Project
PRJNA195909:zebrafish embryo and larva development
Navigation
Downloads
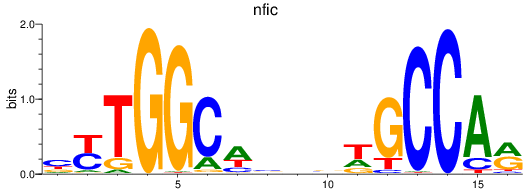
Results for nfic
Z-value: 1.13
Transcription factors associated with nfic
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
nfic
|
ENSDARG00000043210 | nuclear factor I/C |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| nfic | dr11_v1_chr8_+_20776654_20776654 | 0.87 | 2.6e-03 | Click! |
Activity profile of nfic motif
Sorted Z-values of nfic motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr3_-_55139127 | 3.96 |
ENSDART00000115324
|
hbae1.3
|
hemoglobin, alpha embryonic 1.3 |
| chr1_+_7546259 | 3.59 |
ENSDART00000015732
|
mylz3
|
myosin, light polypeptide 3, skeletal muscle |
| chr10_+_44042033 | 3.12 |
ENSDART00000190006
ENSDART00000046172 |
cryba4
|
crystallin, beta A4 |
| chr17_-_125091 | 2.98 |
ENSDART00000158825
|
actc1b
|
actin, alpha, cardiac muscle 1b |
| chr2_+_55982940 | 2.13 |
ENSDART00000097753
ENSDART00000097751 |
nmrk2
|
nicotinamide riboside kinase 2 |
| chr16_+_23397785 | 2.08 |
ENSDART00000148961
|
s100a10b
|
S100 calcium binding protein A10b |
| chr19_-_5332784 | 1.71 |
ENSDART00000010373
|
krt1-19d
|
keratin, type 1, gene 19d |
| chr17_+_52822422 | 1.68 |
ENSDART00000158273
ENSDART00000161414 |
meis2a
|
Meis homeobox 2a |
| chr1_+_4101741 | 1.46 |
ENSDART00000163793
|
slitrk6
|
SLIT and NTRK-like family, member 6 |
| chr1_-_44434707 | 1.34 |
ENSDART00000110148
|
cryba1l2
|
crystallin, beta A1, like 2 |
| chr1_+_49568335 | 1.33 |
ENSDART00000142957
|
col17a1a
|
collagen, type XVII, alpha 1a |
| chr11_+_13630107 | 1.30 |
ENSDART00000172220
|
si:ch211-1a19.3
|
si:ch211-1a19.3 |
| chr23_+_18944388 | 1.22 |
ENSDART00000104487
|
cox4i2
|
cytochrome c oxidase subunit IV isoform 2 |
| chr16_-_43025885 | 1.21 |
ENSDART00000193146
ENSDART00000157302 |
si:dkey-7j14.5
|
si:dkey-7j14.5 |
| chr7_-_26306546 | 1.13 |
ENSDART00000140817
|
zgc:77439
|
zgc:77439 |
| chr20_-_47704973 | 1.09 |
ENSDART00000174808
|
tfap2b
|
transcription factor AP-2 beta |
| chr7_-_26306954 | 1.02 |
ENSDART00000057288
|
zgc:77439
|
zgc:77439 |
| chr2_-_8017579 | 1.00 |
ENSDART00000040209
|
ephb3a
|
eph receptor B3a |
| chr8_+_25616946 | 0.95 |
ENSDART00000133983
|
slc38a5a
|
solute carrier family 38, member 5a |
| chr22_-_10487490 | 0.93 |
ENSDART00000064798
|
aspn
|
asporin (LRR class 1) |
| chr6_-_13114406 | 0.92 |
ENSDART00000188015
|
zgc:194469
|
zgc:194469 |
| chr10_-_8358396 | 0.91 |
ENSDART00000059322
|
csgalnact1a
|
chondroitin sulfate N-acetylgalactosaminyltransferase 1a |
| chr7_-_55675617 | 0.87 |
ENSDART00000021009
ENSDART00000188631 |
cbfa2t3
|
core-binding factor, runt domain, alpha subunit 2; translocated to, 3 |
| chr4_+_22365061 | 0.87 |
ENSDART00000039277
|
lhfpl3
|
LHFPL tetraspan subfamily member 3 |
| chr16_+_50100420 | 0.85 |
ENSDART00000128167
|
nr1d2a
|
nuclear receptor subfamily 1, group D, member 2a |
| chr8_-_6877390 | 0.81 |
ENSDART00000170883
ENSDART00000005321 |
neflb
|
neurofilament, light polypeptide b |
| chr22_-_33679277 | 0.80 |
ENSDART00000169948
|
FO904977.1
|
|
| chr14_+_20893065 | 0.79 |
ENSDART00000079452
|
lygl1
|
lysozyme g-like 1 |
| chr17_-_43466317 | 0.78 |
ENSDART00000155313
|
hspa4l
|
heat shock protein 4 like |
| chr19_+_30388186 | 0.77 |
ENSDART00000103474
|
tspan13b
|
tetraspanin 13b |
| chr4_+_74131530 | 0.74 |
ENSDART00000174125
|
CABZ01072523.1
|
|
| chr12_+_30653047 | 0.74 |
ENSDART00000148562
|
thbs2b
|
thrombospondin 2b |
| chr1_-_28629471 | 0.73 |
ENSDART00000121758
|
ednrba
|
endothelin receptor Ba |
| chr19_+_43780970 | 0.71 |
ENSDART00000063870
|
rpl11
|
ribosomal protein L11 |
| chr2_-_17827983 | 0.69 |
ENSDART00000166518
|
ptprfb
|
protein tyrosine phosphatase, receptor type, f, b |
| chr6_-_35401282 | 0.68 |
ENSDART00000127612
|
rgs5a
|
regulator of G protein signaling 5a |
| chr4_-_17629444 | 0.68 |
ENSDART00000108814
|
nrip2
|
nuclear receptor interacting protein 2 |
| chr1_+_33328857 | 0.65 |
ENSDART00000137151
|
mxra5a
|
matrix-remodelling associated 5a |
| chr2_+_7818368 | 0.64 |
ENSDART00000007068
|
kcnmb2
|
potassium large conductance calcium-activated channel, subfamily M, beta member 2 |
| chr21_-_34658266 | 0.63 |
ENSDART00000023038
|
dacha
|
dachshund a |
| chr5_-_57898008 | 0.63 |
ENSDART00000050945
|
layna
|
layilin a |
| chr12_+_30168342 | 0.62 |
ENSDART00000142756
|
ablim1b
|
actin binding LIM protein 1b |
| chr20_-_25486384 | 0.62 |
ENSDART00000141340
|
si:dkey-183n20.15
|
si:dkey-183n20.15 |
| chr19_+_9786722 | 0.62 |
ENSDART00000138310
|
cacng6a
|
calcium channel, voltage-dependent, gamma subunit 6a |
| chr14_+_8174828 | 0.60 |
ENSDART00000167228
|
psd2
|
pleckstrin and Sec7 domain containing 2 |
| chr6_-_8295657 | 0.60 |
ENSDART00000185135
|
slc44a2
|
solute carrier family 44 (choline transporter), member 2 |
| chr5_-_46329880 | 0.60 |
ENSDART00000156577
|
si:ch211-130m23.5
|
si:ch211-130m23.5 |
| chr6_-_34006917 | 0.58 |
ENSDART00000190702
ENSDART00000184003 |
tie1
|
tyrosine kinase with immunoglobulin-like and EGF-like domains 1 |
| chr17_-_30863252 | 0.55 |
ENSDART00000062811
|
yy1a
|
YY1 transcription factor a |
| chr9_-_48937089 | 0.54 |
ENSDART00000193442
|
cers6
|
ceramide synthase 6 |
| chr10_-_17232372 | 0.53 |
ENSDART00000135679
|
rab36
|
RAB36, member RAS oncogene family |
| chr11_-_43948885 | 0.51 |
ENSDART00000164522
|
CABZ01074203.1
|
|
| chr2_+_47905735 | 0.51 |
ENSDART00000159701
|
ftr23
|
finTRIM family, member 23 |
| chr9_-_48937240 | 0.50 |
ENSDART00000075627
|
cers6
|
ceramide synthase 6 |
| chr10_-_27566481 | 0.50 |
ENSDART00000078920
|
auts2a
|
autism susceptibility candidate 2a |
| chr23_-_45504991 | 0.49 |
ENSDART00000148761
|
col24a1
|
collagen type XXIV alpha 1 |
| chr8_-_44223899 | 0.46 |
ENSDART00000143020
|
stx2b
|
syntaxin 2b |
| chr7_-_29341233 | 0.46 |
ENSDART00000140938
ENSDART00000147251 |
trpm1a
|
transient receptor potential cation channel, subfamily M, member 1a |
| chr10_-_1718395 | 0.45 |
ENSDART00000137620
|
si:ch73-46j18.5
|
si:ch73-46j18.5 |
| chr5_-_68826177 | 0.45 |
ENSDART00000136605
|
si:ch211-283h6.4
|
si:ch211-283h6.4 |
| chr12_+_35654749 | 0.45 |
ENSDART00000169889
ENSDART00000167873 |
baiap2b
|
BAI1-associated protein 2b |
| chr5_+_37517800 | 0.44 |
ENSDART00000048107
|
FP102018.1
|
Danio rerio latent transforming growth factor beta binding protein 3 (ltbp3), mRNA. |
| chr11_+_14284866 | 0.44 |
ENSDART00000163729
|
si:ch211-262i1.3
|
si:ch211-262i1.3 |
| chr24_-_12745222 | 0.44 |
ENSDART00000151836
|
si:ch211-196f5.9
|
si:ch211-196f5.9 |
| chr3_+_57038033 | 0.43 |
ENSDART00000162930
|
bahcc1a
|
BAH domain and coiled-coil containing 1a |
| chr15_-_14469704 | 0.42 |
ENSDART00000185077
|
numbl
|
numb homolog (Drosophila)-like |
| chr25_-_36248053 | 0.42 |
ENSDART00000134928
|
nfatc3b
|
nuclear factor of activated T cells 3b |
| chr7_+_36898850 | 0.41 |
ENSDART00000113342
|
tox3
|
TOX high mobility group box family member 3 |
| chr10_+_7636811 | 0.39 |
ENSDART00000160673
|
hint1
|
histidine triad nucleotide binding protein 1 |
| chr8_-_14375890 | 0.38 |
ENSDART00000090306
|
xpr1a
|
xenotropic and polytropic retrovirus receptor 1a |
| chr15_+_31806303 | 0.37 |
ENSDART00000155902
|
frya
|
furry homolog a (Drosophila) |
| chr2_+_31665836 | 0.37 |
ENSDART00000135411
ENSDART00000143914 |
si:ch211-106h4.12
|
si:ch211-106h4.12 |
| chr9_+_37152564 | 0.36 |
ENSDART00000189497
|
gli2a
|
GLI family zinc finger 2a |
| chr20_-_42100932 | 0.34 |
ENSDART00000191930
|
slc35f1
|
solute carrier family 35, member F1 |
| chr8_-_44223473 | 0.32 |
ENSDART00000098525
|
stx2b
|
syntaxin 2b |
| chr11_-_42980535 | 0.31 |
ENSDART00000181160
ENSDART00000192064 |
CABZ01069998.1
|
|
| chr20_-_33976305 | 0.29 |
ENSDART00000111022
|
sele
|
selectin E |
| chr17_-_10309766 | 0.29 |
ENSDART00000160994
|
ttc6
|
tetratricopeptide repeat domain 6 |
| chr7_+_36898622 | 0.28 |
ENSDART00000190773
|
tox3
|
TOX high mobility group box family member 3 |
| chr23_+_25822742 | 0.27 |
ENSDART00000184436
|
r3hdml
|
R3H domain containing-like |
| chr9_-_23884937 | 0.27 |
ENSDART00000141377
|
FAM124A
|
si:dkeyp-74a11.1 |
| chr21_+_38634296 | 0.26 |
ENSDART00000114467
|
AL935199.1
|
|
| chr10_-_16065185 | 0.26 |
ENSDART00000187266
|
si:dkey-184a18.5
|
si:dkey-184a18.5 |
| chr24_-_31942656 | 0.25 |
ENSDART00000180308
ENSDART00000016879 |
c1ql3a
|
complement component 1, q subcomponent-like 3a |
| chr4_+_61171310 | 0.24 |
ENSDART00000141738
|
si:dkey-9p20.18
|
si:dkey-9p20.18 |
| chr22_-_15693085 | 0.23 |
ENSDART00000141861
|
si:ch1073-396h14.1
|
si:ch1073-396h14.1 |
| chr6_+_41186320 | 0.23 |
ENSDART00000025241
|
opn1mw2
|
opsin 1 (cone pigments), medium-wave-sensitive, 2 |
| chr22_+_32120156 | 0.22 |
ENSDART00000149666
|
dock3
|
dedicator of cytokinesis 3 |
| chr24_-_22756508 | 0.22 |
ENSDART00000035409
ENSDART00000146247 |
zc2hc1a
|
zinc finger, C2HC-type containing 1A |
| chr19_+_9091673 | 0.21 |
ENSDART00000052898
|
si:ch211-81a5.5
|
si:ch211-81a5.5 |
| chr7_+_52135791 | 0.20 |
ENSDART00000098705
|
cyp2x12
|
cytochrome P450, family 2, subfamily X, polypeptide 12 |
| chr4_-_38033800 | 0.20 |
ENSDART00000159662
|
si:dkeyp-82b4.4
|
si:dkeyp-82b4.4 |
| chr9_-_23885220 | 0.18 |
ENSDART00000112058
|
FAM124A
|
si:dkeyp-74a11.1 |
| chr3_+_57997980 | 0.17 |
ENSDART00000168477
ENSDART00000193840 |
pycr1a
|
pyrroline-5-carboxylate reductase 1a |
| chr16_+_33143503 | 0.16 |
ENSDART00000058471
ENSDART00000179385 |
rhbdl2
|
rhomboid, veinlet-like 2 (Drosophila) |
| chr17_+_5846202 | 0.16 |
ENSDART00000189713
|
fndc4b
|
fibronectin type III domain containing 4b |
| chr14_+_23811808 | 0.13 |
ENSDART00000014411
|
kctd16a
|
potassium channel tetramerization domain containing 16a |
| chr3_+_31039923 | 0.13 |
ENSDART00000147706
|
cox6a2
|
cytochrome c oxidase subunit VIa polypeptide 2 |
| chr13_-_22774491 | 0.13 |
ENSDART00000189320
|
si:ch211-150i13.1
|
si:ch211-150i13.1 |
| chr21_+_39361040 | 0.13 |
ENSDART00000085453
|
cluhb
|
clustered mitochondria (cluA/CLU1) homolog b |
| chr1_+_28089703 | 0.12 |
ENSDART00000166989
|
psip1a
|
PC4 and SFRS1 interacting protein 1a |
| chr20_+_51464670 | 0.12 |
ENSDART00000150110
|
thbd
|
thrombomodulin |
| chr24_-_35707552 | 0.12 |
ENSDART00000165199
|
mapre2
|
microtubule-associated protein, RP/EB family, member 2 |
| chr20_+_48739154 | 0.11 |
ENSDART00000100262
|
zgc:110348
|
zgc:110348 |
| chr15_+_21672281 | 0.11 |
ENSDART00000153923
|
si:dkey-40g16.5
|
si:dkey-40g16.5 |
| chr20_-_47188966 | 0.11 |
ENSDART00000152965
|
si:dkeyp-104f11.9
|
si:dkeyp-104f11.9 |
| chr21_+_10866421 | 0.10 |
ENSDART00000137858
|
alpk2
|
alpha-kinase 2 |
| chr3_-_37681824 | 0.08 |
ENSDART00000185858
|
gpatch8
|
G patch domain containing 8 |
| chr17_-_35352690 | 0.05 |
ENSDART00000016053
|
rnf144aa
|
ring finger protein 144aa |
| chr1_+_35194454 | 0.05 |
ENSDART00000165389
|
sc:d189
|
sc:d189 |
| chr6_-_35032792 | 0.05 |
ENSDART00000168256
|
ddr2a
|
discoidin domain receptor tyrosine kinase 2a |
| chr3_-_38777553 | 0.05 |
ENSDART00000193045
|
znf281a
|
zinc finger protein 281a |
| chr3_-_40528333 | 0.05 |
ENSDART00000193047
|
actb2
|
actin, beta 2 |
| chr2_+_6303639 | 0.04 |
ENSDART00000132859
|
otol1a
|
otolin 1a |
| chr22_-_29083070 | 0.04 |
ENSDART00000104812
ENSDART00000172576 |
cbx6a
|
chromobox homolog 6a |
| chr12_+_4160804 | 0.04 |
ENSDART00000152515
|
itgam
|
integrin, alpha M (complement component 3 receptor 3 subunit) |
| chr6_+_7533601 | 0.03 |
ENSDART00000057823
|
pa2g4a
|
proliferation-associated 2G4, a |
| chr15_-_18091157 | 0.02 |
ENSDART00000153848
|
phldb1b
|
pleckstrin homology-like domain, family B, member 1b |
| chr5_+_55225497 | 0.02 |
ENSDART00000144087
|
tmc2a
|
transmembrane channel-like 2a |
| chr2_+_16460321 | 0.02 |
ENSDART00000145107
|
agfg1b
|
ArfGAP with FG repeats 1b |
| chr3_-_34279109 | 0.02 |
ENSDART00000183255
|
tnrc6c1
|
trinucleotide repeat containing 6C1 |
| chr10_+_40684758 | 0.01 |
ENSDART00000133648
|
taar19h
|
trace amine associated receptor 19h |
| chr9_-_46382637 | 0.00 |
ENSDART00000085738
|
lct
|
lactase |
| chr23_-_18572685 | 0.00 |
ENSDART00000047175
|
ssr4
|
signal sequence receptor, delta |
| chr20_-_8110672 | 0.00 |
ENSDART00000113993
|
si:ch211-232i5.1
|
si:ch211-232i5.1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of nfic
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.8 | GO:0016998 | cell wall macromolecule catabolic process(GO:0016998) |
| 0.2 | 1.0 | GO:0089709 | L-histidine transmembrane transport(GO:0089709) L-histidine transport(GO:1902024) |
| 0.2 | 4.0 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.1 | 0.6 | GO:0015871 | choline transport(GO:0015871) |
| 0.1 | 0.6 | GO:0061577 | calcium ion transmembrane transport via high voltage-gated calcium channel(GO:0061577) regulation of calcium ion transmembrane transport via high voltage-gated calcium channel(GO:1902514) |
| 0.1 | 1.0 | GO:0051965 | positive regulation of synapse assembly(GO:0051965) |
| 0.1 | 0.6 | GO:0005513 | detection of calcium ion(GO:0005513) |
| 0.1 | 1.3 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.1 | 0.4 | GO:0048618 | post-embryonic foregut morphogenesis(GO:0048618) |
| 0.1 | 0.5 | GO:0038065 | collagen-activated tyrosine kinase receptor signaling pathway(GO:0038063) collagen-activated signaling pathway(GO:0038065) |
| 0.1 | 0.7 | GO:0097374 | sensory neuron axon guidance(GO:0097374) |
| 0.1 | 0.6 | GO:0090136 | epithelial cell-cell adhesion(GO:0090136) |
| 0.1 | 0.9 | GO:0030206 | chondroitin sulfate biosynthetic process(GO:0030206) |
| 0.0 | 0.4 | GO:2000251 | positive regulation of actin cytoskeleton reorganization(GO:2000251) |
| 0.0 | 4.5 | GO:0002088 | lens development in camera-type eye(GO:0002088) |
| 0.0 | 0.7 | GO:0050935 | iridophore differentiation(GO:0050935) |
| 0.0 | 0.4 | GO:0033173 | calcineurin-NFAT signaling cascade(GO:0033173) |
| 0.0 | 0.6 | GO:0030032 | lamellipodium assembly(GO:0030032) |
| 0.0 | 0.8 | GO:0007340 | acrosome reaction(GO:0007340) |
| 0.0 | 0.6 | GO:0030537 | larval locomotory behavior(GO:0008345) larval behavior(GO:0030537) |
| 0.0 | 0.5 | GO:0021884 | forebrain neuron development(GO:0021884) |
| 0.0 | 0.2 | GO:0055129 | proline biosynthetic process(GO:0006561) L-proline biosynthetic process(GO:0055129) |
| 0.0 | 0.8 | GO:0043524 | negative regulation of neuron apoptotic process(GO:0043524) |
| 0.0 | 1.7 | GO:0009880 | embryonic pattern specification(GO:0009880) |
| 0.0 | 0.7 | GO:0042593 | carbohydrate homeostasis(GO:0033500) glucose homeostasis(GO:0042593) |
| 0.0 | 0.7 | GO:0016525 | negative regulation of angiogenesis(GO:0016525) |
| 0.0 | 1.0 | GO:0046513 | ceramide biosynthetic process(GO:0046513) |
| 0.0 | 1.5 | GO:0071466 | cellular response to xenobiotic stimulus(GO:0071466) |
| 0.0 | 0.5 | GO:0007216 | G-protein coupled glutamate receptor signaling pathway(GO:0007216) |
| 0.0 | 0.4 | GO:0045746 | negative regulation of Notch signaling pathway(GO:0045746) |
| 0.0 | 0.1 | GO:0035372 | protein localization to microtubule(GO:0035372) protein localization to microtubule plus-end(GO:1904825) |
| 0.0 | 0.4 | GO:0006817 | phosphate ion transport(GO:0006817) |
| 0.0 | 0.4 | GO:0009154 | purine ribonucleotide catabolic process(GO:0009154) |
| 0.0 | 0.1 | GO:0048312 | intracellular distribution of mitochondria(GO:0048312) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 4.0 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 0.2 | 3.0 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.2 | 0.5 | GO:0005592 | collagen type XI trimer(GO:0005592) |
| 0.2 | 0.8 | GO:0005883 | neurofilament(GO:0005883) |
| 0.1 | 1.3 | GO:0005751 | mitochondrial respiratory chain complex IV(GO:0005751) |
| 0.1 | 0.6 | GO:1990454 | L-type voltage-gated calcium channel complex(GO:1990454) |
| 0.0 | 0.8 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
| 0.0 | 1.7 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 0.9 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.0 | 0.6 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.0 | 0.6 | GO:0001725 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.0 | 0.7 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.0 | 0.0 | GO:0098651 | collagen type IV trimer(GO:0005587) network-forming collagen trimer(GO:0098642) collagen network(GO:0098645) basement membrane collagen trimer(GO:0098651) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 4.0 | GO:0031720 | haptoglobin binding(GO:0031720) |
| 0.3 | 2.1 | GO:0050262 | ribosylnicotinamide kinase activity(GO:0050262) |
| 0.2 | 2.1 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 0.2 | 0.8 | GO:0003796 | lysozyme activity(GO:0003796) |
| 0.1 | 0.6 | GO:0015220 | choline transmembrane transporter activity(GO:0015220) |
| 0.1 | 0.4 | GO:0000822 | inositol hexakisphosphate binding(GO:0000822) |
| 0.1 | 0.7 | GO:0004962 | endothelin receptor activity(GO:0004962) |
| 0.1 | 0.9 | GO:0047238 | glucuronosyl-N-acetylgalactosaminyl-proteoglycan 4-beta-N-acetylgalactosaminyltransferase activity(GO:0047238) |
| 0.1 | 1.0 | GO:0015186 | L-glutamine transmembrane transporter activity(GO:0015186) |
| 0.1 | 2.1 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.1 | 1.0 | GO:0050291 | sphingosine N-acyltransferase activity(GO:0050291) |
| 0.1 | 1.0 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.1 | 4.5 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.0 | 1.3 | GO:0004129 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors(GO:0016675) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.0 | 0.6 | GO:0015269 | calcium-activated potassium channel activity(GO:0015269) |
| 0.0 | 0.7 | GO:0070001 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.0 | 0.2 | GO:0004735 | pyrroline-5-carboxylate reductase activity(GO:0004735) |
| 0.0 | 1.0 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 0.6 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.0 | 0.6 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.0 | 1.4 | GO:0008201 | heparin binding(GO:0008201) |
| 0.0 | 1.9 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 0.8 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.0 | 0.7 | GO:0031490 | chromatin DNA binding(GO:0031490) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.9 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 0.8 | PID ATF2 PATHWAY | ATF-2 transcription factor network |
| 0.0 | 0.8 | PID ERBB4 PATHWAY | ErbB4 signaling events |
| 0.0 | 0.7 | PID P53 REGULATION PATHWAY | p53 pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |
| 0.0 | 0.2 | REACTOME OPSINS | Genes involved in Opsins |
| 0.0 | 0.6 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.0 | 1.0 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 0.1 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |