Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
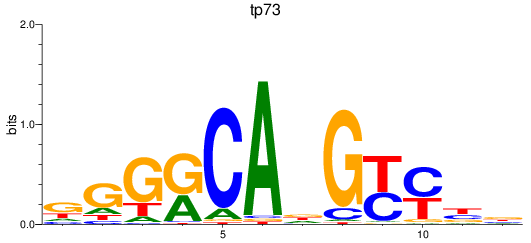
Results for tp73
Z-value: 0.66
Transcription factors associated with tp73
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
tp73
|
ENSDARG00000017953 | tumor protein p73 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| tp73 | dr11_v1_chr8_+_48848200_48848200 | 0.16 | 1.1e-01 | Click! |
Activity profile of tp73 motif
Sorted Z-values of tp73 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr8_+_25767610 | 6.26 |
ENSDART00000062406
|
cacna1sb
|
calcium channel, voltage-dependent, L type, alpha 1S subunit, b |
| chr3_+_16229911 | 6.02 |
ENSDART00000121728
|
rpl19
|
ribosomal protein L19 |
| chr20_-_30376433 | 5.43 |
ENSDART00000190737
|
rps7
|
ribosomal protein S7 |
| chr5_+_40837539 | 4.97 |
ENSDART00000188279
|
si:dkey-3h3.3
|
si:dkey-3h3.3 |
| chr1_-_53714885 | 4.53 |
ENSDART00000026409
|
cct4
|
chaperonin containing TCP1, subunit 4 (delta) |
| chr12_+_22576404 | 4.44 |
ENSDART00000172053
|
capgb
|
capping protein (actin filament), gelsolin-like b |
| chr22_+_8536838 | 4.25 |
ENSDART00000132998
|
si:ch73-27e22.7
|
si:ch73-27e22.7 |
| chr2_+_36691771 | 4.20 |
ENSDART00000131978
|
ptx3b
|
pentraxin 3, long b |
| chr3_-_39488639 | 4.05 |
ENSDART00000161644
|
zgc:100868
|
zgc:100868 |
| chr11_+_12175162 | 4.01 |
ENSDART00000125446
|
si:ch211-156l18.7
|
si:ch211-156l18.7 |
| chr13_-_17464362 | 3.97 |
ENSDART00000145499
|
lrmda
|
leucine rich melanocyte differentiation associated |
| chr21_-_20832482 | 3.92 |
ENSDART00000191928
|
c6
|
complement component 6 |
| chr25_+_34641536 | 3.88 |
ENSDART00000167033
|
CABZ01079011.1
|
|
| chr3_-_39488482 | 3.83 |
ENSDART00000135192
|
zgc:100868
|
zgc:100868 |
| chr15_-_35960250 | 3.78 |
ENSDART00000186765
|
col4a4
|
collagen, type IV, alpha 4 |
| chr18_-_38270077 | 3.78 |
ENSDART00000185546
|
caprin1b
|
cell cycle associated protein 1b |
| chr13_-_4707018 | 3.63 |
ENSDART00000128422
|
oit3
|
oncoprotein induced transcript 3 |
| chr24_+_14859238 | 3.46 |
ENSDART00000113859
ENSDART00000138227 |
crispld1a
|
cysteine-rich secretory protein LCCL domain containing 1a |
| chr14_+_22132896 | 3.45 |
ENSDART00000138274
|
ccng1
|
cyclin G1 |
| chr19_+_4061699 | 3.32 |
ENSDART00000158309
ENSDART00000166512 |
btr25
btr26
|
bloodthirsty-related gene family, member 25 bloodthirsty-related gene family, member 26 |
| chr11_+_14333441 | 3.27 |
ENSDART00000171969
|
ptbp1b
|
polypyrimidine tract binding protein 1b |
| chr3_+_31680592 | 3.19 |
ENSDART00000172456
|
mylk5
|
myosin, light chain kinase 5 |
| chr23_-_20345473 | 3.16 |
ENSDART00000140935
|
si:rp71-17i16.6
|
si:rp71-17i16.6 |
| chr15_-_29387446 | 3.13 |
ENSDART00000145976
ENSDART00000035096 |
serpinh1b
|
serpin peptidase inhibitor, clade H (heat shock protein 47), member 1b |
| chr13_-_17464654 | 3.13 |
ENSDART00000140312
|
lrmda
|
leucine rich melanocyte differentiation associated |
| chr6_-_10902916 | 3.01 |
ENSDART00000122221
|
nfe2l2b
|
nuclear factor, erythroid 2-like 2b |
| chr3_+_8825382 | 2.97 |
ENSDART00000158893
|
si:dkeyp-30d6.2
|
si:dkeyp-30d6.2 |
| chr12_+_5358409 | 2.97 |
ENSDART00000152632
|
plce1
|
phospholipase C, epsilon 1 |
| chr5_+_27898226 | 2.92 |
ENSDART00000098604
ENSDART00000180251 |
adam28
|
ADAM metallopeptidase domain 28 |
| chr14_-_16328331 | 2.86 |
ENSDART00000111262
