Project
PRJNA207719: Tissue specific transcriptome profiling
Navigation
Downloads
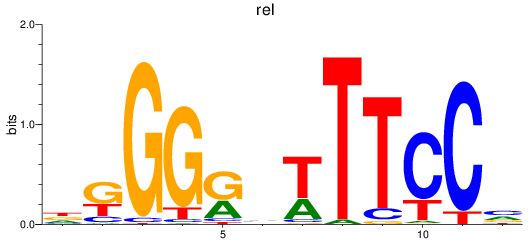
Results for rel
Z-value: 1.13
Transcription factors associated with rel
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
rel
|
ENSDARG00000055276 | v-rel avian reticuloendotheliosis viral oncogene homolog |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| rel | dr11_v1_chr13_-_25819825_25819825 | -0.06 | 9.2e-01 | Click! |
Activity profile of rel motif
Sorted Z-values of rel motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr6_-_54815886 | 1.20 |
ENSDART00000180793
ENSDART00000007498 |
tnni1b
|
troponin I type 1b (skeletal, slow) |
| chr1_-_49225890 | 1.13 |
ENSDART00000111598
|
cxcl18b
|
chemokine (C-X-C motif) ligand 18b |
| chr17_-_6382392 | 0.97 |
ENSDART00000188051
ENSDART00000192560 ENSDART00000137389 ENSDART00000115389 |
txlnbb
|
taxilin beta b |
| chr7_+_12950507 | 0.96 |
ENSDART00000067629
ENSDART00000158004 |
saa
|
serum amyloid A |
| chr22_+_5478353 | 0.95 |
ENSDART00000160596
|
tppp
|
tubulin polymerization promoting protein |
| chr1_-_52447364 | 0.91 |
ENSDART00000140740
|
si:ch211-217k17.10
|
si:ch211-217k17.10 |
| chr22_-_651719 | 0.89 |
ENSDART00000148692
|
apobec2a
|
apolipoprotein B mRNA editing enzyme, catalytic polypeptide-like 2a |
| chr6_+_3334710 | 0.87 |
ENSDART00000132848
|
st3gal3a
|
ST3 beta-galactoside alpha-2,3-sialyltransferase 3a |
| chr7_+_45975537 | 0.86 |
ENSDART00000170253
|
plekhf1
|
pleckstrin homology domain containing, family F (with FYVE domain) member 1 |
| chr6_-_436658 | 0.85 |
ENSDART00000191515
|
grap2b
|
GRB2-related adaptor protein 2b |
| chr2_-_51772438 | 0.85 |
ENSDART00000170241
|
BX908782.2
|
Danio rerio three-finger protein 5 (LOC100003647), mRNA. |
| chr22_+_661711 | 0.74 |
ENSDART00000113795
|
elf3
|
E74-like factor 3 (ets domain transcription factor, epithelial-specific ) |
| chr4_-_12795436 | 0.74 |
ENSDART00000131026
ENSDART00000075127 |
b2m
|
beta-2-microglobulin |
| chr13_+_41819817 | 0.73 |
ENSDART00000185778
|
CABZ01066611.1
|
|
| chr18_-_5527050 | 0.73 |
ENSDART00000145400
ENSDART00000132498 ENSDART00000146209 |
zgc:153317
|
zgc:153317 |
| chr14_+_36889893 | 0.72 |
ENSDART00000124159
|
si:ch211-132p1.3
|
si:ch211-132p1.3 |
| chr2_+_45191049 | 0.70 |
ENSDART00000165392
|
ccl20a.3
|
chemokine (C-C motif) ligand 20a, duplicate 3 |
| chr7_+_39706004 | 0.69 |
ENSDART00000161856
|
ccl36.1
|
chemokine (C-C motif) ligand 36, duplicate 1 |
| chr21_-_42876565 | 0.66 |
ENSDART00000126480
|
zmp:0000001268
|
zmp:0000001268 |
| chr25_-_13188214 | 0.66 |
ENSDART00000187298
|
si:ch211-147m6.1
|
si:ch211-147m6.1 |
| chr11_-_28050559 | 0.65 |
ENSDART00000136859
|
ece1
|
endothelin converting enzyme 1 |
| chr20_+_16881883 | 0.64 |
ENSDART00000130107
|
nfkbiaa
|
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha a |
| chr12_+_27462225 | 0.62 |
ENSDART00000105661
|
meox1
|
mesenchyme homeobox 1 |
| chr20_-_33976937 | 0.62 |
ENSDART00000136834
|
sele
|
selectin E |
| chr24_+_26006730 | 0.62 |
ENSDART00000140384
ENSDART00000139184 |
ccl20b
|
chemokine (C-C motif) ligand 20b |
| chr22_+_6293563 | 0.62 |
ENSDART00000063416
|
rnasel2
|
ribonuclease like 2 |
| chr9_-_98982 | 0.61 |
ENSDART00000147882
|
lims2
|
LIM and senescent cell antigen-like domains 2 |
| chr8_+_24281512 | 0.60 |
ENSDART00000062845
|
mmp9
|
matrix metallopeptidase 9 |
| chr6_+_3334392 | 0.58 |
ENSDART00000133707
ENSDART00000130879 |
st3gal3a
|
ST3 beta-galactoside alpha-2,3-sialyltransferase 3a |
| chr1_-_37377509 | 0.57 |
ENSDART00000113542
|
tnip2
|
TNFAIP3 interacting protein 2 |
| chr20_+_33981946 | 0.57 |
ENSDART00000131775
|
si:dkey-51e6.1
|
si:dkey-51e6.1 |
| chr13_-_36034582 | 0.55 |
ENSDART00000133565
|
si:dkey-157l19.2
|
si:dkey-157l19.2 |
| chr4_-_12790886 | 0.55 |
ENSDART00000182535
ENSDART00000067131 ENSDART00000186426 |
irak3
|
interleukin-1 receptor-associated kinase 3 |
| chr16_+_23403602 | 0.54 |
ENSDART00000159848
|
s100w
|
S100 calcium binding protein W |
| chr7_-_8309505 | 0.54 |
ENSDART00000182530
|
f13a1b
|
coagulation factor XIII, A1 polypeptide b |
| chr16_+_38394371 | 0.53 |
ENSDART00000137954
|
cd83
|
CD83 molecule |
| chr8_-_18667693 | 0.52 |
ENSDART00000100516
|
stap2b
|
signal transducing adaptor family member 2b |
| chr18_-_11729 | 0.51 |
ENSDART00000159781
|
WHAMM
|
WAS protein homolog associated with actin, golgi membranes and microtubules |
| chr2_-_45191319 | 0.51 |
ENSDART00000192272
|
CR407590.2
|
|
| chr6_-_442163 | 0.51 |
ENSDART00000163564
ENSDART00000189134 ENSDART00000169789 |
grap2b
|
GRB2-related adaptor protein 2b |
| chr14_+_36885524 | 0.50 |
ENSDART00000032547
|
lect2l
|
leukocyte cell-derived chemotaxin 2 like |
| chr2_+_42191592 | 0.50 |
ENSDART00000144716
|
cavin4a
|
caveolae associated protein 4a |
| chr19_+_48351624 | 0.50 |
ENSDART00000169729
|
sgo1
|
shugoshin 1 |
| chr12_-_26430507 | 0.50 |
ENSDART00000153214
|
synpo2lb
|
synaptopodin 2-like b |
| chr9_-_23894392 | 0.49 |
ENSDART00000133417
|
asb18
|
ankyrin repeat and SOCS box containing 18 |
| chr2_+_16780643 | 0.49 |
ENSDART00000125647
ENSDART00000108611 ENSDART00000181245 ENSDART00000163194 |
tfa
|
transferrin-a |
| chr13_-_22843562 | 0.49 |
ENSDART00000142738
|
pbld
|
phenazine biosynthesis like protein domain containing |
| chr24_-_33703504 | 0.47 |
ENSDART00000079292
|
cavin4b
|
caveolae associated protein 4b |
| chr3_-_21242460 | 0.47 |
ENSDART00000007293
|
tcap
|
titin-cap (telethonin) |
| chr1_+_12135129 | 0.46 |
ENSDART00000126020
|
spink4
|
serine peptidase inhibitor, Kazal type 4 |
| chr7_+_34786591 | 0.46 |
ENSDART00000173700
|
si:dkey-148a17.5
|
si:dkey-148a17.5 |
| chr19_+_7717449 | 0.46 |
ENSDART00000104719
ENSDART00000146747 |
tuft1b
|
tuftelin 1b |
| chr23_+_19701587 | 0.45 |
ENSDART00000104425
|
dnase1l1
|
deoxyribonuclease I-like 1 |
| chr19_+_23298037 | 0.45 |
ENSDART00000183948
|
irgf1
|
immunity-related GTPase family, f1 |
| chr20_+_47491247 | 0.45 |
ENSDART00000113412
|
lin28b
|
lin-28 homolog B (C. elegans) |
| chr7_-_20241346 | 0.45 |
ENSDART00000173619
ENSDART00000127699 |
si:ch73-335l21.4
|
si:ch73-335l21.4 |
| chr14_-_40821411 | 0.44 |
ENSDART00000166621
|
elf1
|
E74-like ETS transcription factor 1 |
| chr1_+_18716 | 0.43 |
ENSDART00000172454
ENSDART00000161190 |
nfkbiz
|
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, zeta |
| chr20_+_23501535 | 0.43 |
ENSDART00000177922
ENSDART00000058532 |
palld
|
palladin, cytoskeletal associated protein |
| chr13_+_669177 | 0.43 |
ENSDART00000190085
|
CU462915.1
|
|
| chr7_+_4384863 | 0.42 |
ENSDART00000042955
ENSDART00000134653 |
slc12a10.3
|
slc12a10.3 solute carrier family 12 (sodium/potassium/chloride transporters), member 10, tandem duplicate 3 |
| chr13_-_2010191 | 0.42 |
ENSDART00000161021
ENSDART00000124134 |
gfral
|
GDNF family receptor alpha like |
| chr18_+_8937718 | 0.42 |
ENSDART00000138115
|
si:dkey-5i3.1
|
si:dkey-5i3.1 |
| chr14_-_6987649 | 0.42 |
ENSDART00000060990
|
eif4ebp3l
|
eukaryotic translation initiation factor 4E binding protein 3, like |
| chr12_-_11349899 | 0.42 |
ENSDART00000079645
|
zgc:174164
|
zgc:174164 |
| chr14_+_20893065 | 0.42 |
ENSDART00000079452
|
lygl1
|
lysozyme g-like 1 |
| chr5_-_42878178 | 0.41 |
ENSDART00000162981
|
CXCL11 (1 of many)
|
C-X-C motif chemokine ligand 11 |
| chr6_+_28428329 | 0.41 |
ENSDART00000188056
|
lpp
|
LIM domain containing preferred translocation partner in lipoma |
| chr7_+_21275152 | 0.41 |
ENSDART00000173612
|
serpinh2
|
serine (or cysteine) peptidase inhibitor, clade H, member 2 |
| chr15_+_6459847 | 0.41 |
ENSDART00000157250
ENSDART00000065824 |
bace2
|
beta-site APP-cleaving enzyme 2 |
| chr18_-_49286381 | 0.40 |
ENSDART00000174248
ENSDART00000174038 |
si:zfos-464b6.2
|
si:zfos-464b6.2 |
| chr2_+_42344304 | 0.40 |
ENSDART00000136105
|
ftr10
|
finTRIM family, member 10 |
| chr25_-_27843066 | 0.40 |
ENSDART00000179684
ENSDART00000186000 ENSDART00000190065 |
asb15a
|
ankyrin repeat and SOCS box containing 15a |
| chr8_-_12432604 | 0.40 |
ENSDART00000133350
ENSDART00000140699 ENSDART00000101174 |
traf1
|
TNF receptor-associated factor 1 |
