Project
PRJNA195909:zebrafish embryo and larva development
Navigation
Downloads
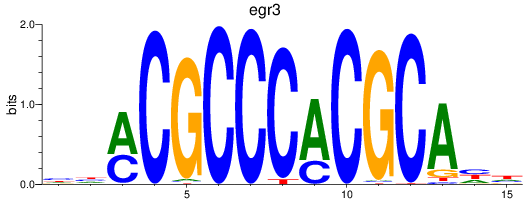
Results for egr3
Z-value: 0.55
Transcription factors associated with egr3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
egr3
|
ENSDARG00000089156 | early growth response 3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| egr3 | dr11_v1_chr8_+_50727220_50727220 | -0.35 | 3.5e-01 | Click! |
Activity profile of egr3 motif
Sorted Z-values of egr3 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr17_-_2584423 | 1.16 |
ENSDART00000013506
|
zp3.2
|
zona pellucida glycoprotein 3, tandem duplicate 2 |
| chr16_-_47381519 | 1.12 |
ENSDART00000032188
ENSDART00000150136 |
si:dkey-256h2.1
|
si:dkey-256h2.1 |
| chr7_+_46019780 | 1.11 |
ENSDART00000163991
|
ccne1
|
cyclin E1 |
| chr22_-_10591876 | 1.10 |
ENSDART00000105846
|
si:dkey-42i9.8
|
si:dkey-42i9.8 |
| chr3_-_49504023 | 0.82 |
ENSDART00000168108
|
prkacaa
|
protein kinase, cAMP-dependent, catalytic, alpha, genome duplicate a |
| chr18_+_3004572 | 0.71 |
ENSDART00000190328
|
CABZ01002690.1
|
|
| chr13_+_35925490 | 0.69 |
ENSDART00000046115
|
mfsd2aa
|
major facilitator superfamily domain containing 2aa |
| chr22_+_34784075 | 0.63 |
ENSDART00000167538
|
lcor
|
ligand dependent nuclear receptor corepressor |
| chr21_-_43398122 | 0.60 |
ENSDART00000050533
|
ccni2
|
cyclin I family, member 2 |
| chr7_-_46019756 | 0.57 |
ENSDART00000162583
|
zgc:162297
|
zgc:162297 |
| chr22_-_16275236 | 0.57 |
ENSDART00000149051
|
cdc14ab
|
cell division cycle 14Ab |
| chr17_+_46387086 | 0.55 |
ENSDART00000157079
|
si:dkey-206p8.1
|
si:dkey-206p8.1 |
| chr18_+_3169579 | 0.54 |
ENSDART00000164724
ENSDART00000186340 ENSDART00000181247 ENSDART00000168056 |
pak1
|
p21 protein (Cdc42/Rac)-activated kinase 1 |
| chr21_-_41369539 | 0.54 |
ENSDART00000187546
|
cpeb4b
|
cytoplasmic polyadenylation element binding protein 4b |
| chr7_+_55518519 | 0.53 |
ENSDART00000098476
ENSDART00000149915 |
cdt1
|
chromatin licensing and DNA replication factor 1 |
| chr2_+_38924975 | 0.53 |
ENSDART00000109219
|
rem2
|
RAS (RAD and GEM)-like GTP binding 2 |
| chr2_+_35880600 | 0.50 |
ENSDART00000004277
|
lamc1
|
laminin, gamma 1 |
| chr21_-_43398457 | 0.49 |
ENSDART00000166530
|
ccni2
|
cyclin I family, member 2 |
| chr2_+_23062085 | 0.48 |
ENSDART00000153745
|
csnk1g2a
|
casein kinase 1, gamma 2a |
| chr20_+_27712714 | 0.47 |
ENSDART00000008306
|
zbtb1
|
zinc finger and BTB domain containing 1 |
| chr2_+_105748 | 0.46 |
ENSDART00000169601
|
CABZ01098670.1
|
|
| chr15_-_14038227 | 0.44 |
ENSDART00000139068
|
zgc:114130
|
zgc:114130 |
| chr2_+_59015878 | 0.44 |
ENSDART00000148816
ENSDART00000122795 |
si:ch1073-391i24.1
|
si:ch1073-391i24.1 |
| chr7_+_67325933 | 0.43 |
ENSDART00000170575
ENSDART00000183342 |
nfat5b
|
nuclear factor of activated T cells 5b |
| chr21_-_41369370 | 0.42 |
ENSDART00000159290
|
cpeb4b
|
cytoplasmic polyadenylation element binding protein 4b |
| chr7_-_6364168 | 0.42 |
ENSDART00000173332
|
zgc:112234
|
zgc:112234 |
| chr20_+_28803642 | 0.40 |
ENSDART00000188526
|
fntb
|
farnesyltransferase, CAAX box, beta |
| chr13_-_49666615 | 0.38 |
ENSDART00000148083
|
tomm20a
|
translocase of outer mitochondrial membrane 20 |
| chr12_-_26537145 | 0.35 |
ENSDART00000138437
ENSDART00000163931 ENSDART00000132737 |
acsf2
|
acyl-CoA synthetase family member 2 |
| chr12_-_37449396 | 0.34 |
ENSDART00000152951
|
cdc42ep4b
|
CDC42 effector protein (Rho GTPase binding) 4b |
| chr5_+_29831235 | 0.31 |
ENSDART00000109660
|
f11r.1
|
F11 receptor, tandem duplicate 1 |
| chr11_+_1565806 | 0.31 |
ENSDART00000137479
|
mybl2b
|
v-myb avian myeloblastosis viral oncogene homolog-like 2b |
| chr23_-_31645760 | 0.30 |
ENSDART00000035031
|
sgk1
|
serum/glucocorticoid regulated kinase 1 |
| chr3_+_22984098 | 0.29 |
ENSDART00000043190
|
lsm12a
|
LSM12 homolog a |
| chr20_+_27298783 | 0.26 |
ENSDART00000013861
|
ppp4r4
|
protein phosphatase 4, regulatory subunit 4 |
| chr11_-_2594045 | 0.26 |
ENSDART00000114079
|
nab2
|
NGFI-A binding protein 2 (EGR1 binding protein 2) |
| chr4_-_73587205 | 0.24 |
ENSDART00000150837
|
znf1015
|
zinc finger protein 1015 |
| chr1_+_12295142 | 0.21 |
ENSDART00000158595
|
gne
|
glucosamine (UDP-N-acetyl)-2-epimerase/N-acetylmannosamine kinase |
| chr6_+_2271559 | 0.21 |
ENSDART00000165223
|
pbx1b
|
pre-B-cell leukemia homeobox 1b |
| chr7_+_41340520 | 0.21 |
ENSDART00000005127
|
garem
|
GRB2 associated, regulator of MAPK1 |
| chr5_+_6617401 | 0.19 |
ENSDART00000060532
|
zgc:110796
|
zgc:110796 |
| chr2_+_16710889 | 0.19 |
ENSDART00000017852
|
ubxn7
|
UBX domain protein 7 |
| chr21_-_24865217 | 0.18 |
ENSDART00000101136
|
igsf9bb
|
immunoglobulin superfamily, member 9Bb |
| chr13_-_36141486 | 0.14 |
ENSDART00000044115
|
med6
|
mediator complex subunit 6 |
| chr25_-_6389713 | 0.14 |
ENSDART00000083539
|
sin3aa
|
SIN3 transcription regulator family member Aa |
| chr3_-_11624694 | 0.14 |
ENSDART00000127157
|
hlfa
|
hepatic leukemia factor a |
| chr21_-_217589 | 0.13 |
ENSDART00000185017
|
CZQB01146713.1
|
|
| chr3_-_7106678 | 0.13 |
ENSDART00000175293
ENSDART00000187732 ENSDART00000158652 |
zgc:113102
BX005085.2
|
zgc:113102 |
| chr13_-_31166544 | 0.12 |
ENSDART00000146250
ENSDART00000132129 ENSDART00000139591 |
mapk8a
|
mitogen-activated protein kinase 8a |
| chr9_-_48036 | 0.09 |
ENSDART00000165230
ENSDART00000171335 |
map4k4
|
mitogen-activated protein kinase kinase kinase kinase 4 |
| chr5_-_2689753 | 0.09 |
ENSDART00000172699
|
gng10
|
guanine nucleotide binding protein (G protein), gamma 10 |
| chr16_+_4138331 | 0.08 |
ENSDART00000192403
ENSDART00000193036 |
mtf1
|
metal-regulatory transcription factor 1 |
| chr12_+_499881 | 0.08 |
ENSDART00000167527
|
mprip
|
myosin phosphatase Rho interacting protein |
| chr21_-_24865454 | 0.08 |
ENSDART00000142907
|
igsf9bb
|
immunoglobulin superfamily, member 9Bb |
| chr22_+_1806235 | 0.08 |
ENSDART00000158966
|
znf1168
|
zinc finger protein 1168 |
| chr24_-_39354829 | 0.06 |
ENSDART00000169108
|
kcnj12a
|
potassium inwardly-rectifying channel, subfamily J, member 12a |
| chr19_+_5480327 | 0.06 |
ENSDART00000148794
|
jupb
|
junction plakoglobin b |
| chr15_+_36105581 | 0.05 |
ENSDART00000184099
|
col4a3
|
collagen, type IV, alpha 3 |
| chr18_+_41549764 | 0.04 |
ENSDART00000059135
|
bcl7bb
|
BCL tumor suppressor 7Bb |
| chr17_+_48164536 | 0.03 |
ENSDART00000161750
ENSDART00000156923 |
plekhd1
|
pleckstrin homology domain containing, family D (with coiled-coil domains) member 1 |
| chr25_+_1304173 | 0.02 |
ENSDART00000155229
|
rxfp3.3b
|
relaxin/insulin-like family peptide receptor 3.3b |
| chr3_-_5228137 | 0.02 |
ENSDART00000137105
|
myh9b
|
myosin, heavy chain 9b, non-muscle |
| chr20_+_13894123 | 0.01 |
ENSDART00000007744
|
slc30a1a
|
solute carrier family 30 (zinc transporter), member 1a |
| chr6_-_53391727 | 0.01 |
ENSDART00000155483
|
si:ch211-161c3.6
|
si:ch211-161c3.6 |
| chr19_-_48039400 | 0.00 |
ENSDART00000166748
ENSDART00000165921 |
csf3b
|
colony stimulating factor 3 (granulocyte) b |
Network of associatons between targets according to the STRING database.
First level regulatory network of egr3
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.6 | GO:0042998 | positive regulation of Golgi to plasma membrane protein transport(GO:0042998) positive regulation of establishment of protein localization to plasma membrane(GO:0090004) |
| 0.2 | 0.7 | GO:0035633 | maintenance of blood-brain barrier(GO:0035633) |
| 0.1 | 0.4 | GO:0018343 | protein farnesylation(GO:0018343) |
| 0.1 | 0.8 | GO:0071331 | cellular response to carbohydrate stimulus(GO:0071322) cellular response to monosaccharide stimulus(GO:0071326) cellular response to hexose stimulus(GO:0071331) cellular response to glucose stimulus(GO:0071333) |
| 0.1 | 0.5 | GO:0003232 | bulbus arteriosus development(GO:0003232) |
| 0.1 | 1.0 | GO:2000766 | negative regulation of cytoplasmic translation(GO:2000766) |
| 0.1 | 1.2 | GO:0007339 | binding of sperm to zona pellucida(GO:0007339) egg coat formation(GO:0035803) regulation of acrosome reaction(GO:0060046) positive regulation of acrosome reaction(GO:2000344) |
| 0.1 | 0.6 | GO:0071850 | mitotic cell cycle arrest(GO:0071850) |
| 0.0 | 2.2 | GO:0000079 | regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0000079) regulation of cyclin-dependent protein kinase activity(GO:1904029) |
| 0.0 | 0.5 | GO:0050908 | detection of light stimulus involved in visual perception(GO:0050908) detection of light stimulus involved in sensory perception(GO:0050962) |
| 0.0 | 0.5 | GO:0000076 | DNA replication checkpoint(GO:0000076) |
| 0.0 | 0.3 | GO:0031274 | pseudopodium organization(GO:0031268) pseudopodium assembly(GO:0031269) regulation of pseudopodium assembly(GO:0031272) positive regulation of pseudopodium assembly(GO:0031274) |
| 0.0 | 0.3 | GO:0032515 | negative regulation of phosphoprotein phosphatase activity(GO:0032515) |
| 0.0 | 0.3 | GO:0050892 | intestinal absorption(GO:0050892) |
| 0.0 | 0.5 | GO:0071542 | dopaminergic neuron differentiation(GO:0071542) |
| 0.0 | 0.1 | GO:0071276 | cellular response to cadmium ion(GO:0071276) |
| 0.0 | 0.1 | GO:0002159 | desmosome assembly(GO:0002159) |
| 0.0 | 0.2 | GO:0006047 | UDP-N-acetylglucosamine metabolic process(GO:0006047) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.1 | GO:0097134 | cyclin E1-CDK2 complex(GO:0097134) |
| 0.1 | 0.4 | GO:0005965 | protein farnesyltransferase complex(GO:0005965) |
| 0.1 | 0.6 | GO:1902636 | kinociliary basal body(GO:1902636) |
| 0.1 | 0.8 | GO:0001669 | acrosomal vesicle(GO:0001669) |
| 0.1 | 1.0 | GO:1990124 | messenger ribonucleoprotein complex(GO:1990124) |
| 0.0 | 0.4 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.0 | 1.1 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.0 | 0.1 | GO:0070847 | core mediator complex(GO:0070847) |
| 0.0 | 0.1 | GO:0016580 | Sin3 complex(GO:0016580) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.1 | GO:0016715 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced ascorbate as one donor, and incorporation of one atom of oxygen(GO:0016715) |
| 0.2 | 0.8 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.1 | 0.6 | GO:0071253 | connexin binding(GO:0071253) |
| 0.1 | 0.5 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.1 | 0.3 | GO:0047760 | medium-chain fatty acid-CoA ligase activity(GO:0031956) butyrate-CoA ligase activity(GO:0047760) |
| 0.1 | 1.2 | GO:0035804 | structural constituent of egg coat(GO:0035804) |
| 0.1 | 0.4 | GO:0004660 | protein farnesyltransferase activity(GO:0004660) |
| 0.1 | 0.2 | GO:0008761 | UDP-N-acetylglucosamine 2-epimerase activity(GO:0008761) N-acylmannosamine kinase activity(GO:0009384) |
| 0.1 | 1.0 | GO:0000900 | translation repressor activity, nucleic acid binding(GO:0000900) |
| 0.0 | 2.2 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.0 | 0.5 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.0 | 0.7 | GO:0005548 | phospholipid transporter activity(GO:0005548) |
| 0.0 | 0.1 | GO:0004705 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.1 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.0 | 0.5 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.0 | 0.5 | PID S1P S1P2 PATHWAY | S1P2 pathway |
| 0.0 | 0.6 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | REACTOME CD28 DEPENDENT VAV1 PATHWAY | Genes involved in CD28 dependent Vav1 pathway |
| 0.1 | 1.6 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |