Project
PRJNA207719: Tissue specific transcriptome profiling
Navigation
Downloads
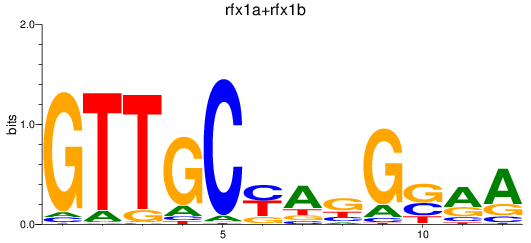
Results for rfx1a+rfx1b
Z-value: 1.02
Transcription factors associated with rfx1a+rfx1b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
rfx1a
|
ENSDARG00000005883 | regulatory factor X, 1a (influences HLA class II expression) |
|
rfx1b
|
ENSDARG00000075904 | regulatory factor X, 1b (influences HLA class II expression) |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| rfx1a | dr11_v1_chr3_-_19200571_19200651 | 0.90 | 3.9e-02 | Click! |
| rfx1b | dr11_v1_chr1_-_54100988_54100988 | 0.06 | 9.3e-01 | Click! |
Activity profile of rfx1a+rfx1b motif
Sorted Z-values of rfx1a+rfx1b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr15_-_47193564 | 1.43 |
ENSDART00000172453
|
LSAMP
|
limbic system-associated membrane protein |
| chr3_-_5664123 | 1.04 |
ENSDART00000145866
|
si:ch211-106h11.1
|
si:ch211-106h11.1 |
| chr15_+_22267847 | 0.95 |
ENSDART00000110665
|
spa17
|
sperm autoantigenic protein 17 |
| chr19_+_2275019 | 0.93 |
ENSDART00000136138
|
itgb8
|
integrin, beta 8 |
| chr25_+_34407740 | 0.91 |
ENSDART00000012677
|
OTUD7A
|
si:dkey-37f18.2 |
| chr6_+_13933464 | 0.90 |
ENSDART00000109144
|
ptprnb
|
protein tyrosine phosphatase, receptor type, Nb |
| chr20_+_31076488 | 0.71 |
ENSDART00000136255
ENSDART00000008840 |
otofa
|
otoferlin a |
| chr11_-_1291012 | 0.68 |
ENSDART00000158390
|
atp2b2
|
ATPase plasma membrane Ca2+ transporting 2 |
| chr17_+_23937262 | 0.66 |
ENSDART00000113276
|
si:ch211-189k9.2
|
si:ch211-189k9.2 |
| chr1_+_8662530 | 0.66 |
ENSDART00000054989
|
fscn1b
|
fascin actin-bundling protein 1b |
| chr22_+_438714 | 0.66 |
ENSDART00000136491
|
celsr2
|
cadherin, EGF LAG seven-pass G-type receptor 2 |
| chr8_+_7144066 | 0.62 |
ENSDART00000146306
|
slc6a6a
|
solute carrier family 6 (neurotransmitter transporter), member 6a |
| chr13_-_36911118 | 0.60 |
ENSDART00000048739
|
trim9
|
tripartite motif containing 9 |
| chr13_-_31470439 | 0.60 |
ENSDART00000076574
|
rtn1a
|
reticulon 1a |
| chr23_+_6272638 | 0.59 |
ENSDART00000190366
|
syt2a
|
synaptotagmin IIa |
| chr24_-_205275 | 0.59 |
ENSDART00000108762
|
vopp1
|
VOPP1, WBP1/VOPP1 family member |
| chr18_-_46763170 | 0.59 |
ENSDART00000171880
|
dner
|
delta/notch-like EGF repeat containing |
| chr7_+_23515966 | 0.58 |
ENSDART00000186893
ENSDART00000186189 |
zgc:109889
|
zgc:109889 |
| chr21_-_43606502 | 0.56 |
ENSDART00000151030
|
si:ch73-362m14.4
|
si:ch73-362m14.4 |
| chr4_+_19534833 | 0.56 |
ENSDART00000140028
|
lrrc4.1
|
leucine rich repeat containing 4.1 |
| chr19_+_21266008 | 0.55 |
ENSDART00000024639
|
tshz1
|
teashirt zinc finger homeobox 1 |
| chr2_+_59527149 | 0.55 |
ENSDART00000168568
|
CU861477.1
|
|
| chr21_+_43669943 | 0.55 |
ENSDART00000136025
|
tmlhe
|
trimethyllysine hydroxylase, epsilon |
| chr16_+_5202042 | 0.54 |
ENSDART00000145368
|
soga3a
|
SOGA family member 3a |
| chr15_+_36115955 | 0.54 |
ENSDART00000032702
|
sst1.2
|
somatostatin 1, tandem duplicate 2 |
| chr14_+_6954579 | 0.53 |
ENSDART00000060998
|
nme5
|
NME/NM23 family member 5 |
| chr11_-_18799827 | 0.53 |
ENSDART00000185438
ENSDART00000189116 ENSDART00000180504 |
pfkfb2b
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 2b |
| chr17_-_29119362 | 0.52 |
ENSDART00000104204
|
foxg1a
|
forkhead box G1a |
| chr13_+_33117528 | 0.52 |
ENSDART00000085719
|
si:ch211-10a23.2
|
si:ch211-10a23.2 |
| chr16_-_16590780 | 0.51 |
ENSDART00000059841
|
si:ch211-257p13.3
|
si:ch211-257p13.3 |
| chr1_+_604127 | 0.51 |
ENSDART00000133165
|
jam2a
|
junctional adhesion molecule 2a |
| chr5_-_38384755 | 0.50 |
ENSDART00000188573
ENSDART00000051233 |
mink1
|
misshapen-like kinase 1 |
| chr21_+_9576176 | 0.48 |
ENSDART00000161289
ENSDART00000159899 ENSDART00000162834 |
mapk10
|
mitogen-activated protein kinase 10 |
| chr18_-_2727764 | 0.48 |
ENSDART00000160841
|
ARHGEF17
|
si:ch211-248g20.5 |
| chr15_-_12113045 | 0.48 |
ENSDART00000159879
|
dscaml1
|
Down syndrome cell adhesion molecule like 1 |
| chr17_+_27176243 | 0.47 |
