Project
PRJNA195909:zebrafish embryo and larva development
Navigation
Downloads
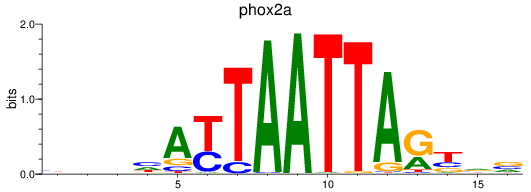
Results for phox2a
Z-value: 0.78
Transcription factors associated with phox2a
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
phox2a
|
ENSDARG00000007406 | paired like homeobox 2A |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| phox2a | dr11_v1_chr15_+_47440477_47440477 | 0.86 | 2.9e-03 | Click! |
Activity profile of phox2a motif
Sorted Z-values of phox2a motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr17_+_15433518 | 2.43 |
ENSDART00000026180
|
fabp7a
|
fatty acid binding protein 7, brain, a |
| chr17_+_15433671 | 2.32 |
ENSDART00000149568
|
fabp7a
|
fatty acid binding protein 7, brain, a |
| chr8_+_16025554 | 1.95 |
ENSDART00000110171
|
elavl4
|
ELAV like neuron-specific RNA binding protein 4 |
| chr12_-_35830625 | 1.52 |
ENSDART00000180028
|
CU459056.1
|
|
| chr4_-_9891874 | 1.27 |
ENSDART00000067193
|
adm2a
|
adrenomedullin 2a |
| chr10_-_22918214 | 1.10 |
ENSDART00000163908
|
rnasekb
|
ribonuclease, RNase K b |
| chr4_+_21129752 | 1.01 |
ENSDART00000169764
|
syt1a
|
synaptotagmin Ia |
| chr2_+_2223837 | 0.89 |
ENSDART00000101038
ENSDART00000129354 |
tmie
|
transmembrane inner ear |
| chr2_+_50608099 | 0.83 |
ENSDART00000185805
ENSDART00000111135 |
neurod6b
|
neuronal differentiation 6b |
| chr11_-_42554290 | 0.83 |
ENSDART00000130573
|
atp6ap1la
|
ATPase H+ transporting accessory protein 1 like a |
| chr21_+_28478663 | 0.80 |
ENSDART00000077887
ENSDART00000134150 |
slc22a6l
|
solute carrier family 22 (organic anion transporter), member 6, like |
| chr13_+_255067 | 0.80 |
ENSDART00000102505
|
foxg1d
|
forkhead box G1d |
| chr14_+_17376940 | 0.80 |
ENSDART00000054590
ENSDART00000010148 |
spon2b
|
spondin 2b, extracellular matrix protein |
| chr22_-_10121880 | 0.72 |
ENSDART00000002348
|
rdh5
|
retinol dehydrogenase 5 (11-cis/9-cis) |
| chr16_-_25568512 | 0.67 |
ENSDART00000149411
|
atxn1b
|
ataxin 1b |
| chr23_+_43255328 | 0.65 |
ENSDART00000102712
|
tgm2a
|
transglutaminase 2, C polypeptide A |
| chr21_-_42100471 | 0.64 |
ENSDART00000166148
|
gabra1
|
gamma-aminobutyric acid (GABA) A receptor, alpha 1 |
| chr9_+_21165484 | 0.54 |
ENSDART00000177286
|
si:rp71-68n21.9
|
si:rp71-68n21.9 |
| chr16_-_11798994 | 0.47 |
ENSDART00000135408
|
cnfn
|
cornifelin |
| chr3_-_56541723 | 0.46 |
ENSDART00000156398
ENSDART00000050576 ENSDART00000184874 |
si:ch211-189a21.1
cyth1a
|
si:ch211-189a21.1 cytohesin 1a |
| chr15_-_22074315 | 0.46 |
ENSDART00000149830
|
drd2a
|
dopamine receptor D2a |
| chr7_-_30174882 | 0.40 |
ENSDART00000110409
|
frmd5
|
FERM domain containing 5 |
| chr2_-_9818640 | 0.40 |
ENSDART00000139499
ENSDART00000165548 ENSDART00000012442 ENSDART00000046587 |
ap2m1b
|
adaptor-related protein complex 2, mu 1 subunit, b |
| chr11_-_2838699 | 0.35 |
ENSDART00000066189
|
lhfpl5a
|
LHFPL tetraspan subfamily member 5a |
| chr5_-_30620625 | 0.34 |
ENSDART00000098273
|
tcnl
|
transcobalamin like |
| chr18_+_28106139 | 0.32 |
ENSDART00000089615
|
kiaa1549lb
|
KIAA1549-like b |
| chr4_+_6032640 | 0.31 |
ENSDART00000157487
|
tfec
|
transcription factor EC |
| chr17_+_49281597 | 0.30 |
ENSDART00000155599
|
zgc:113176
|
zgc:113176 |
| chr6_-_35779348 | 0.28 |
ENSDART00000191159
|
brinp3a.1
|
bone morphogenetic protein/retinoic acid inducible neural-specific 3a, tandem duplicate 1 |
| chr15_-_14552101 | 0.25 |
ENSDART00000171169
|
numbl
|
numb homolog (Drosophila)-like |
| chr20_-_28642061 | 0.25 |
ENSDART00000135513
|
rgs6
|
regulator of G protein signaling 6 |
| chr22_+_7738966 | 0.24 |
ENSDART00000147073
|
si:ch73-44m9.5
|
si:ch73-44m9.5 |
| chr13_+_22476742 | 0.23 |
ENSDART00000078759
ENSDART00000130101 ENSDART00000137220 ENSDART00000133065 ENSDART00000147348 |
ldb3a
|
LIM domain binding 3a |
| chr6_+_13117598 | 0.22 |
ENSDART00000104744
|
casp8l1
|
caspase 8, apoptosis-related cysteine peptidase, like 1 |
| chr1_-_44704261 | 0.22 |
ENSDART00000133210
|
si:dkey-28b4.8
|
si:dkey-28b4.8 |
| chr6_+_24420523 | 0.21 |
ENSDART00000185461
|
tgfbr3
|
transforming growth factor, beta receptor III |
| chr3_+_23692462 | 0.21 |
ENSDART00000145934
|
hoxb7a
|
homeobox B7a |
| chr15_+_5360407 | 0.16 |
ENSDART00000110420
|
or112-1
|
odorant receptor, family A, subfamily 112, member 1 |
| chr21_+_32820175 | 0.16 |
ENSDART00000076903
|
adra2db
|
adrenergic, alpha-2D-, receptor b |
| chr10_+_42423318 | 0.14 |
ENSDART00000134282
|
npy8ar
|
neuropeptide Y receptor Y8a |
| chr1_-_19502322 | 0.14 |
ENSDART00000181888
ENSDART00000044030 |
kitb
|
v-kit Hardy-Zuckerman 4 feline sarcoma viral oncogene homolog b |
| chr17_-_200316 | 0.12 |
ENSDART00000190561
|
CABZ01083778.1
|
|
| chr22_-_8725768 | 0.11 |
ENSDART00000189873
ENSDART00000181819 |
si:ch73-27e22.1
si:ch73-27e22.8
|
si:ch73-27e22.1 si:ch73-27e22.8 |
| chr20_-_9095105 | 0.11 |
ENSDART00000140792
|
oma1
|
OMA1 zinc metallopeptidase |
| chr18_+_34362608 | 0.06 |
ENSDART00000131478
|
kcnab1a
|
potassium voltage-gated channel, shaker-related subfamily, beta member 1 a |
| chr1_-_31171242 | 0.06 |
ENSDART00000190294
|
kcnq5b
|
potassium voltage-gated channel, KQT-like subfamily, member 5b |
| chr9_-_32177117 | 0.02 |
ENSDART00000078568
|
sf3b1
|
splicing factor 3b, subunit 1 |
| chr17_-_10122204 | 0.02 |
ENSDART00000160751
|
BX088587.1
|
|
| chr20_-_29864390 | 0.01 |
ENSDART00000161834
ENSDART00000132278 |
rnf144ab
|
ring finger protein 144ab |
Network of associatons between targets according to the STRING database.
First level regulatory network of phox2a
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | GO:0070208 | protein heterotrimerization(GO:0070208) |
| 0.1 | 0.7 | GO:0042362 | fat-soluble vitamin biosynthetic process(GO:0042362) |
| 0.0 | 0.9 | GO:0060088 | auditory receptor cell stereocilium organization(GO:0060088) |
| 0.0 | 1.0 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.0 | 0.1 | GO:0038093 | Fc receptor signaling pathway(GO:0038093) |
| 0.0 | 0.5 | GO:0051481 | negative regulation of cytosolic calcium ion concentration(GO:0051481) negative regulation of synaptic transmission, glutamatergic(GO:0051967) |
| 0.0 | 0.2 | GO:0071881 | adenylate cyclase-inhibiting adrenergic receptor signaling pathway(GO:0071881) |
| 0.0 | 0.7 | GO:0018149 | peptide cross-linking(GO:0018149) |
| 0.0 | 0.3 | GO:0061484 | hematopoietic stem cell homeostasis(GO:0061484) |
| 0.0 | 0.3 | GO:0071300 | cellular response to retinoic acid(GO:0071300) |
| 0.0 | 0.4 | GO:0060122 | inner ear receptor stereocilium organization(GO:0060122) |
| 0.0 | 0.1 | GO:1903817 | negative regulation of potassium ion transmembrane transporter activity(GO:1901017) negative regulation of potassium ion transmembrane transport(GO:1901380) negative regulation of delayed rectifier potassium channel activity(GO:1902260) negative regulation of voltage-gated potassium channel activity(GO:1903817) |
| 0.0 | 0.1 | GO:1990845 | adaptive thermogenesis(GO:1990845) |
| 0.0 | 0.8 | GO:0030641 | regulation of cellular pH(GO:0030641) |
| 0.0 | 1.1 | GO:0090305 | nucleic acid phosphodiester bond hydrolysis(GO:0090305) |
| 0.0 | 0.5 | GO:0032011 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.0 | GO:0042584 | chromaffin granule membrane(GO:0042584) |
| 0.1 | 0.8 | GO:0033181 | plasma membrane proton-transporting V-type ATPase complex(GO:0033181) |
| 0.0 | 2.0 | GO:0043204 | perikaryon(GO:0043204) |
| 0.0 | 0.4 | GO:0030122 | AP-2 adaptor complex(GO:0030122) |
| 0.0 | 0.6 | GO:1902710 | GABA receptor complex(GO:1902710) GABA-A receptor complex(GO:1902711) |
| 0.0 | 0.5 | GO:0098978 | glutamatergic synapse(GO:0098978) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 4.7 | GO:0005504 | fatty acid binding(GO:0005504) |
| 0.0 | 0.6 | GO:0022851 | benzodiazepine receptor activity(GO:0008503) GABA-gated chloride ion channel activity(GO:0022851) |
| 0.0 | 0.2 | GO:0050431 | transforming growth factor beta binding(GO:0050431) |
| 0.0 | 0.5 | GO:0001591 | dopamine neurotransmitter receptor activity, coupled via Gi/Go(GO:0001591) |
| 0.0 | 0.7 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 0.0 | 0.2 | GO:0004938 | alpha2-adrenergic receptor activity(GO:0004938) |
| 0.0 | 1.0 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.0 | 0.4 | GO:0035615 | clathrin adaptor activity(GO:0035615) endocytic adaptor activity(GO:0098748) |
| 0.0 | 1.1 | GO:0004521 | endoribonuclease activity(GO:0004521) |
| 0.0 | 0.2 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.0 | 0.2 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) alpha-actinin binding(GO:0051393) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.7 | PID CONE PATHWAY | Visual signal transduction: Cones |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |