Project
PRJNA207719: Tissue specific transcriptome profiling
Navigation
Downloads
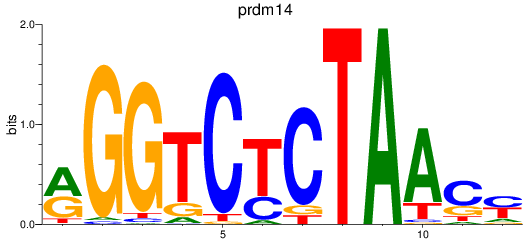
Results for prdm14
Z-value: 0.97
Transcription factors associated with prdm14
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
prdm14
|
ENSDARG00000045371 | PR domain containing 14 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| prdm14 | dr11_v1_chr24_+_14451404_14451404 | 0.89 | 4.1e-02 | Click! |
Activity profile of prdm14 motif
Sorted Z-values of prdm14 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr14_-_30642819 | 1.29 |
ENSDART00000078154
|
npas4a
|
neuronal PAS domain protein 4a |
| chr21_-_43949208 | 1.28 |
ENSDART00000150983
|
camk2a
|
calcium/calmodulin-dependent protein kinase II alpha |
| chr20_+_27020201 | 1.24 |
ENSDART00000126919
ENSDART00000016014 |
chga
|
chromogranin A |
| chr4_-_22207873 | 1.12 |
ENSDART00000142140
|
ppfia2
|
protein tyrosine phosphatase, receptor type, f polypeptide (PTPRF), interacting protein (liprin), alpha 2 |
| chr5_+_49744713 | 0.97 |
ENSDART00000133384
|
nr2f1a
|
nuclear receptor subfamily 2, group F, member 1a |
| chr11_-_23080970 | 0.92 |
ENSDART00000127791
|
atp2b2
|
ATPase plasma membrane Ca2+ transporting 2 |
| chr25_-_8030113 | 0.83 |
ENSDART00000104674
|
camk1db
|
calcium/calmodulin-dependent protein kinase 1Db |
| chr21_-_21089781 | 0.78 |
ENSDART00000144361
|
ank1b
|
ankyrin 1, erythrocytic b |
| chr21_+_44300689 | 0.74 |
ENSDART00000186298
ENSDART00000142810 |
gabra3
|
gamma-aminobutyric acid (GABA) A receptor, alpha 3 |
| chr25_-_8030425 | 0.73 |
ENSDART00000014964
|
camk1db
|
calcium/calmodulin-dependent protein kinase 1Db |
| chr12_+_39685485 | 0.73 |
ENSDART00000163403
|
LO017650.1
|
|
| chr7_+_13418812 | 0.70 |
ENSDART00000191905
ENSDART00000091567 |
dagla
|
diacylglycerol lipase, alpha |
| chr3_-_2593859 | 0.68 |
ENSDART00000143826
|
si:dkey-217f16.5
|
si:dkey-217f16.5 |
| chr22_-_11137268 | 0.65 |
ENSDART00000178882
|
atp6ap2
|
ATPase H+ transporting accessory protein 2 |
| chr5_-_45877387 | 0.64 |
ENSDART00000183714
ENSDART00000041503 |
slc4a4a
|
solute carrier family 4 (sodium bicarbonate cotransporter), member 4a |
| chr1_+_41849152 | 0.61 |
ENSDART00000053685
|
smox
|
spermine oxidase |
| chr25_-_1235457 | 0.61 |
ENSDART00000093093
|
coro2bb
|
coronin, actin binding protein, 2Bb |
| chr22_-_11136625 | 0.59 |
ENSDART00000016873
ENSDART00000125561 |
atp6ap2
|
ATPase H+ transporting accessory protein 2 |
| chr12_-_6818676 | 0.57 |
ENSDART00000106391
|
pcdh15b
|
protocadherin-related 15b |
| chr22_-_26323893 | 0.56 |
ENSDART00000105099
|
capn1b
|
calpain 1, (mu/I) large subunit b |
| chr23_+_39558508 | 0.46 |
ENSDART00000017902
|
camk1gb
|
calcium/calmodulin-dependent protein kinase IGb |
| chr22_+_39074688 | 0.44 |
ENSDART00000153547
|
ip6k1
|
inositol hexakisphosphate kinase 1 |
| chr25_+_16043246 | 0.44 |
ENSDART00000186663
|
sb:cb470
|
sb:cb470 |
| chr7_+_48806420 | 0.44 |
ENSDART00000083431
|
cpt1aa
|
carnitine palmitoyltransferase 1Aa (liver) |
| chr11_-_18791834 | 0.42 |
ENSDART00000156431
|
pfkfb2b
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 2b |
| chr1_+_46509176 | 0.41 |
ENSDART00000166028
|
mcf2la
|
mcf.2 cell line derived transforming sequence-like a |
| chr5_-_69482891 | 0.41 |
ENSDART00000109487
|
CABZ01032476.1
|
|
| chr7_+_48805534 | 0.41 |
ENSDART00000145375
ENSDART00000148744 |
cpt1aa
|
carnitine palmitoyltransferase 1Aa (liver) |
| chr22_-_20342260 | 0.40 |
ENSDART00000161610
ENSDART00000165667 |
tcf3b
|
transcription factor 3b |
| chr7_+_48805725 | 0.39 |
ENSDART00000166543
|
cpt1aa
|
carnitine palmitoyltransferase 1Aa (liver) |
| chr15_-_42760110 | 0.39 |
ENSDART00000152490
|
si:ch211-181d7.3
|
si:ch211-181d7.3 |
| chr8_+_35172594 | 0.37 |
ENSDART00000177146
|
BX897670.1
|
|
| chr5_+_26079178 | 0.36 |
ENSDART00000145920
|
si:dkey-201c13.2
|
si:dkey-201c13.2 |
| chr9_+_33261330 | 0.36 |
ENSDART00000135384
|
usp9
|
ubiquitin specific peptidase 9 |
| chr10_-_32877348 | 0.32 |
ENSDART00000018977
ENSDART00000133421 |
rabgef1
|
RAB guanine nucleotide exchange factor (GEF) 1 |
| chr16_-_26255877 | 0.32 |
ENSDART00000146214
|
erfl1
|
Ets2 repressor factor like 1 |
| chr5_-_67115872 | 0.31 |
ENSDART00000065262
|
rps6ka4
|
ribosomal protein S6 kinase, polypeptide 4 |
| chr20_-_46467280 | 0.30 |
ENSDART00000060702
|
rmdn3
|
regulator of microtubule dynamics 3 |
| chr9_-_33477588 | 0.27 |
ENSDART00000144150
|
caska
|
calcium/calmodulin-dependent serine protein kinase a |
| chr6_+_518979 | 0.25 |
ENSDART00000151012
|
CACNA1I (1 of many)
|
si:ch73-379f7.5 |
| chr20_+_22220988 | 0.25 |
ENSDART00000049204
|
kdr
|
kinase insert domain receptor (a type III receptor tyrosine kinase) |
| chr21_-_14692119 | 0.25 |
ENSDART00000123047
|
ehmt1b
|
euchromatic histone-lysine N-methyltransferase 1b |
| chr24_+_26402110 | 0.25 |
ENSDART00000133684
|
si:ch211-230g15.5
|
si:ch211-230g15.5 |
| chr23_+_21261313 | 0.23 |
ENSDART00000104268
ENSDART00000159046 |
emc1
|
ER membrane protein complex subunit 1 |
| chr16_-_24135508 | 0.22 |
ENSDART00000171819
ENSDART00000103176 |
bcam
|
basal cell adhesion molecule (Lutheran blood group) |
| chr5_+_37903790 | 0.22 |
ENSDART00000162470
|
tmprss4b
|
transmembrane protease, serine 4b |
| chr23_+_9522781 | 0.22 |
ENSDART00000136486
|
osbpl2b
|
oxysterol binding protein-like 2b |
| chr6_-_21988375 | 0.21 |
ENSDART00000161257
|
plxnb1b
|
plexin b1b |
| chr13_+_4871886 | 0.20 |
ENSDART00000132301
|
micu1
|
mitochondrial calcium uptake 1 |
| chr13_+_23157053 | 0.18 |
ENSDART00000162359
|
sorbs1
|
sorbin and SH3 domain containing 1 |
| chr10_-_33343244 | 0.18 |
ENSDART00000164191
|
c2cd2
|
C2 calcium-dependent domain containing 2 |
| chr2_-_36819624 | 0.16 |
ENSDART00000140844
|
slitrk3b
|
SLIT and NTRK-like family, member 3b |
| chr16_+_32014552 | 0.16 |
ENSDART00000047570
|
mboat7
|
membrane bound O-acyltransferase domain containing 7 |
| chr15_-_21837207 | 0.16 |
ENSDART00000089953
|
sik2b
|
salt-inducible kinase 2b |
| chr7_+_16806473 | 0.15 |
ENSDART00000113332
ENSDART00000173541 |
nav2a
|
neuron navigator 2a |
| chr17_+_27162367 | 0.15 |
ENSDART00000193345
|
rps6ka1
|
ribosomal protein S6 kinase a, polypeptide 1 |
| chr15_+_19293744 | 0.15 |
ENSDART00000184994
ENSDART00000123815 |
jam3a
|
junctional adhesion molecule 3a |
| chr18_+_27926839 | 0.15 |
ENSDART00000191835
|
hipk3b
|
homeodomain interacting protein kinase 3b |
| chr2_-_32826108 | 0.14 |
ENSDART00000098834
|
prpf4ba
|
pre-mRNA processing factor 4Ba |
| chr10_-_43721530 | 0.13 |
ENSDART00000025366
|
cetn3
|
centrin 3 |
| chr14_-_30897177 | 0.13 |
ENSDART00000087918
|
slc7a3b
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 3b |
| chr5_-_65081600 | 0.12 |
ENSDART00000160850
|
inpp5e
|
inositol polyphosphate-5-phosphatase E |
| chr5_-_57898008 | 0.12 |
ENSDART00000050945
|
layna
|
layilin a |
| chr8_-_25566347 | 0.11 |
ENSDART00000138289
ENSDART00000078022 |
prex1
|
phosphatidylinositol-3,4,5-trisphosphate-dependent Rac exchange factor 1 |
| chr9_-_34945566 | 0.10 |
ENSDART00000131908
ENSDART00000059861 |
dcun1d2a
|
DCN1, defective in cullin neddylation 1, domain containing 2a |
| chr19_-_4851411 | 0.10 |
ENSDART00000110398
|
fbxl20
|
F-box and leucine-rich repeat protein 20 |
| chr18_+_19883325 | 0.10 |
ENSDART00000182625
|
zgc:162898
|
zgc:162898 |
| chr23_+_9522942 | 0.09 |
ENSDART00000137751
|
osbpl2b
|
oxysterol binding protein-like 2b |
| chr15_+_20363859 | 0.09 |
ENSDART00000166846
|
supt5h
|
SPT5 homolog, DSIF elongation factor subunit |
| chr6_-_24384654 | 0.08 |
ENSDART00000164723
|
brdt
|
bromodomain, testis-specific |
| chr17_+_28675120 | 0.08 |
ENSDART00000159067
|
hectd1
|
HECT domain containing 1 |
| chr20_-_51727860 | 0.08 |
ENSDART00000147044
|
brox
|
BRO1 domain and CAAX motif containing |
| chr21_-_27272657 | 0.07 |
ENSDART00000040754
ENSDART00000175009 |
mark2a
|
MAP/microtubule affinity-regulating kinase 2a |
| chr17_+_12075805 | 0.07 |
ENSDART00000155329
|
cnsta
|
consortin, connexin sorting protein a |
| chr5_+_51833132 | 0.06 |
ENSDART00000167491
|
papd4
|
PAP associated domain containing 4 |
| chr20_+_51061695 | 0.05 |
ENSDART00000134416
|
im:7140055
|
im:7140055 |
| chr17_+_25871304 | 0.05 |
ENSDART00000185143
|
wapla
|
WAPL cohesin release factor a |
| chr23_-_2448234 | 0.05 |
ENSDART00000082097
|
LO018513.1
|
|
| chr6_-_21534301 | 0.05 |
ENSDART00000126186
|
psmd12
|
proteasome 26S subunit, non-ATPase 12 |
| chr7_+_51324834 | 0.04 |
ENSDART00000114429
|
usp12b
|
ubiquitin specific peptidase 12b |
| chr5_+_51833305 | 0.04 |
ENSDART00000165276
ENSDART00000166443 |
papd4
|
PAP associated domain containing 4 |
| chr7_-_17780048 | 0.01 |
ENSDART00000183336
|
si:dkey-106g10.7
|
si:dkey-106g10.7 |
| chr1_+_27690 | 0.01 |
ENSDART00000162928
|
eed
|
embryonic ectoderm development |
| chr2_-_37458527 | 0.01 |
ENSDART00000146820
|
si:dkey-57k2.7
|
si:dkey-57k2.7 |
| chr21_-_45878872 | 0.00 |
ENSDART00000029763
|
sap30l
|
sap30-like |
Network of associatons between targets according to the STRING database.
First level regulatory network of prdm14
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.2 | GO:0002792 | negative regulation of peptide secretion(GO:0002792) negative regulation of peptide hormone secretion(GO:0090278) |
| 0.3 | 1.3 | GO:0060080 | inhibitory postsynaptic potential(GO:0060080) |
| 0.2 | 0.7 | GO:0098921 | endocannabinoid signaling pathway(GO:0071926) retrograde trans-synaptic signaling by lipid(GO:0098920) retrograde trans-synaptic signaling by endocannabinoid(GO:0098921) |
| 0.1 | 1.2 | GO:0031998 | regulation of fatty acid beta-oxidation(GO:0031998) |
| 0.1 | 1.0 | GO:0055011 | atrial cardiac muscle cell differentiation(GO:0055011) |
| 0.1 | 0.5 | GO:0045475 | locomotor rhythm(GO:0045475) |
| 0.1 | 0.3 | GO:0016572 | histone phosphorylation(GO:0016572) |
| 0.0 | 0.6 | GO:0050962 | detection of light stimulus involved in visual perception(GO:0050908) detection of light stimulus involved in sensory perception(GO:0050962) |
| 0.0 | 1.2 | GO:0035622 | intrahepatic bile duct development(GO:0035622) |
| 0.0 | 0.2 | GO:0036149 | phosphatidylinositol acyl-chain remodeling(GO:0036149) |
| 0.0 | 0.2 | GO:0051561 | positive regulation of mitochondrial calcium ion concentration(GO:0051561) |
| 0.0 | 0.7 | GO:0051932 | synaptic transmission, GABAergic(GO:0051932) |
| 0.0 | 0.4 | GO:0001881 | receptor recycling(GO:0001881) |
| 0.0 | 0.6 | GO:0015701 | bicarbonate transport(GO:0015701) |
| 0.0 | 0.1 | GO:0051039 | histone displacement(GO:0001207) regulation of transcription involved in meiotic cell cycle(GO:0051037) positive regulation of transcription involved in meiotic cell cycle(GO:0051039) |
| 0.0 | 0.4 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 0.0 | 0.1 | GO:0090004 | positive regulation of Golgi to plasma membrane protein transport(GO:0042998) positive regulation of establishment of protein localization to plasma membrane(GO:0090004) |
| 0.0 | 0.4 | GO:0032958 | inositol phosphate biosynthetic process(GO:0032958) |
| 0.0 | 0.1 | GO:1902765 | L-arginine import(GO:0043091) amino acid import across plasma membrane(GO:0089718) arginine import(GO:0090467) L-arginine import across plasma membrane(GO:0097638) L-arginine transport(GO:1902023) L-arginine import into cell(GO:1902765) amino acid import into cell(GO:1902837) L-arginine transmembrane transport(GO:1903400) arginine transmembrane transport(GO:1903826) |
| 0.0 | 1.3 | GO:0003146 | heart jogging(GO:0003146) |
| 0.0 | 0.1 | GO:0060623 | regulation of chromosome condensation(GO:0060623) |
| 0.0 | 0.1 | GO:0001961 | positive regulation of cytokine-mediated signaling pathway(GO:0001961) positive regulation of response to cytokine stimulus(GO:0060760) |
| 0.0 | 0.2 | GO:1902285 | semaphorin-plexin signaling pathway involved in neuron projection guidance(GO:1902285) semaphorin-plexin signaling pathway involved in axon guidance(GO:1902287) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.2 | GO:0042583 | chromaffin granule(GO:0042583) |
| 0.1 | 1.2 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.0 | 0.2 | GO:0044233 | ER-mitochondrion membrane contact site(GO:0044233) |
| 0.0 | 0.6 | GO:0032420 | stereocilium(GO:0032420) |
| 0.0 | 0.1 | GO:0032044 | DSIF complex(GO:0032044) |
| 0.0 | 0.7 | GO:0032589 | neuron projection membrane(GO:0032589) dendrite membrane(GO:0032590) |
| 0.0 | 0.3 | GO:0097431 | mitotic spindle pole(GO:0097431) |
| 0.0 | 0.2 | GO:0072546 | ER membrane protein complex(GO:0072546) |
| 0.0 | 0.9 | GO:0098839 | postsynaptic density membrane(GO:0098839) postsynaptic specialization membrane(GO:0099634) |
| 0.0 | 0.2 | GO:1990246 | uniplex complex(GO:1990246) |
| 0.0 | 1.2 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.0 | 0.2 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.0 | 0.2 | GO:0002116 | semaphorin receptor complex(GO:0002116) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.2 | GO:0004095 | carnitine O-palmitoyltransferase activity(GO:0004095) O-palmitoyltransferase activity(GO:0016416) |
| 0.1 | 0.6 | GO:0046592 | polyamine oxidase activity(GO:0046592) |
| 0.1 | 3.3 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.1 | 0.7 | GO:0004465 | lipoprotein lipase activity(GO:0004465) |
| 0.1 | 0.8 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.1 | 0.7 | GO:0008503 | benzodiazepine receptor activity(GO:0008503) GABA-gated chloride ion channel activity(GO:0022851) |
| 0.0 | 0.6 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.0 | 0.9 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.0 | 0.4 | GO:0003873 | 6-phosphofructo-2-kinase activity(GO:0003873) |
| 0.0 | 0.4 | GO:0000828 | inositol hexakisphosphate kinase activity(GO:0000828) |
| 0.0 | 0.2 | GO:0005021 | vascular endothelial growth factor-activated receptor activity(GO:0005021) |
| 0.0 | 0.1 | GO:0071253 | connexin binding(GO:0071253) |
| 0.0 | 0.1 | GO:0015189 | L-lysine transmembrane transporter activity(GO:0015189) |
| 0.0 | 0.6 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.0 | 0.2 | GO:0002039 | p53 binding(GO:0002039) |
| 0.0 | 0.1 | GO:0004711 | ribosomal protein S6 kinase activity(GO:0004711) |
| 0.0 | 0.3 | GO:0015248 | sterol transporter activity(GO:0015248) |
| 0.0 | 0.1 | GO:0004445 | inositol-polyphosphate 5-phosphatase activity(GO:0004445) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.3 | ST WNT CA2 CYCLIC GMP PATHWAY | Wnt/Ca2+/cyclic GMP signaling. |
| 0.1 | 1.2 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.0 | 0.2 | PID VEGF VEGFR PATHWAY | VEGF and VEGFR signaling network |
| 0.0 | 0.7 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.0 | 0.3 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.0 | 0.3 | SIG INSULIN RECEPTOR PATHWAY IN CARDIAC MYOCYTES | Genes related to the insulin receptor pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.3 | REACTOME UNBLOCKING OF NMDA RECEPTOR GLUTAMATE BINDING AND ACTIVATION | Genes involved in Unblocking of NMDA receptor, glutamate binding and activation |
| 0.0 | 0.7 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.0 | 0.9 | REACTOME PLATELET CALCIUM HOMEOSTASIS | Genes involved in Platelet calcium homeostasis |
| 0.0 | 0.6 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.0 | 0.2 | REACTOME ACYL CHAIN REMODELLING OF PI | Genes involved in Acyl chain remodelling of PI |
| 0.0 | 0.2 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.0 | 0.7 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.0 | 0.5 | REACTOME RECYCLING PATHWAY OF L1 | Genes involved in Recycling pathway of L1 |