ENSDART00000162527
|
si:ch211-160f23.7
|
si:ch211-160f23.7 |
| chr10_+_2234283 | 0.47 |
ENSDART00000136363
|
cntnap3
|
contactin associated protein like 3 |
| chr18_+_49411417 | 0.46 |
ENSDART00000028944
|
LRFN3
|
zmp:0000001073 |
| chr14_-_9199968 | 0.45 |
ENSDART00000146113
|
arhgef9b
|
Cdc42 guanine nucleotide exchange factor (GEF) 9b |
| chr18_+_29950233 | 0.45 |
ENSDART00000146431
|
atmin
|
ATM interactor |
| chr12_+_13405445 | 0.44 |
ENSDART00000089042
|
kcnh4b
|
potassium voltage-gated channel, subfamily H (eag-related), member 4b |
| chr2_+_33796207 | 0.44 |
ENSDART00000124647
|
kiss1ra
|
KISS1 receptor a |
| chr22_+_3223489 | 0.44 |
ENSDART00000082011
|
lim2.2
|
lens intrinsic membrane protein 2.2 |
| chr16_+_11483811 | 0.43 |
ENSDART00000169012
ENSDART00000173042 |
grik5
|
glutamate receptor, ionotropic, kainate 5 |
| chr11_-_18800299 | 0.43 |
ENSDART00000156276
|
pfkfb2b
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 2b |
| chr17_-_22067451 | 0.43 |
ENSDART00000156872
|
ttbk1b
|
tau tubulin kinase 1b |
| chr16_+_34493987 | 0.43 |
ENSDART00000138374
|
si:ch211-255i3.4
|
si:ch211-255i3.4 |
| chr19_-_8604429 | 0.42 |
ENSDART00000151165
|
trim46b
|
tripartite motif containing 46b |
| chr17_+_33767890 | 0.42 |
ENSDART00000193177
|
fut8a
|
fucosyltransferase 8a (alpha (1,6) fucosyltransferase) |
| chr6_+_13787855 | 0.41 |
ENSDART00000182899
|
tmem198b
|
transmembrane protein 198b |
| chr22_-_11438627 | 0.41 |
ENSDART00000007649
|
mid1ip1b
|
MID1 interacting protein 1b |
| chr8_-_9570511 | 0.41 |
ENSDART00000044000
|
plxna3
|
plexin A3 |
| chr16_+_46111849 | 0.41 |
ENSDART00000172232
|
sv2a
|
synaptic vesicle glycoprotein 2A |
| chr20_-_27857676 | 0.41 |
ENSDART00000192934
|
syndig1l
|
synapse differentiation inducing 1-like |
| chr9_+_6009077 | 0.41 |
ENSDART00000057484
|
st6gal2a
|
ST6 beta-galactosamide alpha-2,6-sialyltranferase 2a |
| chr2_-_12502495 | 0.41 |
ENSDART00000186781
|
dpp6b
|
dipeptidyl-peptidase 6b |
| chr19_+_2279051 | 0.40 |
ENSDART00000182103
|
itgb8
|
integrin, beta 8 |
| chr10_-_29744921 | 0.39 |
ENSDART00000088605
|
ift46
|
intraflagellar transport 46 homolog (Chlamydomonas) |
| chr8_+_25267903 | 0.39 |
ENSDART00000093090
|
ampd2b
|
adenosine monophosphate deaminase 2b |
| chr11_-_24428237 | 0.38 |
ENSDART00000189107
ENSDART00000103752 |
dvl1b
|
dishevelled segment polarity protein 1b |
| chr11_+_7528599 | 0.38 |
ENSDART00000171813
|
adgrl2a
|
adhesion G protein-coupled receptor L2a |
| chr13_-_18691041 | 0.38 |
ENSDART00000057867
|
sfxn3
|
sideroflexin 3 |
| chr1_+_37752171 | 0.38 |
ENSDART00000183247
ENSDART00000189756 ENSDART00000139448 |
GALNTL6
|
si:ch211-15e22.3 |
| chr16_+_41717313 | 0.38 |
ENSDART00000015170
ENSDART00000075937 |
mtdhb
|
metadherin b |
| chr4_-_9557186 | 0.37 |
ENSDART00000150569
|
shank3b
|
SH3 and multiple ankyrin repeat domains 3b |
| chr20_+_49787584 | 0.37 |
ENSDART00000193458
ENSDART00000181511 ENSDART00000185850 ENSDART00000185613 ENSDART00000191671 |
CABZ01078261.1
|
|
| chr14_-_52542928 | 0.37 |
ENSDART00000075749
|
ppp2r2ba
|
protein phosphatase 2, regulatory subunit B, beta a |
| chr5_-_38384289 | 0.36 |
ENSDART00000135260
|
mink1
|
misshapen-like kinase 1 |
| chr22_+_25049563 | 0.36 |
ENSDART00000078173
|
dzank1
|
double zinc ribbon and ankyrin repeat domains 1 |
| chr17_-_11329959 | 0.36 |
ENSDART00000015418
|
irf2bpl
|
interferon regulatory factor 2 binding protein-like |
| chr2_-_1569250 | 0.35 |
ENSDART00000167202
|
dab1b
|
Dab, reelin signal transducer, homolog 1b (Drosophila) |
| chr6_-_7711349 | 0.35 |
ENSDART00000032494
|
ttc26
|
tetratricopeptide repeat domain 26 |
| chr20_+_36682051 | 0.35 |
ENSDART00000130513
|
ncoa1
|
nuclear receptor coactivator 1 |
| chr18_+_5917625 | 0.35 |
ENSDART00000169100
|
glg1b
|
golgi glycoprotein 1b |
| chr5_+_32260502 | 0.35 |
ENSDART00000149020
|
si:ch211-158m24.12
|
si:ch211-158m24.12 |
| chr20_-_12642685 | 0.35 |
ENSDART00000173257
|
si:dkey-97l20.6
|
si:dkey-97l20.6 |
| chr10_+_15255012 | 0.34 |
ENSDART00000023766
|
vldlr
|
very low density lipoprotein receptor |
| chr13_+_28854438 | 0.33 |
ENSDART00000193407
ENSDART00000189554 |
CU639469.1
|
|
| chr10_+_15255198 | 0.33 |
ENSDART00000139047
ENSDART00000172107 ENSDART00000183413 ENSDART00000185314 |
vldlr
|
very low density lipoprotein receptor |
| chr16_-_9869056 | 0.32 |
ENSDART00000149312
|
ncalda
|
neurocalcin delta a |
| chr1_-_45633955 | 0.31 |
ENSDART00000044057
|
sept3
|
septin 3 |
| chr11_-_30352333 | 0.31 |
ENSDART00000030794
|
tmem169a
|
transmembrane protein 169a |
| chr5_+_61919217 | 0.30 |
ENSDART00000097340
|
rsph4a
|
radial spoke head 4 homolog A |
| chr8_+_29749017 | 0.29 |
ENSDART00000185144
|
mapk4
|
mitogen-activated protein kinase 4 |
| chr25_+_18436301 | 0.29 |
ENSDART00000056180
|
cep41
|
centrosomal protein 41 |
| chr4_-_1801519 | 0.29 |
ENSDART00000188604
ENSDART00000135749 |
nudt4b
|
nudix (nucleoside diphosphate linked moiety X)-type motif 4b |
| chr13_-_36184476 | 0.29 |
ENSDART00000057185
|
map3k9
|
mitogen-activated protein kinase kinase kinase 9 |
| chr24_-_39772045 | 0.29 |
ENSDART00000087441
|
GFOD1
|
si:ch211-276f18.2 |
| chr1_+_594584 | 0.29 |
ENSDART00000135944
|
jam2a
|
junctional adhesion molecule 2a |
| chr15_-_1534232 | 0.29 |
ENSDART00000056763
ENSDART00000133943 |
ift80
|
intraflagellar transport 80 homolog (Chlamydomonas) |
| chr11_-_36341189 | 0.28 |
ENSDART00000159752
|
sort1a
|
sortilin 1a |
| chr8_-_49345388 | 0.28 |
ENSDART00000053203
|
plp2
|
proteolipid protein 2 |
| chr17_+_8988374 | 0.28 |
ENSDART00000109573
|
akap6
|
A kinase (PRKA) anchor protein 6 |
| chr7_-_34505985 | 0.28 |
ENSDART00000173806
ENSDART00000024869 |
tex9
|
testis expressed 9 |
| chr9_-_41401564 | 0.28 |
ENSDART00000059628
|
nab1b
|
NGFI-A binding protein 1b (EGR1 binding protein 1) |
| chr1_+_53279861 | 0.28 |
ENSDART00000035713
|
rnf150a
|
ring finger protein 150a |
| chr6_-_18250857 | 0.28 |
ENSDART00000159486
|
anapc11
|
APC11 anaphase promoting complex subunit 11 homolog (yeast) |
| chr14_+_24283915 | 0.27 |
ENSDART00000172868
|
klhl3
|
kelch-like family member 3 |
| chr2_+_55914699 | 0.27 |
ENSDART00000157905
|
atcayb
|
ataxia, cerebellar, Cayman type b |
| chr21_-_35832548 | 0.27 |
ENSDART00000180840
|
sgcd
|
sarcoglycan, delta (dystrophin-associated glycoprotein) |
| chr2_-_7696287 | 0.27 |
ENSDART00000190769
|
CABZ01055306.1
|
|
| chr4_-_9579299 | 0.27 |
ENSDART00000183079
ENSDART00000192968 ENSDART00000091809 |
shank3b
|
SH3 and multiple ankyrin repeat domains 3b |
| chr4_-_23858900 | 0.26 |
ENSDART00000123199
|
usp6nl
|
USP6 N-terminal like |
| chr14_-_21219659 | 0.26 |
ENSDART00000089867
|
ppp2r2cb
|
protein phosphatase 2, regulatory subunit B, gamma b |
| chr2_-_7696503 | 0.26 |
ENSDART00000169709
|
CABZ01055306.1
|
|
| chr21_+_13387965 | 0.26 |
ENSDART00000134347
|
zgc:113162
|
zgc:113162 |
| chr19_+_17400283 | 0.26 |
ENSDART00000127353
|
nr1d2b
|
nuclear receptor subfamily 1, group D, member 2b |
| chr11_-_28911172 | 0.25 |
ENSDART00000168493
|
igsf21a
|
immunoglobin superfamily, member 21a |
| chr9_+_9112159 | 0.25 |
ENSDART00000165932
|
sik1
|
salt-inducible kinase 1 |
| chr9_+_40874194 | 0.25 |
ENSDART00000141548
|
hibch
|
3-hydroxyisobutyryl-CoA hydrolase |
| chr3_-_46410387 | 0.25 |
ENSDART00000156822
|
cdip1
|
cell death-inducing p53 target 1 |
| chr17_+_53250802 | 0.24 |
ENSDART00000143819
|
VASH1
|
vasohibin 1 |
| chr16_+_9726038 | 0.24 |
ENSDART00000192988
ENSDART00000020859 |
pip5k1ab
|
phosphatidylinositol-4-phosphate 5-kinase, type I, alpha, b |
| chr24_+_39186940 | 0.24 |
ENSDART00000155817
|
spsb3b
|
splA/ryanodine receptor domain and SOCS box containing 3b |
| chr8_-_50525360 | 0.23 |
ENSDART00000175648
|
CABZ01060030.1
|
|
| chr23_-_12158685 | 0.23 |
ENSDART00000135035
|
fam217b
|
family with sequence similarity 217, member B |
| chr21_+_22124736 | 0.23 |
ENSDART00000130179
ENSDART00000172573 |
cul5b
|
cullin 5b |
| chr3_-_61593274 | 0.23 |
ENSDART00000154132
ENSDART00000055071 |
nptx2a
|
neuronal pentraxin 2a |
| chr7_+_27059330 | 0.23 |
ENSDART00000173919
|
plekha7b
|
pleckstrin homology domain containing, family A member 7b |
| chr1_-_49498116 | 0.23 |
ENSDART00000137357
|
zgc:175214
|
zgc:175214 |
| chr17_-_8173380 | 0.22 |
ENSDART00000148520
|
syne1b
|
spectrin repeat containing, nuclear envelope 1b |
| chr1_+_47831619 | 0.21 |
ENSDART00000110335
ENSDART00000101066 |
cfap58
|
cilia and flagella associated protein 58 |
| chr15_+_31735931 | 0.21 |
ENSDART00000185681
ENSDART00000149137 |
rxfp2b
|
relaxin/insulin-like family peptide receptor 2b |
| chr1_-_21538329 | 0.21 |
ENSDART00000182798
ENSDART00000193518 ENSDART00000183873 ENSDART00000183376 |
wdr19
|
WD repeat domain 19 |
| chr21_-_435466 | 0.21 |
ENSDART00000110297
|
klf4
|
Kruppel-like factor 4 |
| chr24_+_38155830 | 0.21 |
ENSDART00000152019
|
si:ch211-234p6.5
|
si:ch211-234p6.5 |
| chr3_-_12026741 | 0.21 |
ENSDART00000132238
|
cfap70
|
cilia and flagella associated protein 70 |
| chr10_-_7988396 | 0.20 |
ENSDART00000141445
ENSDART00000024282 |
ewsr1a
|
EWS RNA-binding protein 1a |
| chr1_+_50538839 | 0.20 |
ENSDART00000020412
|
pkd2
|
polycystic kidney disease 2 |
| chr11_+_77526 | 0.20 |
ENSDART00000193521
|
CABZ01072242.1
|
|
| chr23_+_21638258 | 0.20 |
ENSDART00000104188
|
igsf21b
|
immunoglobin superfamily, member 21b |
| chr17_-_14451718 | 0.20 |
ENSDART00000039438
|
jkamp
|
jnk1/mapk8 associated membrane protein |
| chr1_-_21538673 | 0.20 |
ENSDART00000131257
|
wdr19
|
WD repeat domain 19 |
| chr11_+_24800156 | 0.20 |
ENSDART00000131976
|
adipor1a
|
adiponectin receptor 1a |
| chr20_-_31075972 | 0.20 |
ENSDART00000122927
|
si:ch211-198b3.4
|
si:ch211-198b3.4 |
| chr16_+_13898399 | 0.20 |
ENSDART00000140946
|
si:ch211-149k23.9
|
si:ch211-149k23.9 |
| chr25_-_20523927 | 0.19 |
ENSDART00000114448
|
si:ch211-269c21.2
|
si:ch211-269c21.2 |
| chr23_+_44665230 | 0.19 |
ENSDART00000144958
|
camta2
|
calmodulin binding transcription activator 2 |
| chr18_+_50890749 | 0.19 |
ENSDART00000174109
|
si:ch1073-450f2.1
|
si:ch1073-450f2.1 |
| chr6_+_515181 | 0.19 |
ENSDART00000171374
|
CACNA1I (1 of many)
|
si:ch73-379f7.5 |
| chr17_+_33296852 | 0.19 |
ENSDART00000154580
|
dnajc27
|
DnaJ (Hsp40) homolog, subfamily C, member 27 |
| chr13_+_31205439 | 0.19 |
ENSDART00000132326
|
ptpn20
|
protein tyrosine phosphatase, non-receptor type 20 |
| chr1_-_46718353 | 0.19 |
ENSDART00000074514
|
spryd7a
|
SPRY domain containing 7a |
| chr18_-_50579629 | 0.18 |
ENSDART00000188252
|
zgc:158464
|
zgc:158464 |
| chr13_-_4303641 | 0.18 |
ENSDART00000136262
|
ppp2r5d
|
protein phosphatase 2, regulatory subunit B', delta |
| chr20_-_4766645 | 0.18 |
ENSDART00000147071
ENSDART00000152398 |
ak7a
|
adenylate kinase 7a |
| chr13_+_13805385 | 0.18 |
ENSDART00000166859
|
slc4a11
|
solute carrier family 4, sodium borate transporter, member 11 |
| chr6_-_41138854 | 0.18 |
ENSDART00000128723
ENSDART00000151055 ENSDART00000132484 |
slc6a22.1
|
solute carrier family 6 member 22, tandem duplicate 1 |
| chr17_+_33296436 | 0.17 |
ENSDART00000123114
|
dnajc27
|
DnaJ (Hsp40) homolog, subfamily C, member 27 |
| chr15_-_43768776 | 0.17 |
ENSDART00000170398
|
grm5b
|
glutamate receptor, metabotropic 5b |
| chr21_+_17005737 | 0.17 |
ENSDART00000101246
|
vps29
|
vacuolar protein sorting 29 homolog (S. cerevisiae) |
| chr4_+_74002176 | 0.17 |
ENSDART00000163662
|
si:cabz01069013.3
|
si:cabz01069013.3 |
| chr6_-_58901552 | 0.17 |
ENSDART00000122003
|
mbd6
|
methyl-CpG binding domain protein 6 |
| chr17_-_12758171 | 0.16 |
ENSDART00000131564
|
brms1la
|
breast cancer metastasis-suppressor 1-like a |
| chr23_+_10469955 | 0.16 |
ENSDART00000140557
|
tns2a
|
tensin 2a |
| chr6_+_18418651 | 0.16 |
ENSDART00000171097
ENSDART00000167798 |
rab11fip4b
|
RAB11 family interacting protein 4 (class II) b |
| chr18_-_17531483 | 0.16 |
ENSDART00000061000
|
bbs2
|
Bardet-Biedl syndrome 2 |
| chr20_-_2361226 | 0.16 |
ENSDART00000172130
|
EPB41L2
|
si:ch73-18b11.1 |
| chr24_-_9960290 | 0.16 |
ENSDART00000143390
ENSDART00000092975 ENSDART00000184953 |
vps41
|
vacuolar protein sorting 41 homolog (S. cerevisiae) |
| chr4_+_10616626 | 0.16 |
ENSDART00000067251
ENSDART00000143690 |
cped1
|
cadherin-like and PC-esterase domain containing 1 |
| chr13_+_48482256 | 0.15 |
ENSDART00000053332
|
FOXN2 (1 of many)
|
forkhead box N2 |
| chr12_+_4971515 | 0.15 |
ENSDART00000161076
|
arhgap27
|
Rho GTPase activating protein 27 |
| chr8_-_22838364 | 0.15 |
ENSDART00000187954
ENSDART00000185368 ENSDART00000188210 |
magixa
|
MAGI family member, X-linked a |
| chr18_-_31051847 | 0.15 |
ENSDART00000170982
|
gas8
|
growth arrest-specific 8 |
| chr17_+_27545183 | 0.15 |
ENSDART00000129392
|
pacrg
|
PARK2 co-regulated |
| chr18_+_30028637 | 0.15 |
ENSDART00000139750
|
si:ch211-220f16.1
|
si:ch211-220f16.1 |
| chr25_+_3868438 | 0.15 |
ENSDART00000156438
|
tmem138
|
transmembrane protein 138 |
| chr18_-_44359726 | 0.15 |
ENSDART00000166935
|
prdm10
|
PR domain containing 10 |
| chr14_+_24277556 | 0.14 |
ENSDART00000122660
|
hnrnpa0a
|
heterogeneous nuclear ribonucleoprotein A0a |
| chr20_+_4060839 | 0.14 |
ENSDART00000178565
|
TRIM67
|
tripartite motif containing 67 |
| chr5_+_43807003 | 0.14 |
ENSDART00000097625
|
zgc:158640
|
zgc:158640 |
| chr10_+_37981471 | 0.14 |
ENSDART00000113112
ENSDART00000185102 |
wscd1b
|
WSC domain containing 1b |
| chr7_-_41881177 | 0.14 |
ENSDART00000174258
ENSDART00000018972 |
zgc:92818
|
zgc:92818 |
| chr3_+_24458899 | 0.14 |
ENSDART00000156655
|
cbx6b
|
chromobox homolog 6b |
| chr17_-_33716688 | 0.14 |
ENSDART00000043651
|
dnal1
|
dynein, axonemal, light chain 1 |
| chr23_+_37185247 | 0.14 |
ENSDART00000146269
|
vwa5b1
|
von Willebrand factor A domain containing 5B1 |
| chr19_+_27589201 | 0.14 |
ENSDART00000182060
|
si:dkeyp-46h3.1
|
si:dkeyp-46h3.1 |
| chr8_-_14080534 | 0.13 |
ENSDART00000042867
|
dedd
|
death effector domain containing |
| chr21_+_3244146 | 0.13 |
ENSDART00000127740
|
ctif
|
CBP80/20-dependent translation initiation factor |
| chr10_-_36682509 | 0.13 |
ENSDART00000148093
ENSDART00000063365 |
dnajb13
|
DnaJ (Hsp40) homolog, subfamily B, member 13 |
| chr24_+_744713 | 0.13 |
ENSDART00000067764
|
stk17a
|
serine/threonine kinase 17a |
| chr12_+_28955766 | 0.13 |
ENSDART00000123417
ENSDART00000139347 |
znf668
|
zinc finger protein 668 |
| chr25_+_5755314 | 0.12 |
ENSDART00000172088
ENSDART00000164208 |
appl2
|
adaptor protein, phosphotyrosine interaction, PH domain and leucine zipper containing 2 |
| chr10_+_15608326 | 0.12 |
ENSDART00000188770
|
zfand5b
|
zinc finger, AN1-type domain 5b |
| chr7_-_34192834 | 0.12 |
ENSDART00000125131
|
smad6a
|
SMAD family member 6a |
| chr4_-_30362840 | 0.12 |
ENSDART00000165929
|
znf1083
|
zinc finger protein 1083 |
| chr8_-_19216180 | 0.11 |
ENSDART00000146162
|
si:ch73-222f22.2
|
si:ch73-222f22.2 |
| chr6_-_46560207 | 0.11 |
ENSDART00000187607
|
kcnb1
|
potassium voltage-gated channel, Shab-related subfamily, member 1 |
| chr2_-_36819624 | 0.11 |
ENSDART00000140844
|
slitrk3b
|
SLIT and NTRK-like family, member 3b |
| chr10_-_74408 | 0.11 |
ENSDART00000100073
ENSDART00000141723 |
dyrk1aa
|
dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 1A, a |
| chr16_-_11779508 | 0.11 |
ENSDART00000136329
ENSDART00000060145 ENSDART00000141101 |
pafah1b3
|
platelet-activating factor acetylhydrolase, isoform Ib, gamma subunit |
| chr22_-_20720427 | 0.11 |
ENSDART00000105532
|
oaz1a
|
ornithine decarboxylase antizyme 1a |
| chr18_+_16133595 | 0.11 |
ENSDART00000080423
|
ctsd
|
cathepsin D |
| chr10_+_2529037 | 0.10 |
ENSDART00000123467
|
zmp:0000001301
|
zmp:0000001301 |
| chr12_+_17620067 | 0.10 |
ENSDART00000073599
|
rsph10b
|
radial spoke head 10 homolog B |
| chr4_-_3145359 | 0.10 |
ENSDART00000112210
|
plekha5
|
pleckstrin homology domain containing, family A member 5 |
| chr3_+_60044780 | 0.10 |
ENSDART00000080437
|
zgc:113030
|
zgc:113030 |
| chr23_+_2760573 | 0.10 |
ENSDART00000129719
|
top1
|
DNA topoisomerase I |
| chr10_-_17231917 | 0.10 |
ENSDART00000038577
|
rab36
|
RAB36, member RAS oncogene family |
| chr2_+_41526904 | 0.10 |
ENSDART00000127520
|
acvr1l
|
activin A receptor, type 1 like |
Network of associatons between targets according to the STRING database.
First level regulatory network of rfx1a+rfx1b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.6 | GO:0014014 | negative regulation of gliogenesis(GO:0014014) |
| 0.2 | 0.5 | GO:1902176 | regulation of oxidative stress-induced intrinsic apoptotic signaling pathway(GO:1902175) negative regulation of oxidative stress-induced intrinsic apoptotic signaling pathway(GO:1902176) |
| 0.1 | 0.6 | GO:2000463 | postsynaptic density assembly(GO:0097107) modulation of excitatory postsynaptic potential(GO:0098815) positive regulation of excitatory postsynaptic potential(GO:2000463) |
| 0.1 | 0.3 | GO:0072156 | distal tubule morphogenesis(GO:0072156) |
| 0.1 | 0.5 | GO:0045329 | carnitine biosynthetic process(GO:0045329) |
| 0.1 | 0.4 | GO:2000480 | negative regulation of cAMP-dependent protein kinase activity(GO:2000480) |
| 0.1 | 0.4 | GO:0008594 | photoreceptor cell morphogenesis(GO:0008594) |
| 0.1 | 1.1 | GO:0051127 | positive regulation of actin nucleation(GO:0051127) positive regulation of Arp2/3 complex-mediated actin nucleation(GO:2000601) |
| 0.1 | 0.4 | GO:0048755 | branching morphogenesis of a nerve(GO:0048755) |
| 0.1 | 0.2 | GO:0071498 | cellular response to fluid shear stress(GO:0071498) |
| 0.1 | 0.5 | GO:0043584 | nose development(GO:0043584) |
| 0.1 | 1.0 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 0.1 | 0.3 | GO:1901906 | diphosphoinositol polyphosphate metabolic process(GO:0071543) diadenosine pentaphosphate metabolic process(GO:1901906) diadenosine pentaphosphate catabolic process(GO:1901907) diadenosine hexaphosphate metabolic process(GO:1901908) diadenosine hexaphosphate catabolic process(GO:1901909) adenosine 5'-(hexahydrogen pentaphosphate) metabolic process(GO:1901910) adenosine 5'-(hexahydrogen pentaphosphate) catabolic process(GO:1901911) |
| 0.1 | 0.2 | GO:0099552 | trans-synaptic signaling by lipid, modulating synaptic transmission(GO:0099552) trans-synaptic signaling by endocannabinoid, modulating synaptic transmission(GO:0099553) |
| 0.1 | 0.7 | GO:0001964 | startle response(GO:0001964) |
| 0.1 | 0.4 | GO:0090177 | establishment of planar polarity of embryonic epithelium(GO:0042249) establishment of planar polarity involved in neural tube closure(GO:0090177) regulation of establishment of planar polarity involved in neural tube closure(GO:0090178) planar cell polarity pathway involved in neural tube closure(GO:0090179) |
| 0.1 | 0.2 | GO:0061181 | regulation of chondrocyte development(GO:0061181) |
| 0.1 | 0.7 | GO:0021754 | facial nucleus development(GO:0021754) |
| 0.0 | 0.4 | GO:0097476 | spinal cord motor neuron migration(GO:0097476) lateral motor column neuron migration(GO:0097477) |
| 0.0 | 1.3 | GO:0033627 | cell adhesion mediated by integrin(GO:0033627) |
| 0.0 | 0.2 | GO:0006574 | valine catabolic process(GO:0006574) |
| 0.0 | 0.4 | GO:0035721 | intraciliary retrograde transport(GO:0035721) |
| 0.0 | 0.3 | GO:0006895 | Golgi to endosome transport(GO:0006895) |
| 0.0 | 0.8 | GO:0007520 | myoblast fusion(GO:0007520) |
| 0.0 | 0.3 | GO:0070262 | peptidyl-serine dephosphorylation(GO:0070262) |
| 0.0 | 0.4 | GO:0060294 | cilium movement involved in cell motility(GO:0060294) |
| 0.0 | 0.4 | GO:0032264 | IMP salvage(GO:0032264) |
| 0.0 | 0.6 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.0 | 0.4 | GO:0036065 | fucosylation(GO:0036065) |
| 0.0 | 0.2 | GO:0035889 | otolith tethering(GO:0035889) |
| 0.0 | 0.1 | GO:0090004 | positive regulation of Golgi to plasma membrane protein transport(GO:0042998) positive regulation of establishment of protein localization to plasma membrane(GO:0090004) |
| 0.0 | 0.3 | GO:0006798 | polyphosphate metabolic process(GO:0006797) polyphosphate catabolic process(GO:0006798) |
| 0.0 | 0.5 | GO:0042073 | intraciliary transport(GO:0042073) |
| 0.0 | 0.1 | GO:0035124 | embryonic caudal fin morphogenesis(GO:0035124) |
| 0.0 | 0.3 | GO:1901970 | positive regulation of mitotic metaphase/anaphase transition(GO:0045842) positive regulation of mitotic sister chromatid separation(GO:1901970) positive regulation of metaphase/anaphase transition of cell cycle(GO:1902101) |
| 0.0 | 0.3 | GO:0007525 | somatic muscle development(GO:0007525) |
| 0.0 | 0.2 | GO:0034058 | endosomal vesicle fusion(GO:0034058) |
| 0.0 | 0.2 | GO:0033211 | adiponectin-activated signaling pathway(GO:0033211) |
| 0.0 | 0.1 | GO:0060623 | regulation of chromosome condensation(GO:0060623) |
| 0.0 | 0.3 | GO:0018095 | protein polyglutamylation(GO:0018095) |
| 0.0 | 0.1 | GO:2000096 | positive regulation of Wnt signaling pathway, planar cell polarity pathway(GO:2000096) |
| 0.0 | 0.2 | GO:0007190 | activation of adenylate cyclase activity(GO:0007190) positive regulation of cAMP metabolic process(GO:0030816) positive regulation of cAMP biosynthetic process(GO:0030819) positive regulation of adenylate cyclase activity(GO:0045762) |
| 0.0 | 0.6 | GO:0072348 | sulfur compound transport(GO:0072348) |
| 0.0 | 0.1 | GO:0060394 | negative regulation of pathway-restricted SMAD protein phosphorylation(GO:0060394) |
| 0.0 | 0.5 | GO:0070593 | dendrite self-avoidance(GO:0070593) |
| 0.0 | 0.2 | GO:0051877 | pigment granule aggregation in cell center(GO:0051877) |
| 0.0 | 0.3 | GO:0043507 | activation of JUN kinase activity(GO:0007257) positive regulation of JUN kinase activity(GO:0043507) |
| 0.0 | 0.1 | GO:1904491 | protein localization to ciliary transition zone(GO:1904491) |
| 0.0 | 1.4 | GO:0031098 | stress-activated protein kinase signaling cascade(GO:0031098) |
| 0.0 | 0.1 | GO:0044342 | type B pancreatic cell proliferation(GO:0044342) |
| 0.0 | 0.5 | GO:0046847 | filopodium assembly(GO:0046847) |
| 0.0 | 0.4 | GO:1901379 | regulation of potassium ion transmembrane transport(GO:1901379) |
| 0.0 | 0.1 | GO:0070293 | renal absorption(GO:0070293) |
| 0.0 | 0.5 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.0 | 0.0 | GO:0033540 | fatty acid beta-oxidation using acyl-CoA oxidase(GO:0033540) |
| 0.0 | 0.1 | GO:1904668 | positive regulation of ubiquitin protein ligase activity(GO:1904668) |
| 0.0 | 0.1 | GO:0006004 | fucose metabolic process(GO:0006004) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | GO:0042584 | chromaffin granule membrane(GO:0042584) |
| 0.1 | 0.3 | GO:0001534 | radial spoke(GO:0001534) |
| 0.1 | 0.2 | GO:0031466 | Cul5-RING ubiquitin ligase complex(GO:0031466) |
| 0.1 | 1.1 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.1 | 1.2 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 0.1 | 0.2 | GO:0010369 | chromocenter(GO:0010369) |
| 0.1 | 0.4 | GO:0030990 | intraciliary transport particle(GO:0030990) |
| 0.0 | 0.7 | GO:0034361 | very-low-density lipoprotein particle(GO:0034361) |
| 0.0 | 2.1 | GO:0098636 | protein complex involved in cell adhesion(GO:0098636) |
| 0.0 | 0.3 | GO:0016012 | sarcoglycan complex(GO:0016012) |
| 0.0 | 0.7 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
| 0.0 | 0.8 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.0 | 0.2 | GO:0030897 | HOPS complex(GO:0030897) |
| 0.0 | 0.3 | GO:0016328 | lateral plasma membrane(GO:0016328) |
| 0.0 | 0.2 | GO:0034464 | BBSome(GO:0034464) |
| 0.0 | 0.4 | GO:0002116 | semaphorin receptor complex(GO:0002116) |
| 0.0 | 0.1 | GO:0070724 | BMP receptor complex(GO:0070724) |
| 0.0 | 1.1 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.0 | 0.9 | GO:0098839 | postsynaptic density membrane(GO:0098839) postsynaptic specialization membrane(GO:0099634) |
| 0.0 | 0.2 | GO:0030904 | retromer complex(GO:0030904) |
| 0.0 | 0.1 | GO:1902737 | dendritic spine neck(GO:0044326) dendritic filopodium(GO:1902737) |
| 0.0 | 0.8 | GO:0031514 | motile cilium(GO:0031514) |
| 0.0 | 0.7 | GO:0030426 | growth cone(GO:0030426) |
| 0.0 | 0.6 | GO:0008328 | ionotropic glutamate receptor complex(GO:0008328) neurotransmitter receptor complex(GO:0098878) |
| 0.0 | 0.3 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.0 | 0.2 | GO:0005640 | nuclear outer membrane(GO:0005640) |
| 0.0 | 0.9 | GO:0030141 | secretory granule(GO:0030141) |
| 0.0 | 0.1 | GO:0005779 | integral component of peroxisomal membrane(GO:0005779) intrinsic component of peroxisomal membrane(GO:0031231) |
| 0.0 | 0.1 | GO:0043034 | costamere(GO:0043034) |
| 0.0 | 0.3 | GO:0099569 | presynaptic cytoskeleton(GO:0099569) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:0030228 | lipoprotein particle receptor activity(GO:0030228) |
| 0.1 | 1.1 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 0.1 | 0.6 | GO:0030160 | GKAP/Homer scaffold activity(GO:0030160) |
| 0.1 | 0.4 | GO:0004862 | cAMP-dependent protein kinase inhibitor activity(GO:0004862) |
| 0.1 | 1.0 | GO:0003873 | 6-phosphofructo-2-kinase activity(GO:0003873) |
| 0.1 | 0.4 | GO:0022889 | serine transmembrane transporter activity(GO:0022889) |
| 0.1 | 0.7 | GO:0035612 | AP-2 adaptor complex binding(GO:0035612) |
| 0.1 | 0.3 | GO:0000298 | endopolyphosphatase activity(GO:0000298) diphosphoinositol-polyphosphate diphosphatase activity(GO:0008486) bis(5'-adenosyl)-hexaphosphatase activity(GO:0034431) bis(5'-adenosyl)-pentaphosphatase activity(GO:0034432) |
| 0.1 | 0.4 | GO:0003835 | beta-galactoside alpha-2,6-sialyltransferase activity(GO:0003835) |
| 0.1 | 0.3 | GO:0004127 | cytidylate kinase activity(GO:0004127) |
| 0.0 | 0.5 | GO:0016909 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.0 | 0.4 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.0 | 0.4 | GO:0047623 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 0.0 | 0.3 | GO:0004706 | JUN kinase kinase kinase activity(GO:0004706) |
| 0.0 | 0.1 | GO:0015228 | coenzyme A transmembrane transporter activity(GO:0015228) adenosine 3',5'-bisphosphate transmembrane transporter activity(GO:0071077) AMP transmembrane transporter activity(GO:0080122) |
| 0.0 | 0.5 | GO:0008417 | fucosyltransferase activity(GO:0008417) |
| 0.0 | 0.1 | GO:0008073 | ornithine decarboxylase inhibitor activity(GO:0008073) |
| 0.0 | 0.3 | GO:0004309 | exopolyphosphatase activity(GO:0004309) |
| 0.0 | 0.2 | GO:0099530 | G-protein coupled receptor activity involved in regulation of postsynaptic membrane potential(GO:0099530) |
| 0.0 | 0.7 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.0 | 1.3 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.8 | GO:0005343 | organic acid:sodium symporter activity(GO:0005343) |
| 0.0 | 0.4 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.0 | 0.5 | GO:0004550 | nucleoside diphosphate kinase activity(GO:0004550) |
| 0.0 | 0.3 | GO:0070740 | protein-glutamic acid ligase activity(GO:0070739) tubulin-glutamic acid ligase activity(GO:0070740) |
| 0.0 | 0.1 | GO:0071253 | connexin binding(GO:0071253) |
| 0.0 | 0.3 | GO:0017134 | fibroblast growth factor binding(GO:0017134) |
| 0.0 | 0.5 | GO:0098632 | protein binding involved in cell adhesion(GO:0098631) protein binding involved in cell-cell adhesion(GO:0098632) |
| 0.0 | 0.1 | GO:0008494 | translation activator activity(GO:0008494) |
| 0.0 | 0.2 | GO:0036312 | phosphatidylinositol 3-kinase regulatory subunit binding(GO:0036312) |
| 0.0 | 0.3 | GO:0097602 | cullin family protein binding(GO:0097602) |
| 0.0 | 0.2 | GO:0015271 | outward rectifier potassium channel activity(GO:0015271) |
| 0.0 | 0.1 | GO:1990757 | anaphase-promoting complex binding(GO:0010997) ubiquitin ligase activator activity(GO:1990757) |
| 0.0 | 0.4 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.0 | 0.4 | GO:0099529 | neurotransmitter receptor activity involved in regulation of postsynaptic membrane potential(GO:0099529) transmitter-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1904315) |
| 0.0 | 0.1 | GO:0004692 | cGMP-dependent protein kinase activity(GO:0004692) |
| 0.0 | 0.1 | GO:0010314 | phosphatidylinositol-5-phosphate binding(GO:0010314) |
| 0.0 | 0.6 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.0 | 0.1 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.0 | 0.3 | GO:0004707 | MAP kinase activity(GO:0004707) |
| 0.0 | 0.2 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.3 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.0 | 1.0 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.0 | 0.5 | ST GRANULE CELL SURVIVAL PATHWAY | Granule Cell Survival Pathway is a specific case of more general PAC1 Receptor Pathway. |
| 0.0 | 0.4 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.0 | 0.5 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | REACTOME DSCAM INTERACTIONS | Genes involved in DSCAM interactions |
| 0.1 | 0.5 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 0.1 | 0.4 | REACTOME SEMA3A PAK DEPENDENT AXON REPULSION | Genes involved in Sema3A PAK dependent Axon repulsion |
| 0.0 | 0.7 | REACTOME PLATELET CALCIUM HOMEOSTASIS | Genes involved in Platelet calcium homeostasis |
| 0.0 | 0.4 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.0 | 0.6 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.0 | 0.2 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.0 | 0.4 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.0 | 0.1 | REACTOME CREB PHOSPHORYLATION THROUGH THE ACTIVATION OF CAMKII | Genes involved in CREB phosphorylation through the activation of CaMKII |
| 0.0 | 0.2 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.0 | 1.2 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.0 | 0.3 | REACTOME CONVERSION FROM APC C CDC20 TO APC C CDH1 IN LATE ANAPHASE | Genes involved in Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase |
| 0.0 | 0.2 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.0 | 0.5 | REACTOME NRAGE SIGNALS DEATH THROUGH JNK | Genes involved in NRAGE signals death through JNK |