|
pcdh12
|
protocadherin 12 |
| chr6_-_2154137 | 2.78 |
ENSDART00000162656
|
tgm5l
|
transglutaminase 5, like |
| chr1_+_36651059 | 2.78 |
ENSDART00000187475
|
ednraa
|
endothelin receptor type Aa |
| chr6_-_40885496 | 2.73 |
ENSDART00000189857
|
sirt4
|
sirtuin 4 |
| chr19_+_4068134 | 2.68 |
ENSDART00000158285
|
btr26
|
bloodthirsty-related gene family, member 26 |
| chr20_-_1314355 | 2.56 |
ENSDART00000152436
|
pcmt
|
protein-L-isoaspartate (D-aspartate) O-methyltransferase |
| chr7_+_19495905 | 2.54 |
ENSDART00000125584
ENSDART00000173774 |
si:ch211-212k18.8
|
si:ch211-212k18.8 |
| chr5_+_6854498 | 2.38 |
ENSDART00000148663
|
elac1
|
elaC ribonuclease Z 1 |
| chr21_+_26612777 | 2.23 |
ENSDART00000142667
|
esrra
|
estrogen-related receptor alpha |
| chr1_+_57371447 | 2.22 |
ENSDART00000152229
ENSDART00000181077 |
si:dkey-27j5.3
|
si:dkey-27j5.3 |
| chr16_-_21489514 | 2.14 |
ENSDART00000149525
ENSDART00000148517 ENSDART00000146914 ENSDART00000186493 ENSDART00000193081 ENSDART00000186017 |
mpp6a
|
membrane protein, palmitoylated 6a (MAGUK p55 subfamily member 6) |
| chr18_+_20567542 | 2.13 |
ENSDART00000182585
|
bida
|
BH3 interacting domain death agonist |
| chr1_+_11711503 | 2.09 |
ENSDART00000138355
|
si:dkey-26i13.8
|
si:dkey-26i13.8 |
| chr16_-_19568795 | 2.01 |
ENSDART00000185141
|
abcb5
|
ATP-binding cassette, sub-family B (MDR/TAP), member 5 |
| chr1_+_55600504 | 1.87 |
ENSDART00000123946
|
umod
|
uromodulin |
| chr11_+_30303785 | 1.87 |
ENSDART00000185610
ENSDART00000126815 |
ugt1b5
|
UDP glucuronosyltransferase 1 family, polypeptide B5 |
| chr10_+_2587234 | 1.75 |
ENSDART00000126937
|
wu:fb59d01
|
wu:fb59d01 |
| chr3_+_52545400 | 1.73 |
ENSDART00000184183
|
slc27a1a
|
solute carrier family 27 (fatty acid transporter), member 1a |
| chr7_-_73848458 | 1.69 |
ENSDART00000041385
|
zgc:163061
|
zgc:163061 |
| chr5_-_26936861 | 1.68 |
ENSDART00000191032
|
htra4
|
HtrA serine peptidase 4 |
| chr8_+_30747940 | 1.66 |
ENSDART00000098975
|
p2rx4b
|
purinergic receptor P2X, ligand-gated ion channel, 4b |
| chr7_+_19495379 | 1.62 |
ENSDART00000180514
|
si:ch211-212k18.8
|
si:ch211-212k18.8 |
| chr13_-_50546634 | 1.57 |
ENSDART00000192127
|
CU570781.2
|
|
| chr11_+_14287427 | 1.50 |
ENSDART00000180903
|
si:ch211-262i1.3
|
si:ch211-262i1.3 |
| chr14_+_15484544 | 1.48 |
ENSDART00000188649
|
CR382326.1
|
|
| chr7_-_16596938 | 1.47 |
ENSDART00000134548
|
e2f8
|
E2F transcription factor 8 |
| chr12_-_44151296 | 1.46 |
ENSDART00000168734
|
si:ch73-329n5.3
|
si:ch73-329n5.3 |
| chr15_-_4485828 | 1.46 |
ENSDART00000062868
|
tfdp2
|
transcription factor Dp-2 |
| chr25_+_36292465 | 1.45 |
ENSDART00000152649
|
bmb
|
brambleberry |
| chr7_-_22941472 | 1.40 |
ENSDART00000190334
|
tnfsf10l
|
TNF superfamily member 10, like |
| chr21_-_27338639 | 1.37 |
ENSDART00000130632
|
hif1al2
|
hypoxia-inducible factor 1, alpha subunit, like 2 |
| chr4_-_49607096 | 1.34 |
ENSDART00000154381
|
si:dkey-159n16.4
|
si:dkey-159n16.4 |
| chr7_-_16597130 | 1.34 |
ENSDART00000144118
|
e2f8
|
E2F transcription factor 8 |
| chr20_+_4157815 | 1.32 |
ENSDART00000113132
|
gnpat
|
glyceronephosphate O-acyltransferase |
| chr19_-_35400819 | 1.32 |
ENSDART00000148080
|
rnf19b
|
ring finger protein 19B |
| chr16_+_54829293 | 1.29 |
ENSDART00000024729
|
pabpc1a
|
poly(A) binding protein, cytoplasmic 1a |
| chr6_-_43769332 | 1.27 |
ENSDART00000128174
|
foxp1b
|
forkhead box P1b |
| chr4_+_25680480 | 1.25 |
ENSDART00000100737
|
acot17
|
acyl-CoA thioesterase 17 |
| chr9_-_22240052 | 1.12 |
ENSDART00000111109
|
crygm2d9
|
crystallin, gamma M2d9 |
| chr4_+_25706037 | 1.10 |
ENSDART00000141133
|
lamb1b
|
laminin, beta 1b |
| chr10_-_40479911 | 1.10 |
ENSDART00000136741
|
taar20d1
|
trace amine associated receptor 20d1 |
| chr11_+_16137302 | 1.05 |
ENSDART00000164892
|
pimr204
|
Pim proto-oncogene, serine/threonine kinase, related 204 |
| chr11_-_8271374 | 1.02 |
ENSDART00000168253
|
pimr202
|
Pim proto-oncogene, serine/threonine kinase, related 202 |
| chr23_+_44263986 | 1.00 |
ENSDART00000149194
|
dlg4b
|
discs, large homolog 4b (Drosophila) |
| chr20_-_13673566 | 0.94 |
ENSDART00000181641
|
TAGAP
|
si:ch211-122h15.4 |
| chr1_+_54773877 | 0.88 |
ENSDART00000129831
|
nlrc6
|
NLR family CARD domain containing 6 |
| chr11_-_8208464 | 0.87 |
ENSDART00000161283
|
pimr203
|
Pim proto-oncogene, serine/threonine kinase, related 203 |
| chr8_-_45835056 | 0.82 |
ENSDART00000022242
|
ogdha
|
oxoglutarate (alpha-ketoglutarate) dehydrogenase a (lipoamide) |
| chr13_-_36418921 | 0.81 |
ENSDART00000135804
|
dcaf5
|
ddb1 and cul4 associated factor 5 |
| chr23_-_24047054 | 0.79 |
ENSDART00000184308
ENSDART00000185902 |
CR925720.1
|
|
| chr13_-_9300299 | 0.75 |
ENSDART00000144142
|
HTRA2 (1 of many)
|
si:dkey-33c12.12 |
| chr12_+_26608883 | 0.75 |
ENSDART00000153094
|
si:dkey-148d16.5
|
si:dkey-148d16.5 |
| chr9_-_52847881 | 0.74 |
ENSDART00000166407
|
trpm2
|
transient receptor potential cation channel, subfamily M, member 2 |
| chr6_+_13046720 | 0.73 |
ENSDART00000165896
|
casp8
|
caspase 8, apoptosis-related cysteine peptidase |
| chr20_-_1314537 | 0.65 |
ENSDART00000144288
|
pcmt
|
protein-L-isoaspartate (D-aspartate) O-methyltransferase |
| chr2_-_48539673 | 0.63 |
ENSDART00000168202
|
CR391991.4
|
|
| chr2_+_44528255 | 0.62 |
ENSDART00000193834
|
pask
|
PAS domain containing serine/threonine kinase |
| chr23_+_42254960 | 0.59 |
ENSDART00000102980
|
zcchc11
|
zinc finger, CCHC domain containing 11 |
| chr5_-_1999417 | 0.52 |
ENSDART00000155437
ENSDART00000145781 |
si:ch211-160e1.5
|
si:ch211-160e1.5 |
| chr4_+_71018579 | 0.45 |
ENSDART00000186727
|
si:dkeyp-80d11.10
|
si:dkeyp-80d11.10 |
| chr2_+_23352302 | 0.44 |
ENSDART00000187760
|
rnf2
|
ring finger protein 2 |
| chr8_+_52515188 | 0.42 |
ENSDART00000163668
|
si:ch1073-392o20.2
|
si:ch1073-392o20.2 |
| chr2_-_7185460 | 0.37 |
ENSDART00000092078
|
rc3h1b
|
ring finger and CCCH-type domains 1b |
| chr18_+_44768829 | 0.29 |
ENSDART00000016271
|
ilvbl
|
ilvB (bacterial acetolactate synthase)-like |
| chr7_+_41314862 | 0.25 |
ENSDART00000185198
|
zgc:165532
|
zgc:165532 |
| chr25_+_4581214 | 0.21 |
ENSDART00000185552
|
CABZ01068600.1
|
|
| chr22_+_12477996 | 0.18 |
ENSDART00000177704
|
CR847870.3
|
|
| chr8_+_22485956 | 0.10 |
ENSDART00000099960
|
si:ch211-261n11.7
|
si:ch211-261n11.7 |
| chr17_+_49091661 | 0.08 |
ENSDART00000177166
ENSDART00000177390 ENSDART00000190114 |
tiam2a
|
T cell lymphoma invasion and metastasis 2a |
| chr8_-_34762163 | 0.05 |
ENSDART00000114080
|
setd1bb
|
SET domain containing 1B, b |
Network of associatons between targets according to the STRING database.
First level regulatory network of tp73
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.8 | GO:0033301 | cell cycle comprising mitosis without cytokinesis(GO:0033301) |
| 0.7 | 2.0 | GO:0050787 | response to mercury ion(GO:0046689) detoxification of mercury ion(GO:0050787) |
| 0.6 | 3.0 | GO:0097201 | negative regulation of transcription from RNA polymerase II promoter in response to stress(GO:0097201) |
| 0.6 | 2.4 | GO:0042779 | tRNA 3'-trailer cleavage(GO:0042779) |
| 0.5 | 2.1 | GO:2001244 | positive regulation of intrinsic apoptotic signaling pathway(GO:2001244) |
| 0.5 | 2.8 | GO:0051703 | social behavior(GO:0035176) intraspecies interaction between organisms(GO:0051703) |
| 0.4 | 1.3 | GO:1901503 | ether lipid biosynthetic process(GO:0008611) glycerol ether biosynthetic process(GO:0046504) cellular lipid biosynthetic process(GO:0097384) ether biosynthetic process(GO:1901503) |
| 0.3 | 1.7 | GO:0071072 | heat generation(GO:0031649) regulation of heat generation(GO:0031650) positive regulation of heat generation(GO:0031652) regulation of phospholipid biosynthetic process(GO:0071071) negative regulation of phospholipid biosynthetic process(GO:0071072) |
| 0.3 | 1.0 | GO:0097688 | AMPA glutamate receptor clustering(GO:0097113) glutamate receptor clustering(GO:0097688) |
| 0.3 | 1.5 | GO:0007344 | karyogamy(GO:0000741) pronuclear fusion(GO:0007344) |
| 0.2 | 0.7 | GO:0043903 | regulation of symbiosis, encompassing mutualism through parasitism(GO:0043903) |
| 0.2 | 6.9 | GO:0030199 | collagen fibril organization(GO:0030199) |
| 0.2 | 1.4 | GO:2001238 | positive regulation of extrinsic apoptotic signaling pathway(GO:2001238) |
| 0.1 | 4.4 | GO:0051014 | actin filament severing(GO:0051014) |
| 0.1 | 1.7 | GO:0033198 | response to ATP(GO:0033198) |
| 0.1 | 3.9 | GO:0019835 | cytolysis(GO:0019835) |
| 0.1 | 2.8 | GO:0018149 | peptide cross-linking(GO:0018149) |
| 0.1 | 3.0 | GO:0032836 | glomerular basement membrane development(GO:0032836) |
| 0.1 | 7.1 | GO:0030318 | melanocyte differentiation(GO:0030318) |
| 0.1 | 2.7 | GO:0061035 | regulation of cartilage development(GO:0061035) |
| 0.1 | 0.6 | GO:0045719 | negative regulation of glycogen biosynthetic process(GO:0045719) negative regulation of glycogen metabolic process(GO:0070874) |
| 0.1 | 5.4 | GO:0042274 | ribosomal small subunit biogenesis(GO:0042274) |
| 0.1 | 0.3 | GO:0006549 | isoleucine metabolic process(GO:0006549) isoleucine biosynthetic process(GO:0009097) |
| 0.1 | 3.5 | GO:1904029 | regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0000079) regulation of cyclin-dependent protein kinase activity(GO:1904029) |
| 0.1 | 2.3 | GO:0006471 | protein ADP-ribosylation(GO:0006471) |
| 0.1 | 1.4 | GO:0071456 | cellular response to hypoxia(GO:0071456) |
| 0.1 | 4.5 | GO:0001666 | response to hypoxia(GO:0001666) |
| 0.1 | 6.5 | GO:0070509 | calcium ion import(GO:0070509) |
| 0.0 | 1.1 | GO:0070831 | basement membrane assembly(GO:0070831) |
| 0.0 | 3.8 | GO:0017148 | negative regulation of translation(GO:0017148) |
| 0.0 | 2.9 | GO:0007229 | integrin-mediated signaling pathway(GO:0007229) |
| 0.0 | 1.3 | GO:0032436 | positive regulation of proteasomal ubiquitin-dependent protein catabolic process(GO:0032436) positive regulation of proteasomal protein catabolic process(GO:1901800) |
| 0.0 | 0.8 | GO:0006099 | tricarboxylic acid cycle(GO:0006099) |
| 0.0 | 3.3 | GO:0043484 | regulation of RNA splicing(GO:0043484) |
| 0.0 | 1.7 | GO:0006334 | nucleosome assembly(GO:0006334) |
| 0.0 | 2.5 | GO:0007219 | Notch signaling pathway(GO:0007219) |
| 0.0 | 1.3 | GO:0006417 | regulation of translation(GO:0006417) |
| 0.0 | 2.1 | GO:0044782 | cilium organization(GO:0044782) |
| 0.0 | 2.9 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 4.5 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.2 | 3.9 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.1 | 5.4 | GO:0032040 | small-subunit processome(GO:0032040) |
| 0.1 | 0.3 | GO:0005948 | acetolactate synthase complex(GO:0005948) |
| 0.1 | 6.0 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.1 | 6.3 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.1 | 1.7 | GO:0005639 | integral component of nuclear inner membrane(GO:0005639) intrinsic component of nuclear inner membrane(GO:0031229) nuclear membrane part(GO:0044453) |
| 0.1 | 0.8 | GO:0045252 | oxoglutarate dehydrogenase complex(GO:0045252) |
| 0.1 | 3.5 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.1 | 4.4 | GO:0001726 | ruffle(GO:0001726) |
| 0.1 | 4.9 | GO:0005604 | basement membrane(GO:0005604) |
| 0.0 | 4.0 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 1.0 | GO:0031594 | neuromuscular junction(GO:0031594) |
| 0.0 | 1.3 | GO:0031903 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.0 | 0.8 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.0 | 2.1 | GO:0031514 | motile cilium(GO:0031514) |
| 0.0 | 2.8 | GO:0090575 | RNA polymerase II transcription factor complex(GO:0090575) |
| 0.0 | 0.4 | GO:0071339 | MLL1/2 complex(GO:0044665) MLL1 complex(GO:0071339) |
| 0.0 | 2.1 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.0 | 1.5 | GO:0031965 | nuclear membrane(GO:0031965) |
| 0.0 | 2.7 | GO:0005759 | mitochondrial matrix(GO:0005759) |
| 0.0 | 1.7 | GO:0000786 | nucleosome(GO:0000786) |
| 0.0 | 0.4 | GO:0030140 | trans-Golgi network transport vesicle(GO:0030140) |
| 0.0 | 0.5 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.0 | 15.7 | GO:0005615 | extracellular space(GO:0005615) |
| 0.0 | 2.9 | GO:0005911 | cell-cell junction(GO:0005911) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 2.4 | GO:0042781 | 3'-tRNA processing endoribonuclease activity(GO:0042781) |
| 0.5 | 3.2 | GO:0004719 | protein-L-isoaspartate (D-aspartate) O-methyltransferase activity(GO:0004719) |
| 0.4 | 2.8 | GO:0004962 | endothelin receptor activity(GO:0004962) |
| 0.4 | 2.8 | GO:0001217 | bacterial-type RNA polymerase transcription factor activity, sequence-specific DNA binding(GO:0001130) bacterial-type RNA polymerase transcriptional repressor activity, sequence-specific DNA binding(GO:0001217) |
| 0.2 | 1.7 | GO:0031957 | very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 0.2 | 3.2 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.2 | 6.3 | GO:0008331 | high voltage-gated calcium channel activity(GO:0008331) |
| 0.2 | 0.7 | GO:0072570 | ADP-D-ribose binding(GO:0072570) mono-ADP-D-ribose binding(GO:0072571) |
| 0.2 | 2.7 | GO:0070403 | NAD+ binding(GO:0070403) |
| 0.1 | 2.8 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 0.1 | 1.7 | GO:0035381 | extracellular ATP-gated cation channel activity(GO:0004931) ATP-gated ion channel activity(GO:0035381) |
| 0.1 | 0.8 | GO:0004591 | oxoglutarate dehydrogenase (succinyl-transferring) activity(GO:0004591) |
| 0.1 | 1.3 | GO:0016413 | O-acetyltransferase activity(GO:0016413) |
| 0.1 | 0.3 | GO:0003984 | acetolactate synthase activity(GO:0003984) |
| 0.1 | 1.3 | GO:0008266 | poly(U) RNA binding(GO:0008266) |
| 0.1 | 3.0 | GO:0004629 | phosphatidylinositol phospholipase C activity(GO:0004435) phospholipase C activity(GO:0004629) |
| 0.1 | 3.1 | GO:0005518 | collagen binding(GO:0005518) |
| 0.1 | 8.5 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.1 | 2.1 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.1 | 1.7 | GO:0005520 | insulin-like growth factor binding(GO:0005520) |
| 0.1 | 11.4 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.1 | 3.5 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.1 | 0.7 | GO:0097199 | cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:0097199) |
| 0.0 | 2.2 | GO:0003707 | steroid hormone receptor activity(GO:0003707) |
| 0.0 | 4.5 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.0 | 1.4 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.0 | 0.9 | GO:0047617 | acyl-CoA hydrolase activity(GO:0047617) |
| 0.0 | 7.9 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 1.3 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.0 | 1.9 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.0 | 2.9 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 1.7 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 1.1 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.0 | 1.1 | GO:0001594 | trace-amine receptor activity(GO:0001594) |
| 0.0 | 4.0 | GO:0005198 | structural molecule activity(GO:0005198) |
| 0.0 | 2.0 | GO:0016820 | hydrolase activity, acting on acid anhydrides, catalyzing transmembrane movement of substances(GO:0016820) ATPase activity, coupled to transmembrane movement of substances(GO:0042626) ATPase activity, coupled to movement of substances(GO:0043492) |
| 0.0 | 4.4 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.0 | 3.3 | GO:0044822 | mRNA binding(GO:0003729) poly(A) RNA binding(GO:0044822) |
| 0.0 | 5.9 | GO:0004842 | ubiquitin-protein transferase activity(GO:0004842) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 3.8 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.1 | 3.0 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.1 | 0.7 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.0 | 2.7 | PID HDAC CLASSI PATHWAY | Signaling events mediated by HDAC Class I |
| 0.0 | 4.6 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 3.6 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 1.5 | PID E2F PATHWAY | E2F transcription factor network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 4.5 | REACTOME FORMATION OF TUBULIN FOLDING INTERMEDIATES BY CCT TRIC | Genes involved in Formation of tubulin folding intermediates by CCT/TriC |
| 0.1 | 3.9 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.1 | 5.4 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.1 | 3.8 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.1 | 0.7 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.1 | 6.0 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.1 | 2.0 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.1 | 1.3 | REACTOME PEROXISOMAL LIPID METABOLISM | Genes involved in Peroxisomal lipid metabolism |
| 0.0 | 2.2 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |