Project
PRJNA195909:zebrafish embryo and larva development
Navigation
Downloads
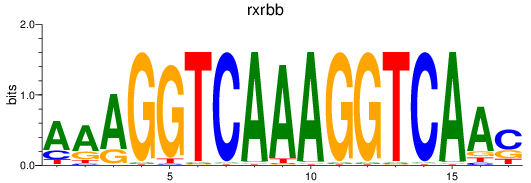
Results for rxrbb
Z-value: 0.40
Transcription factors associated with rxrbb
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
rxrbb
|
ENSDARG00000002006 | retinoid x receptor, beta b |
|
rxrbb
|
ENSDARG00000117079 | retinoid x receptor, beta b |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| rxrbb | dr11_v1_chr16_+_18534834_18534834 | -0.44 | 2.3e-01 | Click! |
Activity profile of rxrbb motif
Sorted Z-values of rxrbb motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_-_16451375 | 0.62 |
ENSDART00000192700
ENSDART00000128835 |
wu:fc23c09
|
wu:fc23c09 |
| chr19_-_3056235 | 0.51 |
ENSDART00000137020
|
bop1
|
block of proliferation 1 |
| chr4_+_45274792 | 0.50 |
ENSDART00000150295
|
znf1138
|
zinc finger protein 1138 |
| chr4_+_37503994 | 0.46 |
ENSDART00000158217
|
znf1102
|
zinc finger protein 1102 |
| chr4_-_37587110 | 0.39 |
ENSDART00000169302
|
znf1100
|
zinc finger protein 1100 |
| chr4_+_47656992 | 0.38 |
ENSDART00000161148
|
znf1040
|
zinc finger protein 1040 |
| chr4_-_64605318 | 0.37 |
ENSDART00000170391
|
znf1099
|
zinc finger protein 1099 |
| chr18_-_43866526 | 0.36 |
ENSDART00000111309
|
treh
|
trehalase (brush-border membrane glycoprotein) |
| chr1_+_12195700 | 0.35 |
ENSDART00000040307
|
tdrd7a
|
tudor domain containing 7 a |
| chr4_-_38421217 | 0.33 |
ENSDART00000164517
|
znf1092
|
szinc finger protein 1092 |
| chr4_-_56157199 | 0.32 |
ENSDART00000169806
|
si:ch211-207e19.15
|
si:ch211-207e19.15 |
| chr5_-_3839285 | 0.31 |
ENSDART00000122292
|
mlxipl
|
MLX interacting protein like |
| chr4_+_64921810 | 0.30 |
ENSDART00000157654
|
BX928743.1
|
|
| chr3_+_39566999 | 0.27 |
ENSDART00000146867
|
aldoaa
|
aldolase a, fructose-bisphosphate, a |
| chr3_-_36419641 | 0.26 |
ENSDART00000173545
|
cog1
|
component of oligomeric golgi complex 1 |
| chr18_-_43866001 | 0.24 |
ENSDART00000150218
|
treh
|
trehalase (brush-border membrane glycoprotein) |
| chr22_-_38274188 | 0.24 |
ENSDART00000139420
ENSDART00000015117 |
elavl2
|
ELAV like neuron-specific RNA binding protein 2 |
| chr7_-_24520866 | 0.23 |
ENSDART00000077039
|
faah2b
|
fatty acid amide hydrolase 2b |
| chr23_+_30730121 | 0.23 |
ENSDART00000134141
|
asxl1
|
additional sex combs like transcriptional regulator 1 |
| chr9_+_22782027 | 0.22 |
ENSDART00000090816
|
rif1
|
replication timing regulatory factor 1 |
| chr24_+_39034090 | 0.21 |
ENSDART00000185763
|
capn15
|
calpain 15 |
| chr23_+_2914577 | 0.21 |
ENSDART00000184897
|
DHX35
|
zgc:158828 |
| chr21_-_1640547 | 0.21 |
ENSDART00000151041
|
zgc:152948
|
zgc:152948 |
| chr17_+_21817382 | 0.20 |
ENSDART00000079011
ENSDART00000189387 |
ikzf5
|
IKAROS family zinc finger 5 |
| chr24_+_39137001 | 0.19 |
ENSDART00000181086
ENSDART00000183724 ENSDART00000193466 |
tbc1d24
|
TBC1 domain family, member 24 |
| chr25_+_36348401 | 0.19 |
ENSDART00000103006
|
hist1h2a3
|
histone cluster 1 H2A family member 3 |
| chr19_+_2631565 | 0.19 |
ENSDART00000171487
|
fam126a
|
family with sequence similarity 126, member A |
| chr12_+_2648043 | 0.19 |
ENSDART00000082220
|
gdf2
|
growth differentiation factor 2 |
| chr7_+_24520518 | 0.19 |
ENSDART00000173604
|
btr09
|
bloodthirsty-related gene family, member 9 |
| chr18_-_158541 | 0.19 |
ENSDART00000188914
ENSDART00000191052 |
trpm7
|
transient receptor potential cation channel, subfamily M, member 7 |
| chr20_-_51656512 | 0.18 |
ENSDART00000129965
|
LO018154.1
|
|
| chr23_-_16737161 | 0.18 |
ENSDART00000132573
|
si:ch211-224l10.4
|
si:ch211-224l10.4 |
| chr22_-_7050 | 0.18 |
ENSDART00000127829
|
atad3
|
ATPase family, AAA domain containing 3 |
| chr8_+_8012570 | 0.18 |
ENSDART00000183429
|
si:ch211-169p10.1
|
si:ch211-169p10.1 |
| chr5_-_3885727 | 0.17 |
ENSDART00000143250
|
mlxipl
|
MLX interacting protein like |
| chr16_-_29146624 | 0.17 |
ENSDART00000159814
ENSDART00000009826 |
mef2d
|
myocyte enhancer factor 2d |
| chr5_-_20491198 | 0.17 |
ENSDART00000183051
ENSDART00000144232 |
ficd
|
FIC domain containing |
| chr23_-_18595020 | 0.17 |
ENSDART00000187032
ENSDART00000191047 |
ikbkg
|
inhibitor of kappa light polypeptide gene enhancer in B-cells, kinase gamma |
| chr25_-_35140746 | 0.17 |
ENSDART00000129969
|
si:ch211-113a14.19
|
si:ch211-113a14.19 |
| chr21_-_21515466 | 0.17 |
ENSDART00000147593
|
nectin3b
|
nectin cell adhesion molecule 3b |
| chr18_+_25546227 | 0.16 |
ENSDART00000085824
|
pex11a
|
peroxisomal biogenesis factor 11 alpha |
| chr18_+_30441740 | 0.16 |
ENSDART00000189074
|
gse1
|
Gse1 coiled-coil protein |
| chr15_+_22448456 | 0.15 |
ENSDART00000040542
ENSDART00000190270 ENSDART00000185897 |
arhgef12a
|
Rho guanine nucleotide exchange factor (GEF) 12a |
| chr18_+_27515640 | 0.15 |
ENSDART00000181593
|
tp53i11b
|
tumor protein p53 inducible protein 11b |
| chr21_+_1119046 | 0.15 |
ENSDART00000184678
|
CABZ01088049.1
|
|
| chr3_-_34561624 | 0.15 |
ENSDART00000129313
|
sept9a
|
septin 9a |
| chr10_+_589501 | 0.15 |
ENSDART00000188415
|
LO018557.1
|
|
| chr9_-_32300611 | 0.14 |
ENSDART00000127938
|
hspd1
|
heat shock 60 protein 1 |
| chr20_+_21595244 | 0.14 |
ENSDART00000010643
|
esr2a
|
estrogen receptor 2a |
| chr12_+_46708920 | 0.14 |
ENSDART00000153089
|
exoc7
|
exocyst complex component 7 |
| chr13_-_36418921 | 0.14 |
ENSDART00000135804
|
dcaf5
|
ddb1 and cul4 associated factor 5 |
| chr2_+_22659787 | 0.14 |
ENSDART00000043956
|
zgc:161973
|
zgc:161973 |
| chr6_+_40952031 | 0.14 |
ENSDART00000189219
|
patz1
|
POZ (BTB) and AT hook containing zinc finger 1 |
| chr5_-_33255759 | 0.14 |
ENSDART00000085531
|
prkaa1
|
protein kinase, AMP-activated, alpha 1 catalytic subunit |
| chr23_-_45319592 | 0.14 |
ENSDART00000189861
|
ccdc171
|
coiled-coil domain containing 171 |
| chr11_+_19433936 | 0.13 |
ENSDART00000162081
|
prickle2b
|
prickle homolog 2b |
| chr15_+_47618221 | 0.13 |
ENSDART00000168722
|
paf1
|
PAF1 homolog, Paf1/RNA polymerase II complex component |
| chr12_+_26670778 | 0.13 |
ENSDART00000144355
|
arhgap12b
|
Rho GTPase activating protein 12b |
| chr5_-_56416188 | 0.13 |
ENSDART00000146773
|
acaca
|
acetyl-CoA carboxylase alpha |
| chr17_+_21817859 | 0.13 |
ENSDART00000143832
ENSDART00000141462 |
ikzf5
|
IKAROS family zinc finger 5 |
| chr9_-_32300783 | 0.12 |
ENSDART00000078596
|
hspd1
|
heat shock 60 protein 1 |
| chr22_-_10752471 | 0.12 |
ENSDART00000081191
|
sass6
|
SAS-6 centriolar assembly protein |
| chr8_-_4097722 | 0.12 |
ENSDART00000135006
|
cux2b
|
cut-like homeobox 2b |
| chr10_-_17745345 | 0.12 |
ENSDART00000132690
ENSDART00000135376 |
si:dkey-200l5.4
|
si:dkey-200l5.4 |
| chr12_-_31457301 | 0.12 |
ENSDART00000043887
ENSDART00000148603 |
acsl5
|
acyl-CoA synthetase long chain family member 5 |
| chr11_-_42230491 | 0.12 |
ENSDART00000164423
|
CABZ01030862.1
|
|
| chr13_+_21826369 | 0.12 |
ENSDART00000165150
ENSDART00000192115 |
zswim8
|
zinc finger, SWIM-type containing 8 |
| chr1_-_58868306 | 0.12 |
ENSDART00000166615
|
dnm2b
|
dynamin 2b |
| chr23_-_44786844 | 0.12 |
ENSDART00000148669
|
si:ch73-269m23.5
|
si:ch73-269m23.5 |
| chr19_+_3056450 | 0.12 |
ENSDART00000141324
ENSDART00000082353 |
hsf1
|
heat shock transcription factor 1 |
| chr19_+_5146460 | 0.11 |
ENSDART00000150740
|
si:dkey-89b17.4
|
si:dkey-89b17.4 |
| chr24_-_7587401 | 0.11 |
ENSDART00000093163
|
galnt11
|
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 11 (GalNAc-T11) |
| chr13_-_4018888 | 0.11 |
ENSDART00000058238
|
tjap1
|
tight junction associated protein 1 (peripheral) |
| chr10_+_45128375 | 0.10 |
ENSDART00000164805
|
camk2b2
|
calcium/calmodulin-dependent protein kinase (CaM kinase) II beta 2 |
| chr24_-_29586082 | 0.10 |
ENSDART00000136763
|
vav3a
|
vav 3 guanine nucleotide exchange factor a |
| chr24_-_1021318 | 0.10 |
ENSDART00000181403
|
ralaa
|
v-ral simian leukemia viral oncogene homolog Aa (ras related) |
| chr17_-_27200634 | 0.10 |
ENSDART00000185332
|
asap3
|
ArfGAP with SH3 domain, ankyrin repeat and PH domain 3 |
| chr17_+_45737992 | 0.10 |
ENSDART00000135073
ENSDART00000143525 ENSDART00000158165 ENSDART00000184167 ENSDART00000109525 |
aspg
|
asparaginase homolog (S. cerevisiae) |
| chr17_-_45733401 | 0.09 |
ENSDART00000185727
|
arf6b
|
ADP-ribosylation factor 6b |
| chr22_-_3595439 | 0.09 |
ENSDART00000083308
|
ptprsa
|
protein tyrosine phosphatase, receptor type, s, a |
| chr19_+_20713424 | 0.09 |
ENSDART00000129730
|
rab5aa
|
RAB5A, member RAS oncogene family, a |
| chr4_-_858434 | 0.09 |
ENSDART00000006961
|
sobpb
|
sine oculis binding protein homolog (Drosophila) b |
| chr17_+_450956 | 0.09 |
ENSDART00000183022
ENSDART00000171386 |
zgc:194887
|
zgc:194887 |
| chr21_-_308852 | 0.09 |
ENSDART00000151613
|
lhfpl2a
|
LHFPL tetraspan subfamily member 2a |
| chr20_-_26937453 | 0.09 |
ENSDART00000139756
|
ftr97
|
finTRIM family, member 97 |
| chr20_+_46286082 | 0.09 |
ENSDART00000073519
|
taar14f
|
trace amine associated receptor 14f |
| chr25_-_35113891 | 0.08 |
ENSDART00000190724
|
zgc:165555
|
zgc:165555 |
| chr9_-_29579908 | 0.08 |
ENSDART00000140876
|
cenpj
|
centromere protein J |
| chr12_-_48671612 | 0.08 |
ENSDART00000007202
|
zgc:92749
|
zgc:92749 |
| chr18_+_27926839 | 0.08 |
ENSDART00000191835
|
hipk3b
|
homeodomain interacting protein kinase 3b |
| chr5_+_43870389 | 0.08 |
ENSDART00000141002
|
zgc:112966
|
zgc:112966 |
| chr7_+_7495034 | 0.08 |
ENSDART00000102629
ENSDART00000180475 |
nit1
|
nitrilase 1 |
| chr14_+_30413312 | 0.08 |
ENSDART00000186864
|
cnot7
|
CCR4-NOT transcription complex, subunit 7 |
| chr6_-_28117995 | 0.08 |
ENSDART00000147253
|
si:ch73-194h10.3
|
si:ch73-194h10.3 |
| chr9_-_37613792 | 0.08 |
ENSDART00000138345
|
sema5ba
|
sema domain, seven thrombospondin repeats (type 1 and type 1-like), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 5Ba |
| chr4_-_75172216 | 0.08 |
ENSDART00000127522
|
naa40
|
N(alpha)-acetyltransferase 40, NatD catalytic subunit, homolog (S. cerevisiae) |
| chr8_-_45835056 | 0.07 |
ENSDART00000022242
|
ogdha
|
oxoglutarate (alpha-ketoglutarate) dehydrogenase a (lipoamide) |
| chr7_+_34296789 | 0.07 |
ENSDART00000052471
ENSDART00000173798 ENSDART00000173778 |
bbox1
|
butyrobetaine (gamma), 2-oxoglutarate dioxygenase (gamma-butyrobetaine hydroxylase) 1 |
| chr7_+_25000060 | 0.07 |
ENSDART00000039265
ENSDART00000141814 |
naa40
|
N(alpha)-acetyltransferase 40, NatD catalytic subunit, homolog (S. cerevisiae) |
| chr12_+_17439580 | 0.07 |
ENSDART00000189257
|
atad1b
|
ATPase family, AAA domain containing 1b |
| chr7_+_34297271 | 0.06 |
ENSDART00000180342
|
bbox1
|
butyrobetaine (gamma), 2-oxoglutarate dioxygenase (gamma-butyrobetaine hydroxylase) 1 |
| chr5_-_27994679 | 0.06 |
ENSDART00000132740
|
ppp3cca
|
protein phosphatase 3, catalytic subunit, gamma isozyme, a |
| chr25_+_15933411 | 0.06 |
ENSDART00000191581
|
ppfibp2b
|
PTPRF interacting protein, binding protein 2b (liprin beta 2) |
| chr10_-_25204034 | 0.06 |
ENSDART00000161100
|
FAT3 (1 of many)
|
si:ch211-214k5.6 |
| chr19_-_9867001 | 0.06 |
ENSDART00000091695
|
cacng7a
|
calcium channel, voltage-dependent, gamma subunit 7a |
| chr9_+_34380299 | 0.06 |
ENSDART00000131705
|
lamp1
|
lysosomal-associated membrane protein 1 |
| chr8_+_8699085 | 0.06 |
ENSDART00000021209
|
uxt
|
ubiquitously-expressed, prefoldin-like chaperone |
| chr11_-_19775182 | 0.06 |
ENSDART00000037894
|
namptb
|
nicotinamide phosphoribosyltransferase b |
| chr4_+_2655358 | 0.06 |
ENSDART00000007638
|
bcap29
|
B cell receptor associated protein 29 |
| chr5_+_24282570 | 0.05 |
ENSDART00000041905
|
zdhhc23a
|
zinc finger, DHHC-type containing 23a |
| chr12_+_19958845 | 0.05 |
ENSDART00000193248
|
ercc4
|
excision repair cross-complementation group 4 |
| chr20_+_25581627 | 0.05 |
ENSDART00000030229
|
cyp2p9
|
cytochrome P450, family 2, subfamily P, polypeptide 9 |
| chr23_-_27050083 | 0.05 |
ENSDART00000142324
ENSDART00000133249 ENSDART00000138751 ENSDART00000128718 |
zgc:66440
|
zgc:66440 |
| chr11_-_23332592 | 0.05 |
ENSDART00000125024
|
golt1a
|
golgi transport 1A |
| chr16_+_25809235 | 0.05 |
ENSDART00000181196
|
irgq1
|
immunity-related GTPase family, q1 |
| chr25_+_31238606 | 0.05 |
ENSDART00000149439
|
tnni2a.2
|
troponin I type 2a (skeletal, fast), tandem duplicate 2 |
| chr12_+_1286642 | 0.05 |
ENSDART00000157467
|
pemt
|
phosphatidylethanolamine N-methyltransferase |
| chr16_-_54978981 | 0.05 |
ENSDART00000154023
|
wdtc1
|
WD and tetratricopeptide repeats 1 |
| chr23_-_44903048 | 0.04 |
ENSDART00000149103
|
fhdc5
|
FH2 domain containing 5 |
| chr14_-_13048355 | 0.04 |
ENSDART00000166434
|
si:dkey-35h6.1
|
si:dkey-35h6.1 |
| chr3_+_25999477 | 0.04 |
ENSDART00000024316
|
mcm5
|
minichromosome maintenance complex component 5 |
| chr6_+_60036767 | 0.04 |
ENSDART00000155009
|
tgm8
|
transglutaminase 8 |
| chr2_-_57110477 | 0.04 |
ENSDART00000181132
|
slc25a42
|
solute carrier family 25, member 42 |
| chr7_-_38183331 | 0.04 |
ENSDART00000149382
|
abcc12
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 12 |
| chr2_-_56649883 | 0.04 |
ENSDART00000191786
|
gpx4b
|
glutathione peroxidase 4b |
| chr23_-_42810664 | 0.04 |
ENSDART00000102328
|
pfkfb2a
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 2a |
| chr3_+_12718100 | 0.04 |
ENSDART00000162343
ENSDART00000192425 |
cyp2k20
|
cytochrome P450, family 2, subfamily k, polypeptide 20 |
| chr9_-_25255490 | 0.04 |
ENSDART00000141502
|
htr2aa
|
5-hydroxytryptamine (serotonin) receptor 2A, genome duplicate a |
| chr4_+_68519945 | 0.03 |
ENSDART00000171148
|
si:dkey-16p6.1
|
si:dkey-16p6.1 |
| chr13_-_3351708 | 0.03 |
ENSDART00000042875
|
si:ch73-193i2.2
|
si:ch73-193i2.2 |
| chr23_+_21405201 | 0.03 |
ENSDART00000144409
|
iffo2a
|
intermediate filament family orphan 2a |
| chr17_-_7371564 | 0.03 |
ENSDART00000060336
|
rab32b
|
RAB32b, member RAS oncogene family |
| chr20_-_18382708 | 0.03 |
ENSDART00000170864
ENSDART00000166762 ENSDART00000191333 |
vipas39
|
VPS33B interacting protein, apical-basolateral polarity regulator, spe-39 homolog |
| chr1_+_58139102 | 0.03 |
ENSDART00000134826
|
si:ch211-15j1.4
|
si:ch211-15j1.4 |
| chr1_+_58303892 | 0.03 |
ENSDART00000147678
|
CR769768.1
|
|
| chr21_-_37435162 | 0.03 |
ENSDART00000133585
|
fam114a2
|
family with sequence similarity 114, member A2 |
| chr24_-_2843107 | 0.02 |
ENSDART00000165290
|
cyb5a
|
cytochrome b5 type A (microsomal) |
| chr12_-_33568174 | 0.02 |
ENSDART00000142329
|
tdrkh
|
tudor and KH domain containing |
| chr19_+_11214007 | 0.02 |
ENSDART00000127362
|
si:ch73-109i22.2
|
si:ch73-109i22.2 |
| chr25_-_31118923 | 0.02 |
ENSDART00000009126
ENSDART00000188286 |
kras
|
v-Ki-ras2 Kirsten rat sarcoma viral oncogene homolog |
| chr3_+_301479 | 0.02 |
ENSDART00000165169
|
CABZ01075611.1
|
|
| chr2_-_22660232 | 0.02 |
ENSDART00000174742
|
thap4
|
THAP domain containing 4 |
| chr24_-_1341543 | 0.02 |
ENSDART00000169341
|
nrp1a
|
neuropilin 1a |
| chr7_-_8156169 | 0.02 |
ENSDART00000182960
ENSDART00000190184 ENSDART00000190885 ENSDART00000189867 ENSDART00000160836 |
si:cabz01030277.1
si:ch211-163c2.3
|
si:cabz01030277.1 si:ch211-163c2.3 |
| chr6_+_13206516 | 0.02 |
ENSDART00000036927
|
ndufs1
|
NADH dehydrogenase (ubiquinone) Fe-S protein 1 |
| chr25_+_18587338 | 0.02 |
ENSDART00000180287
|
met
|
MET proto-oncogene, receptor tyrosine kinase |
| chr11_-_11910225 | 0.01 |
ENSDART00000159922
|
si:ch211-69b7.6
|
si:ch211-69b7.6 |
| chr1_+_58353661 | 0.01 |
ENSDART00000140074
|
si:dkey-222h21.2
|
si:dkey-222h21.2 |
| chr13_-_41546779 | 0.01 |
ENSDART00000163331
|
pcdh15a
|
protocadherin-related 15a |
| chr20_+_43379029 | 0.01 |
ENSDART00000142486
ENSDART00000186486 |
unc93a
|
unc-93 homolog A |
| chr3_-_36750068 | 0.01 |
ENSDART00000173388
|
abcc6b.1
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 6b, tandem duplicate 1 |
| chr1_+_58119117 | 0.01 |
ENSDART00000146788
|
si:ch211-15j1.5
|
si:ch211-15j1.5 |
| chr2_-_57076687 | 0.01 |
ENSDART00000161523
|
slc25a42
|
solute carrier family 25, member 42 |
| chr19_+_19032141 | 0.00 |
ENSDART00000192432
|
cpne4b
|
copine IVb |
| chr1_-_8691969 | 0.00 |
ENSDART00000152153
|
abcg2b
|
ATP-binding cassette, sub-family G (WHITE), member 2b |
| chr11_-_3571605 | 0.00 |
ENSDART00000167863
|
si:dkey-33m11.7
|
si:dkey-33m11.7 |
| chr23_+_41679586 | 0.00 |
ENSDART00000067662
|
CU914487.1
|
|
| chr1_+_58260886 | 0.00 |
ENSDART00000110453
|
si:dkey-222h21.8
|
si:dkey-222h21.8 |
| chr16_-_46645396 | 0.00 |
ENSDART00000131485
|
tmem176l.2
|
transmembrane protein 176l.2 |
| chr1_+_9994811 | 0.00 |
ENSDART00000143719
ENSDART00000110749 |
si:dkeyp-75b4.10
|
si:dkeyp-75b4.10 |
| chr1_+_58094551 | 0.00 |
ENSDART00000146316
|
si:ch211-114l13.1
|
si:ch211-114l13.1 |
| chr9_-_46382637 | 0.00 |
ENSDART00000085738
|
lct
|
lactase |
| chr1_+_58182400 | 0.00 |
ENSDART00000144784
|
si:ch211-15j1.1
|
si:ch211-15j1.1 |
| chr1_+_57050899 | 0.00 |
ENSDART00000152601
|
si:ch211-1f22.14
|
si:ch211-1f22.14 |
Network of associatons between targets according to the STRING database.
First level regulatory network of rxrbb
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.6 | GO:0005991 | trehalose metabolic process(GO:0005991) |
| 0.1 | 0.3 | GO:0032755 | positive regulation of interleukin-6 production(GO:0032755) protein import into mitochondrial intermembrane space(GO:0045041) |
| 0.1 | 0.3 | GO:0071908 | determination of intestine left/right asymmetry(GO:0071908) |
| 0.1 | 0.2 | GO:0010961 | cellular magnesium ion homeostasis(GO:0010961) |
| 0.0 | 0.4 | GO:0051673 | membrane disruption in other organism(GO:0051673) |
| 0.0 | 0.1 | GO:2000193 | regulation of plasma membrane long-chain fatty acid transport(GO:0010746) positive regulation of plasma membrane long-chain fatty acid transport(GO:0010747) plasma membrane long-chain fatty acid transport(GO:0015911) fatty acid transmembrane transport(GO:1902001) positive regulation of anion transmembrane transport(GO:1903961) regulation of fatty acid transport(GO:2000191) positive regulation of fatty acid transport(GO:2000193) |
| 0.0 | 0.2 | GO:0007252 | I-kappaB phosphorylation(GO:0007252) |
| 0.0 | 0.1 | GO:2001295 | malonyl-CoA biosynthetic process(GO:2001295) |
| 0.0 | 0.3 | GO:0030719 | P granule organization(GO:0030719) |
| 0.0 | 0.1 | GO:0048169 | regulation of long-term neuronal synaptic plasticity(GO:0048169) |
| 0.0 | 0.5 | GO:0000463 | maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000463) |
| 0.0 | 0.2 | GO:0016559 | peroxisome fission(GO:0016559) |
| 0.0 | 0.1 | GO:0045329 | carnitine biosynthetic process(GO:0045329) |
| 0.0 | 0.1 | GO:1901255 | nucleotide-excision repair involved in interstrand cross-link repair(GO:1901255) |
| 0.0 | 0.1 | GO:0040016 | embryonic cleavage(GO:0040016) |
| 0.0 | 0.1 | GO:0071392 | cellular response to estradiol stimulus(GO:0071392) |
| 0.0 | 0.2 | GO:0030241 | skeletal muscle myosin thick filament assembly(GO:0030241) |
| 0.0 | 0.3 | GO:0001966 | thigmotaxis(GO:0001966) |
| 0.0 | 0.1 | GO:0061511 | centriole elongation(GO:0061511) |
| 0.0 | 0.0 | GO:0045922 | negative regulation of fatty acid metabolic process(GO:0045922) |
| 0.0 | 0.1 | GO:0070973 | protein localization to endoplasmic reticulum exit site(GO:0070973) |
| 0.0 | 0.1 | GO:0006474 | N-terminal protein amino acid acetylation(GO:0006474) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.5 | GO:0070545 | PeBoW complex(GO:0070545) |
| 0.0 | 0.1 | GO:0098536 | deuterosome(GO:0098536) |
| 0.0 | 0.1 | GO:0035327 | transcriptionally active chromatin(GO:0035327) |
| 0.0 | 0.2 | GO:0035517 | PR-DUB complex(GO:0035517) |
| 0.0 | 0.2 | GO:0008385 | IkappaB kinase complex(GO:0008385) |
| 0.0 | 0.1 | GO:0098842 | postsynaptic early endosome(GO:0098842) |
| 0.0 | 0.2 | GO:0005779 | integral component of peroxisomal membrane(GO:0005779) intrinsic component of peroxisomal membrane(GO:0031231) |
| 0.0 | 0.3 | GO:0017119 | Golgi transport complex(GO:0017119) |
| 0.0 | 0.1 | GO:0000110 | nucleotide-excision repair factor 1 complex(GO:0000110) |
| 0.0 | 0.1 | GO:0005955 | calcineurin complex(GO:0005955) |
| 0.0 | 0.3 | GO:0043186 | P granule(GO:0043186) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.6 | GO:0004555 | alpha,alpha-trehalase activity(GO:0004555) trehalase activity(GO:0015927) |
| 0.0 | 0.2 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.0 | 0.1 | GO:0008336 | gamma-butyrobetaine dioxygenase activity(GO:0008336) |
| 0.0 | 0.1 | GO:0043998 | H2A histone acetyltransferase activity(GO:0043998) |
| 0.0 | 0.3 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 0.0 | 0.1 | GO:0003989 | acetyl-CoA carboxylase activity(GO:0003989) |
| 0.0 | 0.1 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.0 | 0.1 | GO:1903924 | estradiol binding(GO:1903924) |
| 0.0 | 0.0 | GO:0071077 | coenzyme A transmembrane transporter activity(GO:0015228) adenosine 3',5'-bisphosphate transmembrane transporter activity(GO:0071077) AMP transmembrane transporter activity(GO:0080122) |
| 0.0 | 0.2 | GO:0047631 | ADP-ribose diphosphatase activity(GO:0047631) |
| 0.0 | 0.1 | GO:0004067 | asparaginase activity(GO:0004067) |
| 0.0 | 0.2 | GO:0070530 | K63-linked polyubiquitin binding(GO:0070530) |
| 0.0 | 0.1 | GO:0047676 | arachidonate-CoA ligase activity(GO:0047676) |
| 0.0 | 0.1 | GO:0033192 | calcium-dependent protein serine/threonine phosphatase activity(GO:0004723) calmodulin-dependent protein phosphatase activity(GO:0033192) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 0.0 | 0.1 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.0 | 0.2 | REACTOME IKK COMPLEX RECRUITMENT MEDIATED BY RIP1 | Genes involved in IKK complex recruitment mediated by RIP1 |
| 0.0 | 0.3 | REACTOME FATTY ACYL COA BIOSYNTHESIS | Genes involved in Fatty Acyl-CoA Biosynthesis |