Project
PRJNA195909:zebrafish embryo and larva development
Navigation
Downloads
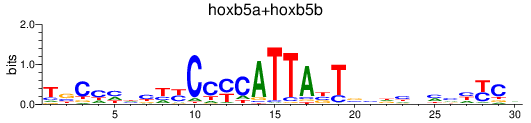
Results for hoxb5a+hoxb5b
Z-value: 0.63
Transcription factors associated with hoxb5a+hoxb5b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
hoxb5a
|
ENSDARG00000013057 | homeobox B5a |
|
hoxb5b
|
ENSDARG00000054030 | homeobox B5b |
|
hoxb5b
|
ENSDARG00000113174 | homeobox B5b |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| hoxb5a | dr11_v1_chr3_+_23707691_23707691 | -0.85 | 3.7e-03 | Click! |
| hoxb5b | dr11_v1_chr12_+_27129659_27129659 | -0.85 | 3.9e-03 | Click! |
Activity profile of hoxb5a+hoxb5b motif
Sorted Z-values of hoxb5a+hoxb5b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr16_-_29387215 | 1.10 |
ENSDART00000148787
|
s100a1
|
S100 calcium binding protein A1 |
| chr4_-_77551860 | 1.02 |
ENSDART00000188176
|
AL935186.6
|
|
| chr4_-_77563411 | 1.00 |
ENSDART00000186841
|
AL935186.8
|
|
| chr21_-_9769500 | 0.77 |
ENSDART00000170710
|
arhgap24
|
Rho GTPase activating protein 24 |
| chr13_+_534453 | 0.70 |
ENSDART00000147909
|
wu:fc17b08
|
wu:fc17b08 |
| chr6_+_4255319 | 0.62 |
ENSDART00000170351
|
nbeal1
|
neurobeachin-like 1 |
| chr15_-_25099679 | 0.58 |
ENSDART00000154628
|
rflnb
|
refilin B |
| chr19_-_27827744 | 0.57 |
ENSDART00000181620
|
papd7
|
PAP associated domain containing 7 |
| chr22_+_25352530 | 0.52 |
ENSDART00000176742
ENSDART00000113186 |
si:dkey-240e12.6
|
si:dkey-240e12.6 |
| chr19_+_15441022 | 0.52 |
ENSDART00000098970
ENSDART00000140276 |
lin28a
|
lin-28 homolog A (C. elegans) |
| chr5_-_16791223 | 0.52 |
ENSDART00000192744
|
atad1a
|
ATPase family, AAA domain containing 1a |
| chr11_-_2297832 | 0.51 |
ENSDART00000158266
|
znf740a
|
zinc finger protein 740a |
| chr2_-_15324837 | 0.50 |
ENSDART00000015655
|
tecrl2b
|
trans-2,3-enoyl-CoA reductase-like 2b |
| chr23_+_22293682 | 0.48 |
ENSDART00000187304
|
CR354612.1
|
|
| chr20_+_48543492 | 0.48 |
ENSDART00000175480
|
LO018254.1
|
|
| chr10_+_18557102 | 0.47 |
ENSDART00000160157
|
CR388008.1
|
|
| chr8_-_36554675 | 0.46 |
ENSDART00000132804
ENSDART00000078746 |
ccdc157
|
coiled-coil domain containing 157 |
| chr22_+_12477996 | 0.42 |
ENSDART00000177704
|
CR847870.3
|
|
| chr4_+_77966055 | 0.40 |
ENSDART00000174203
ENSDART00000130100 ENSDART00000080665 ENSDART00000174317 ENSDART00000190123 |
zgc:113921
|
zgc:113921 |
| chr9_+_48755638 | 0.39 |
ENSDART00000047401
|
abcb11a
|
ATP-binding cassette, sub-family B (MDR/TAP), member 11a |
| chr19_+_43341424 | 0.38 |
ENSDART00000134815
|
sesn2
|
sestrin 2 |
| chr22_+_17536989 | 0.34 |
ENSDART00000149531
|
hnrnpm
|
heterogeneous nuclear ribonucleoprotein M |
| chr15_-_952891 | 0.34 |
ENSDART00000153780
|
alox5b.1
|
arachidonate 5-lipoxygenase b, tandem duplicate 1 |
| chr19_+_43341115 | 0.33 |
ENSDART00000145846
ENSDART00000102384 |
sesn2
|
sestrin 2 |
| chr13_-_36134317 | 0.32 |
ENSDART00000015664
ENSDART00000188301 ENSDART00000191073 ENSDART00000184192 ENSDART00000139593 |
pcnx
|
pecanex homolog (Drosophila) |
| chr17_+_20569806 | 0.32 |
ENSDART00000113936
|
SAMD8
|
zgc:162183 |
| chr13_-_44819254 | 0.30 |
ENSDART00000131673
|
si:dkeyp-2e4.8
|
si:dkeyp-2e4.8 |
| chr14_-_25078569 | 0.30 |
ENSDART00000172802
ENSDART00000173345 ENSDART00000135004 |
matr3l1.1
|
matrin 3-like 1.1 |
| chr9_+_6082793 | 0.29 |
ENSDART00000192045
|
st6gal2a
|
ST6 beta-galactosamide alpha-2,6-sialyltranferase 2a |
| chr13_-_86847 | 0.29 |
ENSDART00000158062
|
pole2
|
polymerase (DNA directed), epsilon 2 |
| chr19_+_48060464 | 0.27 |
ENSDART00000123163
|
zgc:85936
|
zgc:85936 |
| chr8_+_25962833 | 0.26 |
ENSDART00000086583
|
si:dkey-72l14.4
|
si:dkey-72l14.4 |
| chr3_-_4911989 | 0.25 |
ENSDART00000158197
|
si:dkey-73p2.2
|
si:dkey-73p2.2 |
| chr9_-_41817712 | 0.25 |
ENSDART00000182657
|
si:dkeyp-30e7.2
|
si:dkeyp-30e7.2 |
| chr19_+_17642356 | 0.25 |
ENSDART00000176431
|
CR382334.1
|
|
| chr5_-_9625459 | 0.24 |
ENSDART00000143347
|
sh2b3
|
SH2B adaptor protein 3 |
| chr9_-_12034444 | 0.24 |
ENSDART00000038651
|
znf804a
|
zinc finger protein 804A |
| chr15_-_684911 | 0.24 |
ENSDART00000161685
|
zgc:171901
|
zgc:171901 |
| chr6_-_15603675 | 0.24 |
ENSDART00000143502
|
lrrfip1b
|
leucine rich repeat (in FLII) interacting protein 1b |
| chr22_-_7461603 | 0.23 |
ENSDART00000170630
|
BX511034.6
|
|
| chr11_-_44030962 | 0.23 |
ENSDART00000171910
|
FP016005.1
|
|
| chr19_+_46259619 | 0.22 |
ENSDART00000158032
|
grinaa
|
glutamate receptor, ionotropic, N-methyl D-aspartate-associated protein 1a (glutamate binding) |
| chr8_+_247163 | 0.21 |
ENSDART00000122378
|
cep120
|
centrosomal protein 120 |
| chr25_+_35212919 | 0.21 |
ENSDART00000180127
|
ano5a
|
anoctamin 5a |
| chr8_-_43847138 | 0.20 |
ENSDART00000148358
|
adgrd1
|
adhesion G protein-coupled receptor D1 |
| chr12_+_17436904 | 0.20 |
ENSDART00000079130
|
atad1b
|
ATPase family, AAA domain containing 1b |
| chr5_-_33769211 | 0.19 |
ENSDART00000133504
|
dab2ipb
|
DAB2 interacting protein b |
| chr25_+_36316280 | 0.18 |
ENSDART00000152735
|
si:ch211-113a14.11
|
si:ch211-113a14.11 |
| chr9_+_16854121 | 0.18 |
ENSDART00000110866
|
cln5
|
CLN5, intracellular trafficking protein |
| chr3_+_54776475 | 0.16 |
ENSDART00000142710
|
si:ch211-74m13.1
|
si:ch211-74m13.1 |
| chr3_+_54776117 | 0.15 |
ENSDART00000189047
ENSDART00000115076 |
si:ch211-74m13.1
|
si:ch211-74m13.1 |
| chr23_-_41965557 | 0.14 |
ENSDART00000144183
|
slc1a7b
|
solute carrier family 1 (glutamate transporter), member 7b |
| chr6_+_41191482 | 0.13 |
ENSDART00000000877
|
opn1mw3
|
opsin 1 (cone pigments), medium-wave-sensitive, 3 |
| chr8_+_36554816 | 0.13 |
ENSDART00000126687
|
sf3a1
|
splicing factor 3a, subunit 1 |
| chr18_-_506082 | 0.13 |
ENSDART00000169974
|
sdr42e1
|
short chain dehydrogenase/reductase family 42E, member 1 |
| chr22_-_15385442 | 0.13 |
ENSDART00000090975
|
tmem264
|
transmembrane protein 264 |
| chr5_-_41142467 | 0.12 |
ENSDART00000129415
|
zfr
|
zinc finger RNA binding protein |
| chr6_+_54888493 | 0.12 |
ENSDART00000113331
|
nav1b
|
neuron navigator 1b |
| chr21_-_308852 | 0.11 |
ENSDART00000151613
|
lhfpl2a
|
LHFPL tetraspan subfamily member 2a |
| chr3_-_1520657 | 0.11 |
ENSDART00000053201
|
polr2f
|
polymerase (RNA) II (DNA directed) polypeptide F |
| chr2_-_42065069 | 0.10 |
ENSDART00000140188
|
cspp1b
|
centrosome and spindle pole associated protein 1b |
| chr14_+_8476769 | 0.10 |
ENSDART00000131773
|
dnajc4
|
DnaJ (Hsp40) homolog, subfamily C, member 4 |
| chr20_+_51730658 | 0.10 |
ENSDART00000010271
|
aida
|
axin interactor, dorsalization associated |
| chr10_+_26944418 | 0.10 |
ENSDART00000135493
|
frmd8
|
FERM domain containing 8 |
| chr23_+_3513201 | 0.10 |
ENSDART00000092258
|
spata2
|
spermatogenesis associated 2 |
| chr22_+_1680628 | 0.09 |
ENSDART00000166672
|
si:dkey-1b17.6
|
si:dkey-1b17.6 |
| chr3_+_28831450 | 0.08 |
ENSDART00000055422
|
flr
|
fleer |
| chr7_+_62320134 | 0.07 |
ENSDART00000012089
|
stim2b
|
stromal interaction molecule 2b |
| chr6_-_10927766 | 0.07 |
ENSDART00000134327
|
ccr7
|
chemokine (C-C motif) receptor 7 |
| chr1_-_56596701 | 0.07 |
ENSDART00000133693
|
LO018605.1
|
|
| chr7_+_56889879 | 0.07 |
ENSDART00000039810
|
myo9aa
|
myosin IXAa |
| chr4_+_38547203 | 0.07 |
ENSDART00000171349
|
si:ch211-209n20.1
|
si:ch211-209n20.1 |
| chr4_-_25812329 | 0.07 |
ENSDART00000146658
|
tmcc3
|
transmembrane and coiled-coil domain family 3 |
| chr24_-_29822913 | 0.06 |
ENSDART00000160929
|
aglb
|
amylo-alpha-1, 6-glucosidase, 4-alpha-glucanotransferase b |
| chr4_+_42295479 | 0.06 |
ENSDART00000162279
|
znf1118
|
zinc finger protein 1118 |
| chr3_-_5413018 | 0.05 |
ENSDART00000063138
|
vmo1a
|
vitelline membrane outer layer 1 homolog a |
| chr5_+_59392183 | 0.05 |
ENSDART00000082983
ENSDART00000180882 |
clip2
|
CAP-GLY domain containing linker protein 2 |
| chr20_+_53181017 | 0.05 |
ENSDART00000189692
ENSDART00000177109 |
FIG4
|
FIG4 phosphoinositide 5-phosphatase |
| chr22_-_17729778 | 0.05 |
ENSDART00000192132
|
si:ch73-63e15.2
|
si:ch73-63e15.2 |
| chr11_-_6070192 | 0.05 |
ENSDART00000162776
|
babam1
|
BRISC and BRCA1 A complex member 1 |
| chr4_-_59094319 | 0.05 |
ENSDART00000162560
|
si:dkey-28i19.1
|
si:dkey-28i19.1 |
| chr2_+_22823952 | 0.04 |
ENSDART00000147935
|
aire
|
autoimmune regulator |
| chr1_+_55703120 | 0.04 |
ENSDART00000141089
|
adgre6
|
adhesion G protein-coupled receptor E6 |
| chr4_-_9549693 | 0.04 |
ENSDART00000160242
|
FQ377934.1
|
|
| chr3_+_39579393 | 0.03 |
ENSDART00000055170
|
cln3
|
ceroid-lipofuscinosis, neuronal 3 |
| chr16_+_16826504 | 0.03 |
ENSDART00000108691
|
kcnj14
|
potassium inwardly-rectifying channel, subfamily J, member 14 |
| chr15_-_42283830 | 0.01 |
ENSDART00000015979
|
farsb
|
phenylalanyl-tRNA synthetase, beta subunit |
| chr15_-_35031523 | 0.01 |
ENSDART00000154848
|
si:ch211-272b8.6
|
si:ch211-272b8.6 |
| chr23_+_39331216 | 0.00 |
ENSDART00000160957
|
kcng1
|
potassium voltage-gated channel, subfamily G, member 1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of hoxb5a+hoxb5b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.6 | GO:1900158 | negative regulation of chondrocyte differentiation(GO:0032331) negative regulation of cartilage development(GO:0061037) negative regulation of chondrocyte development(GO:0061182) regulation of bone mineralization involved in bone maturation(GO:1900157) negative regulation of bone mineralization involved in bone maturation(GO:1900158) negative regulation of bone development(GO:1903011) |
| 0.1 | 0.8 | GO:1900028 | wound healing, spreading of epidermal cells(GO:0035313) negative regulation of ruffle assembly(GO:1900028) |
| 0.1 | 0.7 | GO:1901031 | response to leucine(GO:0043201) cellular response to leucine(GO:0071233) regulation of response to reactive oxygen species(GO:1901031) |
| 0.1 | 0.4 | GO:0032782 | canalicular bile acid transport(GO:0015722) bile acid secretion(GO:0032782) |
| 0.1 | 0.3 | GO:0010610 | regulation of mRNA stability involved in response to stress(GO:0010610) regulation of mRNA stability involved in response to oxidative stress(GO:2000815) |
| 0.1 | 0.2 | GO:0010825 | positive regulation of centrosome duplication(GO:0010825) |
| 0.0 | 0.3 | GO:0042276 | error-prone translesion synthesis(GO:0042276) |
| 0.0 | 0.3 | GO:0051122 | lipoxygenase pathway(GO:0019372) linoleic acid metabolic process(GO:0043651) hepoxilin metabolic process(GO:0051121) hepoxilin biosynthetic process(GO:0051122) |
| 0.0 | 0.2 | GO:0038026 | negative regulation of vascular endothelial growth factor receptor signaling pathway(GO:0030948) reelin-mediated signaling pathway(GO:0038026) |
| 0.0 | 0.0 | GO:0002513 | tolerance induction(GO:0002507) tolerance induction to self antigen(GO:0002513) |
| 0.0 | 0.2 | GO:1902108 | regulation of mitochondrial membrane permeability involved in apoptotic process(GO:1902108) |
| 0.0 | 0.1 | GO:0043508 | negative regulation of JUN kinase activity(GO:0043508) |
| 0.0 | 0.1 | GO:0010939 | regulation of necrotic cell death(GO:0010939) regulation of necroptotic process(GO:0060544) |
| 0.0 | 0.3 | GO:0032418 | lysosome localization(GO:0032418) |
| 0.0 | 0.5 | GO:0042761 | very long-chain fatty acid biosynthetic process(GO:0042761) |
| 0.0 | 0.1 | GO:0098529 | neuromuscular junction development, skeletal muscle fiber(GO:0098529) |
| 0.0 | 0.1 | GO:0046546 | development of primary male sexual characteristics(GO:0046546) |
| 0.0 | 0.1 | GO:0032237 | activation of store-operated calcium channel activity(GO:0032237) positive regulation of store-operated calcium channel activity(GO:1901341) |
| 0.0 | 0.1 | GO:0036372 | opsin transport(GO:0036372) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.7 | GO:0061700 | GATOR2 complex(GO:0061700) |
| 0.1 | 0.3 | GO:0008622 | epsilon DNA polymerase complex(GO:0008622) |
| 0.0 | 0.3 | GO:0071014 | post-mRNA release spliceosomal complex(GO:0071014) |
| 0.0 | 0.1 | GO:0043194 | axon initial segment(GO:0043194) |
| 0.0 | 0.1 | GO:0005879 | axonemal microtubule(GO:0005879) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | GO:0031005 | filamin binding(GO:0031005) |
| 0.1 | 0.7 | GO:0070728 | leucine binding(GO:0070728) |
| 0.1 | 0.4 | GO:0015126 | canalicular bile acid transmembrane transporter activity(GO:0015126) bile acid-exporting ATPase activity(GO:0015432) |
| 0.0 | 0.3 | GO:0004051 | arachidonate 5-lipoxygenase activity(GO:0004051) |
| 0.0 | 0.2 | GO:0005068 | transmembrane receptor protein tyrosine kinase adaptor activity(GO:0005068) |
| 0.0 | 1.1 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.0 | 0.3 | GO:0003835 | beta-galactoside alpha-2,6-sialyltransferase activity(GO:0003835) |
| 0.0 | 0.1 | GO:0070735 | protein-glycine ligase activity(GO:0070735) tubulin-glycine ligase activity(GO:0070738) |
| 0.0 | 0.1 | GO:0004134 | glycogen debranching enzyme activity(GO:0004133) 4-alpha-glucanotransferase activity(GO:0004134) amylo-alpha-1,6-glucosidase activity(GO:0004135) |
| 0.0 | 0.1 | GO:0003854 | 3-beta-hydroxy-delta5-steroid dehydrogenase activity(GO:0003854) |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | REACTOME REPAIR SYNTHESIS FOR GAP FILLING BY DNA POL IN TC NER | Genes involved in Repair synthesis for gap-filling by DNA polymerase in TC-NER |
| 0.0 | 0.1 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.0 | 0.2 | REACTOME REGULATION OF KIT SIGNALING | Genes involved in Regulation of KIT signaling |