| chr6_+_19933763 | 0.39 |
ENSDART00000166192
|
pik3r5
|
phosphoinositide-3-kinase, regulatory subunit 5 |
| chr18_-_18543358 | 0.39 |
ENSDART00000126460
|
il34
|
interleukin 34 |
| chr13_-_39947335 | 0.39 |
ENSDART00000056996
|
sfrp5
|
secreted frizzled-related protein 5 |
| chr16_-_41762983 | 0.38 |
ENSDART00000192936
|
si:dkey-199f5.8
|
si:dkey-199f5.8 |
| chr3_+_19216567 | 0.38 |
ENSDART00000134433
|
il12rb2l
|
interleukin 12 receptor, beta 2a, like |
| chr16_-_25739331 | 0.38 |
ENSDART00000189455
|
bcl3
|
B cell CLL/lymphoma 3 |
| chr3_+_34670076 | 0.38 |
ENSDART00000133457
|
dlx4a
|
distal-less homeobox 4a |
| chr20_+_33991801 | 0.37 |
ENSDART00000061744
|
zp3a.1
|
zona pellucida glycoprotein 3a, tandem duplicate 1 |
| chr8_-_25771474 | 0.37 |
ENSDART00000193883
|
suv39h1b
|
suppressor of variegation 3-9 homolog 1b |
| chr21_+_43328685 | 0.37 |
ENSDART00000109620
ENSDART00000139668 |
sept8a
|
septin 8a |
| chr5_+_13373593 | 0.37 |
ENSDART00000051668
ENSDART00000183883 |
ccl19a.2
|
chemokine (C-C motif) ligand 19a, tandem duplicate 2 |
| chr4_-_33525219 | 0.36 |
ENSDART00000150491
|
CT027801.2
|
|
| chr16_-_24819860 | 0.36 |
ENSDART00000183874
|
kcnn4
|
potassium intermediate/small conductance calcium-activated channel, subfamily N, member 4 |
| chr16_-_29714540 | 0.36 |
ENSDART00000067854
|
tnfaip8l2b
|
tumor necrosis factor, alpha-induced protein 8-like 2b |
| chr22_-_23067859 | 0.36 |
ENSDART00000137873
|
ptprc
|
protein tyrosine phosphatase, receptor type, C |
| chr12_-_30760971 | 0.36 |
ENSDART00000066257
|
entpd1
|
ectonucleoside triphosphate diphosphohydrolase 1 |
| chr21_-_22357545 | 0.35 |
ENSDART00000134320
|
skp2
|
S-phase kinase-associated protein 2, E3 ubiquitin protein ligase |
| chr14_+_3038473 | 0.35 |
ENSDART00000026021
ENSDART00000150000 |
cd74a
|
CD74 molecule, major histocompatibility complex, class II invariant chain a |
| chr1_+_18550864 | 0.35 |
ENSDART00000142515
|
si:dkey-192k22.2
|
si:dkey-192k22.2 |
| chr7_+_4538334 | 0.35 |
ENSDART00000139326
ENSDART00000146915 |
si:dkey-83f18.9
|
si:dkey-83f18.9 |
| chr25_-_28926330 | 0.35 |
ENSDART00000155173
|
etfbkmt
|
electron transfer flavoprotein beta subunit lysine methyltransferase |
| chr5_+_25596234 | 0.34 |
ENSDART00000143949
ENSDART00000167193 |
si:dkey-96l17.6
|
si:dkey-96l17.6 |
| chr20_-_51307815 | 0.33 |
ENSDART00000098833
|
nfkbie
|
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, epsilon |
| chr9_-_5263947 | 0.33 |
ENSDART00000088342
|
cytip
|
cytohesin 1 interacting protein |
| chr8_+_18101608 | 0.33 |
ENSDART00000140941
|
glis1b
|
GLIS family zinc finger 1b |
| chr11_+_14333441 | 0.33 |
ENSDART00000171969
|
ptbp1b
|
polypyrimidine tract binding protein 1b |
| chr12_-_4206869 | 0.33 |
ENSDART00000106572
|
si:dkey-32n7.9
|
si:dkey-32n7.9 |
| chr19_+_28187480 | 0.32 |
ENSDART00000183825
|
irx4b
|
iroquois homeobox 4b |
| chr5_-_30984271 | 0.32 |
ENSDART00000051392
|
spns3
|
spinster homolog 3 (Drosophila) |
| chr19_+_7043634 | 0.32 |
ENSDART00000133954
|
mhc1uka
|
major histocompatibility complex class I UKA |
| chr14_+_30773764 | 0.32 |
ENSDART00000186961
|
atl3
|
atlastin 3 |
| chr18_-_17077596 | 0.32 |
ENSDART00000111821
|
il17c
|
interleukin 17c |
| chr8_-_46572298 | 0.32 |
ENSDART00000030470
|
sult1st3
|
sulfotransferase family 1, cytosolic sulfotransferase 3 |
| chr17_-_10315644 | 0.32 |
ENSDART00000168572
|
ttc6
|
tetratricopeptide repeat domain 6 |
| chr13_+_22717366 | 0.32 |
ENSDART00000134122
|
nfkb2
|
nuclear factor of kappa light polypeptide gene enhancer in B-cells 2 (p49/p100) |
| chr13_+_22717939 | 0.31 |
ENSDART00000188288
|
nfkb2
|
nuclear factor of kappa light polypeptide gene enhancer in B-cells 2 (p49/p100) |
| chr20_-_39367895 | 0.31 |
ENSDART00000136476
ENSDART00000021788 ENSDART00000180784 |
pbk
|
PDZ binding kinase |
| chr11_+_35050253 | 0.31 |
ENSDART00000124800
|
fam212aa
|
family with sequence similarity 212, member Aa |
| chr6_-_52675630 | 0.31 |
ENSDART00000083830
|
sdc4
|
syndecan 4 |
| chr20_-_51917771 | 0.31 |
ENSDART00000147240
|
dusp10
|
dual specificity phosphatase 10 |
| chr4_-_18954001 | 0.30 |
ENSDART00000144814
|
si:dkey-31f5.8
|
si:dkey-31f5.8 |
| chr8_+_22955478 | 0.30 |
ENSDART00000099911
|
zgc:136605
|
zgc:136605 |
| chr2_+_2737422 | 0.30 |
ENSDART00000032459
|
aqp1a.1
|
aquaporin 1a (Colton blood group), tandem duplicate 1 |
| chr21_+_35289469 | 0.30 |
ENSDART00000155140
|
il12bb
|
interleukin 12B, b |
| chr14_-_40850481 | 0.30 |
ENSDART00000173236
|
elf1
|
E74-like ETS transcription factor 1 |
| chr18_-_17077419 | 0.30 |
ENSDART00000148714
|
il17c
|
interleukin 17c |
| chr11_+_8129536 | 0.30 |
ENSDART00000158112
ENSDART00000011183 |
prkacba
|
protein kinase, cAMP-dependent, catalytic, beta a |
| chr3_-_32590164 | 0.29 |
ENSDART00000151151
|
tspan4b
|
tetraspanin 4b |
| chr16_+_27444098 | 0.29 |
ENSDART00000157690
|
invs
|
inversin |
| chr23_-_33738570 | 0.29 |
ENSDART00000131680
|
si:ch211-210c8.7
|
si:ch211-210c8.7 |
| chr4_-_11081961 | 0.28 |
ENSDART00000150307
|
si:dkey-21h14.3
|
si:dkey-21h14.3 |
| chr20_-_54014539 | 0.28 |
ENSDART00000060466
|
si:dkey-241l7.6
|
si:dkey-241l7.6 |
| chr7_+_29293452 | 0.28 |
ENSDART00000127358
|
si:ch211-112g6.4
|
si:ch211-112g6.4 |
| chr1_+_17892944 | 0.28 |
ENSDART00000013021
|
tlr3
|
toll-like receptor 3 |
| chr2_-_29485408 | 0.28 |
ENSDART00000013411
|
cahz
|
carbonic anhydrase |
| chr7_+_4523051 | 0.28 |
ENSDART00000140860
ENSDART00000143778 |
si:dkey-83f18.10
|
si:dkey-83f18.10 |
| chr4_+_18806251 | 0.27 |
ENSDART00000138662
|
slc26a3.2
|
solute carrier family 26 (anion exchanger), member 3, tandem duplicate 2 |
| chr11_-_40101246 | 0.27 |
ENSDART00000161083
|
tnfrsf9b
|
tumor necrosis factor receptor superfamily, member 9b |
| chr14_+_6182346 | 0.27 |
ENSDART00000075525
|
si:ch73-22a13.3
|
si:ch73-22a13.3 |
| chr14_-_33218418 | 0.27 |
ENSDART00000163046
|
si:dkey-31j3.11
|
si:dkey-31j3.11 |
| chr22_+_25733817 | 0.26 |
ENSDART00000176962
ENSDART00000187865 |
si:dkeyp-98a7.9
|
si:dkeyp-98a7.9 |
| chr8_+_39674707 | 0.26 |
ENSDART00000126301
ENSDART00000040330 |
prkab1b
|
protein kinase, AMP-activated, beta 1 non-catalytic subunit, b |
| chr4_-_22311610 | 0.26 |
ENSDART00000137814
|
hcls1
|
hematopoietic cell-specific Lyn substrate 1 |
| chr25_+_8955530 | 0.26 |
ENSDART00000156444
|
si:ch211-256a21.4
|
si:ch211-256a21.4 |
| chr21_+_21817975 | 0.25 |
ENSDART00000151292
ENSDART00000166704 |
neu3.5
|
sialidase 3 (membrane sialidase), tandem duplicate 5 |
| chr7_-_3792972 | 0.25 |
ENSDART00000146304
|
si:dkey-28d5.12
|
si:dkey-28d5.12 |
| chr3_-_31619463 | 0.25 |
ENSDART00000124559
|
moto
|
minamoto |
| chr23_+_3591690 | 0.25 |
ENSDART00000180822
|
si:dkey-9l20.3
|
si:dkey-9l20.3 |
| chr8_+_22355909 | 0.24 |
ENSDART00000146457
ENSDART00000142883 |
zgc:153631
|
zgc:153631 |
| chr1_+_57004178 | 0.24 |
ENSDART00000152690
|
si:ch211-1f22.13
|
si:ch211-1f22.13 |
| chr8_-_26937225 | 0.24 |
ENSDART00000191787
|
slc16a1b
|
solute carrier family 16 (monocarboxylate transporter), member 1b |
| chr24_-_18477672 | 0.24 |
ENSDART00000126928
ENSDART00000158393 |
si:dkey-73n8.3
|
si:dkey-73n8.3 |
| chr23_+_3620618 | 0.24 |
ENSDART00000190126
|
si:dkey-9l20.3
|
si:dkey-9l20.3 |
| chr25_-_28926631 | 0.24 |
ENSDART00000112850
|
etfbkmt
|
electron transfer flavoprotein beta subunit lysine methyltransferase |
| chr22_+_26263290 | 0.23 |
ENSDART00000184840
|
BX649315.1
|
|
| chr23_+_3618454 | 0.23 |
ENSDART00000189393
|
si:dkey-9l20.3
|
si:dkey-9l20.3 |
| chr14_+_14806692 | 0.23 |
ENSDART00000193050
|
fhdc2
|
FH2 domain containing 2 |
| chr3_+_54744069 | 0.23 |
ENSDART00000134958
ENSDART00000114443 |
si:ch211-74m13.3
|
si:ch211-74m13.3 |
| chr25_-_35960229 | 0.23 |
ENSDART00000073434
|
snx20
|
sorting nexin 20 |
| chr18_+_30847237 | 0.22 |
ENSDART00000012374
|
foxf1
|
forkhead box F1 |
| chr20_-_20410029 | 0.22 |
ENSDART00000192177
ENSDART00000063483 |
prkchb
|
protein kinase C, eta, b |
| chr14_-_1646360 | 0.22 |
ENSDART00000186528
|
LO018106.2
|
|
| chr5_-_38643188 | 0.22 |
ENSDART00000144972
|
si:ch211-271e10.1
|
si:ch211-271e10.1 |
| chr5_+_27525477 | 0.22 |
ENSDART00000051491
|
sfrp1a
|
secreted frizzled-related protein 1a |
| chr16_-_55259199 | 0.22 |
ENSDART00000161130
|
iqgap3
|
IQ motif containing GTPase activating protein 3 |
| chr14_+_14806851 | 0.22 |
ENSDART00000169235
|
fhdc2
|
FH2 domain containing 2 |
| chr23_+_3594171 | 0.22 |
ENSDART00000159609
|
si:dkey-9l20.3
|
si:dkey-9l20.3 |
| chr2_-_14798295 | 0.22 |
ENSDART00000143430
ENSDART00000145869 |
si:ch73-366i20.1
|
si:ch73-366i20.1 |
| chr2_-_53592532 | 0.21 |
ENSDART00000184066
|
ccl25a
|
chemokine (C-C motif) ligand 25a |
| chr3_-_58733718 | 0.21 |
ENSDART00000154603
|
si:ch73-281f12.4
|
si:ch73-281f12.4 |
| chr13_-_9335891 | 0.21 |
ENSDART00000080637
|
BX901922.1
|
|
| chr23_+_3596401 | 0.21 |
ENSDART00000185908
|
si:dkey-9l20.3
|
si:dkey-9l20.3 |
| chr19_-_10432134 | 0.20 |
ENSDART00000081440
|
il11b
|
interleukin 11b |
| chr22_+_18156000 | 0.20 |
ENSDART00000143483
ENSDART00000136133 |
nr2c2ap
|
nuclear receptor 2C2-associated protein |
| chr3_-_54544612 | 0.20 |
ENSDART00000018044
|
angptl6
|
angiopoietin-like 6 |
| chr10_+_2799285 | 0.20 |
ENSDART00000030709
|
pnx
|
posterior neuron-specific homeobox |
| chr23_-_38054 | 0.20 |
ENSDART00000170393
|
CABZ01074076.1
|
|
| chr24_-_14711597 | 0.19 |
ENSDART00000131830
|
jph1a
|
junctophilin 1a |
| chr8_-_48704050 | 0.19 |
ENSDART00000163916
|
pimr182
|
Pim proto-oncogene, serine/threonine kinase, related 182 |
| chr7_-_41915312 | 0.19 |
ENSDART00000159869
|
dnaja2
|
DnaJ (Hsp40) homolog, subfamily A, member 2 |
| chr10_+_20630444 | 0.19 |
ENSDART00000191182
|
prnpb
|
prion protein b |
| chr13_-_25819825 | 0.19 |
ENSDART00000077612
|
rel
|
v-rel avian reticuloendotheliosis viral oncogene homolog |
| chr18_-_16922905 | 0.18 |
ENSDART00000187165
|
wee1
|
WEE1 G2 checkpoint kinase |
| chr5_+_37729207 | 0.18 |
ENSDART00000184378
|
cdc42ep2
|
CDC42 effector protein (Rho GTPase binding) 2 |
| chr23_-_20361971 | 0.18 |
ENSDART00000133373
|
si:rp71-17i16.5
|
si:rp71-17i16.5 |
| chr17_-_39761086 | 0.18 |
ENSDART00000032410
|
gpr132a
|
G protein-coupled receptor 132a |
| chr18_-_16924221 | 0.18 |
ENSDART00000122102
|
wee1
|
WEE1 G2 checkpoint kinase |
| chr25_-_7723685 | 0.18 |
ENSDART00000146882
|
phf21ab
|
PHD finger protein 21Ab |
| chr22_-_4439311 | 0.18 |
ENSDART00000169317
|
uhrf1
|
ubiquitin-like with PHD and ring finger domains 1 |
| chr22_-_8499778 | 0.17 |
ENSDART00000142219
|
si:ch73-27e22.4
|
si:ch73-27e22.4 |
| chr4_+_36489448 | 0.17 |
ENSDART00000143181
|
znf1149
|
zinc finger protein 1149 |
| chr13_+_21601149 | 0.17 |
ENSDART00000179369
|
sh2d4ba
|
SH2 domain containing 4Ba |
| chr4_-_76488581 | 0.17 |
ENSDART00000174291
|
ftr51
|
finTRIM family, member 51 |
| chr20_-_3290791 | 0.17 |
ENSDART00000146870
|
si:ch1073-412h12.3
|
si:ch1073-412h12.3 |
| chr4_+_69888328 | 0.16 |
ENSDART00000170163
|
si:ch211-145h19.5
|
si:ch211-145h19.5 |
| chr14_-_11529311 | 0.16 |
ENSDART00000127208
|
si:ch211-153b23.7
|
si:ch211-153b23.7 |
| chr16_+_52999778 | 0.16 |
ENSDART00000011506
|
nkd2a
|
naked cuticle homolog 2a |
| chr8_-_13010011 | 0.15 |
ENSDART00000140969
|
dennd2da
|
DENN/MADD domain containing 2Da |
| chr13_+_21600946 | 0.15 |
ENSDART00000144045
|
sh2d4ba
|
SH2 domain containing 4Ba |
| chr1_-_8553165 | 0.15 |
ENSDART00000135197
ENSDART00000054981 |
zgc:112980
|
zgc:112980 |
| chr23_-_27589754 | 0.15 |
ENSDART00000138381
ENSDART00000133721 |
si:ch211-156j22.4
|
si:ch211-156j22.4 |
| chr4_+_57580303 | 0.15 |
ENSDART00000166492
ENSDART00000103025 ENSDART00000170786 |
il17ra1b
il17ra2
|
interleukin 17 receptor A1b interleukin 17 receptor A2 |
| chr2_+_36939749 | 0.14 |
ENSDART00000005382
|
gadd45bb
|
growth arrest and DNA-damage-inducible, beta b |
| chr4_+_75575252 | 0.14 |
ENSDART00000166536
|
si:ch211-227e10.1
|
si:ch211-227e10.1 |
| chr4_-_71708855 | 0.14 |
ENSDART00000158819
|
si:dkeyp-4f2.1
|
si:dkeyp-4f2.1 |
| chr10_+_22536693 | 0.14 |
ENSDART00000185037
ENSDART00000146040 |
efnb3a
|
ephrin-B3a |
Network of associatons between targets according to the STRING database.
First level regulatory network of rel
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.9 | GO:0043388 | cytidine to uridine editing(GO:0016554) positive regulation of DNA binding(GO:0043388) |
| 0.3 | 0.9 | GO:0002468 | dendritic cell antigen processing and presentation(GO:0002468) |
| 0.3 | 0.9 | GO:0034138 | toll-like receptor 3 signaling pathway(GO:0034138) |
| 0.2 | 0.6 | GO:1901533 | negative regulation of hematopoietic progenitor cell differentiation(GO:1901533) |
| 0.2 | 0.6 | GO:0002432 | granuloma formation(GO:0002432) chronic inflammatory response(GO:0002544) regulation of granuloma formation(GO:0002631) regulation of chronic inflammatory response(GO:0002676) |
| 0.2 | 0.6 | GO:0050995 | negative regulation of lipid catabolic process(GO:0050995) |
| 0.2 | 0.6 | GO:0035971 | peptidyl-histidine dephosphorylation(GO:0035971) |
| 0.2 | 0.5 | GO:0032640 | tumor necrosis factor production(GO:0032640) tumor necrosis factor superfamily cytokine production(GO:0071706) |
| 0.2 | 1.0 | GO:0044364 | killing of cells of other organism(GO:0031640) disruption of cells of other organism(GO:0044364) disruption of cells of other organism involved in symbiotic interaction(GO:0051818) killing of cells in other organism involved in symbiotic interaction(GO:0051883) |
| 0.1 | 0.6 | GO:0015670 | carbon dioxide transport(GO:0015670) |
| 0.1 | 0.4 | GO:0016998 | cell wall macromolecule catabolic process(GO:0016998) |
| 0.1 | 0.4 | GO:0045649 | regulation of macrophage differentiation(GO:0045649) |
| 0.1 | 0.4 | GO:0044333 | Wnt signaling pathway involved in digestive tract morphogenesis(GO:0044333) |
| 0.1 | 0.5 | GO:0071962 | mitotic sister chromatid cohesion, centromeric(GO:0071962) |
| 0.1 | 0.9 | GO:0033292 | T-tubule organization(GO:0033292) |
| 0.1 | 0.4 | GO:0014743 | regulation of muscle hypertrophy(GO:0014743) |
| 0.1 | 0.3 | GO:0036445 | neuronal stem cell division(GO:0036445) somatic stem cell division(GO:0048103) |
| 0.1 | 0.6 | GO:1900026 | positive regulation of substrate adhesion-dependent cell spreading(GO:1900026) |
| 0.1 | 0.3 | GO:0072116 | kidney rudiment formation(GO:0072003) pronephros formation(GO:0072116) |
| 0.1 | 0.2 | GO:0072679 | thymocyte migration(GO:0072679) |
| 0.1 | 3.6 | GO:0070098 | chemokine-mediated signaling pathway(GO:0070098) |
| 0.1 | 0.6 | GO:0061056 | sclerotome development(GO:0061056) |
| 0.1 | 0.3 | GO:0043508 | negative regulation of JUN kinase activity(GO:0043508) |
| 0.1 | 0.4 | GO:0071405 | response to methanol(GO:0033986) cellular response to methanol(GO:0071405) |
| 0.1 | 0.7 | GO:0034122 | regulation of toll-like receptor signaling pathway(GO:0034121) negative regulation of toll-like receptor signaling pathway(GO:0034122) negative regulation of innate immune response(GO:0045824) |
| 0.1 | 0.5 | GO:0019731 | antibacterial humoral response(GO:0019731) |
| 0.0 | 0.7 | GO:0016486 | peptide hormone processing(GO:0016486) |
| 0.0 | 0.9 | GO:0050829 | defense response to Gram-negative bacterium(GO:0050829) |
| 0.0 | 0.2 | GO:0016338 | calcium-independent cell-cell adhesion via plasma membrane cell-adhesion molecules(GO:0016338) |
| 0.0 | 0.4 | GO:0050435 | beta-amyloid metabolic process(GO:0050435) |
| 0.0 | 0.2 | GO:0010447 | response to acidic pH(GO:0010447) |
| 0.0 | 0.5 | GO:0072378 | blood coagulation, fibrin clot formation(GO:0072378) |
| 0.0 | 0.1 | GO:1901004 | ubiquinone-6 metabolic process(GO:1901004) ubiquinone-6 biosynthetic process(GO:1901006) |
| 0.0 | 0.2 | GO:0007141 | male meiosis I(GO:0007141) |
| 0.0 | 0.2 | GO:0031507 | heterochromatin assembly(GO:0031507) |
| 0.0 | 0.1 | GO:0032780 | negative regulation of ATPase activity(GO:0032780) |
| 0.0 | 0.1 | GO:0090161 | Golgi ribbon formation(GO:0090161) |
| 0.0 | 0.5 | GO:0000737 | DNA catabolic process, endonucleolytic(GO:0000737) |
| 0.0 | 1.2 | GO:0003009 | skeletal muscle contraction(GO:0003009) |
| 0.0 | 0.4 | GO:0035803 | binding of sperm to zona pellucida(GO:0007339) sperm-egg recognition(GO:0035036) egg coat formation(GO:0035803) regulation of acrosome reaction(GO:0060046) positive regulation of acrosome reaction(GO:2000344) |
| 0.0 | 0.6 | GO:0071222 | cellular response to molecule of bacterial origin(GO:0071219) cellular response to lipopolysaccharide(GO:0071222) |
| 0.0 | 0.1 | GO:0036371 | protein localization to T-tubule(GO:0036371) |
| 0.0 | 0.6 | GO:0030513 | positive regulation of BMP signaling pathway(GO:0030513) |
| 0.0 | 0.6 | GO:0055092 | cholesterol homeostasis(GO:0042632) sterol homeostasis(GO:0055092) |
| 0.0 | 0.3 | GO:0009712 | catecholamine metabolic process(GO:0006584) catechol-containing compound metabolic process(GO:0009712) |
| 0.0 | 0.5 | GO:0032233 | positive regulation of actin filament bundle assembly(GO:0032233) |
| 0.0 | 0.2 | GO:0044030 | regulation of DNA methylation(GO:0044030) |
| 0.0 | 0.4 | GO:0006884 | cell volume homeostasis(GO:0006884) |
| 0.0 | 0.3 | GO:0033209 | tumor necrosis factor-mediated signaling pathway(GO:0033209) |
| 0.0 | 0.3 | GO:0032958 | inositol phosphate biosynthetic process(GO:0032958) |
| 0.0 | 0.4 | GO:0060030 | dorsal convergence(GO:0060030) |
| 0.0 | 0.3 | GO:0015701 | bicarbonate transport(GO:0015701) |
| 0.0 | 0.1 | GO:0051601 | exocyst localization(GO:0051601) |
| 0.0 | 0.1 | GO:0035677 | posterior lateral line neuromast hair cell development(GO:0035677) |
| 0.0 | 0.1 | GO:0019883 | antigen processing and presentation of endogenous peptide antigen(GO:0002483) antigen processing and presentation of endogenous antigen(GO:0019883) antigen processing and presentation of endogenous peptide antigen via MHC class I(GO:0019885) |
| 0.0 | 0.1 | GO:0008207 | C21-steroid hormone metabolic process(GO:0008207) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.7 | GO:0042612 | MHC class I protein complex(GO:0042612) |
| 0.1 | 0.3 | GO:0070743 | interleukin-12 complex(GO:0043514) interleukin-23 complex(GO:0070743) |
| 0.1 | 1.0 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.1 | 0.7 | GO:0005751 | mitochondrial respiratory chain complex IV(GO:0005751) |
| 0.0 | 0.4 | GO:0042613 | MHC protein complex(GO:0042611) MHC class II protein complex(GO:0042613) |
| 0.0 | 0.6 | GO:0005944 | phosphatidylinositol 3-kinase complex, class IB(GO:0005944) phosphatidylinositol 3-kinase complex, class I(GO:0097651) |
| 0.0 | 0.2 | GO:0030314 | junctional membrane complex(GO:0030314) |
| 0.0 | 1.2 | GO:0005861 | troponin complex(GO:0005861) |
| 0.0 | 1.0 | GO:0044853 | caveola(GO:0005901) plasma membrane raft(GO:0044853) |
| 0.0 | 0.3 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.0 | 0.1 | GO:0000839 | Hrd1p ubiquitin ligase ERAD-L complex(GO:0000839) |
| 0.0 | 0.2 | GO:0000792 | heterochromatin(GO:0000792) |
| 0.0 | 0.3 | GO:0031594 | neuromuscular junction(GO:0031594) |
| 0.0 | 0.5 | GO:0019814 | immunoglobulin complex(GO:0019814) |
| 0.0 | 1.3 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.0 | 1.0 | GO:0030018 | Z disc(GO:0030018) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.1 | GO:0051916 | granulocyte colony-stimulating factor binding(GO:0051916) |
| 0.2 | 1.5 | GO:0008118 | N-acetyllactosaminide alpha-2,3-sialyltransferase activity(GO:0008118) |
| 0.1 | 0.9 | GO:0004126 | cytidine deaminase activity(GO:0004126) |
| 0.1 | 0.6 | GO:0101006 | protein histidine phosphatase activity(GO:0101006) |
| 0.1 | 0.4 | GO:0003796 | lysozyme activity(GO:0003796) |
| 0.1 | 0.3 | GO:0019972 | interleukin-12 binding(GO:0019972) interleukin-12 alpha subunit binding(GO:0042164) |
| 0.1 | 2.7 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.1 | 0.4 | GO:0008190 | eukaryotic initiation factor 4E binding(GO:0008190) |
| 0.1 | 0.3 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.0 | 0.4 | GO:0008511 | sodium:potassium:chloride symporter activity(GO:0008511) |
| 0.0 | 0.5 | GO:0004530 | deoxyribonuclease I activity(GO:0004530) |
| 0.0 | 0.4 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) |
| 0.0 | 0.4 | GO:0016286 | small conductance calcium-activated potassium channel activity(GO:0016286) |
| 0.0 | 0.3 | GO:0015168 | glycerol transmembrane transporter activity(GO:0015168) |
| 0.0 | 0.5 | GO:0070530 | K63-linked polyubiquitin binding(GO:0070530) |
| 0.0 | 0.3 | GO:0004062 | aryl sulfotransferase activity(GO:0004062) |
| 0.0 | 0.4 | GO:0035804 | structural constituent of egg coat(GO:0035804) |
| 0.0 | 0.1 | GO:0047777 | (3S)-citramalyl-CoA lyase activity(GO:0047777) |
| 0.0 | 0.5 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 0.0 | 0.7 | GO:0016676 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors(GO:0016675) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.0 | 0.1 | GO:1990381 | ubiquitin-specific protease binding(GO:1990381) |
| 0.0 | 0.5 | GO:0035591 | signaling adaptor activity(GO:0035591) |
| 0.0 | 0.1 | GO:0034714 | type III transforming growth factor beta receptor binding(GO:0034714) |
| 0.0 | 0.4 | GO:0004382 | guanosine-diphosphatase activity(GO:0004382) uridine-diphosphatase activity(GO:0045134) |
| 0.0 | 0.1 | GO:0042030 | ATPase inhibitor activity(GO:0042030) |
| 0.0 | 0.2 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.0 | 0.5 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.0 | 0.6 | GO:0017147 | Wnt-protein binding(GO:0017147) |
| 0.0 | 0.3 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.0 | 0.3 | GO:0008330 | protein tyrosine/threonine phosphatase activity(GO:0008330) |
| 0.0 | 0.5 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.0 | 0.3 | GO:0000828 | inositol hexakisphosphate kinase activity(GO:0000828) |
| 0.0 | 0.4 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 0.0 | 0.3 | GO:0019531 | oxalate transmembrane transporter activity(GO:0019531) |
| 0.0 | 0.3 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.0 | 0.4 | GO:0008378 | galactosyltransferase activity(GO:0008378) |
| 0.0 | 0.5 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.0 | 0.4 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.0 | 0.1 | GO:0030368 | interleukin-17 receptor activity(GO:0030368) |
| 0.0 | 0.2 | GO:0035005 | 1-phosphatidylinositol-4-phosphate 3-kinase activity(GO:0035005) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.0 | PID TXA2PATHWAY | Thromboxane A2 receptor signaling |
| 0.0 | 0.6 | PID NFKAPPAB ATYPICAL PATHWAY | Atypical NF-kappaB pathway |
| 0.0 | 0.8 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.0 | 0.4 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.0 | 0.7 | PID CD8 TCR DOWNSTREAM PATHWAY | Downstream signaling in naïve CD8+ T cells |
| 0.0 | 1.3 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.0 | 0.8 | ST TUMOR NECROSIS FACTOR PATHWAY | Tumor Necrosis Factor Pathway. |
| 0.0 | 0.3 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.0 | 0.3 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 0.4 | PID HIV NEF PATHWAY | HIV-1 Nef: Negative effector of Fas and TNF-alpha |
| 0.0 | 0.6 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.0 | 0.4 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.0 | 0.2 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.0 | 0.5 | PID FOXO PATHWAY | FoxO family signaling |
| 0.0 | 0.4 | SIG BCR SIGNALING PATHWAY | Members of the BCR signaling pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.7 | REACTOME ENDOSOMAL VACUOLAR PATHWAY | Genes involved in Endosomal/Vacuolar pathway |
| 0.1 | 0.6 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.1 | 0.4 | REACTOME PHOSPHORYLATION OF CD3 AND TCR ZETA CHAINS | Genes involved in Phosphorylation of CD3 and TCR zeta chains |
| 0.0 | 0.7 | REACTOME RIP MEDIATED NFKB ACTIVATION VIA DAI | Genes involved in RIP-mediated NFkB activation via DAI |
| 0.0 | 0.4 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 0.0 | 1.1 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.0 | 0.3 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.0 | 0.4 | REACTOME G BETA GAMMA SIGNALLING THROUGH PI3KGAMMA | Genes involved in G beta:gamma signalling through PI3Kgamma |
| 0.0 | 0.5 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.0 | 0.5 | REACTOME G1 PHASE | Genes involved in G1 Phase |
| 0.0 | 0.5 | REACTOME ACTIVATION OF NF KAPPAB IN B CELLS | Genes involved in Activation of NF-kappaB in B Cells |