Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
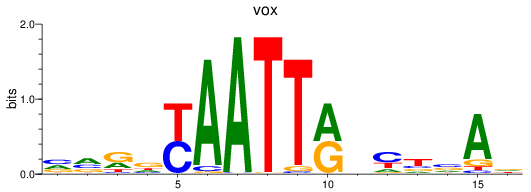
Results for vox
Z-value: 0.52
Transcription factors associated with vox
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
vox
|
ENSDARG00000099761 | ventral homeobox |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| vox | dr11_v1_chr13_-_50624173_50624173 | -0.33 | 1.0e-03 | Click! |
Activity profile of vox motif
Sorted Z-values of vox motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr16_-_17197546 | 6.53 |
ENSDART00000139939
ENSDART00000135146 ENSDART00000063800 ENSDART00000163606 |
gapdh
|
glyceraldehyde-3-phosphate dehydrogenase |
| chr6_-_54815886 | 5.95 |
ENSDART00000180793
ENSDART00000007498 |
tnni1b
|
troponin I type 1b (skeletal, slow) |
| chr23_+_20110086 | 4.37 |
ENSDART00000054664
|
tnnc1b
|
troponin C type 1b (slow) |
| chr18_+_8346920 | 3.84 |
ENSDART00000083421
|
cpt1b
|
carnitine palmitoyltransferase 1B (muscle) |
| chr6_-_7720332 | 3.65 |
ENSDART00000135945
|
rpsa
|
ribosomal protein SA |
| chr22_+_16308450 | 3.27 |
ENSDART00000105678
|
lrrc39
|
leucine rich repeat containing 39 |
| chr25_+_5035343 | 3.05 |
ENSDART00000011751
|
parvb
|
parvin, beta |
| chr10_+_32104305 | 2.95 |
ENSDART00000099880
|
wnt11r
|
wingless-type MMTV integration site family, member 11, related |
| chr5_+_19343880 | 2.59 |
ENSDART00000148130
|
acacb
|
acetyl-CoA carboxylase beta |
| chr7_-_45999333 | 2.56 |
ENSDART00000158603
|
si:ch211-260e23.8
|
si:ch211-260e23.8 |
| chr21_-_32097908 | 2.55 |
ENSDART00000147387
|
si:ch211-160j14.3
|
si:ch211-160j14.3 |
| chr12_+_22580579 | 2.46 |
ENSDART00000171725
ENSDART00000192290 |
capgb
|
capping protein (actin filament), gelsolin-like b |
| chr18_+_48423973 | 2.39 |
ENSDART00000184233
ENSDART00000147074 |
fli1a
|
Fli-1 proto-oncogene, ETS transcription factor a |
| chr5_-_42878178 | 2.31 |
ENSDART00000162981
|
CXCL11 (1 of many)
|
C-X-C motif chemokine ligand 11 |
| chr15_+_41027466 | 2.16 |
ENSDART00000075940
|
mtnr1ba
|
melatonin receptor type 1Ba |
| chr11_-_7380674 | 2.09 |
ENSDART00000014979
ENSDART00000103418 |
vtg3
|
vitellogenin 3, phosvitinless |
| chr6_-_35401282 | 1.96 |
ENSDART00000127612
|
rgs5a
|
regulator of G protein signaling 5a |
| chr2_-_15324837 | 1.61 |
ENSDART00000015655
|
tecrl2b
|
trans-2,3-enoyl-CoA reductase-like 2b |
| chr2_-_48171112 | 1.55 |
ENSDART00000156258
|
pfkpb
|
phosphofructokinase, platelet b |
| chr20_+_26966725 | 1.45 |
ENSDART00000029781
|
ahsa1a
|
AHA1, activator of heat shock protein ATPase homolog 1a |
| chr25_+_19149241 | 1.37 |
ENSDART00000184982
ENSDART00000067324 |
mfge8b
|
milk fat globule-EGF factor 8 protein b |
| chr23_-_7674902 | 1.35 |
ENSDART00000185612
ENSDART00000180524 |
plagl2
|
pleiomorphic adenoma gene-like 2 |
| chr19_+_7735157 | 1.35 |
ENSDART00000186717
|
tuft1b
|
tuftelin 1b |
| chr24_+_19415124 | 1.34 |
ENSDART00000186931
|
sulf1
|
sulfatase 1 |
| chr18_+_1837668 | 1.31 |
ENSDART00000164210
|
CABZ01079192.1
|
|
| chr20_-_47188966 | 1.27 |
ENSDART00000152965
|
si:dkeyp-104f11.9
|
si:dkeyp-104f11.9 |
| chr6_+_28051978 | 1.26 |
ENSDART00000143218
|
si:ch73-194h10.2
|
si:ch73-194h10.2 |
| chr1_+_52392511 | 1.26 |
ENSDART00000144025
|
si:ch211-217k17.8
|
si:ch211-217k17.8 |
| chr21_-_17296789 | 1.23 |
ENSDART00000192180
|
gfi1b
|
growth factor independent 1B transcription repressor |
| chr6_+_19948043 | 1.23 |
ENSDART00000182636
|
pik3r5
|
phosphoinositide-3-kinase, regulatory subunit 5 |
| chr20_-_34663209 | 1.22 |
ENSDART00000132545
|
slc25a24
|
solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 24 |
| chr16_-_7443388 | 1.22 |
ENSDART00000017445
|
prdm1a
|
PR domain containing 1a, with ZNF domain |
| chr5_+_38752287 | 1.20 |
ENSDART00000133571
|
cxcl11.8
|
chemokine (C-X-C motif) ligand 11, duplicate 8 |
| chr3_-_23643751 | 1.20 |
ENSDART00000078425
ENSDART00000140264 |
eve1
|
even-skipped-like1 |
| chr6_-_39649504 | 1.20 |
ENSDART00000179960
ENSDART00000190951 |
larp4ab
|
La ribonucleoprotein domain family, member 4Ab |
| chr8_+_11471350 | 1.18 |
ENSDART00000092355
ENSDART00000136184 |
tjp2b
|
tight junction protein 2b (zona occludens 2) |
| chr5_-_51484156 | 1.16 |
ENSDART00000162064
|
CR388055.1
|
|
| chr17_+_44697604 | 1.14 |
ENSDART00000156625
|
pgfb
|
placental growth factor b |
| chr20_-_49704915 | 1.13 |
ENSDART00000189232
|
COX7A2 (1 of many)
|
cytochrome c oxidase subunit 7A2 |
| chr10_+_21867307 | 1.13 |
ENSDART00000126629
|
cbln17
|
cerebellin 17 |
| chr3_-_15444396 | 1.11 |
ENSDART00000104361
|
si:dkey-56d12.4
|
si:dkey-56d12.4 |
| chr21_+_15713097 | 1.09 |
ENSDART00000015841
|
gstt1b
|
glutathione S-transferase theta 1b |
| chr23_-_35082494 | 1.06 |
ENSDART00000189809
|
BX294434.1
|
|
| chr20_+_35484070 | 1.05 |
ENSDART00000026234
ENSDART00000141675 |
mep1a.2
|
meprin A, alpha (PABA peptide hydrolase), tandem duplicate 2 |
| chr10_-_35149513 | 1.04 |
ENSDART00000063434
ENSDART00000131291 |
ripk4
|
receptor-interacting serine-threonine kinase 4 |
| chr12_+_23812530 | 1.04 |
ENSDART00000066331
|
svila
|
supervillin a |
| chr11_+_2391649 | 1.03 |
ENSDART00000104571
|
igfbp6a
|
insulin-like growth factor binding protein 6a |
| chr17_+_50701748 | 1.02 |
ENSDART00000191938
ENSDART00000183220 ENSDART00000049464 |
fermt2
|
fermitin family member 2 |
| chr15_-_2497568 | 1.00 |
ENSDART00000080398
|
neu4
|
sialidase 4 |
| chr11_+_24703108 | 0.99 |
ENSDART00000159173
|
gpr25
|
G protein-coupled receptor 25 |
| chr20_+_37295006 | 0.96 |
ENSDART00000153137
|
cx23
|
connexin 23 |
| chr20_-_43771871 | 0.94 |
ENSDART00000153304
|
matn3a
|
matrilin 3a |
| chr7_-_34262080 | 0.93 |
ENSDART00000183246
|
si:ch211-98n17.5
|
si:ch211-98n17.5 |
| chr22_-_7457247 | 0.91 |
ENSDART00000106081
|
BX511034.2
|
|
| chr17_+_33311784 | 0.91 |
ENSDART00000156229
|
si:ch211-132f19.7
|
si:ch211-132f19.7 |
| chr9_+_21402863 | 0.90 |
ENSDART00000125357
|
cx30.3
|
connexin 30.3 |
| chr11_+_2391469 | 0.90 |
ENSDART00000182121
|
igfbp6a
|
insulin-like growth factor binding protein 6a |
| chr2_-_48171441 | 0.89 |
ENSDART00000123040
|
pfkpb
|
phosphofructokinase, platelet b |
| chr7_-_23768234 | 0.88 |
ENSDART00000173981
|
si:ch211-200p22.4
|
si:ch211-200p22.4 |
| chr12_-_19119176 | 0.88 |
ENSDART00000149180
|
aco2
|
aconitase 2, mitochondrial |
| chr7_+_48667081 | 0.88 |
ENSDART00000083473
|
trpm5
|
transient receptor potential cation channel, subfamily M, member 5 |
| chr21_+_21812311 | 0.88 |
ENSDART00000151253
|
neu3.4
|
sialidase 3 (membrane sialidase), tandem duplicate 4 |
| chr5_-_29671586 | 0.87 |
ENSDART00000098336
|
spaca9
|
sperm acrosome associated 9 |
| chr6_-_39344259 | 0.86 |
ENSDART00000104074
|
zgc:158846
|
zgc:158846 |
| chr16_-_21799407 | 0.85 |
ENSDART00000123717
|
FP015862.4
|
|
| chr15_-_18200358 | 0.85 |
ENSDART00000158569
|
si:ch211-247l8.8
|
si:ch211-247l8.8 |
| chr1_-_17715493 | 0.81 |
ENSDART00000133027
|
si:dkey-256e7.8
|
si:dkey-256e7.8 |
| chr16_+_54263921 | 0.81 |
ENSDART00000002856
|
drd2l
|
dopamine receptor D2 like |
| chr4_-_67800414 | 0.80 |
ENSDART00000160213
|
si:ch211-66c13.1
|
si:ch211-66c13.1 |
| chr19_+_46158078 | 0.79 |
ENSDART00000183933
ENSDART00000164055 |
cap2
|
CAP, adenylate cyclase-associated protein, 2 (yeast) |
| chr2_-_23411368 | 0.78 |
ENSDART00000159495
|
si:ch73-129a22.11
|
si:ch73-129a22.11 |
| chr22_+_7738966 | 0.77 |
ENSDART00000147073
|
si:ch73-44m9.5
|
si:ch73-44m9.5 |
| chr23_+_41679586 | 0.77 |
ENSDART00000067662
|
CU914487.1
|
|
| chr15_-_11955485 | 0.77 |
ENSDART00000160286
|
si:dkey-202l22.3
|
si:dkey-202l22.3 |
| chr11_-_33868881 | 0.76 |
ENSDART00000163295
ENSDART00000172633 ENSDART00000171439 |
si:ch211-227n13.3
|
si:ch211-227n13.3 |
| chr20_-_53435483 | 0.76 |
ENSDART00000135091
|
mettl24
|
methyltransferase like 24 |
| chr6_-_52348562 | 0.76 |
ENSDART00000142565
ENSDART00000145369 ENSDART00000016890 |
eif6
|
eukaryotic translation initiation factor 6 |
| chr20_+_45853154 | 0.74 |
ENSDART00000181109
|
AL929237.4
|
|
| chr19_-_7690975 | 0.73 |
ENSDART00000151384
|
si:dkey-204a24.10
|
si:dkey-204a24.10 |
| chr4_-_50930346 | 0.73 |
ENSDART00000184245
|
si:ch211-208f21.3
|
si:ch211-208f21.3 |
| chr1_-_49250490 | 0.73 |
ENSDART00000150386
|
si:ch73-6k14.2
|
si:ch73-6k14.2 |
| chr15_-_2493771 | 0.73 |
ENSDART00000184906
|
neu4
|
sialidase 4 |
| chr2_-_10896745 | 0.72 |
ENSDART00000114609
|
cdcp2
|
CUB domain containing protein 2 |
| chr21_-_3672343 | 0.72 |
ENSDART00000086492
|
atp8b1
|
ATPase phospholipid transporting 8B1 |
| chr11_+_31864921 | 0.72 |
ENSDART00000180252
|
diaph3
|
diaphanous-related formin 3 |
| chr3_+_16724614 | 0.71 |
ENSDART00000182135
|
gys1
|
glycogen synthase 1 (muscle) |
| chr2_+_49799470 | 0.69 |
ENSDART00000146325
|
si:ch211-190k17.19
|
si:ch211-190k17.19 |
| chr2_+_33747509 | 0.66 |
ENSDART00000134187
|
si:dkey-31m5.5
|
si:dkey-31m5.5 |
| chr25_-_18953322 | 0.65 |
ENSDART00000155927
|
si:ch211-68a17.7
|
si:ch211-68a17.7 |
| chr7_+_6652967 | 0.65 |
ENSDART00000102681
|
pnp5a
|
purine nucleoside phosphorylase 5a |
| chr4_+_32384950 | 0.64 |
ENSDART00000151910
|
zgc:174698
|
zgc:174698 |
| chr23_+_35713557 | 0.62 |
ENSDART00000123518
|
tuba1c
|
tubulin, alpha 1c |
| chr12_-_7157527 | 0.61 |
ENSDART00000152274
|
si:dkey-187i8.2
|
si:dkey-187i8.2 |
| chr5_-_64883082 | 0.61 |
ENSDART00000064983
ENSDART00000139066 |
krt1-c5
|
keratin, type 1, gene c5 |
| chr1_-_7582859 | 0.60 |
ENSDART00000110696
|
mxb
|
myxovirus (influenza) resistance B |
| chr7_+_25033924 | 0.59 |
ENSDART00000170873
|
sb:cb1058
|
sb:cb1058 |
| chr10_+_28160265 | 0.58 |
ENSDART00000022484
|
rnft1
|
ring finger protein, transmembrane 1 |
| chr5_+_7989210 | 0.58 |
ENSDART00000168071
|
gdnfb
|
glial cell derived neurotrophic factor b |
| chr3_-_36690348 | 0.57 |
ENSDART00000192513
|
myh11b
|
myosin, heavy chain 11b, smooth muscle |
| chr2_-_2451181 | 0.55 |
ENSDART00000101033
|
pth1ra
|
parathyroid hormone 1 receptor a |
| chr1_-_11596829 | 0.55 |
ENSDART00000140725
|
si:dkey-26i13.7
|
si:dkey-26i13.7 |
| chr10_+_11265387 | 0.55 |
ENSDART00000038888
|
hsdl2
|
hydroxysteroid dehydrogenase like 2 |
| chr24_+_6107901 | 0.54 |
ENSDART00000156419
|
si:ch211-37e10.2
|
si:ch211-37e10.2 |
| chr4_+_32385395 | 0.54 |
ENSDART00000109176
|
zgc:174698
|
zgc:174698 |
| chr24_+_38301080 | 0.54 |
ENSDART00000105672
|
mybpc2b
|
myosin binding protein C, fast type b |
| chr20_-_14925281 | 0.51 |
ENSDART00000152641
|
dnm3a
|
dynamin 3a |
| chr5_-_25733745 | 0.51 |
ENSDART00000051566
|
zgc:101016
|
zgc:101016 |
| chr3_-_15999501 | 0.50 |
ENSDART00000160668
|
nme3
|
NME/NM23 nucleoside diphosphate kinase 3 |
| chr10_+_35257651 | 0.50 |
ENSDART00000028940
|
styxl1
|
serine/threonine/tyrosine interacting-like 1 |
| chr6_-_39275793 | 0.49 |
ENSDART00000180477
ENSDART00000148531 |
arhgef25b
|
Rho guanine nucleotide exchange factor (GEF) 25b |
| chr19_-_10971230 | 0.48 |
ENSDART00000166196
|
LO018584.1
|
|
| chr13_+_33688474 | 0.47 |
ENSDART00000161465
|
CABZ01087953.1
|
|
| chr3_-_34586403 | 0.47 |
ENSDART00000151515
|
sept9a
|
septin 9a |
| chr1_-_22691182 | 0.47 |
ENSDART00000076766
|
fgfbp2b
|
fibroblast growth factor binding protein 2b |
| chr22_-_17474781 | 0.46 |
ENSDART00000186817
|
si:ch211-197g15.8
|
si:ch211-197g15.8 |
| chr7_-_48667056 | 0.46 |
ENSDART00000006378
|
cdkn1ca
|
cyclin-dependent kinase inhibitor 1Ca |
| chr22_-_17474583 | 0.45 |
ENSDART00000148027
|
si:ch211-197g15.8
|
si:ch211-197g15.8 |
| chr3_+_22334012 | 0.45 |
ENSDART00000193526
|
ifnphi3
|
interferon phi 3 |
| chr16_+_45425181 | 0.45 |
ENSDART00000168591
|
phf1
|
PHD finger protein 1 |
| chr22_-_22337382 | 0.43 |
ENSDART00000144684
|
si:ch211-129c21.1
|
si:ch211-129c21.1 |
| chr2_+_54798689 | 0.43 |
ENSDART00000183426
|
twsg1a
|
twisted gastrulation BMP signaling modulator 1a |
| chr17_+_17764979 | 0.42 |
ENSDART00000105013
|
alkbh1
|
alkB homolog 1, histone H2A dioxygenase |
| chr11_-_6877973 | 0.42 |
ENSDART00000160271
|
ddx49
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 49 |
| chr25_+_31868268 | 0.42 |
ENSDART00000022325
|
parp16
|
poly (ADP-ribose) polymerase family, member 16 |
| chr15_+_5360407 | 0.42 |
ENSDART00000110420
|
or112-1
|
odorant receptor, family A, subfamily 112, member 1 |
| chr13_+_8677166 | 0.42 |
ENSDART00000181016
ENSDART00000135028 |
prop1
|
PROP paired-like homeobox 1 |
| chr4_+_54645654 | 0.42 |
ENSDART00000192864
|
si:ch211-227e10.1
|
si:ch211-227e10.1 |
| chr14_+_30568961 | 0.41 |
ENSDART00000184303
|
mrpl11
|
mitochondrial ribosomal protein L11 |
| chr7_-_12464412 | 0.40 |
ENSDART00000178723
|
adamtsl3
|
ADAMTS-like 3 |
| chr3_-_34084387 | 0.40 |
ENSDART00000155365
|
ighv4-3
|
immunoglobulin heavy variable 4-3 |
| chr2_-_22363460 | 0.40 |
ENSDART00000158486
|
selenof
|
selenoprotein F |
| chr2_-_39558643 | 0.39 |
ENSDART00000139860
ENSDART00000145231 ENSDART00000141721 |
cbln7
|
cerebellin 7 |
| chr23_-_9991060 | 0.38 |
ENSDART00000111518
|
plxnb1a
|
plexin b1a |
| chr1_-_55263736 | 0.38 |
ENSDART00000152504
ENSDART00000152687 |
si:ch211-286b5.4
|
si:ch211-286b5.4 |
| chr10_-_35257458 | 0.38 |
ENSDART00000143890
ENSDART00000139107 ENSDART00000082445 |
prr11
|
proline rich 11 |
| chr15_-_38154616 | 0.36 |
ENSDART00000099392
|
irgq2
|
immunity-related GTPase family, q2 |
| chr15_+_20344670 | 0.36 |
ENSDART00000158986
|
plekhg2
|
pleckstrin homology domain containing, family G (with RhoGef domain) member 2 |
| chr6_+_30091811 | 0.33 |
ENSDART00000088403
|
meltf
|
melanotransferrin |
| chr10_-_8046764 | 0.32 |
ENSDART00000099031
|
zgc:136254
|
zgc:136254 |
| chr12_-_314899 | 0.32 |
ENSDART00000066579
|
pts
|
6-pyruvoyltetrahydropterin synthase |
| chr5_+_28160503 | 0.32 |
ENSDART00000051516
|
tacr1a
|
tachykinin receptor 1a |
| chr23_+_45966436 | 0.32 |
ENSDART00000172160
|
CABZ01069338.1
|
|
| chr4_-_41712014 | 0.31 |
ENSDART00000138165
|
znf976
|
zinc finger protein 976 |
| chr13_-_45811137 | 0.30 |
ENSDART00000190512
|
fndc5a
|
fibronectin type III domain containing 5a |
| chr10_+_3049636 | 0.30 |
ENSDART00000081794
ENSDART00000183167 ENSDART00000191634 ENSDART00000183514 |
rasgrf2a
|
Ras protein-specific guanine nucleotide-releasing factor 2a |
| chr25_-_21507676 | 0.29 |
ENSDART00000010706
|
immp2l
|
inner mitochondrial membrane peptidase subunit 2 |
| chr7_-_44957503 | 0.29 |
ENSDART00000165077
|
cdh16
|
cadherin 16, KSP-cadherin |
| chr20_-_14924858 | 0.28 |
ENSDART00000047039
|
dnm3a
|
dynamin 3a |
| chr16_+_42471455 | 0.28 |
ENSDART00000166640
|
si:ch211-215k15.5
|
si:ch211-215k15.5 |
| chr16_+_45424907 | 0.28 |
ENSDART00000134471
|
phf1
|
PHD finger protein 1 |
| chr7_-_22699716 | 0.27 |
ENSDART00000193773
|
BX470211.5
|
|
| chr18_+_35173683 | 0.25 |
ENSDART00000192545
|
cfap45
|
cilia and flagella associated protein 45 |
| chr20_-_37813863 | 0.23 |
ENSDART00000147529
|
batf3
|
basic leucine zipper transcription factor, ATF-like 3 |
| chr22_+_6905210 | 0.21 |
ENSDART00000167530
|
CABZ01065328.2
|
|
| chr1_+_57311901 | 0.19 |
ENSDART00000149397
|
mycbpap
|
mycbp associated protein |
| chr8_+_17078692 | 0.19 |
ENSDART00000023206
|
plk2b
|
polo-like kinase 2b (Drosophila) |
| chr6_+_19383267 | 0.19 |
ENSDART00000166549
|
mchr1a
|
melanin-concentrating hormone receptor 1a |
| chr9_+_43799829 | 0.18 |
ENSDART00000186240
|
ube2e3
|
ubiquitin-conjugating enzyme E2E 3 (UBC4/5 homolog, yeast) |
| chr10_-_5847655 | 0.18 |
ENSDART00000192773
|
ankrd55
|
ankyrin repeat domain 55 |
| chr1_+_27024068 | 0.16 |
ENSDART00000102322
|
bnc2
|
basonuclin 2 |
| chr4_+_49322310 | 0.13 |
ENSDART00000184154
ENSDART00000167162 |
BX942819.1
|
|
| chr19_-_10323845 | 0.08 |
ENSDART00000151259
ENSDART00000151821 |
u2af2b
|
U2 small nuclear RNA auxiliary factor 2b |
| chr14_+_30762131 | 0.08 |
ENSDART00000145039
|
si:ch211-145o7.3
|
si:ch211-145o7.3 |
| chr5_+_56268436 | 0.08 |
ENSDART00000021159
|
lhx1b
|
LIM homeobox 1b |
| chr17_+_16046314 | 0.04 |
ENSDART00000154554
ENSDART00000154338 ENSDART00000155336 |
si:ch73-204p21.2
|
si:ch73-204p21.2 |
| chr11_+_11303458 | 0.02 |
ENSDART00000162486
ENSDART00000160703 |
si:dkey-23f9.4
|
si:dkey-23f9.4 |
| chr8_-_39822917 | 0.01 |
ENSDART00000067843
|
zgc:162025
|
zgc:162025 |
| chr21_-_40317035 | 0.00 |
ENSDART00000143648
ENSDART00000013359 |
or101-1
|
odorant receptor, family B, subfamily 101, member 1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of vox
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 2.6 | GO:2001295 | malonyl-CoA biosynthetic process(GO:2001295) |
| 0.3 | 0.8 | GO:0042256 | mature ribosome assembly(GO:0042256) assembly of large subunit precursor of preribosome(GO:1902626) |
| 0.2 | 2.4 | GO:0061620 | glucose catabolic process(GO:0006007) NADH regeneration(GO:0006735) glycolytic process through fructose-6-phosphate(GO:0061615) glycolytic process through glucose-6-phosphate(GO:0061620) canonical glycolysis(GO:0061621) glucose catabolic process to pyruvate(GO:0061718) |
| 0.2 | 2.6 | GO:0009313 | oligosaccharide catabolic process(GO:0009313) |
| 0.2 | 3.8 | GO:0009437 | carnitine metabolic process(GO:0009437) |
| 0.2 | 10.3 | GO:0003009 | skeletal muscle contraction(GO:0003009) |
| 0.2 | 0.6 | GO:0061213 | regulation of morphogenesis of a branching structure(GO:0060688) positive regulation of mesonephros development(GO:0061213) regulation of mesonephros development(GO:0061217) regulation of kidney development(GO:0090183) positive regulation of kidney development(GO:0090184) regulation of branching involved in ureteric bud morphogenesis(GO:0090189) positive regulation of branching involved in ureteric bud morphogenesis(GO:0090190) |
| 0.2 | 6.5 | GO:0050821 | protein stabilization(GO:0050821) |
| 0.2 | 1.9 | GO:0043567 | regulation of insulin-like growth factor receptor signaling pathway(GO:0043567) |
| 0.2 | 1.3 | GO:2000290 | smoothened signaling pathway involved in ventral spinal cord patterning(GO:0021910) regulation of myotome development(GO:2000290) |
| 0.2 | 3.7 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 0.1 | 4.3 | GO:0055008 | cardiac muscle tissue morphogenesis(GO:0055008) |
| 0.1 | 1.2 | GO:0042661 | regulation of mesodermal cell fate specification(GO:0042661) |
| 0.1 | 1.4 | GO:0032781 | positive regulation of ATPase activity(GO:0032781) |
| 0.1 | 3.1 | GO:0030032 | lamellipodium assembly(GO:0030032) |
| 0.1 | 0.5 | GO:2001244 | positive regulation of intrinsic apoptotic signaling pathway(GO:2001244) |
| 0.1 | 1.2 | GO:0001709 | cell fate determination(GO:0001709) |
| 0.1 | 0.6 | GO:1904292 | regulation of ERAD pathway(GO:1904292) |
| 0.1 | 3.5 | GO:0051014 | actin filament severing(GO:0051014) |
| 0.1 | 0.6 | GO:0097033 | respiratory chain complex III assembly(GO:0017062) mitochondrial respiratory chain complex III assembly(GO:0034551) mitochondrial respiratory chain complex III biogenesis(GO:0097033) |
| 0.1 | 0.3 | GO:0071314 | response to cocaine(GO:0042220) cellular response to alkaloid(GO:0071312) cellular response to cocaine(GO:0071314) |
| 0.1 | 1.1 | GO:0060754 | mast cell chemotaxis(GO:0002551) regulation of mast cell chemotaxis(GO:0060753) positive regulation of mast cell chemotaxis(GO:0060754) mast cell migration(GO:0097531) |
| 0.1 | 0.4 | GO:0010693 | regulation of alkaline phosphatase activity(GO:0010692) negative regulation of alkaline phosphatase activity(GO:0010693) |
| 0.1 | 0.4 | GO:0035513 | oxidative RNA demethylation(GO:0035513) |
| 0.1 | 0.9 | GO:0050909 | sensory perception of taste(GO:0050909) |
| 0.1 | 2.3 | GO:0016185 | synaptic vesicle budding from presynaptic endocytic zone membrane(GO:0016185) |
| 0.1 | 0.5 | GO:0045736 | negative regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0045736) negative regulation of cyclin-dependent protein kinase activity(GO:1904030) |
| 0.1 | 0.8 | GO:0051481 | negative regulation of cytosolic calcium ion concentration(GO:0051481) negative regulation of synaptic transmission, glutamatergic(GO:0051967) |
| 0.1 | 2.1 | GO:0032355 | response to estradiol(GO:0032355) |
| 0.1 | 1.2 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.0 | 1.6 | GO:0042761 | very long-chain fatty acid biosynthetic process(GO:0042761) |
| 0.0 | 0.6 | GO:2000574 | regulation of microtubule motor activity(GO:2000574) |
| 0.0 | 0.7 | GO:0031060 | regulation of histone methylation(GO:0031060) |
| 0.0 | 0.3 | GO:0006627 | protein processing involved in protein targeting to mitochondrion(GO:0006627) |
| 0.0 | 1.1 | GO:0006749 | glutathione metabolic process(GO:0006749) |
| 0.0 | 0.3 | GO:0006729 | tetrahydrobiopterin biosynthetic process(GO:0006729) tetrahydrobiopterin metabolic process(GO:0046146) |
| 0.0 | 0.5 | GO:0046051 | UTP biosynthetic process(GO:0006228) UTP metabolic process(GO:0046051) |
| 0.0 | 0.8 | GO:0019933 | cAMP-mediated signaling(GO:0019933) |
| 0.0 | 1.3 | GO:0030050 | vesicle transport along actin filament(GO:0030050) actin filament-based transport(GO:0099515) |
| 0.0 | 0.9 | GO:0006099 | tricarboxylic acid cycle(GO:0006099) |
| 0.0 | 1.1 | GO:0045766 | positive regulation of angiogenesis(GO:0045766) |
| 0.0 | 1.2 | GO:0034599 | cellular response to oxidative stress(GO:0034599) |
| 0.0 | 0.7 | GO:0045332 | phospholipid translocation(GO:0045332) |
| 0.0 | 1.4 | GO:0007338 | single fertilization(GO:0007338) |
| 0.0 | 1.2 | GO:0070098 | chemokine-mediated signaling pathway(GO:0070098) |
| 0.0 | 0.4 | GO:0021984 | adenohypophysis development(GO:0021984) |
| 0.0 | 0.4 | GO:1902287 | semaphorin-plexin signaling pathway involved in neuron projection guidance(GO:1902285) semaphorin-plexin signaling pathway involved in axon guidance(GO:1902287) |
| 0.0 | 0.2 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
| 0.0 | 1.2 | GO:0043406 | positive regulation of MAP kinase activity(GO:0043406) |
| 0.0 | 0.7 | GO:0009250 | glycogen biosynthetic process(GO:0005978) glucan biosynthetic process(GO:0009250) |
| 0.0 | 0.4 | GO:0006471 | protein ADP-ribosylation(GO:0006471) |
| 0.0 | 0.3 | GO:0006826 | iron ion transport(GO:0006826) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 10.3 | GO:0005861 | troponin complex(GO:0005861) |
| 0.2 | 2.4 | GO:0005945 | 6-phosphofructokinase complex(GO:0005945) |
| 0.2 | 1.0 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.2 | 3.3 | GO:0031430 | M band(GO:0031430) |
| 0.1 | 1.2 | GO:0005944 | phosphatidylinositol 3-kinase complex, class IB(GO:0005944) phosphatidylinositol 3-kinase complex, class I(GO:0097651) |
| 0.1 | 0.9 | GO:0001669 | acrosomal vesicle(GO:0001669) |
| 0.1 | 0.9 | GO:0098894 | presynaptic endocytic zone(GO:0098833) presynaptic endocytic zone membrane(GO:0098835) extrinsic component of presynaptic membrane(GO:0098888) extrinsic component of presynaptic endocytic zone(GO:0098894) |
| 0.1 | 0.9 | GO:0045095 | keratin filament(GO:0045095) |
| 0.1 | 3.7 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.1 | 1.4 | GO:0098844 | postsynaptic endocytic zone membrane(GO:0098844) |
| 0.1 | 0.3 | GO:0042720 | mitochondrial inner membrane peptidase complex(GO:0042720) |
| 0.1 | 5.5 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.0 | 0.3 | GO:0097225 | sperm midpiece(GO:0097225) |
| 0.0 | 0.4 | GO:0071782 | endoplasmic reticulum tubular network(GO:0071782) endoplasmic reticulum subcompartment(GO:0098827) |
| 0.0 | 3.5 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.0 | 0.8 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.0 | 1.3 | GO:0005902 | microvillus(GO:0005902) |
| 0.0 | 0.8 | GO:0098978 | glutamatergic synapse(GO:0098978) |
| 0.0 | 0.4 | GO:0002116 | semaphorin receptor complex(GO:0002116) |
| 0.0 | 0.5 | GO:0032156 | septin ring(GO:0005940) septin complex(GO:0031105) septin cytoskeleton(GO:0032156) |
| 0.0 | 4.4 | GO:0031966 | mitochondrial membrane(GO:0031966) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 6.5 | GO:0043891 | glyceraldehyde-3-phosphate dehydrogenase (NAD+) (phosphorylating) activity(GO:0004365) glyceraldehyde-3-phosphate dehydrogenase (NAD(P)+) (phosphorylating) activity(GO:0043891) |
| 0.6 | 2.6 | GO:0003989 | acetyl-CoA carboxylase activity(GO:0003989) |
| 0.5 | 3.8 | GO:0004095 | carnitine O-palmitoyltransferase activity(GO:0004095) O-palmitoyltransferase activity(GO:0016416) |
| 0.4 | 2.2 | GO:0008502 | melatonin receptor activity(GO:0008502) |
| 0.3 | 2.1 | GO:0045735 | nutrient reservoir activity(GO:0045735) |
| 0.3 | 0.9 | GO:0003994 | aconitate hydratase activity(GO:0003994) |
| 0.3 | 0.8 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 0.2 | 2.6 | GO:0052795 | exo-alpha-(2->3)-sialidase activity(GO:0052794) exo-alpha-(2->6)-sialidase activity(GO:0052795) exo-alpha-(2->8)-sialidase activity(GO:0052796) |
| 0.2 | 2.4 | GO:0003872 | 6-phosphofructokinase activity(GO:0003872) |
| 0.2 | 1.0 | GO:0060182 | apelin receptor activity(GO:0060182) |
| 0.2 | 0.7 | GO:0004373 | glycogen (starch) synthase activity(GO:0004373) |
| 0.2 | 1.9 | GO:0031994 | insulin-like growth factor I binding(GO:0031994) insulin-like growth factor II binding(GO:0031995) |
| 0.2 | 3.7 | GO:0043236 | laminin binding(GO:0043236) |
| 0.2 | 4.4 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.1 | 0.8 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 0.1 | 0.6 | GO:0030116 | glial cell-derived neurotrophic factor receptor binding(GO:0030116) |
| 0.1 | 1.2 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.1 | 0.3 | GO:0016496 | substance P receptor activity(GO:0016496) |
| 0.1 | 0.4 | GO:0035516 | oxidative DNA demethylase activity(GO:0035516) |
| 0.1 | 1.1 | GO:0005172 | vascular endothelial growth factor receptor binding(GO:0005172) |
| 0.1 | 1.4 | GO:0051879 | Hsp90 protein binding(GO:0051879) |
| 0.1 | 0.4 | GO:0008430 | selenium binding(GO:0008430) |
| 0.1 | 1.3 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 0.1 | 0.6 | GO:0004731 | purine-nucleoside phosphorylase activity(GO:0004731) |
| 0.1 | 0.9 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.1 | 0.4 | GO:0070180 | large ribosomal subunit rRNA binding(GO:0070180) |
| 0.1 | 0.2 | GO:0030273 | melanin-concentrating hormone receptor activity(GO:0030273) |
| 0.1 | 0.5 | GO:0004861 | cyclin-dependent protein serine/threonine kinase inhibitor activity(GO:0004861) |
| 0.1 | 0.8 | GO:0001591 | dopamine neurotransmitter receptor activity, coupled via Gi/Go(GO:0001591) |
| 0.0 | 0.8 | GO:0004983 | neuropeptide Y receptor activity(GO:0004983) |
| 0.0 | 1.2 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.0 | 0.3 | GO:0016838 | carbon-oxygen lyase activity, acting on phosphates(GO:0016838) |
| 0.0 | 0.9 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.0 | 1.1 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.0 | 0.9 | GO:0005227 | calcium activated cation channel activity(GO:0005227) |
| 0.0 | 2.6 | GO:0004722 | protein serine/threonine phosphatase activity(GO:0004722) |
| 0.0 | 1.0 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.0 | 1.2 | GO:0042826 | histone deacetylase binding(GO:0042826) |
| 0.0 | 1.3 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.0 | 1.2 | GO:0005347 | ATP transmembrane transporter activity(GO:0005347) |
| 0.0 | 0.7 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.0 | 1.0 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.4 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.0 | 0.4 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.0 | 0.5 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.0 | 0.6 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.0 | 6.8 | GO:0003779 | actin binding(GO:0003779) |
| 0.0 | 0.3 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.0 | 0.7 | GO:0016627 | oxidoreductase activity, acting on the CH-CH group of donors(GO:0016627) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 3.7 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.1 | 6.5 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.1 | 3.8 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 3.1 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.0 | 1.2 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.0 | 0.7 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.0 | 0.2 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.0 | 0.7 | PID CDC42 PATHWAY | CDC42 signaling events |
| 0.0 | 1.7 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 6.4 | REACTOME ACTIVATED AMPK STIMULATES FATTY ACID OXIDATION IN MUSCLE | Genes involved in Activated AMPK stimulates fatty-acid oxidation in muscle |
| 0.3 | 4.1 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.3 | 5.9 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.1 | 3.7 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.1 | 1.2 | REACTOME G BETA GAMMA SIGNALLING THROUGH PI3KGAMMA | Genes involved in G beta:gamma signalling through PI3Kgamma |
| 0.1 | 0.3 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.0 | 1.7 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.0 | 0.9 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.0 | 0.6 | REACTOME POST CHAPERONIN TUBULIN FOLDING PATHWAY | Genes involved in Post-chaperonin tubulin folding pathway |
| 0.0 | 0.7 | REACTOME GLUCOSE METABOLISM | Genes involved in Glucose metabolism |
| 0.0 | 0.8 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |