Project
PRJNA207719: Tissue specific transcriptome profiling
Navigation
Downloads
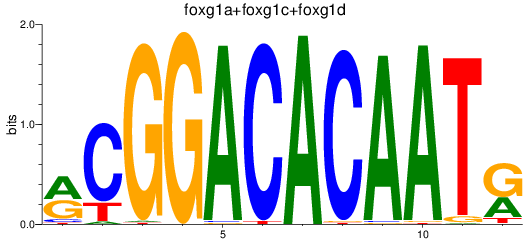
Results for foxg1a+foxg1c+foxg1d
Z-value: 0.81
Transcription factors associated with foxg1a+foxg1c+foxg1d
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
foxg1c
|
ENSDARG00000068380 | forkhead box G1c |
|
foxg1d
|
ENSDARG00000070053 | forkhead box G1d |
|
foxg1a
|
ENSDARG00000070769 | forkhead box G1a |
|
foxg1c
|
ENSDARG00000114414 | forkhead box G1c |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| foxg1a | dr11_v1_chr17_-_29119362_29119362 | 0.91 | 3.1e-02 | Click! |
| foxg1d | dr11_v1_chr13_+_255067_255067 | -0.63 | 2.5e-01 | Click! |
| foxg1c | dr11_v1_chr18_-_40901707_40901707 | -0.41 | 5.0e-01 | Click! |
Activity profile of foxg1a+foxg1c+foxg1d motif
Sorted Z-values of foxg1a+foxg1c+foxg1d motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr18_+_18104235 | 1.12 |
ENSDART00000145342
|
cbln1
|
cerebellin 1 precursor |
| chr18_-_38087875 | 1.03 |
ENSDART00000111301
|
luzp2
|
leucine zipper protein 2 |
| chr3_-_15264698 | 0.93 |
ENSDART00000111948
ENSDART00000142594 |
sez6l2
|
seizure related 6 homolog (mouse)-like 2 |
| chr11_+_13058613 | 0.91 |
ENSDART00000161532
|
zfyve9b
|
zinc finger, FYVE domain containing 9b |
| chr14_-_2361692 | 0.88 |
ENSDART00000167696
|
PCDHGC3 (1 of many)
|
si:ch73-233f7.4 |
| chr18_-_38088099 | 0.87 |
ENSDART00000146120
|
luzp2
|
leucine zipper protein 2 |
| chr24_+_11106402 | 0.78 |
ENSDART00000146697
|
prlh2
|
prolactin releasing hormone 2 |
| chr15_-_19250543 | 0.76 |
ENSDART00000092705
ENSDART00000138895 |
igsf9ba
|
immunoglobulin superfamily, member 9Ba |
| chr17_-_15657029 | 0.66 |
ENSDART00000153925
|
fut9a
|
fucosyltransferase 9a |
| chr22_-_38543630 | 0.57 |
ENSDART00000172029
|
si:ch211-126j24.1
|
si:ch211-126j24.1 |
| chr6_+_36942966 | 0.56 |
ENSDART00000028895
|
negr1
|
neuronal growth regulator 1 |
| chr21_+_43669943 | 0.54 |
ENSDART00000136025
|
tmlhe
|
trimethyllysine hydroxylase, epsilon |
| chr25_-_35664817 | 0.52 |
ENSDART00000148718
|
lrrk2
|
leucine-rich repeat kinase 2 |
| chr24_+_29382109 | 0.50 |
ENSDART00000184620
ENSDART00000188414 ENSDART00000186132 ENSDART00000191489 |
ntng1a
|
netrin g1a |
| chr11_-_42554290 | 0.49 |
ENSDART00000130573
|
atp6ap1la
|
ATPase H+ transporting accessory protein 1 like a |
| chr3_+_23248542 | 0.48 |
ENSDART00000185765
ENSDART00000192332 |
ppp1r9ba
|
protein phosphatase 1, regulatory subunit 9Ba |
| chr16_-_18702249 | 0.48 |
ENSDART00000191595
|
fhod3b
|
formin homology 2 domain containing 3b |
| chr3_+_23248704 | 0.47 |
ENSDART00000156032
|
ppp1r9ba
|
protein phosphatase 1, regulatory subunit 9Ba |
| chr4_+_3287819 | 0.46 |
ENSDART00000168633
|
CABZ01085700.1
|
|
| chr4_+_14343706 | 0.43 |
ENSDART00000142845
|
prl2
|
prolactin 2 |
| chr24_+_29381946 | 0.42 |
ENSDART00000189551
|
ntng1a
|
netrin g1a |
| chr23_+_5977965 | 0.41 |
ENSDART00000115403
ENSDART00000183147 |
NAV1 (1 of many)
|
neuron navigator 1 |
| chr25_-_26673570 | 0.40 |
ENSDART00000154917
|
ciartb
|
circadian associated repressor of transcription b |
| chr4_-_8902406 | 0.40 |
ENSDART00000192962
|
mpped1
|
metallophosphoesterase domain containing 1 |
| chr12_+_39685485 | 0.39 |
ENSDART00000163403
|
LO017650.1
|
|
| chr5_-_21044693 | 0.39 |
ENSDART00000140298
|
si:dkey-13n15.2
|
si:dkey-13n15.2 |
| chr20_+_38032143 | 0.38 |
ENSDART00000032161
|
galnt14
|
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 14 (GalNAc-T14) |
| chr6_-_20952187 | 0.37 |
ENSDART00000074327
|
igfbp2a
|
insulin-like growth factor binding protein 2a |
| chr24_+_19591893 | 0.35 |
ENSDART00000152026
|
slco5a1a
|
solute carrier organic anion transporter family member 5A1a |
| chr9_+_38163876 | 0.34 |
ENSDART00000137955
|
clasp1a
|
cytoplasmic linker associated protein 1a |
| chr12_-_17897134 | 0.33 |
ENSDART00000066407
|
nptx2b
|
neuronal pentraxin IIb |
| chr4_+_3980247 | 0.32 |
ENSDART00000049194
|
gpr37b
|
G protein-coupled receptor 37b |
| chr19_-_30904590 | 0.31 |
ENSDART00000137633
|
si:ch211-194e15.5
|
si:ch211-194e15.5 |
| chr3_+_41726360 | 0.31 |
ENSDART00000154401
|
chst12a
|
carbohydrate (chondroitin 4) sulfotransferase 12a |
| chr14_-_30452218 | 0.26 |
ENSDART00000011480
|
zdhhc2
|
zinc finger, DHHC-type containing 2 |
| chr17_-_31719071 | 0.25 |
ENSDART00000136199
|
dtd2
|
D-tyrosyl-tRNA deacylase 2 |
| chr17_-_4252221 | 0.25 |
ENSDART00000152020
|
gdf3
|
growth differentiation factor 3 |
| chr18_+_16246806 | 0.25 |
ENSDART00000142584
|
alx1
|
ALX homeobox 1 |
| chr2_-_7246848 | 0.25 |
ENSDART00000146434
|
zgc:153115
|
zgc:153115 |
| chr7_-_8881514 | 0.24 |
ENSDART00000081620
|
vax2
|
ventral anterior homeobox 2 |
| chr5_+_22791686 | 0.22 |
ENSDART00000014806
|
npas2
|
neuronal PAS domain protein 2 |
| chr4_+_1283068 | 0.22 |
ENSDART00000167233
|
chrm2a
|
cholinergic receptor, muscarinic 2a |
| chr15_-_30815826 | 0.22 |
ENSDART00000156160
ENSDART00000145918 |
msi2b
|
musashi RNA-binding protein 2b |
| chr1_-_50710468 | 0.21 |
ENSDART00000080389
|
fam13a
|
family with sequence similarity 13, member A |
| chr2_-_7246338 | 0.21 |
ENSDART00000186735
|
zgc:153115
|
zgc:153115 |
| chr19_-_19025998 | 0.19 |
ENSDART00000186156
ENSDART00000163359 ENSDART00000167951 |
dync1li1
|
dynein, cytoplasmic 1, light intermediate chain 1 |
| chr18_-_44935174 | 0.18 |
ENSDART00000081025
|
pex16
|
peroxisomal biogenesis factor 16 |
| chr7_+_47243564 | 0.17 |
ENSDART00000098942
ENSDART00000162237 |
znf507
|
zinc finger protein 507 |
| chr16_-_13613475 | 0.16 |
ENSDART00000139102
|
dbpb
|
D site albumin promoter binding protein b |
| chr15_-_16155729 | 0.14 |
ENSDART00000192212
|
si:ch211-259g3.4
|
si:ch211-259g3.4 |
| chr19_-_82504 | 0.13 |
ENSDART00000027864
ENSDART00000160560 |
hnrnpr
|
heterogeneous nuclear ribonucleoprotein R |
| chr20_+_34320635 | 0.12 |
ENSDART00000153207
|
ivns1abpa
|
influenza virus NS1A binding protein a |
| chr14_+_2095394 | 0.11 |
ENSDART00000186847
|
CT573264.2
|
|
| chr7_+_9189547 | 0.09 |
ENSDART00000169783
|
pcsk6
|
proprotein convertase subtilisin/kexin type 6 |
| chr9_+_2002701 | 0.09 |
ENSDART00000082329
|
evx2
|
even-skipped homeobox 2 |
| chr18_-_34170918 | 0.07 |
ENSDART00000015079
|
slc33a1
|
solute carrier family 33 (acetyl-CoA transporter), member 1 |
| chr23_-_15090782 | 0.06 |
ENSDART00000133624
|
si:ch211-218g4.2
|
si:ch211-218g4.2 |
| chr3_+_28860283 | 0.05 |
ENSDART00000077235
|
alg1
|
ALG1, chitobiosyldiphosphodolichol beta-mannosyltransferase |
| chr5_-_38122126 | 0.04 |
ENSDART00000141791
ENSDART00000170528 |
si:ch211-284e13.6
|
si:ch211-284e13.6 |
| chr18_+_20482369 | 0.04 |
ENSDART00000100668
|
kbtbd4
|
kelch repeat and BTB (POZ) domain containing 4 |
| chr18_-_34171280 | 0.02 |
ENSDART00000122321
|
slc33a1
|
solute carrier family 33 (acetyl-CoA transporter), member 1 |
| chr3_+_35406998 | 0.02 |
ENSDART00000102994
|
rbbp6
|
retinoblastoma binding protein 6 |
| chr1_-_46875493 | 0.01 |
ENSDART00000115081
|
agpat3
|
1-acylglycerol-3-phosphate O-acyltransferase 3 |
| chr20_-_14665002 | 0.00 |
ENSDART00000152816
|
scrn2
|
secernin 2 |
Network of associatons between targets according to the STRING database.
First level regulatory network of foxg1a+foxg1c+foxg1d
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0060159 | regulation of dopamine receptor signaling pathway(GO:0060159) |
| 0.1 | 0.5 | GO:0045329 | carnitine biosynthetic process(GO:0045329) |
| 0.1 | 0.2 | GO:0003161 | cardiac conduction system development(GO:0003161) negative regulation of heart rate(GO:0010459) |
| 0.1 | 0.3 | GO:1900145 | nodal signaling pathway involved in determination of left/right asymmetry(GO:0038107) regulation of nodal signaling pathway involved in determination of left/right asymmetry(GO:1900145) |
| 0.1 | 0.3 | GO:0010718 | positive regulation of epithelial to mesenchymal transition(GO:0010718) |
| 0.0 | 0.7 | GO:0036065 | fucosylation(GO:0036065) |
| 0.0 | 0.2 | GO:0009794 | regulation of mitotic cell cycle, embryonic(GO:0009794) mitotic cell cycle, embryonic(GO:0045448) |
| 0.0 | 0.4 | GO:0040014 | regulation of multicellular organism growth(GO:0040014) |
| 0.0 | 0.9 | GO:0070831 | basement membrane assembly(GO:0070831) |
| 0.0 | 0.2 | GO:0090259 | regulation of retinal ganglion cell axon guidance(GO:0090259) |
| 0.0 | 0.4 | GO:0090110 | cargo loading into COPII-coated vesicle(GO:0090110) |
| 0.0 | 0.3 | GO:0016188 | synaptic vesicle maturation(GO:0016188) |
| 0.0 | 0.5 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.0 | 0.3 | GO:0043252 | sodium-independent organic anion transport(GO:0043252) |
| 0.0 | 0.4 | GO:0046427 | positive regulation of JAK-STAT cascade(GO:0046427) positive regulation of STAT cascade(GO:1904894) |
| 0.0 | 0.6 | GO:0032922 | circadian regulation of gene expression(GO:0032922) |
| 0.0 | 0.3 | GO:0048168 | regulation of neuronal synaptic plasticity(GO:0048168) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0005828 | kinetochore microtubule(GO:0005828) |
| 0.1 | 0.5 | GO:0033181 | plasma membrane proton-transporting V-type ATPase complex(GO:0033181) |
| 0.0 | 0.2 | GO:1990513 | CLOCK-BMAL transcription complex(GO:1990513) |
| 0.0 | 0.3 | GO:0098837 | postsynaptic recycling endosome(GO:0098837) |
| 0.0 | 0.9 | GO:0005605 | basal lamina(GO:0005605) laminin complex(GO:0043256) |
| 0.0 | 0.9 | GO:0070971 | endoplasmic reticulum exit site(GO:0070971) |
| 0.0 | 0.2 | GO:0005868 | cytoplasmic dynein complex(GO:0005868) |
| 0.0 | 0.7 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0051500 | D-aminoacyl-tRNA deacylase activity(GO:0051499) D-tyrosyl-tRNA(Tyr) deacylase activity(GO:0051500) |
| 0.1 | 0.7 | GO:0046920 | alpha-(1->3)-fucosyltransferase activity(GO:0046920) |
| 0.0 | 0.3 | GO:0043515 | kinetochore binding(GO:0043515) |
| 0.0 | 0.4 | GO:0031994 | insulin-like growth factor I binding(GO:0031994) insulin-like growth factor II binding(GO:0031995) |
| 0.0 | 0.9 | GO:0098632 | protein binding involved in cell-cell adhesion(GO:0098632) |
| 0.0 | 1.0 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.0 | 0.2 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 0.0 | 0.3 | GO:0015347 | sodium-independent organic anion transmembrane transporter activity(GO:0015347) |
| 0.0 | 0.5 | GO:0051018 | protein kinase A binding(GO:0051018) |
| 0.0 | 0.1 | GO:0051185 | coenzyme transporter activity(GO:0051185) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |