Project
PRJNA195909:zebrafish embryo and larva development
Navigation
Downloads
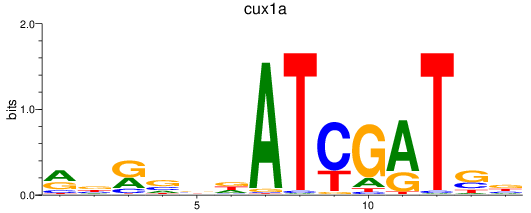
Results for cux1a
Z-value: 0.74
Transcription factors associated with cux1a
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
cux1a
|
ENSDARG00000078459 | cut-like homeobox 1a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| cux1a | dr11_v1_chr10_-_32920690_32920690 | -0.84 | 4.4e-03 | Click! |
Activity profile of cux1a motif
Sorted Z-values of cux1a motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr17_+_24597001 | 0.94 |
ENSDART00000191834
|
rlf
|
rearranged L-myc fusion |
| chr17_-_7792376 | 0.73 |
ENSDART00000064655
|
zbtb2a
|
zinc finger and BTB domain containing 2a |
| chr7_+_32695954 | 0.66 |
ENSDART00000184425
|
slc39a13
|
solute carrier family 39 (zinc transporter), member 13 |
| chr21_-_19919918 | 0.64 |
ENSDART00000137307
ENSDART00000142523 ENSDART00000065670 |
ppp1r3b
|
protein phosphatase 1, regulatory subunit 3B |
| chr2_-_32513538 | 0.55 |
ENSDART00000056640
|
abcf2a
|
ATP-binding cassette, sub-family F (GCN20), member 2a |
| chr13_-_22961605 | 0.55 |
ENSDART00000143112
ENSDART00000057641 |
tspan15
|
tetraspanin 15 |
| chr21_+_21012210 | 0.53 |
ENSDART00000190317
|
ndr1
|
nodal-related 1 |
| chr2_+_23007675 | 0.52 |
ENSDART00000163649
|
mknk2a
|
MAP kinase interacting serine/threonine kinase 2a |
| chr14_-_7045009 | 0.50 |
ENSDART00000112082
|
rufy1
|
RUN and FYVE domain containing 1 |
| chr16_+_1383914 | 0.50 |
ENSDART00000185089
|
cers2b
|
ceramide synthase 2b |
| chr18_+_45571378 | 0.47 |
ENSDART00000077251
|
kifc3
|
kinesin family member C3 |
| chr12_-_9700605 | 0.47 |
ENSDART00000161063
|
heatr1
|
HEAT repeat containing 1 |
| chr18_+_7611298 | 0.46 |
ENSDART00000062156
|
odf3b
|
outer dense fiber of sperm tails 3B |
| chr16_-_25380903 | 0.46 |
ENSDART00000086375
ENSDART00000188587 |
adnp2a
|
ADNP homeobox 2a |
| chr10_+_44057177 | 0.44 |
ENSDART00000164610
|
grk3
|
G protein-coupled receptor kinase 3 |
| chr11_-_42396302 | 0.44 |
ENSDART00000165624
|
slmapa
|
sarcolemma associated protein a |
| chr5_+_37729207 | 0.43 |
ENSDART00000184378
|
cdc42ep2
|
CDC42 effector protein (Rho GTPase binding) 2 |
| chr16_+_25126935 | 0.43 |
ENSDART00000058945
|
zgc:92590
|
zgc:92590 |
| chr6_+_38896158 | 0.42 |
ENSDART00000029930
ENSDART00000131347 |
slc48a1b
|
solute carrier family 48 (heme transporter), member 1b |
| chr2_-_37352514 | 0.41 |
ENSDART00000140498
ENSDART00000186422 |
skila
|
SKI-like proto-oncogene a |
| chr8_+_16758304 | 0.40 |
ENSDART00000133514
|
elovl7a
|
ELOVL fatty acid elongase 7a |
| chr17_+_53435279 | 0.37 |
ENSDART00000126630
|
cx52.9
|
connexin 52.9 |
| chr3_-_30152836 | 0.37 |
ENSDART00000165920
|
nucb1
|
nucleobindin 1 |
| chr3_-_30153242 | 0.36 |
ENSDART00000077089
|
nucb1
|
nucleobindin 1 |
| chr2_-_37353098 | 0.35 |
ENSDART00000056522
|
skila
|
SKI-like proto-oncogene a |
| chr12_+_10115964 | 0.35 |
ENSDART00000152369
|
si:dkeyp-118b1.2
|
si:dkeyp-118b1.2 |
| chr7_-_25132592 | 0.34 |
ENSDART00000192214
|
badb
|
BCL2 associated agonist of cell death b |
| chr21_-_20120543 | 0.32 |
ENSDART00000065664
|
dusp4
|
dual specificity phosphatase 4 |
| chr7_+_49272702 | 0.29 |
ENSDART00000083389
|
abtb2a
|
ankyrin repeat and BTB (POZ) domain containing 2a |
| chr12_+_38770654 | 0.28 |
ENSDART00000155367
|
kif19
|
kinesin family member 19 |
| chr15_-_16704417 | 0.27 |
ENSDART00000155163
|
caln1
|
calneuron 1 |
| chr13_-_4664403 | 0.27 |
ENSDART00000023803
ENSDART00000177957 |
c1d
|
C1D nuclear receptor corepressor |
| chr7_+_29863548 | 0.25 |
ENSDART00000136555
ENSDART00000134068 |
tln2a
|
talin 2a |
| chr20_-_54075136 | 0.25 |
ENSDART00000074255
|
mgat2
|
mannosyl (alpha-1,6-)-glycoprotein beta-1,2-N-acetylglucosaminyltransferase |
| chr16_-_12095144 | 0.25 |
ENSDART00000145106
|
pex5
|
peroxisomal biogenesis factor 5 |
| chr15_-_7598294 | 0.23 |
ENSDART00000165898
|
gbe1b
|
glucan (1,4-alpha-), branching enzyme 1b |
| chr14_-_33083539 | 0.23 |
ENSDART00000160173
|
dlg3
|
discs, large homolog 3 (Drosophila) |
| chr4_-_52189449 | 0.23 |
ENSDART00000192426
|
BX649431.4
|
|
| chr12_+_14149686 | 0.23 |
ENSDART00000123741
|
kbtbd2
|
kelch repeat and BTB (POZ) domain containing 2 |
| chr14_-_32486757 | 0.22 |
ENSDART00000148830
|
mcf2a
|
MCF.2 cell line derived transforming sequence a |
| chr7_-_12789251 | 0.22 |
ENSDART00000052750
|
adamtsl3
|
ADAMTS-like 3 |
| chr19_+_14059349 | 0.22 |
ENSDART00000166230
|
tpbga
|
trophoblast glycoprotein a |
| chr8_-_29851706 | 0.22 |
ENSDART00000149297
|
slc20a2
|
solute carrier family 20 (phosphate transporter), member 2 |
| chr3_-_5555500 | 0.20 |
ENSDART00000176133
ENSDART00000057464 |
trim35-31
|
tripartite motif containing 35-31 |
| chr2_-_56649883 | 0.20 |
ENSDART00000191786
|
gpx4b
|
glutathione peroxidase 4b |
| chr16_+_39242339 | 0.20 |
ENSDART00000102510
|
zgc:77056
|
zgc:77056 |
| chr4_-_14915268 | 0.20 |
ENSDART00000067040
|
si:dkey-180p18.9
|
si:dkey-180p18.9 |
| chr17_+_45454943 | 0.19 |
ENSDART00000074838
|
kcnk3b
|
potassium channel, subfamily K, member 3b |
| chr3_-_31079186 | 0.17 |
ENSDART00000145636
ENSDART00000140569 |
ELOB (1 of many)
elob
|
elongin B elongin B |
| chr17_+_37249736 | 0.15 |
ENSDART00000189686
|
CR318625.1
|
|
| chr15_-_7598542 | 0.15 |
ENSDART00000173092
|
gbe1b
|
glucan (1,4-alpha-), branching enzyme 1b |
| chr22_-_25078650 | 0.15 |
ENSDART00000131811
ENSDART00000141546 |
kat14
|
lysine acetyltransferase 14 |
| chr10_-_11397590 | 0.14 |
ENSDART00000064212
|
srek1ip1
|
SREK1-interacting protein 1 |
| chr11_+_41089574 | 0.14 |
ENSDART00000170023
|
phf13
|
PHD finger protein 13 |
| chr23_-_36446307 | 0.14 |
ENSDART00000136623
|
zgc:174906
|
zgc:174906 |
| chr3_-_37082618 | 0.13 |
ENSDART00000026701
ENSDART00000110716 |
tubg1
|
tubulin, gamma 1 |
| chr8_-_24135171 | 0.13 |
ENSDART00000112926
|
adora1b
|
adenosine A1 receptor b |
| chr16_+_4658250 | 0.13 |
ENSDART00000006212
|
LIN28A
|
si:ch1073-284b18.2 |
| chr22_+_2844865 | 0.12 |
ENSDART00000139123
|
si:dkey-20i20.4
|
si:dkey-20i20.4 |
| chr2_+_17451656 | 0.12 |
ENSDART00000163620
|
BX640408.1
|
|
| chr8_-_19451487 | 0.12 |
ENSDART00000184037
|
zgc:92140
|
zgc:92140 |
| chr23_-_24047054 | 0.12 |
ENSDART00000184308
ENSDART00000185902 |
CR925720.1
|
|
| chr22_+_21516689 | 0.12 |
ENSDART00000105550
ENSDART00000136374 |
mier2
|
mesoderm induction early response 1, family member 2 |
| chr11_-_20956309 | 0.11 |
ENSDART00000188659
|
CABZ01008739.1
|
|
| chr11_+_1522136 | 0.10 |
ENSDART00000002318
|
srsf6b
|
serine/arginine-rich splicing factor 6b |
| chr5_+_41996889 | 0.10 |
ENSDART00000097580
|
pigl
|
phosphatidylinositol glycan anchor biosynthesis, class L |
| chr11_+_16152316 | 0.10 |
ENSDART00000081054
|
tada3l
|
transcriptional adaptor 3 (NGG1 homolog, yeast)-like |
| chr22_+_29067388 | 0.08 |
ENSDART00000133673
|
pimr100
|
Pim proto-oncogene, serine/threonine kinase, related 100 |
| chr25_-_35182347 | 0.07 |
ENSDART00000115210
|
ano9a
|
anoctamin 9a |
| chr1_-_49932040 | 0.07 |
ENSDART00000048281
|
cyp2u1
|
cytochrome P450, family 2, subfamily U, polypeptide 1 |
| chr3_+_57820913 | 0.07 |
ENSDART00000168101
|
CU571328.1
|
|
| chr2_+_59041081 | 0.06 |
ENSDART00000067736
|
stk11
|
serine/threonine kinase 11 |
| chr4_-_51460013 | 0.06 |
ENSDART00000193382
|
CR628395.1
|
|
| chr15_-_9419472 | 0.05 |
ENSDART00000122117
|
sacs
|
sacsin molecular chaperone |
| chr8_-_14121634 | 0.05 |
ENSDART00000184946
|
bgna
|
biglycan a |
| chr24_+_31374324 | 0.05 |
ENSDART00000172335
ENSDART00000163162 |
cpne3
|
copine III |
| chr11_-_12705608 | 0.05 |
ENSDART00000158937
ENSDART00000114391 |
zgc:174353
|
zgc:174353 |
| chr12_-_29305220 | 0.04 |
ENSDART00000153458
|
sh2d4bb
|
SH2 domain containing 4Bb |
| chr22_+_28969071 | 0.04 |
ENSDART00000163427
|
pimr95
|
Pim proto-oncogene, serine/threonine kinase, related 95 |
| chr2_-_51275873 | 0.04 |
ENSDART00000168019
|
si:ch211-215e19.4
|
si:ch211-215e19.4 |
| chrM_+_9735 | 0.03 |
ENSDART00000093613
|
mt-co3
|
cytochrome c oxidase III, mitochondrial |
| chr1_+_55703120 | 0.03 |
ENSDART00000141089
|
adgre6
|
adhesion G protein-coupled receptor E6 |
| chr11_-_12673637 | 0.02 |
ENSDART00000186058
|
CR450764.7
|
|
| chr7_-_56766973 | 0.02 |
ENSDART00000020967
|
csnk2a2a
|
casein kinase 2, alpha prime polypeptide a |
| chr2_+_7715810 | 0.02 |
ENSDART00000130781
|
eif4a2
|
eukaryotic translation initiation factor 4A, isoform 2 |
| chr12_-_19091214 | 0.01 |
ENSDART00000153225
|
si:ch73-139e5.4
|
si:ch73-139e5.4 |
Network of associatons between targets according to the STRING database.
First level regulatory network of cux1a
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.6 | GO:0005981 | regulation of glycogen catabolic process(GO:0005981) |
| 0.1 | 0.5 | GO:0045943 | positive regulation of transcription from RNA polymerase I promoter(GO:0045943) |
| 0.1 | 0.2 | GO:0060829 | negative regulation of canonical Wnt signaling pathway involved in neural plate anterior/posterior pattern formation(GO:0060829) |
| 0.0 | 0.1 | GO:0000212 | meiotic spindle organization(GO:0000212) |
| 0.0 | 0.4 | GO:0031274 | pseudopodium organization(GO:0031268) pseudopodium assembly(GO:0031269) regulation of pseudopodium assembly(GO:0031272) positive regulation of pseudopodium assembly(GO:0031274) |
| 0.0 | 0.4 | GO:0015886 | heme transport(GO:0015886) iron coordination entity transport(GO:1901678) |
| 0.0 | 0.4 | GO:0003315 | heart rudiment formation(GO:0003315) |
| 0.0 | 0.3 | GO:0009312 | oligosaccharide biosynthetic process(GO:0009312) |
| 0.0 | 0.5 | GO:0045743 | positive regulation of fibroblast growth factor receptor signaling pathway(GO:0045743) |
| 0.0 | 0.7 | GO:0071577 | zinc II ion transmembrane transport(GO:0071577) |
| 0.0 | 0.3 | GO:0070373 | negative regulation of ERK1 and ERK2 cascade(GO:0070373) |
| 0.0 | 0.3 | GO:0071479 | cellular response to ionizing radiation(GO:0071479) |
| 0.0 | 0.8 | GO:0030512 | negative regulation of transforming growth factor beta receptor signaling pathway(GO:0030512) negative regulation of cellular response to transforming growth factor beta stimulus(GO:1903845) |
| 0.0 | 0.2 | GO:0016560 | protein import into peroxisome matrix, docking(GO:0016560) protein to membrane docking(GO:0022615) |
| 0.0 | 0.4 | GO:0019368 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.0 | 0.5 | GO:0043249 | erythrocyte maturation(GO:0043249) |
| 0.0 | 0.1 | GO:0035588 | adenosine receptor signaling pathway(GO:0001973) purinergic receptor signaling pathway(GO:0035587) G-protein coupled purinergic receptor signaling pathway(GO:0035588) |
| 0.0 | 0.1 | GO:0007076 | mitotic chromosome condensation(GO:0007076) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.0 | 0.2 | GO:0030891 | VCB complex(GO:0030891) |
| 0.0 | 0.2 | GO:0005671 | Ada2/Gcn5/Ada3 transcription activator complex(GO:0005671) |
| 0.0 | 0.3 | GO:0000178 | exosome (RNase complex)(GO:0000178) |
| 0.0 | 0.5 | GO:0030686 | 90S preribosome(GO:0030686) |
| 0.0 | 0.1 | GO:0000930 | gamma-tubulin complex(GO:0000930) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0003844 | 1,4-alpha-glucan branching enzyme activity(GO:0003844) |
| 0.1 | 0.5 | GO:0070698 | type I activin receptor binding(GO:0070698) |
| 0.1 | 0.2 | GO:0005252 | open rectifier potassium channel activity(GO:0005252) |
| 0.1 | 0.4 | GO:0015232 | heme transporter activity(GO:0015232) |
| 0.0 | 0.6 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.0 | 0.2 | GO:0047066 | phospholipid-hydroperoxide glutathione peroxidase activity(GO:0047066) |
| 0.0 | 0.5 | GO:0050291 | sphingosine N-acyltransferase activity(GO:0050291) |
| 0.0 | 0.2 | GO:0005052 | peroxisome matrix targeting signal-1 binding(GO:0005052) |
| 0.0 | 0.4 | GO:0004703 | G-protein coupled receptor kinase activity(GO:0004703) |
| 0.0 | 0.7 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 0.5 | GO:0030515 | snoRNA binding(GO:0030515) |
| 0.0 | 0.3 | GO:0008330 | protein tyrosine/threonine phosphatase activity(GO:0008330) |
| 0.0 | 0.4 | GO:0009922 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.0 | 0.1 | GO:0001609 | G-protein coupled adenosine receptor activity(GO:0001609) |
| 0.0 | 0.2 | GO:0005315 | inorganic phosphate transmembrane transporter activity(GO:0005315) |
| 0.0 | 0.5 | GO:0008569 | ATP-dependent microtubule motor activity, minus-end-directed(GO:0008569) |
| 0.0 | 0.5 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 0.0 | 0.8 | GO:0046332 | SMAD binding(GO:0046332) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.0 | 0.3 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.0 | 0.2 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.0 | 0.1 | REACTOME RECRUITMENT OF NUMA TO MITOTIC CENTROSOMES | Genes involved in Recruitment of NuMA to mitotic centrosomes |
| 0.0 | 0.5 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |