Project
PRJNA195909:zebrafish embryo and larva development
Navigation
Downloads
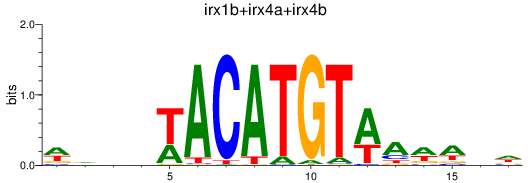
Results for irx1b+irx4a+irx4b
Z-value: 0.64
Transcription factors associated with irx1b+irx4a+irx4b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
irx4a
|
ENSDARG00000035648 | iroquois homeobox 4a |
|
irx4b
|
ENSDARG00000036051 | iroquois homeobox 4b |
|
irx1b
|
ENSDARG00000056594 | iroquois homeobox 1b |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| irx4a | dr11_v1_chr16_-_383664_383664 | -0.84 | 4.2e-03 | Click! |
| irx4b | dr11_v1_chr19_+_28116410_28116410 | -0.77 | 1.6e-02 | Click! |
| irx1b | dr11_v1_chr19_-_28130658_28130658 | 0.23 | 5.6e-01 | Click! |
Activity profile of irx1b+irx4a+irx4b motif
Sorted Z-values of irx1b+irx4a+irx4b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr7_+_56577906 | 1.06 |
ENSDART00000184023
|
hp
|
haptoglobin |
| chr10_-_45002196 | 0.94 |
ENSDART00000170767
|
ved
|
ventrally expressed dharma/bozozok antagonist |
| chr25_-_29072162 | 0.74 |
ENSDART00000169269
|
arid3b
|
AT rich interactive domain 3B (BRIGHT-like) |
| chr5_+_66170479 | 0.73 |
ENSDART00000172117
|
gldc
|
glycine dehydrogenase (decarboxylating) |
| chr7_+_56577522 | 0.71 |
ENSDART00000149130
ENSDART00000149624 |
hp
|
haptoglobin |
| chr18_+_22109379 | 0.70 |
ENSDART00000147230
|
zgc:158868
|
zgc:158868 |
| chr10_+_8629275 | 0.57 |
ENSDART00000129643
|
aplnrb
|
apelin receptor b |
| chr21_-_45871866 | 0.55 |
ENSDART00000161716
|
larp1
|
La ribonucleoprotein domain family, member 1 |
| chr14_-_10617127 | 0.54 |
ENSDART00000154299
|
si:dkey-92i17.2
|
si:dkey-92i17.2 |
| chr17_-_5352924 | 0.36 |
ENSDART00000167275
|
supt3h
|
SPT3 homolog, SAGA and STAGA complex component |
| chr3_-_402714 | 0.35 |
ENSDART00000134062
ENSDART00000105659 |
mhc1zja
|
major histocompatibility complex class I ZJA |
| chr10_-_28117740 | 0.34 |
ENSDART00000134491
|
med13a
|
mediator complex subunit 13a |
| chr2_-_7185460 | 0.33 |
ENSDART00000092078
|
rc3h1b
|
ring finger and CCCH-type domains 1b |
| chr8_+_23784471 | 0.33 |
ENSDART00000189457
|
si:ch211-163l21.8
|
si:ch211-163l21.8 |
| chr5_-_58996324 | 0.31 |
ENSDART00000033923
|
mis12
|
MIS12 kinetochore complex component |
| chr20_-_27190393 | 0.30 |
ENSDART00000149024
|
btbd7
|
BTB (POZ) domain containing 7 |
| chr14_-_26392146 | 0.29 |
ENSDART00000037999
|
b4galt7
|
xylosylprotein beta 1,4-galactosyltransferase, polypeptide 7 (galactosyltransferase I) |
| chr5_-_35159379 | 0.27 |
ENSDART00000144105
|
fcho2
|
FCH domain only 2 |
| chr8_-_23783633 | 0.20 |
ENSDART00000132657
|
si:ch211-163l21.7
|
si:ch211-163l21.7 |
| chr23_+_40275400 | 0.18 |
ENSDART00000184259
|
fam46ab
|
family with sequence similarity 46, member Ab |
| chr23_+_19701587 | 0.17 |
ENSDART00000104425
|
dnase1l1
|
deoxyribonuclease I-like 1 |
| chr8_+_26410197 | 0.15 |
ENSDART00000145836
ENSDART00000053447 |
ifrd2
|
interferon-related developmental regulator 2 |
| chr23_+_45845159 | 0.15 |
ENSDART00000023944
|
lmnl3
|
lamin L3 |
| chr22_+_1940595 | 0.15 |
ENSDART00000163506
|
znf1167
|
zinc finger protein 1167 |
| chr6_-_34838397 | 0.14 |
ENSDART00000060169
ENSDART00000169605 |
mier1a
|
mesoderm induction early response 1a, transcriptional regulator |
| chr23_+_40275601 | 0.13 |
ENSDART00000076876
|
fam46ab
|
family with sequence similarity 46, member Ab |
| chr20_-_23440955 | 0.12 |
ENSDART00000153386
|
slc10a4
|
solute carrier family 10, member 4 |
| chr21_+_12036238 | 0.12 |
ENSDART00000102463
ENSDART00000155426 |
zgc:162344
|
zgc:162344 |
| chr9_-_2945008 | 0.12 |
ENSDART00000183452
|
zak
|
sterile alpha motif and leucine zipper containing kinase AZK |
| chr4_+_33012407 | 0.12 |
ENSDART00000151873
|
si:dkey-26h11.2
|
si:dkey-26h11.2 |
| chr5_-_41103583 | 0.12 |
ENSDART00000051070
ENSDART00000074781 |
golph3
|
golgi phosphoprotein 3 |
| chr13_-_43149063 | 0.11 |
ENSDART00000099601
|
vsir
|
V-set immunoregulatory receptor |
| chr4_-_5691257 | 0.10 |
ENSDART00000110497
|
tmem63a
|
transmembrane protein 63A |
| chr17_-_14705039 | 0.10 |
ENSDART00000154281
ENSDART00000123550 |
ptp4a2a
|
protein tyrosine phosphatase type IVA, member 2a |
| chr21_+_20549395 | 0.10 |
ENSDART00000181633
|
efna5a
|
ephrin-A5a |
| chr18_+_5549672 | 0.09 |
ENSDART00000184970
|
nnt2
|
nicotinamide nucleotide transhydrogenase 2 |
| chr1_+_53919110 | 0.09 |
ENSDART00000020680
|
nup133
|
nucleoporin 133 |
| chr15_+_17100412 | 0.09 |
ENSDART00000154418
|
relb
|
v-rel avian reticuloendotheliosis viral oncogene homolog B |
| chr21_+_43882274 | 0.08 |
ENSDART00000075672
|
sra1
|
steroid receptor RNA activator 1 |
| chr15_-_41734639 | 0.07 |
ENSDART00000154230
ENSDART00000167443 |
ftr90
|
finTRIM family, member 90 |
| chr23_+_45845423 | 0.07 |
ENSDART00000183404
|
lmnl3
|
lamin L3 |
| chr10_-_22912255 | 0.05 |
ENSDART00000131992
|
si:ch1073-143l10.2
|
si:ch1073-143l10.2 |
| chr22_-_7778265 | 0.05 |
ENSDART00000097276
|
si:ch73-44m9.3
|
si:ch73-44m9.3 |
| chr1_-_7951002 | 0.04 |
ENSDART00000138187
|
si:dkey-79f11.8
|
si:dkey-79f11.8 |
| chr6_-_43262400 | 0.04 |
ENSDART00000156267
|
frmd4ba
|
FERM domain containing 4Ba |
| chr1_+_961607 | 0.04 |
ENSDART00000184660
|
n6amt1
|
N-6 adenine-specific DNA methyltransferase 1 |
| chr8_-_11832771 | 0.03 |
ENSDART00000193412
ENSDART00000184720 ENSDART00000139947 |
rapgef1a
|
Rap guanine nucleotide exchange factor (GEF) 1a |
| chr16_-_44709832 | 0.02 |
ENSDART00000156784
|
si:ch211-151m7.6
|
si:ch211-151m7.6 |
| chr15_+_37331585 | 0.02 |
ENSDART00000170715
|
zgc:171592
|
zgc:171592 |
| chr5_+_34578479 | 0.01 |
ENSDART00000016314
|
gfm2
|
G elongation factor, mitochondrial 2 |
| chr15_+_11381532 | 0.00 |
ENSDART00000124172
|
si:ch73-321d9.2
|
si:ch73-321d9.2 |
Network of associatons between targets according to the STRING database.
First level regulatory network of irx1b+irx4a+irx4b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:0019464 | glycine catabolic process(GO:0006546) glycine decarboxylation via glycine cleavage system(GO:0019464) |
| 0.1 | 0.6 | GO:1903589 | regulation of blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:1903587) positive regulation of blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:1903589) |
| 0.1 | 0.9 | GO:0042661 | regulation of mesodermal cell fate specification(GO:0042661) |
| 0.0 | 0.3 | GO:0044090 | positive regulation of vacuole organization(GO:0044090) |
| 0.0 | 0.3 | GO:0034501 | protein localization to kinetochore(GO:0034501) protein localization to chromosome, centromeric region(GO:0071459) |
| 0.0 | 0.1 | GO:0006740 | NADPH regeneration(GO:0006740) |
| 0.0 | 0.1 | GO:0000972 | transcription-dependent tethering of RNA polymerase II gene DNA at nuclear periphery(GO:0000972) |
| 0.0 | 0.1 | GO:0039703 | viral genome replication(GO:0019079) negative stranded viral RNA replication(GO:0039689) viral RNA genome replication(GO:0039694) RNA replication(GO:0039703) multi-organism biosynthetic process(GO:0044034) |
| 0.0 | 0.2 | GO:0000737 | DNA catabolic process, endonucleolytic(GO:0000737) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:0005960 | glycine cleavage complex(GO:0005960) |
| 0.1 | 0.3 | GO:0000818 | nuclear MIS12/MIND complex(GO:0000818) |
| 0.1 | 0.4 | GO:0030914 | STAGA complex(GO:0030914) |
| 0.0 | 0.1 | GO:0031233 | intrinsic component of external side of plasma membrane(GO:0031233) anchored component of external side of plasma membrane(GO:0031362) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | GO:0060182 | apelin receptor activity(GO:0060182) |
| 0.1 | 0.7 | GO:0016594 | glycine binding(GO:0016594) |
| 0.0 | 0.3 | GO:0035613 | RNA stem-loop binding(GO:0035613) |
| 0.0 | 0.1 | GO:0016652 | NAD(P)+ transhydrogenase activity(GO:0008746) oxidoreductase activity, acting on NAD(P)H, NAD(P) as acceptor(GO:0016652) |
| 0.0 | 0.3 | GO:0035250 | UDP-galactosyltransferase activity(GO:0035250) |
| 0.0 | 0.1 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.0 | 0.2 | GO:0004530 | deoxyribonuclease I activity(GO:0004530) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | PID MYC PATHWAY | C-MYC pathway |
| 0.0 | 0.1 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | REACTOME A TETRASACCHARIDE LINKER SEQUENCE IS REQUIRED FOR GAG SYNTHESIS | Genes involved in A tetrasaccharide linker sequence is required for GAG synthesis |