSDART00000054891 ENSDART00000183268 |
pcdh2ab2
pcdh2ab2
|
protocadherin 2 alpha b2 protocadherin 2 alpha b2 |
| chr12_+_44610912 | 1.00 |
ENSDART00000179030
ENSDART00000178008 |
FAM196A (1 of many)
|
family with sequence similarity 196 member A |
| chr8_+_25267903 | 0.98 |
ENSDART00000093090
|
ampd2b
|
adenosine monophosphate deaminase 2b |
| chr15_-_11956981 | 0.98 |
ENSDART00000164163
|
si:dkey-202l22.3
|
si:dkey-202l22.3 |
| chr5_+_53482597 | 0.97 |
ENSDART00000180333
|
BX323994.1
|
|
| chr18_-_44356656 | 0.97 |
ENSDART00000193084
|
prdm10
|
PR domain containing 10 |
| chr20_-_54435287 | 0.96 |
ENSDART00000148632
|
yy1b
|
YY1 transcription factor b |
| chr12_+_18971100 | 0.95 |
ENSDART00000164212
|
l3mbtl2
|
l(3)mbt-like 2 (Drosophila) |
| chr5_+_12888260 | 0.94 |
ENSDART00000175916
|
gal3st1a
|
galactose-3-O-sulfotransferase 1a |
| chr24_-_14711597 | 0.94 |
ENSDART00000131830
|
jph1a
|
junctophilin 1a |
| chr4_-_2014406 | 0.93 |
ENSDART00000180463
|
si:dkey-97m3.1
|
si:dkey-97m3.1 |
| chr17_-_38887424 | 0.93 |
ENSDART00000141177
|
slc24a4a
|
solute carrier family 24 (sodium/potassium/calcium exchanger), member 4a |
| chr20_+_66857 | 0.93 |
ENSDART00000114999
|
lrfn2b
|
leucine rich repeat and fibronectin type III domain containing 2b |
| chr3_-_23596809 | 0.93 |
ENSDART00000156897
|
ube2z
|
ubiquitin-conjugating enzyme E2Z |
| chr21_-_21790372 | 0.91 |
ENSDART00000151094
|
xrra1
|
X-ray radiation resistance associated 1 |
| chr22_+_24157807 | 0.90 |
ENSDART00000159165
|
b3galt2
|
UDP-Gal:betaGlcNAc beta 1,3-galactosyltransferase, polypeptide 2 |
| chr8_+_13106760 | 0.89 |
ENSDART00000029308
|
itgb4
|
integrin, beta 4 |
| chr19_+_43256986 | 0.89 |
ENSDART00000182336
|
DGKD
|
diacylglycerol kinase delta |
| chr16_-_50203058 | 0.88 |
ENSDART00000154570
|
vsig10l
|
V-set and immunoglobulin domain containing 10 like |
| chr16_+_12240605 | 0.88 |
ENSDART00000060056
|
tpi1b
|
triosephosphate isomerase 1b |
| chr12_-_7806007 | 0.87 |
ENSDART00000190359
|
ank3b
|
ankyrin 3b |
| chr23_+_2740741 | 0.87 |
ENSDART00000134938
|
zgc:114123
|
zgc:114123 |
| chr23_-_9965148 | 0.86 |
ENSDART00000137308
|
plxnb1a
|
plexin b1a |
| chr4_-_4250317 | 0.86 |
ENSDART00000103316
|
cd9b
|
CD9 molecule b |
| chr20_-_1560724 | 0.85 |
ENSDART00000179193
ENSDART00000092508 ENSDART00000133369 |
ptprk
|
protein tyrosine phosphatase, receptor type, K |
| chr17_+_49081828 | 0.84 |
ENSDART00000156492
|
tiam2a
|
T cell lymphoma invasion and metastasis 2a |
| chr1_-_7893808 | 0.83 |
ENSDART00000110154
|
radil
|
Ras association and DIL domains |
| chr16_-_47483142 | 0.83 |
ENSDART00000147072
|
cthrc1b
|
collagen triple helix repeat containing 1b |
| chr13_+_41819817 | 0.83 |
ENSDART00000185778
|
CABZ01066611.1
|
|
| chr8_-_22157301 | 0.82 |
ENSDART00000158383
|
nphp4
|
nephronophthisis 4 |
| chr14_-_30808174 | 0.82 |
ENSDART00000173262
|
prss23
|
protease, serine, 23 |
Network of associatons between targets according to the STRING database.
First level regulatory network of rcor1+rcor2+rcor3
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.6 | 7.8 | GO:0045671 | negative regulation of osteoclast differentiation(GO:0045671) |
| 1.5 | 4.4 | GO:1901223 | negative regulation of NIK/NF-kappaB signaling(GO:1901223) |
| 1.1 | 3.3 | GO:0015824 | proline transport(GO:0015824) |
| 1.0 | 2.9 | GO:1901890 | positive regulation of cell junction assembly(GO:1901890) |
| 0.9 | 6.3 | GO:0098971 | dendritic transport(GO:0098935) anterograde dendritic transport(GO:0098937) anterograde dendritic transport of neurotransmitter receptor complex(GO:0098971) |
| 0.7 | 2.9 | GO:0038093 | Fc receptor signaling pathway(GO:0038093) |
| 0.7 | 2.2 | GO:0031630 | regulation of synaptic vesicle fusion to presynaptic membrane(GO:0031630) |
| 0.7 | 3.6 | GO:0010591 | regulation of lamellipodium assembly(GO:0010591) |
| 0.7 | 1.4 | GO:0035477 | regulation of angioblast cell migration involved in selective angioblast sprouting(GO:0035477) |
| 0.6 | 2.4 | GO:0035095 | behavioral response to nicotine(GO:0035095) |
| 0.6 | 1.8 | GO:0071435 | potassium ion export(GO:0071435) potassium ion export across plasma membrane(GO:0097623) |
| 0.5 | 1.6 | GO:0015874 | norepinephrine transport(GO:0015874) |
| 0.4 | 3.1 | GO:0030511 | positive regulation of transforming growth factor beta receptor signaling pathway(GO:0030511) positive regulation of cellular response to transforming growth factor beta stimulus(GO:1903846) |
| 0.4 | 1.7 | GO:1901881 | positive regulation of protein depolymerization(GO:1901881) |
| 0.4 | 1.6 | GO:0021557 | oculomotor nerve development(GO:0021557) |
| 0.4 | 1.1 | GO:1904983 | transmembrane glycine transport from cytosol to mitochondrion(GO:1904983) |
| 0.4 | 1.1 | GO:0036135 | activation of transmembrane receptor protein tyrosine kinase activity(GO:0007171) Schwann cell migration(GO:0036135) cardiac neuron differentiation(GO:0060945) cardiac neuron development(GO:0060959) |
| 0.4 | 2.1 | GO:0003232 | bulbus arteriosus development(GO:0003232) |
| 0.4 | 7.8 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.3 | 1.0 | GO:0072156 | distal tubule morphogenesis(GO:0072156) |
| 0.3 | 2.4 | GO:0045050 | protein insertion into ER membrane by stop-transfer membrane-anchor sequence(GO:0045050) |
| 0.3 | 1.9 | GO:0006086 | acetyl-CoA biosynthetic process from pyruvate(GO:0006086) |
| 0.3 | 1.2 | GO:2000815 | regulation of mRNA stability involved in response to stress(GO:0010610) regulation of mRNA stability involved in response to oxidative stress(GO:2000815) |
| 0.3 | 1.8 | GO:0048755 | branching morphogenesis of a nerve(GO:0048755) |
| 0.3 | 5.4 | GO:0030007 | cellular potassium ion homeostasis(GO:0030007) sodium ion export from cell(GO:0036376) sodium ion export(GO:0071436) |
| 0.3 | 2.5 | GO:0034605 | cellular response to heat(GO:0034605) |
| 0.3 | 6.5 | GO:0007168 | receptor guanylyl cyclase signaling pathway(GO:0007168) |
| 0.2 | 1.6 | GO:1905097 | regulation of guanyl-nucleotide exchange factor activity(GO:1905097) regulation of Rho guanyl-nucleotide exchange factor activity(GO:2001106) |
| 0.2 | 1.4 | GO:0038065 | collagen-activated tyrosine kinase receptor signaling pathway(GO:0038063) collagen-activated signaling pathway(GO:0038065) |
| 0.2 | 1.4 | GO:0010801 | regulation of peptidyl-threonine phosphorylation(GO:0010799) negative regulation of peptidyl-threonine phosphorylation(GO:0010801) |
| 0.2 | 1.4 | GO:0035332 | positive regulation of hippo signaling(GO:0035332) |
| 0.2 | 0.9 | GO:0046166 | methylglyoxal biosynthetic process(GO:0019242) glyceraldehyde-3-phosphate biosynthetic process(GO:0046166) |
| 0.2 | 2.6 | GO:1990504 | dense core granule exocytosis(GO:1990504) |
| 0.2 | 1.4 | GO:0045604 | regulation of epidermal cell differentiation(GO:0045604) |
| 0.2 | 5.4 | GO:0032233 | positive regulation of actin filament bundle assembly(GO:0032233) |
| 0.2 | 2.6 | GO:0044154 | histone H3-K14 acetylation(GO:0044154) |
| 0.2 | 1.3 | GO:0032515 | negative regulation of phosphoprotein phosphatase activity(GO:0032515) |
| 0.2 | 0.8 | GO:2000096 | positive regulation of Wnt signaling pathway, planar cell polarity pathway(GO:2000096) |
| 0.2 | 0.5 | GO:0006864 | pyrimidine nucleotide transport(GO:0006864) mitochondrial pyrimidine nucleotide import(GO:1990519) |
| 0.2 | 0.6 | GO:0060829 | negative regulation of canonical Wnt signaling pathway involved in neural plate anterior/posterior pattern formation(GO:0060829) |
| 0.2 | 1.8 | GO:0031111 | negative regulation of microtubule depolymerization(GO:0007026) negative regulation of microtubule polymerization or depolymerization(GO:0031111) |
| 0.1 | 5.2 | GO:0007019 | microtubule depolymerization(GO:0007019) |
| 0.1 | 1.2 | GO:0016024 | CDP-diacylglycerol biosynthetic process(GO:0016024) CDP-diacylglycerol metabolic process(GO:0046341) |
| 0.1 | 1.0 | GO:0006682 | galactosylceramide biosynthetic process(GO:0006682) galactolipid biosynthetic process(GO:0019375) |
| 0.1 | 2.8 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.1 | 2.0 | GO:0051127 | positive regulation of actin nucleation(GO:0051127) positive regulation of Arp2/3 complex-mediated actin nucleation(GO:2000601) |
| 0.1 | 1.3 | GO:0036058 | filtration diaphragm assembly(GO:0036058) slit diaphragm assembly(GO:0036060) |
| 0.1 | 0.8 | GO:0006542 | glutamine biosynthetic process(GO:0006542) |
| 0.1 | 0.5 | GO:1903589 | regulation of blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:1903587) positive regulation of blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:1903589) |
| 0.1 | 2.5 | GO:0010962 | regulation of glycogen biosynthetic process(GO:0005979) regulation of glucan biosynthetic process(GO:0010962) |
| 0.1 | 1.9 | GO:0098943 | neurotransmitter receptor transport, postsynaptic endosome to lysosome(GO:0098943) |
| 0.1 | 0.8 | GO:2001045 | negative regulation of integrin-mediated signaling pathway(GO:2001045) |
| 0.1 | 9.2 | GO:0006612 | protein targeting to membrane(GO:0006612) |
| 0.1 | 0.9 | GO:0033206 | meiotic cytokinesis(GO:0033206) polar body extrusion after meiotic divisions(GO:0040038) |
| 0.1 | 0.4 | GO:1903723 | negative regulation of centriole elongation(GO:1903723) |
| 0.1 | 3.3 | GO:0006893 | Golgi to plasma membrane transport(GO:0006893) |
| 0.1 | 1.2 | GO:0015014 | heparan sulfate proteoglycan biosynthetic process, polysaccharide chain biosynthetic process(GO:0015014) |
| 0.1 | 1.2 | GO:0021681 | cerebellar granular layer development(GO:0021681) cerebellar granular layer morphogenesis(GO:0021683) cerebellar granular layer formation(GO:0021684) cerebellar granule cell differentiation(GO:0021707) |
| 0.1 | 2.8 | GO:0048752 | semicircular canal morphogenesis(GO:0048752) |
| 0.1 | 0.4 | GO:0001774 | microglial cell activation(GO:0001774) |
| 0.1 | 0.6 | GO:0000395 | mRNA 5'-splice site recognition(GO:0000395) |
| 0.1 | 2.7 | GO:0070593 | dendrite self-avoidance(GO:0070593) |
| 0.1 | 1.5 | GO:0010738 | regulation of protein kinase A signaling(GO:0010738) |
| 0.1 | 1.3 | GO:0045880 | positive regulation of smoothened signaling pathway(GO:0045880) |
| 0.1 | 0.6 | GO:0031507 | heterochromatin assembly(GO:0031507) |
| 0.1 | 2.0 | GO:0032008 | positive regulation of TOR signaling(GO:0032008) |
| 0.1 | 2.7 | GO:0007631 | feeding behavior(GO:0007631) |
| 0.1 | 2.5 | GO:0030513 | positive regulation of BMP signaling pathway(GO:0030513) |
| 0.1 | 0.8 | GO:0003160 | endocardium morphogenesis(GO:0003160) |
| 0.1 | 1.0 | GO:0055090 | acylglycerol homeostasis(GO:0055090) triglyceride homeostasis(GO:0070328) |
| 0.1 | 3.0 | GO:0048247 | lymphocyte chemotaxis(GO:0048247) |
| 0.1 | 2.5 | GO:0060219 | camera-type eye photoreceptor cell differentiation(GO:0060219) |
| 0.1 | 0.9 | GO:0030309 | poly-N-acetyllactosamine metabolic process(GO:0030309) poly-N-acetyllactosamine biosynthetic process(GO:0030311) |
| 0.1 | 1.8 | GO:0003323 | type B pancreatic cell development(GO:0003323) |
| 0.1 | 0.9 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.1 | 0.7 | GO:0006646 | phosphatidylethanolamine biosynthetic process(GO:0006646) |
| 0.1 | 3.1 | GO:0015914 | phospholipid transport(GO:0015914) |
| 0.1 | 1.3 | GO:0007064 | mitotic sister chromatid cohesion(GO:0007064) |
| 0.0 | 0.3 | GO:1904491 | protein localization to ciliary transition zone(GO:1904491) |
| 0.0 | 3.1 | GO:0006414 | translational elongation(GO:0006414) |
| 0.0 | 1.6 | GO:0050708 | regulation of protein secretion(GO:0050708) |
| 0.0 | 0.1 | GO:0099553 | trans-synaptic signaling by lipid, modulating synaptic transmission(GO:0099552) trans-synaptic signaling by endocannabinoid, modulating synaptic transmission(GO:0099553) |
| 0.0 | 2.6 | GO:0006096 | glycolytic process(GO:0006096) |
| 0.0 | 0.9 | GO:1902287 | semaphorin-plexin signaling pathway involved in neuron projection guidance(GO:1902285) semaphorin-plexin signaling pathway involved in axon guidance(GO:1902287) |
| 0.0 | 1.0 | GO:0090502 | RNA phosphodiester bond hydrolysis, endonucleolytic(GO:0090502) |
| 0.0 | 9.3 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.0 | 0.2 | GO:0097355 | protein localization to heterochromatin(GO:0097355) |
| 0.0 | 5.6 | GO:0050769 | positive regulation of neurogenesis(GO:0050769) |
| 0.0 | 0.7 | GO:0032467 | positive regulation of cytokinesis(GO:0032467) |
| 0.0 | 0.3 | GO:0036372 | opsin transport(GO:0036372) |
| 0.0 | 2.0 | GO:0016441 | posttranscriptional gene silencing(GO:0016441) posttranscriptional gene silencing by RNA(GO:0035194) |
| 0.0 | 0.5 | GO:0030206 | chondroitin sulfate biosynthetic process(GO:0030206) |
| 0.0 | 1.0 | GO:0006829 | zinc II ion transport(GO:0006829) |
| 0.0 | 3.0 | GO:0007218 | neuropeptide signaling pathway(GO:0007218) |
| 0.0 | 0.3 | GO:0060114 | vestibular receptor cell differentiation(GO:0060114) vestibular receptor cell development(GO:0060118) |
| 0.0 | 0.8 | GO:0016082 | synaptic vesicle priming(GO:0016082) |
| 0.0 | 0.1 | GO:0006212 | uracil catabolic process(GO:0006212) beta-alanine metabolic process(GO:0019482) beta-alanine biosynthetic process(GO:0019483) uracil metabolic process(GO:0019860) 'de novo' UMP biosynthetic process(GO:0044205) |
| 0.0 | 1.8 | GO:0030866 | cortical actin cytoskeleton organization(GO:0030866) |
| 0.0 | 1.3 | GO:0031532 | actin cytoskeleton reorganization(GO:0031532) |
| 0.0 | 5.1 | GO:0000209 | protein polyubiquitination(GO:0000209) |
| 0.0 | 0.4 | GO:0030324 | respiratory tube development(GO:0030323) lung development(GO:0030324) |
| 0.0 | 1.4 | GO:0034446 | substrate adhesion-dependent cell spreading(GO:0034446) |
| 0.0 | 0.1 | GO:1902808 | positive regulation of cell cycle G1/S phase transition(GO:1902808) |
| 0.0 | 0.5 | GO:0030322 | stabilization of membrane potential(GO:0030322) |
| 0.0 | 0.3 | GO:0006346 | methylation-dependent chromatin silencing(GO:0006346) positive regulation of chromatin silencing(GO:0031937) regulation of methylation-dependent chromatin silencing(GO:0090308) positive regulation of methylation-dependent chromatin silencing(GO:0090309) |
| 0.0 | 0.1 | GO:0071043 | cerebellar Purkinje cell layer structural organization(GO:0021693) cerebellar cortex structural organization(GO:0021698) CUT catabolic process(GO:0071034) CUT metabolic process(GO:0071043) |
| 0.0 | 0.7 | GO:0060037 | pharyngeal system development(GO:0060037) |
| 0.0 | 1.1 | GO:1904029 | regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0000079) regulation of cyclin-dependent protein kinase activity(GO:1904029) |
| 0.0 | 0.7 | GO:0030574 | collagen catabolic process(GO:0030574) multicellular organism catabolic process(GO:0044243) |
| 0.0 | 1.5 | GO:1902476 | chloride transmembrane transport(GO:1902476) |
| 0.0 | 3.0 | GO:0060828 | regulation of canonical Wnt signaling pathway(GO:0060828) |
| 0.0 | 0.6 | GO:0036297 | interstrand cross-link repair(GO:0036297) |
| 0.0 | 1.3 | GO:0018108 | peptidyl-tyrosine phosphorylation(GO:0018108) |
| 0.0 | 1.5 | GO:0000381 | regulation of alternative mRNA splicing, via spliceosome(GO:0000381) |
| 0.0 | 1.6 | GO:0007051 | spindle organization(GO:0007051) |
| 0.0 | 1.1 | GO:0030042 | actin filament depolymerization(GO:0030042) |
| 0.0 | 0.7 | GO:0048484 | enteric nervous system development(GO:0048484) |
| 0.0 | 0.3 | GO:0043491 | protein kinase B signaling(GO:0043491) |
| 0.0 | 1.1 | GO:0008016 | regulation of heart contraction(GO:0008016) |
| 0.0 | 0.2 | GO:0006265 | DNA topological change(GO:0006265) |
| 0.0 | 0.2 | GO:0010569 | regulation of double-strand break repair via homologous recombination(GO:0010569) |
| 0.0 | 0.4 | GO:0060319 | primitive erythrocyte differentiation(GO:0060319) |
| 0.0 | 0.9 | GO:0048916 | posterior lateral line development(GO:0048916) |
| 0.0 | 1.3 | GO:0048545 | response to steroid hormone(GO:0048545) |
| 0.0 | 0.3 | GO:0015701 | bicarbonate transport(GO:0015701) |
| 0.0 | 0.1 | GO:0006363 | termination of RNA polymerase I transcription(GO:0006363) |
| 0.0 | 1.2 | GO:0060047 | heart process(GO:0003015) heart contraction(GO:0060047) |
| 0.0 | 0.6 | GO:0042147 | retrograde transport, endosome to Golgi(GO:0042147) |
| 0.0 | 0.2 | GO:0071108 | protein K48-linked deubiquitination(GO:0071108) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.5 | 9.2 | GO:0005854 | nascent polypeptide-associated complex(GO:0005854) |
| 0.9 | 4.5 | GO:0042584 | chromaffin granule membrane(GO:0042584) |
| 0.8 | 3.2 | GO:0008290 | F-actin capping protein complex(GO:0008290) |
| 0.7 | 2.9 | GO:0016600 | flotillin complex(GO:0016600) |
| 0.7 | 2.8 | GO:0008622 | epsilon DNA polymerase complex(GO:0008622) |
| 0.5 | 5.4 | GO:0090533 | sodium:potassium-exchanging ATPase complex(GO:0005890) cation-transporting ATPase complex(GO:0090533) |
| 0.5 | 3.3 | GO:0045254 | pyruvate dehydrogenase complex(GO:0045254) |
| 0.5 | 1.4 | GO:0005592 | collagen type XI trimer(GO:0005592) |
| 0.4 | 1.8 | GO:0089717 | spanning component of plasma membrane(GO:0044214) spanning component of membrane(GO:0089717) |
| 0.4 | 3.1 | GO:0005853 | eukaryotic translation elongation factor 1 complex(GO:0005853) |
| 0.4 | 6.9 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.3 | 2.4 | GO:0097648 | G-protein coupled receptor dimeric complex(GO:0038037) G-protein coupled receptor heterodimeric complex(GO:0038039) G-protein coupled receptor complex(GO:0097648) |
| 0.3 | 1.5 | GO:0005797 | Golgi medial cisterna(GO:0005797) |
| 0.2 | 2.4 | GO:0072546 | ER membrane protein complex(GO:0072546) |
| 0.2 | 0.9 | GO:0030314 | junctional membrane complex(GO:0030314) |
| 0.2 | 1.3 | GO:0016012 | sarcoglycan complex(GO:0016012) |
| 0.2 | 1.8 | GO:0036449 | microtubule minus-end(GO:0036449) |
| 0.2 | 1.5 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.1 | 2.5 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.1 | 2.6 | GO:0070775 | H3 histone acetyltransferase complex(GO:0070775) MOZ/MORF histone acetyltransferase complex(GO:0070776) |
| 0.1 | 1.1 | GO:0016586 | RSC complex(GO:0016586) |
| 0.1 | 2.1 | GO:0098563 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.1 | 2.1 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.1 | 3.3 | GO:0000145 | exocyst(GO:0000145) |
| 0.1 | 3.3 | GO:0048788 | cytoskeleton of presynaptic active zone(GO:0048788) |
| 0.1 | 4.3 | GO:0044306 | axon terminus(GO:0043679) neuron projection terminus(GO:0044306) |
| 0.1 | 1.6 | GO:0044298 | neuronal cell body membrane(GO:0032809) cell body membrane(GO:0044298) |
| 0.1 | 8.7 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.1 | 1.3 | GO:0000940 | condensed chromosome outer kinetochore(GO:0000940) |
| 0.1 | 3.7 | GO:0043204 | perikaryon(GO:0043204) |
| 0.1 | 0.2 | GO:0031213 | RSF complex(GO:0031213) |
| 0.1 | 2.6 | GO:0098978 | glutamatergic synapse(GO:0098978) |
| 0.1 | 1.2 | GO:0071014 | post-mRNA release spliceosomal complex(GO:0071014) |
| 0.1 | 8.4 | GO:0016459 | myosin complex(GO:0016459) |
| 0.1 | 2.0 | GO:0043186 | P granule(GO:0043186) |
| 0.1 | 5.4 | GO:0030018 | Z disc(GO:0030018) |
| 0.1 | 1.1 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.1 | 7.0 | GO:0005765 | lysosomal membrane(GO:0005765) lytic vacuole membrane(GO:0098852) |
| 0.1 | 2.2 | GO:0031970 | mitochondrial intermembrane space(GO:0005758) organelle envelope lumen(GO:0031970) |
| 0.0 | 1.5 | GO:1902711 | GABA receptor complex(GO:1902710) GABA-A receptor complex(GO:1902711) |
| 0.0 | 1.9 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.0 | 0.8 | GO:0097546 | ciliary base(GO:0097546) |
| 0.0 | 2.5 | GO:0031201 | SNARE complex(GO:0031201) |
| 0.0 | 0.9 | GO:0002116 | semaphorin receptor complex(GO:0002116) |
| 0.0 | 2.2 | GO:0008076 | voltage-gated potassium channel complex(GO:0008076) potassium channel complex(GO:0034705) |
| 0.0 | 0.6 | GO:0036038 | MKS complex(GO:0036038) |
| 0.0 | 3.5 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 5.3 | GO:0008021 | synaptic vesicle(GO:0008021) |
| 0.0 | 1.0 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.0 | 2.4 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.0 | 0.2 | GO:0000126 | transcription factor TFIIIB complex(GO:0000126) |
| 0.0 | 0.9 | GO:0031903 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.0 | 0.6 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.0 | 1.0 | GO:0031519 | PcG protein complex(GO:0031519) |
| 0.0 | 0.4 | GO:0071144 | SMAD2-SMAD3 protein complex(GO:0071144) |
| 0.0 | 0.0 | GO:0070209 | ASTRA complex(GO:0070209) |
| 0.0 | 2.0 | GO:0030424 | axon(GO:0030424) |
| 0.0 | 0.6 | GO:0030139 | endocytic vesicle(GO:0030139) |
| 0.0 | 2.7 | GO:0005874 | microtubule(GO:0005874) |
| 0.0 | 1.1 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 4.7 | GO:0072591 | N-acetyl-L-aspartate-L-glutamate ligase activity(GO:0072590) citrate-L-glutamate ligase activity(GO:0072591) |
| 0.5 | 2.7 | GO:0031843 | neuropeptide Y receptor binding(GO:0031841) type 2 neuropeptide Y receptor binding(GO:0031843) |
| 0.5 | 1.6 | GO:0005329 | dopamine transmembrane transporter activity(GO:0005329) dopamine:sodium symporter activity(GO:0005330) norepinephrine transmembrane transporter activity(GO:0005333) norepinephrine:sodium symporter activity(GO:0005334) |
| 0.5 | 2.6 | GO:0004619 | bisphosphoglycerate mutase activity(GO:0004082) phosphoglycerate mutase activity(GO:0004619) |
| 0.5 | 3.3 | GO:1990050 | phosphatidic acid transporter activity(GO:1990050) |
| 0.5 | 3.3 | GO:0004738 | pyruvate dehydrogenase activity(GO:0004738) |
| 0.4 | 2.8 | GO:0098809 | nitrite reductase activity(GO:0098809) |
| 0.4 | 4.9 | GO:0031628 | opioid receptor binding(GO:0031628) |
| 0.3 | 2.4 | GO:0004965 | G-protein coupled GABA receptor activity(GO:0004965) |
| 0.3 | 1.8 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 0.3 | 5.4 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.3 | 2.4 | GO:0032977 | membrane insertase activity(GO:0032977) |
| 0.3 | 1.6 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.2 | 2.1 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 0.2 | 0.9 | GO:0004807 | triose-phosphate isomerase activity(GO:0004807) methylglyoxal synthase activity(GO:0008929) |
| 0.2 | 1.5 | GO:0008603 | cAMP-dependent protein kinase regulator activity(GO:0008603) |
| 0.2 | 6.3 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.2 | 2.6 | GO:0010484 | H3 histone acetyltransferase activity(GO:0010484) histone acetyltransferase activity (H3-K23 specific)(GO:0043994) |
| 0.2 | 2.5 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.2 | 2.4 | GO:0004169 | dolichyl-phosphate-mannose-protein mannosyltransferase activity(GO:0004169) |
| 0.2 | 0.5 | GO:0015218 | pyrimidine nucleotide transmembrane transporter activity(GO:0015218) |
| 0.2 | 1.1 | GO:0004045 | aminoacyl-tRNA hydrolase activity(GO:0004045) |
| 0.2 | 1.2 | GO:0004366 | glycerol-3-phosphate O-acyltransferase activity(GO:0004366) |
| 0.1 | 1.2 | GO:0015016 | [heparan sulfate]-glucosamine N-sulfotransferase activity(GO:0015016) |
| 0.1 | 0.7 | GO:0004307 | diacylglycerol cholinephosphotransferase activity(GO:0004142) ethanolaminephosphotransferase activity(GO:0004307) |
| 0.1 | 1.8 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.1 | 1.6 | GO:0015280 | ligand-gated sodium channel activity(GO:0015280) |
| 0.1 | 0.9 | GO:0080019 | fatty-acyl-CoA reductase (alcohol-forming) activity(GO:0080019) |
| 0.1 | 2.1 | GO:0000828 | inositol hexakisphosphate kinase activity(GO:0000828) |
| 0.1 | 0.8 | GO:0016211 | glutamate-ammonia ligase activity(GO:0004356) ammonia ligase activity(GO:0016211) acid-ammonia (or amide) ligase activity(GO:0016880) |
| 0.1 | 1.8 | GO:0038191 | neuropilin binding(GO:0038191) |
| 0.1 | 1.0 | GO:0001733 | galactosylceramide sulfotransferase activity(GO:0001733) galactose 3-O-sulfotransferase activity(GO:0050694) |
| 0.1 | 4.5 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.1 | 0.9 | GO:0008273 | calcium, potassium:sodium antiporter activity(GO:0008273) |
| 0.1 | 1.7 | GO:0045159 | myosin II binding(GO:0045159) |
| 0.1 | 5.2 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.1 | 0.5 | GO:0060182 | apelin receptor activity(GO:0060182) |
| 0.1 | 1.2 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.1 | 4.7 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.1 | 1.1 | GO:0030296 | protein tyrosine kinase activator activity(GO:0030296) |
| 0.1 | 1.3 | GO:0051721 | protein phosphatase 2A binding(GO:0051721) |
| 0.1 | 6.5 | GO:0051427 | hormone receptor binding(GO:0051427) |
| 0.1 | 7.9 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.1 | 2.7 | GO:0098632 | protein binding involved in cell-cell adhesion(GO:0098632) |
| 0.1 | 1.1 | GO:0015187 | glycine transmembrane transporter activity(GO:0015187) |
| 0.1 | 2.7 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 0.1 | 2.7 | GO:0019003 | GDP binding(GO:0019003) |
| 0.1 | 3.1 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.1 | 3.0 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.1 | 0.9 | GO:0008532 | N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase activity(GO:0008532) |
| 0.1 | 1.1 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.1 | 3.3 | GO:0044325 | ion channel binding(GO:0044325) |
| 0.1 | 0.5 | GO:0015271 | outward rectifier potassium channel activity(GO:0015271) |
| 0.1 | 0.4 | GO:0016316 | phosphatidylinositol-3,4-bisphosphate 4-phosphatase activity(GO:0016316) |
| 0.1 | 0.5 | GO:0047238 | glucuronosyl-N-acetylgalactosaminyl-proteoglycan 4-beta-N-acetylgalactosaminyltransferase activity(GO:0047238) |
| 0.1 | 1.0 | GO:0004983 | neuropeptide Y receptor activity(GO:0004983) |
| 0.1 | 1.8 | GO:0051020 | GTPase binding(GO:0051020) |
| 0.1 | 0.2 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.0 | 1.7 | GO:0004707 | MAP kinase activity(GO:0004707) |
| 0.0 | 1.5 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 0.0 | 9.4 | GO:0003774 | motor activity(GO:0003774) |
| 0.0 | 1.4 | GO:0004629 | phosphatidylinositol phospholipase C activity(GO:0004435) phospholipase C activity(GO:0004629) |
| 0.0 | 0.8 | GO:0019211 | phosphatase activator activity(GO:0019211) protein phosphatase activator activity(GO:0072542) |
| 0.0 | 2.3 | GO:0016709 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of one atom of oxygen(GO:0016709) |
| 0.0 | 0.1 | GO:0016635 | oxidoreductase activity, acting on the CH-CH group of donors, quinone or related compound as acceptor(GO:0016635) |
| 0.0 | 1.7 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.0 | 0.2 | GO:1990380 | Lys48-specific deubiquitinase activity(GO:1990380) |
| 0.0 | 0.1 | GO:0048487 | beta-tubulin binding(GO:0048487) |
| 0.0 | 0.2 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.0 | 0.2 | GO:0000995 | transcription factor activity, core RNA polymerase III binding(GO:0000995) |
| 0.0 | 0.9 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.0 | 2.3 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.0 | 0.4 | GO:0032041 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.0 | 5.2 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.0 | 13.7 | GO:0003779 | actin binding(GO:0003779) |
| 0.0 | 0.4 | GO:0001608 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.0 | 21.0 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.0 | 0.9 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.1 | GO:0099530 | G-protein coupled receptor activity involved in regulation of postsynaptic membrane potential(GO:0099530) |
| 0.0 | 0.3 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.0 | 0.4 | GO:0070411 | I-SMAD binding(GO:0070411) |
| 0.0 | 1.5 | GO:0016278 | lysine N-methyltransferase activity(GO:0016278) protein-lysine N-methyltransferase activity(GO:0016279) |
| 0.0 | 1.1 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.0 | 0.2 | GO:0003917 | DNA topoisomerase type I activity(GO:0003917) |
| 0.0 | 0.3 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.0 | 2.3 | GO:0003714 | transcription corepressor activity(GO:0003714) |
| 0.0 | 0.7 | GO:0019707 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.0 | 0.4 | GO:0005104 | fibroblast growth factor receptor binding(GO:0005104) |
| 0.0 | 3.7 | GO:0015631 | tubulin binding(GO:0015631) |
| 0.0 | 2.5 | GO:0046982 | protein heterodimerization activity(GO:0046982) |
| 0.0 | 1.3 | GO:0004714 | transmembrane receptor protein tyrosine kinase activity(GO:0004714) |
| 0.0 | 0.1 | GO:0051879 | Hsp90 protein binding(GO:0051879) |
| 0.0 | 0.8 | GO:0019905 | syntaxin binding(GO:0019905) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.4 | PID S1P S1P4 PATHWAY | S1P4 pathway |
| 0.1 | 2.3 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.1 | 2.4 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.1 | 0.9 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.1 | 1.5 | PID GLYPICAN 1PATHWAY | Glypican 1 network |
| 0.1 | 1.1 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.1 | 1.6 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
| 0.0 | 1.6 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 0.0 | 1.5 | PID ARF6 TRAFFICKING PATHWAY | Arf6 trafficking events |
| 0.0 | 0.9 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.0 | 1.4 | PID ATF2 PATHWAY | ATF-2 transcription factor network |
| 0.0 | 0.8 | PID ERBB4 PATHWAY | ErbB4 signaling events |
| 0.0 | 0.8 | PID AR PATHWAY | Coregulation of Androgen receptor activity |
| 0.0 | 0.3 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 2.7 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.4 | 2.7 | REACTOME DSCAM INTERACTIONS | Genes involved in DSCAM interactions |
| 0.4 | 1.4 | REACTOME ROLE OF SECOND MESSENGERS IN NETRIN1 SIGNALING | Genes involved in Role of second messengers in netrin-1 signaling |
| 0.3 | 3.3 | REACTOME REGULATION OF PYRUVATE DEHYDROGENASE PDH COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |
| 0.1 | 1.4 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.1 | 1.6 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.1 | 0.8 | REACTOME RAP1 SIGNALLING | Genes involved in Rap1 signalling |
| 0.1 | 1.5 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.1 | 0.4 | REACTOME SIGNALING BY ACTIVATED POINT MUTANTS OF FGFR1 | Genes involved in Signaling by activated point mutants of FGFR1 |
| 0.1 | 1.2 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.1 | 1.2 | REACTOME SYNTHESIS OF PA | Genes involved in Synthesis of PA |
| 0.1 | 1.4 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.1 | 1.8 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.0 | 0.8 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.0 | 0.7 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.0 | 0.9 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.0 | 0.4 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.0 | 0.4 | REACTOME SYNTHESIS OF PIPS AT THE EARLY ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the early endosome membrane |
| 0.0 | 1.1 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.0 | 3.9 | REACTOME ANTIGEN PROCESSING UBIQUITINATION PROTEASOME DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |
| 0.0 | 2.3 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |
| 0.0 | 0.9 | REACTOME CELL JUNCTION ORGANIZATION | Genes involved in Cell junction organization |
| 0.0 | 1.5 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |