Project
PRJNA207719: Tissue specific transcriptome profiling
Navigation
Downloads
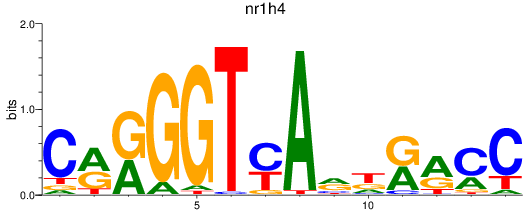
Results for nr1h4
Z-value: 1.51
Transcription factors associated with nr1h4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
nr1h4
|
ENSDARG00000057741 | nuclear receptor subfamily 1, group H, member 4 |
|
nr1h4
|
ENSDARG00000113600 | nuclear receptor subfamily 1, group H, member 4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| nr1h4 | dr11_v1_chr18_+_15644559_15644559 | 0.21 | 7.3e-01 | Click! |
Activity profile of nr1h4 motif
Sorted Z-values of nr1h4 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr3_+_4346854 | 1.57 |
ENSDART00000004273
|
si:dkey-73p2.3
|
si:dkey-73p2.3 |
| chr3_-_34027178 | 1.37 |
ENSDART00000170201
ENSDART00000151408 |
ighv1-4
ighv14-1
|
immunoglobulin heavy variable 1-4 immunoglobulin heavy variable 14-1 |
| chr9_+_48007081 | 1.17 |
ENSDART00000060593
ENSDART00000099835 |
zgc:92380
|
zgc:92380 |
| chr23_+_19790962 | 1.16 |
ENSDART00000142228
|
flna
|
filamin A, alpha (actin binding protein 280) |
| chr18_+_46151505 | 1.07 |
ENSDART00000015034
ENSDART00000141287 |
blvrb
|
biliverdin reductase B |
| chr1_-_58900851 | 1.02 |
ENSDART00000183085
ENSDART00000188855 ENSDART00000182567 |
CABZ01084501.3
|
Danio rerio microfibril-associated glycoprotein 4-like (LOC100334800), transcript variant 2, mRNA. |
| chr18_-_16791331 | 0.94 |
ENSDART00000148222
|
ampd3b
|
adenosine monophosphate deaminase 3b |
| chr3_-_4303262 | 0.92 |
ENSDART00000112819
|
si:dkey-73p2.2
|
si:dkey-73p2.2 |
| chr20_-_19864131 | 0.89 |
ENSDART00000057819
|
ptk2bb
|
protein tyrosine kinase 2 beta, b |
| chr6_-_27139396 | 0.87 |
ENSDART00000055848
|
zgc:103559
|
zgc:103559 |
| chr1_-_43920371 | 0.84 |
ENSDART00000109283
|
scpp7
|
secretory calcium-binding phosphoprotein 7 |
| chr23_+_7471072 | 0.81 |
ENSDART00000135551
|
si:ch211-200e2.1
|
si:ch211-200e2.1 |
| chr24_-_21689146 | 0.75 |
ENSDART00000105917
|
urad
|
ureidoimidazoline (2-oxo-4-hydroxy-4-carboxy-5-) decarboxylase |
| chr15_+_6652396 | 0.74 |
ENSDART00000192813
ENSDART00000157678 |
nop53
|
NOP53 ribosome biogenesis factor |
| chr6_-_609880 | 0.74 |
ENSDART00000149248
ENSDART00000148867 ENSDART00000149414 ENSDART00000148552 ENSDART00000148391 |
lgals2b
|
lectin, galactoside-binding, soluble, 2b |
| chr23_+_42810055 | 0.73 |
ENSDART00000186647
|
myl9a
|
myosin, light chain 9a, regulatory |
| chr1_+_127250 | 0.72 |
ENSDART00000003463
|
f7i
|
coagulation factor VIIi |
| chr4_-_13518381 | 0.69 |
ENSDART00000067153
|
ifng1-1
|
interferon, gamma 1-1 |
| chr10_+_28428222 | 0.68 |
ENSDART00000135003
|
si:ch211-222e20.4
|
si:ch211-222e20.4 |
| chr23_-_44723102 | 0.66 |
ENSDART00000129138
|
mogat3a
|
monoacylglycerol O-acyltransferase 3a |
| chr23_-_21515182 | 0.66 |
ENSDART00000142000
|
rnf207b
|
ring finger protein 207b |
| chr19_-_32493866 | 0.62 |
ENSDART00000052090
|
fuca1.2
|
alpha-L-fucosidase 1, tandem duplicate 2 |
| chr21_-_19918286 | 0.61 |
ENSDART00000180816
|
ppp1r3b
|
protein phosphatase 1, regulatory subunit 3B |
| chr8_-_36469117 | 0.61 |
ENSDART00000111240
|
mhc2dab
|
major histocompatibility complex class II DAB gene |
| chr14_+_904850 | 0.60 |
ENSDART00000161847
|
si:ch73-208h1.1
|
si:ch73-208h1.1 |
| chr2_-_42234484 | 0.60 |
ENSDART00000132617
ENSDART00000136690 ENSDART00000141358 |
apom
|
apolipoprotein M |
| chr5_+_28260158 | 0.59 |
ENSDART00000181434
|
ncaph
|
non-SMC condensin I complex, subunit H |
| chr15_-_19705707 | 0.59 |
ENSDART00000047643
|
sytl2b
|
synaptotagmin-like 2b |
| chr21_-_5879897 | 0.58 |
ENSDART00000184034
|
rpl35
|
ribosomal protein L35 |
| chr1_-_39859626 | 0.56 |
ENSDART00000053763
|
dctd
|
dCMP deaminase |
| chr19_+_30990815 | 0.56 |
ENSDART00000134645
|
sync
|
syncoilin, intermediate filament protein |
| chr7_+_24523017 | 0.54 |
ENSDART00000077047
|
btr09
|
bloodthirsty-related gene family, member 9 |
| chr3_-_34060153 | 0.54 |
ENSDART00000151610
|
ighv1-1
|
immunoglobulin heavy variable 1-1 |
| chr19_+_32166702 | 0.53 |
ENSDART00000021798
|
fabp11a
|
fatty acid binding protein 11a |
| chr7_-_32980017 | 0.52 |
ENSDART00000113744
|
pkp3b
|
plakophilin 3b |
| chr17_-_36529449 | 0.52 |
ENSDART00000187252
|
colec11
|
collectin sub-family member 11 |
| chr9_-_34269066 | 0.52 |
ENSDART00000059955
|
ildr1b
|
immunoglobulin-like domain containing receptor 1b |
| chr25_+_5012791 | 0.50 |
ENSDART00000156970
|
si:ch73-265h17.5
|
si:ch73-265h17.5 |
| chr16_-_20312146 | 0.50 |
ENSDART00000134980
|
si:dkeyp-86h10.3
|
si:dkeyp-86h10.3 |
| chr21_+_40092301 | 0.49 |
ENSDART00000145150
|
serpinf2a
|
serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 2a |
| chr13_+_29238850 | 0.49 |
ENSDART00000026000
|
myofl
|
myoferlin like |
| chr3_+_24062748 | 0.48 |
ENSDART00000188574
|
cbx1a
|
chromobox homolog 1a (HP1 beta homolog Drosophila) |
| chr17_-_36529016 | 0.47 |
ENSDART00000025019
|
colec11
|
collectin sub-family member 11 |
| chr3_-_4253214 | 0.47 |
ENSDART00000155807
|
si:dkey-175d9.2
|
si:dkey-175d9.2 |
| chr15_-_462783 | 0.47 |
ENSDART00000161049
|
CABZ01033205.1
|
Danio rerio uncharacterized LOC100002960 (LOC100002960), mRNA. |
| chr9_-_31090593 | 0.47 |
ENSDART00000089136
|
gpr18
|
G protein-coupled receptor 18 |
| chr1_-_44413629 | 0.46 |
ENSDART00000192747
|
CT583626.1
|
|
| chr21_+_261490 | 0.46 |
ENSDART00000177919
|
jak2a
|
Janus kinase 2a |
| chr5_-_69716501 | 0.46 |
ENSDART00000158956
|
mob1a
|
MOB kinase activator 1A |
| chr8_+_7315625 | 0.46 |
ENSDART00000135655
ENSDART00000181048 |
selenoh
|
selenoprotein H |
| chr8_+_10305400 | 0.46 |
ENSDART00000172400
|
pim1
|
Pim-1 proto-oncogene, serine/threonine kinase |
| chr2_-_37862380 | 0.45 |
ENSDART00000186005
|
si:ch211-284o19.8
|
si:ch211-284o19.8 |
| chr3_+_4113551 | 0.44 |
ENSDART00000192309
|
LO018551.1
|
|
| chr5_-_19833310 | 0.43 |
ENSDART00000138186
|
trpv4
|
transient receptor potential cation channel, subfamily V, member 4 |
| chr9_-_27717006 | 0.43 |
ENSDART00000146860
|
gtf2e1
|
general transcription factor IIE, polypeptide 1, alpha |
| chr16_-_4769877 | 0.43 |
ENSDART00000149421
ENSDART00000054078 |
rpa2
|
replication protein A2 |
| chr2_-_40191603 | 0.42 |
ENSDART00000180691
|
si:ch211-122l24.6
|
si:ch211-122l24.6 |
| chr16_-_7228276 | 0.41 |
ENSDART00000149030
|
nt5c3a
|
5'-nucleotidase, cytosolic IIIA |
| chr15_+_20530649 | 0.41 |
ENSDART00000186312
|
tnfaip1
|
tumor necrosis factor, alpha-induced protein 1 (endothelial) |
| chr9_-_135774 | 0.41 |
ENSDART00000160435
|
FQ377903.1
|
|
| chr14_+_34514336 | 0.41 |
ENSDART00000024440
|
foxi3b
|
forkhead box I3b |
| chr19_+_30990129 | 0.40 |
ENSDART00000052169
ENSDART00000193376 |
sync
|
syncoilin, intermediate filament protein |
| chr7_-_8014186 | 0.40 |
ENSDART00000190012
|
si:cabz01030277.1
|
si:cabz01030277.1 |
| chr23_+_22597624 | 0.39 |
ENSDART00000054337
|
gpr157
|
G protein-coupled receptor 157 |
| chr22_+_9294336 | 0.39 |
ENSDART00000193694
|
si:ch211-250k18.7
|
si:ch211-250k18.7 |
| chr8_-_13013123 | 0.39 |
ENSDART00000147802
|
dennd2da
|
DENN/MADD domain containing 2Da |
| chr9_+_15890558 | 0.38 |
ENSDART00000144032
|
si:dkey-14o1.20
|
si:dkey-14o1.20 |
| chr11_-_544187 | 0.38 |
ENSDART00000173161
|
CU695207.1
|
|
| chr2_+_371759 | 0.38 |
ENSDART00000153788
|
si:dkey-33c14.7
|
si:dkey-33c14.7 |
| chr15_+_24563504 | 0.37 |
ENSDART00000140658
ENSDART00000130589 ENSDART00000045549 |
dhrs13b
|
dehydrogenase/reductase (SDR family) member 13b |
| chr5_+_28830388 | 0.37 |
ENSDART00000149150
|
serpina7
|
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 7 |
| chr13_-_37200120 | 0.36 |
ENSDART00000135001
|
si:dkeyp-77c8.3
|
si:dkeyp-77c8.3 |
| chr22_-_36926342 | 0.36 |
ENSDART00000151804
|
si:dkey-37m8.11
|
si:dkey-37m8.11 |
| chr3_+_24595922 | 0.35 |
ENSDART00000169405
|
si:dkey-68o6.5
|
si:dkey-68o6.5 |
| chr17_+_14711765 | 0.35 |
ENSDART00000012889
|
cx28.6
|
connexin 28.6 |
| chr5_-_38777852 | 0.33 |
ENSDART00000131603
|
si:dkey-58f10.4
|
si:dkey-58f10.4 |
| chr7_+_22792132 | 0.33 |
ENSDART00000135207
ENSDART00000146801 |
rbm4.3
|
RNA binding motif protein 4.3 |
| chr6_-_1514767 | 0.32 |
ENSDART00000067586
|
chchd6b
|
coiled-coil-helix-coiled-coil-helix domain containing 6b |
| chr17_-_50018133 | 0.32 |
ENSDART00000112267
|
filip1a
|
filamin A interacting protein 1a |
| chr24_+_25467465 | 0.32 |
ENSDART00000189933
|
smpx
|
small muscle protein, X-linked |
| chr25_-_255971 | 0.32 |
ENSDART00000162006
|
CU855936.1
|
|
| chr5_+_54400971 | 0.32 |
ENSDART00000169695
|
bspry
|
B-box and SPRY domain containing |
| chr9_+_2499627 | 0.30 |
ENSDART00000160782
|
wipf1a
|
WAS/WASL interacting protein family, member 1a |
| chr22_-_17474583 | 0.30 |
ENSDART00000148027
|
si:ch211-197g15.8
|
si:ch211-197g15.8 |
| chr15_-_31000275 | 0.30 |
ENSDART00000132493
ENSDART00000100183 |
lgals9l6
lgals9l5
|
lectin, galactoside-binding, soluble, 9 (galectin 9)-like 6 lectin, galactoside-binding, soluble, 9 (galectin 9)-like 5 |
| chr21_-_2360906 | 0.30 |
ENSDART00000171645
|
si:ch73-299h12.9
|
si:ch73-299h12.9 |
| chr21_-_20733615 | 0.30 |
ENSDART00000145544
|
si:ch211-22d5.2
|
si:ch211-22d5.2 |
| chr4_-_77114795 | 0.29 |
ENSDART00000144849
|
CU467646.2
|
|
| chr15_+_37973197 | 0.29 |
ENSDART00000156661
|
si:dkey-238d18.9
|
si:dkey-238d18.9 |
| chr10_-_26202766 | 0.28 |
ENSDART00000136393
|
fhdc3
|
FH2 domain containing 3 |
| chr20_-_13625588 | 0.28 |
ENSDART00000078893
|
sytl3
|
synaptotagmin-like 3 |
| chr1_-_9130136 | 0.28 |
ENSDART00000113013
|
palb2
|
partner and localizer of BRCA2 |
| chr4_+_70013627 | 0.27 |
ENSDART00000180432
|
si:dkey-190j3.4
|
si:dkey-190j3.4 |
| chr9_-_28616436 | 0.27 |
ENSDART00000136985
|
si:ch73-7i4.2
|
si:ch73-7i4.2 |
| chr15_+_2857556 | 0.27 |
ENSDART00000157758
|
mre11a
|
MRE11 homolog A, double strand break repair nuclease |
| chr4_+_60492313 | 0.26 |
ENSDART00000191043
ENSDART00000191631 |
si:dkey-211i20.2
|
si:dkey-211i20.2 |
| chr8_+_1769475 | 0.26 |
ENSDART00000079073
|
serpind1
|
serpin peptidase inhibitor, clade D (heparin cofactor), member 1 |
| chr16_-_11789108 | 0.25 |
ENSDART00000060142
|
tlr21
|
toll-like receptor 21 |
| chr1_+_58139102 | 0.25 |
ENSDART00000134826
|
si:ch211-15j1.4
|
si:ch211-15j1.4 |
| chr12_+_695619 | 0.25 |
ENSDART00000161691
|
abcc3
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 3 |
| chr15_+_37331585 | 0.25 |
ENSDART00000170715
|
zgc:171592
|
zgc:171592 |
| chr23_+_20431388 | 0.24 |
ENSDART00000132920
ENSDART00000102963 ENSDART00000109899 ENSDART00000140219 |
slc35c2
|
solute carrier family 35 (GDP-fucose transporter), member C2 |
| chr1_+_58303892 | 0.23 |
ENSDART00000147678
|
CR769768.1
|
|
| chr10_+_20070178 | 0.23 |
ENSDART00000027612
ENSDART00000145264 ENSDART00000172713 |
xpo7
|
exportin 7 |
| chr7_-_59159253 | 0.23 |
ENSDART00000159285
|
haus6
|
HAUS augmin-like complex, subunit 6 |
| chr22_+_25590391 | 0.23 |
ENSDART00000178133
|
aars2
|
alanyl-tRNA synthetase 2, mitochondrial (putative) |
| chr13_-_23665580 | 0.23 |
ENSDART00000144282
|
map3k21
|
mitogen-activated protein kinase kinase kinase 21 |
| chr1_+_58323980 | 0.23 |
ENSDART00000145743
|
si:dkey-222h21.3
|
si:dkey-222h21.3 |
| chr21_+_44112914 | 0.22 |
ENSDART00000062836
|
fgf1b
|
fibroblast growth factor 1b |
| chr7_+_38667066 | 0.22 |
ENSDART00000013394
|
mtch2
|
mitochondrial carrier homolog 2 |
| chr21_-_22664554 | 0.21 |
ENSDART00000161438
|
gig2k
|
grass carp reovirus (GCRV)-induced gene 2k |
| chr22_+_10645088 | 0.21 |
ENSDART00000142265
|
rassf1
|
Ras association (RalGDS/AF-6) domain family 1 |
| chr2_+_10127762 | 0.21 |
ENSDART00000100726
|
insl5b
|
insulin-like 5b |
| chr22_+_21252790 | 0.21 |
ENSDART00000079046
|
cpamd8
|
C3 and PZP-like, alpha-2-macroglobulin domain containing 8 |
| chr21_+_29179887 | 0.20 |
ENSDART00000161941
|
si:ch211-57b15.1
|
si:ch211-57b15.1 |
| chr4_-_38281909 | 0.20 |
ENSDART00000162448
|
si:ch211-229l10.9
|
si:ch211-229l10.9 |
| chr17_+_30843881 | 0.20 |
ENSDART00000149600
ENSDART00000148547 |
tpp1
|
tripeptidyl peptidase I |
| chr15_+_45563656 | 0.20 |
ENSDART00000157501
|
cldn15lb
|
claudin 15-like b |
| chr21_-_45186431 | 0.19 |
ENSDART00000193449
|
si:ch73-269m14.2
|
si:ch73-269m14.2 |
| chr11_+_691734 | 0.19 |
ENSDART00000191463
|
timp4.1
|
TIMP metallopeptidase inhibitor 4, tandem duplicate 1 |
| chr16_-_17175731 | 0.19 |
ENSDART00000183057
|
opn9
|
opsin 9 |
| chr15_+_37992275 | 0.19 |
ENSDART00000154928
|
si:dkey-238d18.7
|
si:dkey-238d18.7 |
| chr10_+_44363195 | 0.18 |
ENSDART00000161744
|
kmt5aa
|
lysine methyltransferase 5Aa |
| chr7_+_3597413 | 0.18 |
ENSDART00000064281
|
si:dkey-192d15.2
|
si:dkey-192d15.2 |
| chr7_+_36539124 | 0.18 |
ENSDART00000173653
|
chd9
|
chromodomain helicase DNA binding protein 9 |
| chr14_+_22297257 | 0.18 |
ENSDART00000113676
|
atp10b
|
ATPase phospholipid transporting 10B |
| chr23_+_28809002 | 0.17 |
ENSDART00000134121
ENSDART00000183661 |
pex14
|
peroxisomal biogenesis factor 14 |
| chr13_-_37108310 | 0.17 |
ENSDART00000163845
|
syne2b
|
spectrin repeat containing, nuclear envelope 2b |
| chr7_+_34688527 | 0.17 |
ENSDART00000108473
|
plekhg4
|
pleckstrin homology domain containing, family G (with RhoGef domain) member 4 |
| chr5_-_16218777 | 0.17 |
ENSDART00000141698
|
kremen1
|
kringle containing transmembrane protein 1 |
| chr12_-_25201576 | 0.17 |
ENSDART00000077188
|
cox7a3
|
cytochrome c oxidase subunit VIIa polypeptide 3 |
| chr21_-_22547496 | 0.16 |
ENSDART00000166835
ENSDART00000089030 |
myo5b
|
myosin VB |
| chr12_-_14143344 | 0.16 |
ENSDART00000152742
|
buc2l
|
bucky ball 2-like |
| chr11_-_14924138 | 0.16 |
ENSDART00000168348
|
si:dkey-6d5.1
|
si:dkey-6d5.1 |
| chr19_+_2685779 | 0.16 |
ENSDART00000160533
ENSDART00000097531 |
tomm7
|
translocase of outer mitochondrial membrane 7 homolog (yeast) |
| chr13_-_337318 | 0.16 |
ENSDART00000166175
|
zgc:171534
|
zgc:171534 |
| chr7_-_24838367 | 0.16 |
ENSDART00000139455
ENSDART00000012483 ENSDART00000131530 |
fam113
|
family with sequence similarity 113 |
| chr11_+_11200550 | 0.16 |
ENSDART00000181339
ENSDART00000187116 |
myom2a
|
myomesin 2a |
| chr7_+_39399747 | 0.15 |
ENSDART00000147037
|
tnni2b.1
|
troponin I type 2b (skeletal, fast), tandem duplicate 1 |
| chr2_+_49799470 | 0.14 |
ENSDART00000146325
|
si:ch211-190k17.19
|
si:ch211-190k17.19 |
| chr16_+_14684916 | 0.14 |
ENSDART00000138611
|
col14a1a
|
collagen, type XIV, alpha 1a |
| chr15_-_37752236 | 0.14 |
ENSDART00000154263
|
si:dkey-42l23.5
|
si:dkey-42l23.5 |
| chr13_-_30713236 | 0.14 |
ENSDART00000112372
ENSDART00000142221 |
tmem72
|
transmembrane protein 72 |
| chr3_-_58582663 | 0.13 |
ENSDART00000180055
|
CABZ01038521.1
|
|
| chr14_-_33308138 | 0.13 |
ENSDART00000136442
ENSDART00000139615 |
sept6
|
septin 6 |
| chr20_-_7547080 | 0.13 |
ENSDART00000146135
|
usp24
|
ubiquitin specific peptidase 24 |
| chr21_+_12056934 | 0.12 |
ENSDART00000125380
|
zgc:162344
|
zgc:162344 |
| chr22_-_17474781 | 0.12 |
ENSDART00000186817
|
si:ch211-197g15.8
|
si:ch211-197g15.8 |
| chr5_+_50913357 | 0.12 |
ENSDART00000092938
|
col4a3bpa
|
collagen, type IV, alpha 3 (Goodpasture antigen) binding protein a |
| chr18_+_31056645 | 0.12 |
ENSDART00000159316
|
mvda
|
mevalonate (diphospho) decarboxylase a |
| chr1_-_44940830 | 0.11 |
ENSDART00000097500
ENSDART00000134464 ENSDART00000137216 |
tmem176
|
transmembrane protein 176 |
| chr22_+_8313513 | 0.11 |
ENSDART00000181169
ENSDART00000103911 |
CABZ01077217.1
|
|
| chr10_-_21542702 | 0.11 |
ENSDART00000146761
ENSDART00000134502 |
zgc:165539
|
zgc:165539 |
| chr14_-_8787525 | 0.11 |
ENSDART00000160848
|
tgfb2l
|
transforming growth factor, beta 2, like |
| chr22_+_24318131 | 0.11 |
ENSDART00000187360
ENSDART00000165618 |
ccdc50
|
coiled-coil domain containing 50 |
| chr20_-_28352352 | 0.11 |
ENSDART00000128806
|
ino80
|
INO80 complex subunit |
| chr12_-_4370585 | 0.11 |
ENSDART00000129502
|
CA4 (1 of many)
|
si:ch211-173d10.4 |
| chr17_+_16192486 | 0.10 |
ENSDART00000156832
|
kif13ba
|
kinesin family member 13Ba |
| chr2_-_37281748 | 0.10 |
ENSDART00000137598
|
nadkb
|
NAD kinase b |
| chr25_+_245018 | 0.10 |
ENSDART00000155344
|
zgc:92481
|
zgc:92481 |
| chr2_+_49644803 | 0.10 |
ENSDART00000160342
|
si:ch211-209f23.7
|
si:ch211-209f23.7 |
| chr19_-_1929310 | 0.10 |
ENSDART00000144113
|
si:ch211-149a19.3
|
si:ch211-149a19.3 |
| chr10_-_18463934 | 0.09 |
ENSDART00000133116
ENSDART00000113422 |
si:dkey-28o19.1
|
si:dkey-28o19.1 |
| chr24_-_25184553 | 0.09 |
ENSDART00000166917
|
plcxd2
|
phosphatidylinositol-specific phospholipase C, X domain containing 2 |
| chr25_+_32473433 | 0.08 |
ENSDART00000152326
|
sqor
|
sulfide quinone oxidoreductase |
| chr4_-_31784872 | 0.08 |
ENSDART00000182911
|
si:dkey-19c16.12
|
si:dkey-19c16.12 |
| chr15_-_17619306 | 0.08 |
ENSDART00000184011
|
adamts15b
|
ADAM metallopeptidase with thrombospondin type 1 motif, 15b |
| chr25_+_23591990 | 0.08 |
ENSDART00000151938
ENSDART00000089199 |
cpt1ab
|
carnitine palmitoyltransferase 1Ab (liver) |
| chr3_+_12835370 | 0.08 |
ENSDART00000166089
|
si:ch211-8c17.3
|
si:ch211-8c17.3 |
| chr22_+_28888781 | 0.08 |
ENSDART00000145629
|
pimr98
|
Pim proto-oncogene, serine/threonine kinase, related 98 |
| chr11_-_18017287 | 0.08 |
ENSDART00000155443
|
qrich1
|
glutamine-rich 1 |
| chr9_-_48184823 | 0.08 |
ENSDART00000180264
|
klhl23
|
kelch-like family member 23 |
| chr2_-_27651674 | 0.08 |
ENSDART00000177402
|
tgs1
|
trimethylguanosine synthase 1 |
| chr9_+_38158570 | 0.08 |
ENSDART00000059549
ENSDART00000133060 |
nifk
|
nucleolar protein interacting with the FHA domain of MKI67 |
| chr11_+_12175162 | 0.07 |
ENSDART00000125446
|
si:ch211-156l18.7
|
si:ch211-156l18.7 |
| chr25_+_32473277 | 0.07 |
ENSDART00000146451
|
sqor
|
sulfide quinone oxidoreductase |
| chr21_-_14811058 | 0.07 |
ENSDART00000143100
|
phpt1
|
phosphohistidine phosphatase 1 |
| chr18_+_34181655 | 0.07 |
ENSDART00000130831
ENSDART00000109535 |
gmps
|
guanine monophosphate synthase |
| chr17_+_35431724 | 0.07 |
ENSDART00000190293
|
BX571665.1
|
|
| chr17_+_996509 | 0.07 |
ENSDART00000158830
|
cyp1c2
|
cytochrome P450, family 1, subfamily C, polypeptide 2 |
| chr15_-_34322915 | 0.06 |
ENSDART00000190543
ENSDART00000019651 ENSDART00000193601 |
dgkb
|
diacylglycerol kinase, beta |
| chr21_+_27558381 | 0.06 |
ENSDART00000024287
ENSDART00000144869 |
zgc:165604
|
zgc:165604 |
| chr5_+_13385837 | 0.06 |
ENSDART00000191190
|
ccl19a.1
|
chemokine (C-C motif) ligand 19a, tandem duplicate 1 |
| chr5_+_65946222 | 0.05 |
ENSDART00000190969
ENSDART00000161578 |
mymk
|
myomaker, myoblast fusion factor |
| chr21_+_27370671 | 0.05 |
ENSDART00000009234
ENSDART00000142071 |
rbm14a
|
RNA binding motif protein 14a |
| chr8_-_19342111 | 0.05 |
ENSDART00000138881
|
rgl1
|
ral guanine nucleotide dissociation stimulator-like 1 |
| chr19_-_20199167 | 0.05 |
ENSDART00000155527
|
si:ch211-155k24.9
|
si:ch211-155k24.9 |
| chr11_+_24340166 | 0.04 |
ENSDART00000188719
|
rbm39a
|
RNA binding motif protein 39a |
| chr23_+_43954809 | 0.04 |
ENSDART00000164080
|
corin
|
corin, serine peptidase |
| chr21_+_12036238 | 0.04 |
ENSDART00000102463
ENSDART00000155426 |
zgc:162344
|
zgc:162344 |
| chr6_+_27418541 | 0.04 |
ENSDART00000187410
|
crocc2
|
ciliary rootlet coiled-coil, rootletin family member 2 |
| chr17_+_20173882 | 0.04 |
ENSDART00000155379
|
si:ch211-248a14.8
|
si:ch211-248a14.8 |
| chr22_+_7486867 | 0.04 |
ENSDART00000034586
|
CELA1 (1 of many)
|
zgc:112302 |
| chr2_-_52020049 | 0.04 |
ENSDART00000128198
ENSDART00000135331 |
scn12aa
|
sodium channel, voltage gated, type XII, alpha a |
| chr18_-_21188768 | 0.04 |
ENSDART00000060166
|
got2a
|
glutamic-oxaloacetic transaminase 2a, mitochondrial |
| chr4_-_18455480 | 0.03 |
ENSDART00000150221
|
sco2
|
SCO2 cytochrome c oxidase assembly protein |
| chr17_-_15149192 | 0.03 |
ENSDART00000180511
ENSDART00000103405 |
gch1
|
GTP cyclohydrolase 1 |
| chr24_-_36337395 | 0.03 |
ENSDART00000154402
|
si:ch211-40k21.9
|
si:ch211-40k21.9 |
| chr6_+_38626684 | 0.03 |
ENSDART00000086533
|
atp10a
|
ATPase phospholipid transporting 10A |
| chr24_+_28528000 | 0.03 |
ENSDART00000155924
|
arhgap29a
|
Rho GTPase activating protein 29a |
Network of associatons between targets according to the STRING database.
First level regulatory network of nr1h4
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.6 | GO:0000255 | allantoin metabolic process(GO:0000255) |
| 0.2 | 0.7 | GO:1903779 | regulation of cardiac conduction(GO:1903779) |
| 0.2 | 0.6 | GO:0005981 | regulation of glycogen catabolic process(GO:0005981) |
| 0.2 | 0.6 | GO:0009162 | deoxyribonucleoside monophosphate metabolic process(GO:0009162) |
| 0.1 | 0.4 | GO:0071498 | cellular response to osmotic stress(GO:0071470) cellular response to fluid shear stress(GO:0071498) |
| 0.1 | 0.4 | GO:0001112 | DNA-templated transcriptional open complex formation(GO:0001112) transcriptional open complex formation at RNA polymerase II promoter(GO:0001113) protein-DNA complex remodeling(GO:0001120) |
| 0.1 | 0.6 | GO:0006004 | fucose metabolic process(GO:0006004) |
| 0.1 | 0.7 | GO:0030195 | negative regulation of blood coagulation(GO:0030195) negative regulation of hemostasis(GO:1900047) |
| 0.1 | 0.4 | GO:0009957 | epidermal cell fate specification(GO:0009957) |
| 0.1 | 1.2 | GO:0019730 | antimicrobial humoral response(GO:0019730) |
| 0.1 | 0.2 | GO:0006419 | alanyl-tRNA aminoacylation(GO:0006419) |
| 0.1 | 0.9 | GO:0007172 | signal complex assembly(GO:0007172) |
| 0.1 | 0.9 | GO:0032264 | IMP salvage(GO:0032264) |
| 0.1 | 0.3 | GO:2000562 | negative regulation of interferon-gamma production(GO:0032689) CD4-positive, alpha-beta T cell proliferation(GO:0035739) regulation of CD4-positive, alpha-beta T cell proliferation(GO:2000561) negative regulation of CD4-positive, alpha-beta T cell proliferation(GO:2000562) |
| 0.1 | 0.2 | GO:0070221 | sulfide oxidation(GO:0019418) sulfide oxidation, using sulfide:quinone oxidoreductase(GO:0070221) |
| 0.0 | 0.6 | GO:0007076 | mitotic chromosome condensation(GO:0007076) |
| 0.0 | 0.2 | GO:0003241 | growth involved in heart morphogenesis(GO:0003241) cardiac chamber ballooning(GO:0003242) cardiac muscle tissue growth involved in heart morphogenesis(GO:0003245) |
| 0.0 | 1.9 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.0 | 0.7 | GO:0000463 | maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000463) |
| 0.0 | 0.2 | GO:0048903 | anterior lateral line neuromast hair cell differentiation(GO:0048903) |
| 0.0 | 0.3 | GO:0042138 | meiotic DNA double-strand break formation(GO:0042138) |
| 0.0 | 0.4 | GO:0070212 | protein poly-ADP-ribosylation(GO:0070212) |
| 0.0 | 0.7 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 0.0 | 0.4 | GO:0035024 | negative regulation of Rho protein signal transduction(GO:0035024) |
| 0.0 | 0.2 | GO:0051045 | negative regulation of membrane protein ectodomain proteolysis(GO:0051045) |
| 0.0 | 0.1 | GO:0019287 | isopentenyl diphosphate biosynthetic process, mevalonate pathway(GO:0019287) |
| 0.0 | 0.5 | GO:0048922 | posterior lateral line neuromast deposition(GO:0048922) |
| 0.0 | 0.1 | GO:0035971 | peptidyl-histidine dephosphorylation(GO:0035971) |
| 0.0 | 0.5 | GO:0007520 | myoblast fusion(GO:0007520) |
| 0.0 | 0.1 | GO:0042766 | nucleosome mobilization(GO:0042766) |
| 0.0 | 0.2 | GO:0045444 | fat cell differentiation(GO:0045444) |
| 0.0 | 0.2 | GO:0006611 | protein export from nucleus(GO:0006611) |
| 0.0 | 0.7 | GO:0019432 | triglyceride biosynthetic process(GO:0019432) |
| 0.0 | 0.5 | GO:0046427 | positive regulation of JAK-STAT cascade(GO:0046427) positive regulation of STAT cascade(GO:1904894) |
| 0.0 | 0.2 | GO:0016560 | protein import into peroxisome matrix, docking(GO:0016560) protein to membrane docking(GO:0022615) |
| 0.0 | 0.5 | GO:0045454 | cell redox homeostasis(GO:0045454) |
| 0.0 | 0.5 | GO:0007634 | optokinetic behavior(GO:0007634) |
| 0.0 | 0.5 | GO:0019882 | antigen processing and presentation(GO:0019882) |
| 0.0 | 0.1 | GO:0006741 | NADP biosynthetic process(GO:0006741) |
| 0.0 | 1.1 | GO:0010951 | negative regulation of endopeptidase activity(GO:0010951) |
| 0.0 | 0.2 | GO:0030323 | respiratory tube development(GO:0030323) lung development(GO:0030324) |
| 0.0 | 0.4 | GO:0040023 | nuclear migration(GO:0007097) establishment of nucleus localization(GO:0040023) |
| 0.0 | 0.5 | GO:0035025 | positive regulation of Rho protein signal transduction(GO:0035025) |
| 0.0 | 0.3 | GO:0007007 | inner mitochondrial membrane organization(GO:0007007) |
| 0.0 | 0.1 | GO:0072383 | plus-end-directed vesicle transport along microtubule(GO:0072383) plus-end-directed organelle transport along microtubule(GO:0072386) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0005673 | transcription factor TFIIE complex(GO:0005673) |
| 0.1 | 0.6 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.1 | 0.6 | GO:0000796 | condensin complex(GO:0000796) |
| 0.1 | 0.3 | GO:0043034 | costamere(GO:0043034) |
| 0.1 | 0.4 | GO:0005662 | DNA replication factor A complex(GO:0005662) |
| 0.1 | 0.2 | GO:1990429 | Pex17p-Pex14p docking complex(GO:1990415) peroxisomal importomer complex(GO:1990429) |
| 0.1 | 0.2 | GO:1990498 | mitotic spindle microtubule(GO:1990498) |
| 0.0 | 0.3 | GO:0030870 | Mre11 complex(GO:0030870) |
| 0.0 | 1.9 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.0 | 0.6 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.0 | 0.3 | GO:0061617 | MICOS complex(GO:0061617) |
| 0.0 | 0.5 | GO:0030057 | desmosome(GO:0030057) |
| 0.0 | 2.2 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 0.4 | GO:0034992 | microtubule organizing center attachment site(GO:0034992) LINC complex(GO:0034993) |
| 0.0 | 0.1 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.0 | 0.2 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.0 | 0.4 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.0 | GO:0042806 | fucose binding(GO:0042806) |
| 0.2 | 0.9 | GO:0050897 | cobalt ion binding(GO:0050897) |
| 0.2 | 0.6 | GO:0015928 | alpha-L-fucosidase activity(GO:0004560) fucosidase activity(GO:0015928) |
| 0.1 | 0.7 | GO:0008097 | 5S rRNA binding(GO:0008097) |
| 0.1 | 1.0 | GO:0016936 | galactoside binding(GO:0016936) |
| 0.1 | 0.9 | GO:0047623 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 0.1 | 0.2 | GO:0004813 | alanine-tRNA ligase activity(GO:0004813) |
| 0.1 | 0.7 | GO:0004144 | diacylglycerol O-acyltransferase activity(GO:0004144) |
| 0.1 | 0.2 | GO:0070224 | sulfide:quinone oxidoreductase activity(GO:0070224) |
| 0.0 | 0.3 | GO:0000014 | single-stranded DNA endodeoxyribonuclease activity(GO:0000014) |
| 0.0 | 0.7 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.0 | 1.9 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.0 | 0.6 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.0 | 0.9 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.0 | 0.4 | GO:1990404 | protein ADP-ribosylase activity(GO:1990404) |
| 0.0 | 0.6 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.0 | 0.5 | GO:0033691 | sialic acid binding(GO:0033691) |
| 0.0 | 0.2 | GO:0004706 | JUN kinase kinase kinase activity(GO:0004706) |
| 0.0 | 0.2 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.0 | 0.1 | GO:0050649 | testosterone 6-beta-hydroxylase activity(GO:0050649) |
| 0.0 | 0.4 | GO:0030552 | cAMP binding(GO:0030552) |
| 0.0 | 0.4 | GO:0008253 | 5'-nucleotidase activity(GO:0008253) |
| 0.0 | 0.9 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.0 | 0.1 | GO:0034714 | type III transforming growth factor beta receptor binding(GO:0034714) |
| 0.0 | 0.1 | GO:0101006 | protein histidine phosphatase activity(GO:0101006) |
| 0.0 | 0.6 | GO:0019239 | deaminase activity(GO:0019239) |
| 0.0 | 1.4 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 0.2 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.0 | 0.1 | GO:0016416 | carnitine O-palmitoyltransferase activity(GO:0004095) O-palmitoyltransferase activity(GO:0016416) |
| 0.0 | 1.3 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.2 | PID WNT NONCANONICAL PATHWAY | Noncanonical Wnt signaling pathway |
| 0.0 | 0.5 | PID IL5 PATHWAY | IL5-mediated signaling events |
| 0.0 | 1.0 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.0 | 0.6 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.0 | 1.0 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.1 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.0 | 0.2 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 0.0 | 0.4 | REACTOME REMOVAL OF THE FLAP INTERMEDIATE FROM THE C STRAND | Genes involved in Removal of the Flap Intermediate from the C-strand |
| 0.0 | 0.6 | REACTOME PYRIMIDINE METABOLISM | Genes involved in Pyrimidine metabolism |
| 0.0 | 0.5 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.0 | 0.3 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.0 | 0.3 | REACTOME HOMOLOGOUS RECOMBINATION REPAIR OF REPLICATION INDEPENDENT DOUBLE STRAND BREAKS | Genes involved in Homologous recombination repair of replication-independent double-strand breaks |
| 0.0 | 0.2 | REACTOME MITOCHONDRIAL TRNA AMINOACYLATION | Genes involved in Mitochondrial tRNA aminoacylation |