Project
PRJNA207719: Tissue specific transcriptome profiling
Navigation
Downloads
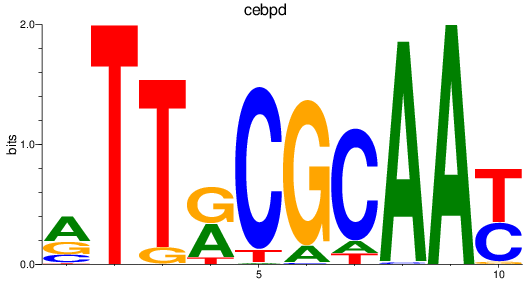
Results for cebpd
Z-value: 1.27
Transcription factors associated with cebpd
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
cebpd
|
ENSDARG00000087303 | CCAAT enhancer binding protein delta |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| cebpd | dr11_v1_chr24_+_35564668_35564668 | 0.03 | 9.7e-01 | Click! |
Activity profile of cebpd motif
Sorted Z-values of cebpd motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_-_63509581 | 1.05 |
ENSDART00000097325
|
c5
|
complement component 5 |
| chr6_-_11780070 | 0.92 |
ENSDART00000151195
|
march7
|
membrane-associated ring finger (C3HC4) 7 |
| chr20_+_23440632 | 0.92 |
ENSDART00000180685
ENSDART00000042820 |
si:dkey-90m5.4
|
si:dkey-90m5.4 |
| chr14_+_16345003 | 0.90 |
ENSDART00000003040
ENSDART00000165193 |
itln3
|
intelectin 3 |
| chr12_+_46960579 | 0.89 |
ENSDART00000149032
|
oat
|
ornithine aminotransferase |
| chr2_-_30460293 | 0.86 |
ENSDART00000113193
|
cbln2a
|
cerebellin 2a precursor |
| chr21_+_17051478 | 0.85 |
ENSDART00000047201
ENSDART00000161650 ENSDART00000167298 |
atp2a2a
|
ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 2a |
| chr1_-_10071422 | 0.84 |
ENSDART00000135522
ENSDART00000033118 |
fga
|
fibrinogen alpha chain |
| chr5_-_43935119 | 0.80 |
ENSDART00000142271
|
si:ch211-204c21.1
|
si:ch211-204c21.1 |
| chr2_-_54387550 | 0.79 |
ENSDART00000097388
|
napgb
|
N-ethylmaleimide-sensitive factor attachment protein, gamma b |
| chr12_+_35119762 | 0.78 |
ENSDART00000085774
|
si:ch73-127m5.1
|
si:ch73-127m5.1 |
| chr25_-_19443421 | 0.76 |
ENSDART00000067362
|
cart2
|
cocaine- and amphetamine-regulated transcript 2 |
| chr7_-_52963493 | 0.71 |
ENSDART00000052029
|
cart3
|
cocaine- and amphetamine-regulated transcript 3 |
| chr5_-_43935460 | 0.67 |
ENSDART00000166152
ENSDART00000188969 |
si:ch211-204c21.1
|
si:ch211-204c21.1 |
| chr19_+_1370504 | 0.66 |
ENSDART00000158946
|
dgat1a
|
diacylglycerol O-acyltransferase 1a |
| chr16_+_14029283 | 0.65 |
ENSDART00000146165
ENSDART00000132075 |
rusc1
|
RUN and SH3 domain containing 1 |
| chr8_+_4368534 | 0.65 |
ENSDART00000015214
|
pptc7a
|
PTC7 protein phosphatase homolog a |
| chr8_-_39739627 | 0.64 |
ENSDART00000135422
ENSDART00000067844 |
si:ch211-170d8.5
|
si:ch211-170d8.5 |
| chr16_+_50289916 | 0.64 |
ENSDART00000168861
ENSDART00000167332 |
hamp
|
hepcidin antimicrobial peptide |
| chr19_-_9829965 | 0.63 |
ENSDART00000136842
ENSDART00000142766 |
cacng8a
|
calcium channel, voltage-dependent, gamma subunit 8a |
| chr19_+_46259619 | 0.63 |
ENSDART00000158032
|
grinaa
|
glutamate receptor, ionotropic, N-methyl D-aspartate-associated protein 1a (glutamate binding) |
| chr5_+_32162684 | 0.62 |
ENSDART00000134472
|
taok3b
|
TAO kinase 3b |
| chr24_-_32665283 | 0.62 |
ENSDART00000038364
|
ca2
|
carbonic anhydrase II |
| chr15_-_17960228 | 0.61 |
ENSDART00000155898
|
phldb1b
|
pleckstrin homology-like domain, family B, member 1b |
| chr17_-_43466317 | 0.58 |
ENSDART00000155313
|
hspa4l
|
heat shock protein 4 like |
| chr21_-_43949208 | 0.57 |
ENSDART00000150983
|
camk2a
|
calcium/calmodulin-dependent protein kinase II alpha |
| chr21_+_29227224 | 0.57 |
ENSDART00000013641
|
adra1ba
|
adrenoceptor alpha 1Ba |
| chr16_+_46111849 | 0.57 |
ENSDART00000172232
|
sv2a
|
synaptic vesicle glycoprotein 2A |
| chr24_-_15648636 | 0.55 |
ENSDART00000136200
|
cbln2b
|
cerebellin 2b precursor |
| chr20_-_45661049 | 0.54 |
ENSDART00000124582
ENSDART00000131251 |
napbb
|
N-ethylmaleimide-sensitive factor attachment protein, beta b |
| chr2_-_50298337 | 0.53 |
ENSDART00000155125
|
cntnap2b
|
contactin associated protein like 2b |
| chr5_-_43682930 | 0.52 |
ENSDART00000075017
|
si:dkey-40c11.1
|
si:dkey-40c11.1 |
| chr11_-_29082175 | 0.51 |
ENSDART00000123245
|
igsf21a
|
immunoglobin superfamily, member 21a |
| chr12_+_25775734 | 0.51 |
ENSDART00000024415
ENSDART00000149198 |
epas1a
|
endothelial PAS domain protein 1a |
| chr9_+_219124 | 0.49 |
ENSDART00000161484
|
map3k12
|
mitogen-activated protein kinase kinase kinase 12 |
| chr3_-_46818001 | 0.49 |
ENSDART00000166505
|
elavl3
|
ELAV like neuron-specific RNA binding protein 3 |
| chr4_-_17409533 | 0.48 |
ENSDART00000011943
|
pah
|
phenylalanine hydroxylase |
| chr11_-_29082429 | 0.47 |
ENSDART00000041443
|
igsf21a
|
immunoglobin superfamily, member 21a |
| chr14_-_8890437 | 0.47 |
ENSDART00000167242
|
ATG2A
|
si:ch73-45o6.2 |
| chr20_-_40750953 | 0.47 |
ENSDART00000061256
|
cx28.9
|
connexin 28.9 |
| chr25_+_7784582 | 0.46 |
ENSDART00000155016
|
dgkzb
|
diacylglycerol kinase, zeta b |
| chr9_+_3388099 | 0.45 |
ENSDART00000019910
|
dlx1a
|
distal-less homeobox 1a |
| chr5_+_8196264 | 0.45 |
ENSDART00000174564
ENSDART00000161261 |
lmbrd2a
|
LMBR1 domain containing 2a |
| chr12_+_4920451 | 0.43 |
ENSDART00000171525
ENSDART00000159986 |
plekhm1
|
pleckstrin homology domain containing, family M (with RUN domain) member 1 |
| chr20_-_40755614 | 0.42 |
ENSDART00000061247
|
cx32.3
|
connexin 32.3 |
| chr22_+_27090136 | 0.42 |
ENSDART00000136770
|
si:dkey-246e1.3
|
si:dkey-246e1.3 |
| chr8_+_42917515 | 0.42 |
ENSDART00000021715
|
slc23a2
|
solute carrier family 23 (ascorbic acid transporter), member 2 |
| chr4_-_5302866 | 0.41 |
ENSDART00000138590
|
SNAP91 (1 of many)
|
si:ch211-214j24.9 |
| chr25_-_12805295 | 0.40 |
ENSDART00000157629
|
ca5a
|
carbonic anhydrase Va |
| chr24_-_39772045 | 0.40 |
ENSDART00000087441
|
GFOD1
|
si:ch211-276f18.2 |
| chr4_+_12113105 | 0.40 |
ENSDART00000182399
|
tmem178b
|
transmembrane protein 178B |
| chr22_-_26175237 | 0.37 |
ENSDART00000108737
|
c3b.2
|
complement component c3b, tandem duplicate 2 |
| chr4_-_5302162 | 0.37 |
ENSDART00000177099
|
SNAP91 (1 of many)
|
si:ch211-214j24.9 |
| chr1_+_25801648 | 0.36 |
ENSDART00000129471
|
gucy1b1
|
guanylate cyclase 1 soluble subunit beta 1 |
| chr14_+_21106444 | 0.35 |
ENSDART00000075744
ENSDART00000132363 |
aldob
|
aldolase b, fructose-bisphosphate |
| chr9_+_38292947 | 0.35 |
ENSDART00000146663
|
tfcp2l1
|
transcription factor CP2-like 1 |
| chr5_-_45894802 | 0.35 |
ENSDART00000097648
|
crfb6
|
cytokine receptor family member b6 |
| chr22_+_15973122 | 0.35 |
ENSDART00000144545
|
rc3h1a
|
ring finger and CCCH-type domains 1a |
| chr16_-_27138478 | 0.33 |
ENSDART00000147438
|
tmem245
|
transmembrane protein 245 |
| chr12_-_26383242 | 0.33 |
ENSDART00000152941
|
usp54b
|
ubiquitin specific peptidase 54b |
| chr23_+_44732863 | 0.33 |
ENSDART00000160044
ENSDART00000172268 |
atp1b2a
|
ATPase Na+/K+ transporting subunit beta 2a |
| chr12_-_19862912 | 0.33 |
ENSDART00000145788
|
shisa9a
|
shisa family member 9a |
| chr12_-_22238004 | 0.32 |
ENSDART00000038310
|
ormdl3
|
ORMDL sphingolipid biosynthesis regulator 3 |
| chr24_-_31123365 | 0.32 |
ENSDART00000182947
|
tmem56a
|
transmembrane protein 56a |
| chr5_-_29488245 | 0.31 |
ENSDART00000047719
ENSDART00000141154 ENSDART00000171165 |
cacna1ba
|
calcium channel, voltage-dependent, N type, alpha 1B subunit, a |
| chr2_-_51772438 | 0.31 |
ENSDART00000170241
|
BX908782.2
|
Danio rerio three-finger protein 5 (LOC100003647), mRNA. |
| chr13_+_28495419 | 0.31 |
ENSDART00000025583
|
fgf8a
|
fibroblast growth factor 8a |
| chr11_+_23933016 | 0.30 |
ENSDART00000000486
|
cntn2
|
contactin 2 |
| chr14_+_21107032 | 0.30 |
ENSDART00000138319
ENSDART00000139103 ENSDART00000184735 |
aldob
|
aldolase b, fructose-bisphosphate |
| chr22_-_16377960 | 0.30 |
ENSDART00000168170
|
ttc39c
|
tetratricopeptide repeat domain 39C |
| chr21_-_43550120 | 0.29 |
ENSDART00000151627
|
si:ch73-362m14.2
|
si:ch73-362m14.2 |
| chr19_+_41080240 | 0.29 |
ENSDART00000087295
|
ppp1r9a
|
protein phosphatase 1, regulatory subunit 9A |
| chr15_-_29348212 | 0.29 |
ENSDART00000133117
|
tsku
|
tsukushi small leucine rich proteoglycan homolog (Xenopus laevis) |
| chr3_-_43821191 | 0.29 |
ENSDART00000186816
|
snn
|
stannin |
| chr9_-_12034444 | 0.29 |
ENSDART00000038651
|
znf804a
|
zinc finger protein 804A |
| chr17_+_5623514 | 0.28 |
ENSDART00000171220
ENSDART00000176083 |
CU571310.1
|
|
| chr22_-_17595310 | 0.28 |
ENSDART00000099056
|
gpx4a
|
glutathione peroxidase 4a |
| chr10_+_44940693 | 0.28 |
ENSDART00000157515
|
cnnm4a
|
cyclin and CBS domain divalent metal cation transport mediator 4a |
| chr10_+_45248326 | 0.27 |
ENSDART00000159797
|
zmiz2
|
zinc finger, MIZ-type containing 2 |
| chr7_+_63325819 | 0.27 |
ENSDART00000085612
ENSDART00000161436 |
pcdh7b
|
protocadherin 7b |
| chr22_-_16377666 | 0.27 |
ENSDART00000161878
|
ttc39c
|
tetratricopeptide repeat domain 39C |
| chr18_+_16330025 | 0.27 |
ENSDART00000142353
|
nts
|
neurotensin |
| chr9_-_9225980 | 0.26 |
ENSDART00000180301
|
cbsb
|
cystathionine-beta-synthase b |
| chr13_+_40437550 | 0.26 |
ENSDART00000057090
ENSDART00000167859 |
got1
|
glutamic-oxaloacetic transaminase 1, soluble |
| chr9_-_7238839 | 0.26 |
ENSDART00000142726
|
creg2
|
cellular repressor of E1A-stimulated genes 2 |
| chr4_-_4795205 | 0.25 |
ENSDART00000039313
|
zgc:162331
|
zgc:162331 |
| chr4_+_8670662 | 0.25 |
ENSDART00000168768
|
adipor2
|
adiponectin receptor 2 |
| chr6_+_11397269 | 0.25 |
ENSDART00000114260
|
senp2
|
SUMO1/sentrin/SMT3 specific peptidase 2 |
| chr2_-_7246848 | 0.25 |
ENSDART00000146434
|
zgc:153115
|
zgc:153115 |
| chr15_-_17868870 | 0.25 |
ENSDART00000170950
|
atf5b
|
activating transcription factor 5b |
| chr9_+_9502610 | 0.24 |
ENSDART00000061525
ENSDART00000125174 |
nr1i2
|
nuclear receptor subfamily 1, group I, member 2 |
| chr5_+_3501859 | 0.24 |
ENSDART00000080486
|
ywhag1
|
3-monooxygenase/tryptophan 5-monooxygenase activation protein, gamma polypeptide 1 |
| chr20_-_19511700 | 0.22 |
ENSDART00000040191
|
snx17
|
sorting nexin 17 |
| chr5_+_51111343 | 0.21 |
ENSDART00000092002
|
pomt1
|
protein-O-mannosyltransferase 1 |
| chr15_-_17869115 | 0.21 |
ENSDART00000112838
|
atf5b
|
activating transcription factor 5b |
| chr2_+_31476065 | 0.21 |
ENSDART00000049219
|
cacnb2b
|
calcium channel, voltage-dependent, beta 2b |
| chr5_+_25585869 | 0.20 |
ENSDART00000138060
|
si:dkey-229d2.7
|
si:dkey-229d2.7 |
| chr20_+_2950005 | 0.20 |
ENSDART00000135919
|
amd1
|
adenosylmethionine decarboxylase 1 |
| chr12_+_32368574 | 0.20 |
ENSDART00000086389
|
ANKFN1
|
si:ch211-277e21.2 |
| chr16_+_25245857 | 0.20 |
ENSDART00000155220
|
klhl38b
|
kelch-like family member 38b |
| chr11_+_43751263 | 0.20 |
ENSDART00000163843
|
zgc:153431
|
zgc:153431 |
| chr2_+_11029138 | 0.20 |
ENSDART00000138737
ENSDART00000081058 ENSDART00000153662 |
acot11a
|
acyl-CoA thioesterase 11a |
| chr8_+_17884569 | 0.19 |
ENSDART00000134660
|
slc44a5b
|
solute carrier family 44, member 5b |
| chr18_+_507835 | 0.19 |
ENSDART00000189701
|
nedd4a
|
neural precursor cell expressed, developmentally down-regulated 4a |
| chr9_-_35069645 | 0.19 |
ENSDART00000122679
ENSDART00000077908 ENSDART00000077894 ENSDART00000125536 |
appb
|
amyloid beta (A4) precursor protein b |
| chr2_-_36819624 | 0.18 |
ENSDART00000140844
|
slitrk3b
|
SLIT and NTRK-like family, member 3b |
| chr5_-_69948099 | 0.18 |
ENSDART00000034639
ENSDART00000191111 |
ugt2a4
|
UDP glucuronosyltransferase 2 family, polypeptide A4 |
| chr24_+_119680 | 0.18 |
ENSDART00000061973
|
tgfbr1b
|
transforming growth factor, beta receptor 1 b |
| chr21_-_39058490 | 0.18 |
ENSDART00000114885
|
aldh3a2b
|
aldehyde dehydrogenase 3 family, member A2b |
| chr11_-_12998400 | 0.18 |
ENSDART00000018614
|
chrna4b
|
cholinergic receptor, nicotinic, alpha 4b |
| chr23_-_36003441 | 0.17 |
ENSDART00000164699
|
calcoco1a
|
calcium binding and coiled-coil domain 1a |
| chr19_-_25113660 | 0.17 |
ENSDART00000035538
|
ptp4a3
|
protein tyrosine phosphatase type IVA, member 3 |
| chr9_-_30808073 | 0.17 |
ENSDART00000146300
|
klf5b
|
Kruppel-like factor 5b |
| chr10_+_38526496 | 0.17 |
ENSDART00000144329
|
acer3
|
alkaline ceramidase 3 |
| chr23_-_36003282 | 0.17 |
ENSDART00000103150
|
calcoco1a
|
calcium binding and coiled-coil domain 1a |
| chr17_+_9040165 | 0.16 |
ENSDART00000181221
ENSDART00000181846 |
akap6
|
A kinase (PRKA) anchor protein 6 |
| chr7_+_23580082 | 0.16 |
ENSDART00000183245
|
si:dkey-172j4.3
|
si:dkey-172j4.3 |
| chr16_-_42770064 | 0.16 |
ENSDART00000112879
|
slc4a7
|
solute carrier family 4, sodium bicarbonate cotransporter, member 7 |
| chr9_+_2574122 | 0.15 |
ENSDART00000166326
ENSDART00000191822 |
CIR1
|
si:ch73-167c12.2 |
| chr13_+_28705143 | 0.15 |
ENSDART00000183338
|
ldb1a
|
LIM domain binding 1a |
| chr9_+_48415043 | 0.15 |
ENSDART00000159930
|
lrp2a
|
low density lipoprotein receptor-related protein 2a |
| chr8_-_25011157 | 0.14 |
ENSDART00000078795
|
ahcyl1
|
adenosylhomocysteinase-like 1 |
| chr22_+_18469004 | 0.14 |
ENSDART00000061430
|
cilp2
|
cartilage intermediate layer protein 2 |
| chr25_-_3469576 | 0.14 |
ENSDART00000186738
|
hbp1
|
HMG-box transcription factor 1 |
| chr3_+_41731527 | 0.14 |
ENSDART00000049007
ENSDART00000187866 |
chst12a
|
carbohydrate (chondroitin 4) sulfotransferase 12a |
| chr15_-_20939579 | 0.13 |
ENSDART00000152371
|
usp2a
|
ubiquitin specific peptidase 2a |
| chr13_-_18195942 | 0.13 |
ENSDART00000079902
|
slc25a16
|
solute carrier family 25 (mitochondrial carrier; Graves disease autoantigen), member 16 |
| chr11_+_31323746 | 0.13 |
ENSDART00000180220
ENSDART00000189937 |
sipa1l2
|
signal-induced proliferation-associated 1 like 2 |
| chr7_-_66452792 | 0.13 |
ENSDART00000170701
ENSDART00000082604 |
galnt18b
|
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 18b |
| chr18_-_15551360 | 0.13 |
ENSDART00000159915
ENSDART00000172690 |
ppfibp1b
|
PTPRF interacting protein, binding protein 1b (liprin beta 1) |
| chr17_+_28670132 | 0.13 |
ENSDART00000076344
ENSDART00000164981 ENSDART00000182851 |
hectd1
|
HECT domain containing 1 |
| chr12_+_34953038 | 0.13 |
ENSDART00000187022
ENSDART00000123988 ENSDART00000027034 |
qki2
|
QKI, KH domain containing, RNA binding 2 |
| chr6_-_39649504 | 0.13 |
ENSDART00000179960
ENSDART00000190951 |
larp4ab
|
La ribonucleoprotein domain family, member 4Ab |
| chr9_-_6927587 | 0.13 |
ENSDART00000059092
|
tmem182a
|
transmembrane protein 182a |
| chr5_+_51111766 | 0.13 |
ENSDART00000188552
|
pomt1
|
protein-O-mannosyltransferase 1 |
| chr21_+_45510448 | 0.12 |
ENSDART00000160494
ENSDART00000167914 |
fnip1
|
folliculin interacting protein 1 |
| chr3_+_36972586 | 0.12 |
ENSDART00000102784
|
si:ch211-18i17.2
|
si:ch211-18i17.2 |
| chr8_+_26253252 | 0.12 |
ENSDART00000142031
|
slc26a6
|
solute carrier family 26, member 6 |
| chr8_-_35960987 | 0.12 |
ENSDART00000160503
|
slc15a4
|
solute carrier family 15 (oligopeptide transporter), member 4 |
| chr24_+_9881219 | 0.12 |
ENSDART00000036204
|
cdv3
|
carnitine deficiency-associated gene expressed in ventricle 3 |
| chr8_+_26874924 | 0.11 |
ENSDART00000141794
|
rimkla
|
ribosomal modification protein rimK-like family member A |
| chr17_-_6076084 | 0.11 |
ENSDART00000058890
|
ephx2
|
epoxide hydrolase 2, cytoplasmic |
| chr5_-_54395488 | 0.11 |
ENSDART00000160781
|
zmynd19
|
zinc finger, MYND-type containing 19 |
| chr3_+_36972298 | 0.11 |
ENSDART00000150917
|
si:ch211-18i17.2
|
si:ch211-18i17.2 |
| chr13_-_25548733 | 0.10 |
ENSDART00000168099
ENSDART00000135788 ENSDART00000077655 |
mcmbp
|
minichromosome maintenance complex binding protein |
| chr7_+_67733759 | 0.10 |
ENSDART00000172015
|
cyb5b
|
cytochrome b5 type B |
| chr15_+_21882419 | 0.10 |
ENSDART00000157216
|
si:dkey-103g5.4
|
si:dkey-103g5.4 |
| chr4_+_33012407 | 0.09 |
ENSDART00000151873
|
si:dkey-26h11.2
|
si:dkey-26h11.2 |
| chr14_-_30945515 | 0.09 |
ENSDART00000161540
|
si:zfos-80g12.1
|
si:zfos-80g12.1 |
| chr21_+_11969603 | 0.09 |
ENSDART00000142247
ENSDART00000140652 |
mlnl
|
motilin-like |
| chr16_-_27749172 | 0.09 |
ENSDART00000145198
|
steap4
|
STEAP family member 4 |
| chr6_+_8314451 | 0.09 |
ENSDART00000147793
ENSDART00000183688 |
gcdha
|
glutaryl-CoA dehydrogenase a |
| chr5_-_15283509 | 0.08 |
ENSDART00000052712
|
gnb1l
|
guanine nucleotide binding protein (G protein), beta polypeptide 1-like |
| chr1_-_9231952 | 0.08 |
ENSDART00000166515
|
si:dkeyp-57d7.4
|
si:dkeyp-57d7.4 |
| chr12_-_1085227 | 0.08 |
ENSDART00000169936
|
sox8a
|
SRY (sex determining region Y)-box 8a |
| chr10_+_9595575 | 0.08 |
ENSDART00000091780
ENSDART00000184287 |
rc3h2
|
ring finger and CCCH-type domains 2 |
| chr21_-_36948 | 0.08 |
ENSDART00000181230
|
JMY
|
junction mediating and regulatory protein, p53 cofactor |
| chr8_+_30699429 | 0.08 |
ENSDART00000005345
|
upb1
|
ureidopropionase, beta |
| chr21_-_5007109 | 0.08 |
ENSDART00000187042
ENSDART00000097796 ENSDART00000146766 |
rnf165a
|
ring finger protein 165a |
| chr12_-_17592215 | 0.07 |
ENSDART00000134597
|
usp42
|
ubiquitin specific peptidase 42 |
| chr6_+_8315050 | 0.07 |
ENSDART00000189987
|
gcdha
|
glutaryl-CoA dehydrogenase a |
| chr17_+_18031899 | 0.07 |
ENSDART00000022758
|
setd3
|
SET domain containing 3 |
| chr6_+_23810529 | 0.07 |
ENSDART00000166921
|
glulb
|
glutamate-ammonia ligase (glutamine synthase) b |
| chr15_-_4314042 | 0.07 |
ENSDART00000173311
ENSDART00000170562 |
ECE2
|
si:ch211-117a13.2 |
| chr2_-_58257624 | 0.06 |
ENSDART00000098940
|
foxl2b
|
forkhead box L2b |
| chr11_+_37251825 | 0.06 |
ENSDART00000169804
|
il17rc
|
interleukin 17 receptor C |
| chr16_+_23984179 | 0.06 |
ENSDART00000175879
|
apoc2
|
apolipoprotein C-II |
| chr22_+_31244728 | 0.06 |
ENSDART00000163309
|
grip2b
|
glutamate receptor interacting protein 2b |
| chr4_-_4261673 | 0.06 |
ENSDART00000150694
|
cd9b
|
CD9 molecule b |
| chr15_-_24960730 | 0.06 |
ENSDART00000109990
ENSDART00000186706 |
abhd15a
|
abhydrolase domain containing 15a |
| chr10_-_20523405 | 0.06 |
ENSDART00000114824
|
ddhd2
|
DDHD domain containing 2 |
| chr8_+_23703464 | 0.05 |
ENSDART00000132584
|
ppardb
|
peroxisome proliferator-activated receptor delta b |
| chr5_-_26893310 | 0.05 |
ENSDART00000126669
|
lman2lb
|
lectin, mannose-binding 2-like b |
| chr17_+_10318071 | 0.05 |
ENSDART00000161844
|
foxa1
|
forkhead box A1 |
| chr5_+_36439405 | 0.05 |
ENSDART00000102973
|
eda
|
ectodysplasin A |
| chr15_+_20543770 | 0.04 |
ENSDART00000092357
|
sgsm2
|
small G protein signaling modulator 2 |
| chr1_+_50538839 | 0.04 |
ENSDART00000020412
|
pkd2
|
polycystic kidney disease 2 |
| chr5_+_53482597 | 0.04 |
ENSDART00000180333
|
BX323994.1
|
|
| chr2_+_54387710 | 0.04 |
ENSDART00000127124
|
rab12
|
RAB12, member RAS oncogene family |
| chr5_-_69437422 | 0.04 |
ENSDART00000073676
|
isca1
|
iron-sulfur cluster assembly 1 |
| chr5_-_23574234 | 0.04 |
ENSDART00000002453
|
cwc15
|
CWC15 spliceosome-associated protein homolog (S. cerevisiae) |
| chr16_-_7828838 | 0.03 |
ENSDART00000191434
ENSDART00000108653 |
tcaim
|
T cell activation inhibitor, mitochondrial |
| chr13_+_30035253 | 0.03 |
ENSDART00000181303
ENSDART00000057525 ENSDART00000136622 |
dnajb12a
|
DnaJ (Hsp40) homolog, subfamily B, member 12a |
| chr12_-_13205572 | 0.03 |
ENSDART00000152670
|
pelo
|
pelota mRNA surveillance and ribosome rescue factor |
| chr3_+_23703704 | 0.03 |
ENSDART00000024256
|
hoxb6a
|
homeobox B6a |
| chr17_-_42799104 | 0.03 |
ENSDART00000154755
|
prkd3
|
protein kinase D3 |
| chr9_-_53062083 | 0.03 |
ENSDART00000122155
|
zmp:0000000936
|
zmp:0000000936 |
| chr19_+_19600297 | 0.03 |
ENSDART00000160134
ENSDART00000183493 |
hibadha
|
3-hydroxyisobutyrate dehydrogenase a |
| chr7_-_46019756 | 0.02 |
ENSDART00000162583
|
zgc:162297
|
zgc:162297 |
| chr2_-_24348948 | 0.02 |
ENSDART00000136559
|
ano8a
|
anoctamin 8a |
| chr12_-_13205854 | 0.02 |
ENSDART00000077829
|
pelo
|
pelota mRNA surveillance and ribosome rescue factor |
| chr16_+_44298902 | 0.02 |
ENSDART00000114795
|
dpys
|
dihydropyrimidinase |
| chr2_-_24348642 | 0.02 |
ENSDART00000181739
|
ano8a
|
anoctamin 8a |
| chr20_-_33961697 | 0.01 |
ENSDART00000061765
|
selp
|
selectin P |
| chr17_-_17759138 | 0.01 |
ENSDART00000157128
ENSDART00000123845 |
adck1
|
aarF domain containing kinase 1 |
| chr9_-_32300783 | 0.01 |
ENSDART00000078596
|
hspd1
|
heat shock 60 protein 1 |
| chr6_+_10333920 | 0.01 |
ENSDART00000151667
ENSDART00000151477 |
cobll1a
|
cordon-bleu WH2 repeat protein-like 1a |
| chr13_-_293250 | 0.01 |
ENSDART00000138581
|
chs1
|
chitin synthase 1 |
| chr5_-_58840971 | 0.01 |
ENSDART00000050932
|
tmem136b
|
transmembrane protein 136b |
| chr5_-_48680580 | 0.00 |
ENSDART00000031194
|
lysmd3
|
LysM, putative peptidoglycan-binding, domain containing 3 |
Network of associatons between targets according to the STRING database.
First level regulatory network of cebpd
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.5 | GO:0008343 | adult feeding behavior(GO:0008343) |
| 0.2 | 0.6 | GO:0015670 | carbon dioxide transport(GO:0015670) |
| 0.1 | 0.4 | GO:0015882 | L-ascorbic acid transport(GO:0015882) |
| 0.1 | 0.5 | GO:0010807 | regulation of synaptic vesicle priming(GO:0010807) |
| 0.1 | 0.5 | GO:0071205 | protein localization to juxtaparanode region of axon(GO:0071205) protein localization to axon(GO:0099612) |
| 0.1 | 0.3 | GO:0048677 | axon extension involved in regeneration(GO:0048677) sprouting of injured axon(GO:0048682) |
| 0.1 | 0.9 | GO:0055129 | proline biosynthetic process(GO:0006561) L-proline biosynthetic process(GO:0055129) |
| 0.1 | 0.6 | GO:0032570 | response to progesterone(GO:0032570) |
| 0.1 | 0.4 | GO:0099543 | trans-synaptic signaling by soluble gas(GO:0099543) trans-synaptic signaling by nitric oxide(GO:0099548) |
| 0.1 | 0.3 | GO:0031335 | regulation of sulfur amino acid metabolic process(GO:0031335) regulation of homocysteine metabolic process(GO:0050666) |
| 0.1 | 1.5 | GO:0045807 | positive regulation of endocytosis(GO:0045807) |
| 0.1 | 0.6 | GO:0003321 | positive regulation of heart rate by epinephrine-norepinephrine(GO:0001996) positive regulation of blood pressure by epinephrine-norepinephrine(GO:0003321) positive regulation of heart rate(GO:0010460) |
| 0.1 | 0.3 | GO:1900060 | regulation of sphingolipid biosynthetic process(GO:0090153) negative regulation of sphingolipid biosynthetic process(GO:0090155) negative regulation of ceramide biosynthetic process(GO:1900060) regulation of membrane lipid metabolic process(GO:1905038) regulation of ceramide biosynthetic process(GO:2000303) |
| 0.1 | 0.5 | GO:0035881 | amacrine cell differentiation(GO:0035881) |
| 0.1 | 0.5 | GO:0031106 | septin ring organization(GO:0031106) septin cytoskeleton organization(GO:0032185) |
| 0.1 | 0.3 | GO:0003418 | chondrocyte differentiation involved in endochondral bone morphogenesis(GO:0003413) growth plate cartilage chondrocyte differentiation(GO:0003418) |
| 0.1 | 0.3 | GO:0006531 | aspartate metabolic process(GO:0006531) |
| 0.1 | 0.8 | GO:0072378 | blood coagulation, fibrin clot formation(GO:0072378) |
| 0.1 | 0.3 | GO:0061194 | rhombomere 5 development(GO:0021571) rhombomere 6 development(GO:0021572) tongue development(GO:0043586) tongue morphogenesis(GO:0043587) taste bud development(GO:0061193) taste bud morphogenesis(GO:0061194) taste bud formation(GO:0061195) |
| 0.0 | 0.5 | GO:0000186 | activation of MAPKK activity(GO:0000186) |
| 0.0 | 0.5 | GO:1902221 | L-phenylalanine metabolic process(GO:0006558) L-phenylalanine catabolic process(GO:0006559) erythrose 4-phosphate/phosphoenolpyruvate family amino acid metabolic process(GO:1902221) erythrose 4-phosphate/phosphoenolpyruvate family amino acid catabolic process(GO:1902222) |
| 0.0 | 0.6 | GO:0098943 | neurotransmitter receptor transport, postsynaptic endosome to lysosome(GO:0098943) |
| 0.0 | 0.3 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.0 | 1.0 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.0 | 0.7 | GO:0030388 | fructose 1,6-bisphosphate metabolic process(GO:0030388) |
| 0.0 | 0.6 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.0 | 0.3 | GO:0033211 | adiponectin-activated signaling pathway(GO:0033211) |
| 0.0 | 0.3 | GO:0010960 | magnesium ion homeostasis(GO:0010960) |
| 0.0 | 0.1 | GO:0033353 | S-adenosylmethionine cycle(GO:0033353) |
| 0.0 | 0.1 | GO:0015677 | copper ion import(GO:0015677) |
| 0.0 | 0.2 | GO:0008216 | spermidine metabolic process(GO:0008216) |
| 0.0 | 0.1 | GO:0048496 | maintenance of organ identity(GO:0048496) |
| 0.0 | 0.3 | GO:0035269 | protein O-linked mannosylation(GO:0035269) |
| 0.0 | 0.1 | GO:0045852 | pH elevation(GO:0045852) intracellular pH elevation(GO:0051454) |
| 0.0 | 0.5 | GO:0071456 | cellular response to hypoxia(GO:0071456) |
| 0.0 | 0.2 | GO:0033539 | fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:0033539) |
| 0.0 | 0.6 | GO:0032474 | otolith morphogenesis(GO:0032474) |
| 0.0 | 0.7 | GO:0019432 | triglyceride biosynthetic process(GO:0019432) |
| 0.0 | 0.3 | GO:0048096 | chromatin-mediated maintenance of transcription(GO:0048096) |
| 0.0 | 0.3 | GO:0071436 | cellular potassium ion homeostasis(GO:0030007) sodium ion export from cell(GO:0036376) sodium ion export(GO:0071436) |
| 0.0 | 0.1 | GO:0070293 | renal absorption(GO:0070293) |
| 0.0 | 0.2 | GO:0035094 | response to nicotine(GO:0035094) |
| 0.0 | 0.3 | GO:0016926 | protein desumoylation(GO:0016926) |
| 0.0 | 0.1 | GO:0019483 | beta-alanine metabolic process(GO:0019482) beta-alanine biosynthetic process(GO:0019483) |
| 0.0 | 0.0 | GO:0071498 | cellular response to fluid shear stress(GO:0071498) |
| 0.0 | 0.2 | GO:0046520 | sphingoid biosynthetic process(GO:0046520) |
| 0.0 | 0.1 | GO:0032790 | ribosome disassembly(GO:0032790) |
| 0.0 | 0.1 | GO:0006857 | oligopeptide transport(GO:0006857) |
| 0.0 | 0.8 | GO:0031638 | zymogen activation(GO:0031638) |
| 0.0 | 0.1 | GO:1902108 | regulation of mitochondrial membrane permeability involved in apoptotic process(GO:1902108) |
| 0.0 | 0.3 | GO:1990126 | retrograde transport, endosome to plasma membrane(GO:1990126) |
| 0.0 | 0.5 | GO:0071875 | adrenergic receptor signaling pathway(GO:0071875) |
| 0.0 | 0.9 | GO:0008016 | regulation of heart contraction(GO:0008016) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.8 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.1 | 0.5 | GO:0070044 | synaptobrevin 2-SNAP-25-syntaxin-1a complex(GO:0070044) |
| 0.1 | 0.3 | GO:0035339 | SPOTS complex(GO:0035339) |
| 0.1 | 0.5 | GO:0005826 | actomyosin contractile ring(GO:0005826) |
| 0.1 | 0.5 | GO:0033010 | paranodal junction(GO:0033010) |
| 0.0 | 0.9 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.0 | 0.3 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) cation-transporting ATPase complex(GO:0090533) |
| 0.0 | 0.6 | GO:0045180 | basal cortex(GO:0045180) |
| 0.0 | 1.0 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.0 | 0.2 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.0 | 0.6 | GO:0044853 | caveola(GO:0005901) plasma membrane raft(GO:0044853) |
| 0.0 | 0.2 | GO:0048179 | activin receptor complex(GO:0048179) |
| 0.0 | 0.4 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.3 | GO:0005483 | soluble NSF attachment protein activity(GO:0005483) |
| 0.2 | 0.6 | GO:0070891 | lipoteichoic acid binding(GO:0070891) |
| 0.1 | 0.7 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 0.1 | 0.7 | GO:0004144 | diacylglycerol O-acyltransferase activity(GO:0004144) |
| 0.1 | 0.6 | GO:0004937 | alpha1-adrenergic receptor activity(GO:0004937) |
| 0.1 | 0.3 | GO:0004069 | L-aspartate:2-oxoglutarate aminotransferase activity(GO:0004069) |
| 0.1 | 0.4 | GO:0035613 | RNA stem-loop binding(GO:0035613) |
| 0.1 | 0.3 | GO:0047066 | phospholipid-hydroperoxide glutathione peroxidase activity(GO:0047066) |
| 0.1 | 0.5 | GO:0004706 | JUN kinase kinase kinase activity(GO:0004706) |
| 0.1 | 0.3 | GO:0004122 | cystathionine beta-synthase activity(GO:0004122) |
| 0.1 | 0.9 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.0 | 0.4 | GO:0016714 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced pteridine as one donor, and incorporation of one atom of oxygen(GO:0016714) |
| 0.0 | 0.1 | GO:0004013 | adenosylhomocysteinase activity(GO:0004013) trialkylsulfonium hydrolase activity(GO:0016802) |
| 0.0 | 1.7 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.0 | 0.2 | GO:0004361 | glutaryl-CoA dehydrogenase activity(GO:0004361) |
| 0.0 | 0.9 | GO:0070492 | oligosaccharide binding(GO:0070492) |
| 0.0 | 0.3 | GO:0005105 | type 1 fibroblast growth factor receptor binding(GO:0005105) |
| 0.0 | 0.1 | GO:0072590 | N-acetyl-L-aspartate-L-glutamate ligase activity(GO:0072590) citrate-L-glutamate ligase activity(GO:0072591) |
| 0.0 | 1.0 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.0 | 0.9 | GO:0008483 | transaminase activity(GO:0008483) |
| 0.0 | 0.2 | GO:0016361 | activin receptor activity, type I(GO:0016361) |
| 0.0 | 0.2 | GO:0036042 | long-chain fatty acyl-CoA binding(GO:0036042) |
| 0.0 | 0.2 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.0 | 0.6 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.0 | 0.1 | GO:0042030 | ATPase inhibitor activity(GO:0042030) |
| 0.0 | 0.3 | GO:0004169 | dolichyl-phosphate-mannose-protein mannosyltransferase activity(GO:0004169) |
| 0.0 | 0.1 | GO:0004301 | epoxide hydrolase activity(GO:0004301) |
| 0.0 | 0.1 | GO:0008823 | cupric reductase activity(GO:0008823) ferric-chelate reductase (NADPH) activity(GO:0052851) |
| 0.0 | 0.1 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.0 | 0.4 | GO:0004383 | guanylate cyclase activity(GO:0004383) |
| 0.0 | 0.2 | GO:0004028 | 3-chloroallyl aldehyde dehydrogenase activity(GO:0004028) |
| 0.0 | 0.3 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.0 | 0.3 | GO:0016929 | SUMO-specific protease activity(GO:0016929) |
| 0.0 | 1.2 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.0 | 0.3 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.0 | 0.2 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.0 | 0.8 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.0 | 0.5 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.8 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.0 | 0.6 | ST WNT CA2 CYCLIC GMP PATHWAY | Wnt/Ca2+/cyclic GMP signaling. |
| 0.0 | 0.7 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 0.3 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.0 | 0.4 | PID MAPK TRK PATHWAY | Trk receptor signaling mediated by the MAPK pathway |
| 0.0 | 0.2 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.6 | REACTOME CREB PHOSPHORYLATION THROUGH THE ACTIVATION OF CAMKII | Genes involved in CREB phosphorylation through the activation of CaMKII |
| 0.1 | 1.1 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.1 | 1.0 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.1 | 0.8 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.0 | 0.7 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.0 | 1.3 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.0 | 0.2 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.0 | 0.1 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.0 | 0.1 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |