Project
PRJNA207719: Tissue specific transcriptome profiling
Navigation
Downloads
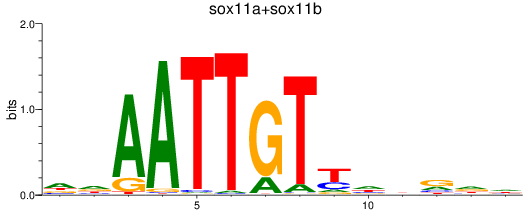
Results for sox11a+sox11b
Z-value: 0.30
Transcription factors associated with sox11a+sox11b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
sox11a
|
ENSDARG00000077811 | SRY-box transcription factor 11a |
|
sox11b
|
ENSDARG00000095743 | SRY-box transcription factor 11b |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| sox11a | dr11_v1_chr17_-_35881841_35881841 | 0.52 | 3.7e-01 | Click! |
| sox11b | dr11_v1_chr20_-_30035326_30035326 | 0.46 | 4.3e-01 | Click! |
Activity profile of sox11a+sox11b motif
Sorted Z-values of sox11a+sox11b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_-_40135942 | 0.10 |
ENSDART00000176951
ENSDART00000098632 ENSDART00000148563 ENSDART00000149895 |
epha4a
|
eph receptor A4a |
| chr8_-_50888806 | 0.10 |
ENSDART00000053750
|
acsl2
|
acyl-CoA synthetase long chain family member 2 |
| chr8_+_24861264 | 0.09 |
ENSDART00000099607
|
slc6a17
|
solute carrier family 6 (neutral amino acid transporter), member 17 |
| chr20_-_14718801 | 0.08 |
ENSDART00000137605
|
suco
|
SUN domain containing ossification factor |
| chr11_+_25596038 | 0.07 |
ENSDART00000140856
|
ccdc120
|
coiled-coil domain containing 120 |
| chr21_+_11401247 | 0.06 |
ENSDART00000143952
|
cel.1
|
carboxyl ester lipase, tandem duplicate 1 |
| chr11_+_30282141 | 0.06 |
ENSDART00000122756
|
si:dkey-163f14.6
|
si:dkey-163f14.6 |
| chr8_-_1838315 | 0.06 |
ENSDART00000114476
ENSDART00000140077 |
pi4kab
|
phosphatidylinositol 4-kinase, catalytic, alpha b |
| chr17_-_48915427 | 0.06 |
ENSDART00000054781
|
lgals8b
|
galectin 8b |
| chr13_-_11644806 | 0.06 |
ENSDART00000169953
|
dctn1b
|
dynactin 1b |
| chr9_+_17984358 | 0.06 |
ENSDART00000192399
|
akap11
|
A kinase (PRKA) anchor protein 11 |
| chr5_+_61657010 | 0.06 |
ENSDART00000050916
|
ap2b1
|
adaptor-related protein complex 2, beta 1 subunit |
| chr8_-_39978767 | 0.06 |
ENSDART00000083066
|
asphd2
|
aspartate beta-hydroxylase domain containing 2 |
| chr13_-_40499296 | 0.06 |
ENSDART00000158338
|
CNNM1
|
Danio rerio cyclin and CBS domain divalent metal cation transport mediator 1 (cnnm1), mRNA. |
| chr1_+_26445615 | 0.05 |
ENSDART00000180810
|
g3bp2
|
GTPase activating protein (SH3 domain) binding protein 2 |
| chr23_+_11669109 | 0.05 |
ENSDART00000091416
|
cntn3a.1
|
contactin 3a, tandem duplicate 1 |
| chr25_-_7764083 | 0.05 |
ENSDART00000179800
ENSDART00000181858 |
phf21ab
|
PHD finger protein 21Ab |
| chr10_-_28761454 | 0.05 |
ENSDART00000129400
|
alcama
|
activated leukocyte cell adhesion molecule a |
| chr3_-_18710009 | 0.05 |
ENSDART00000142478
|
grid2ipa
|
glutamate receptor, ionotropic, delta 2 (Grid2) interacting protein, a |
| chr11_+_30513656 | 0.05 |
ENSDART00000008594
|
tmem178
|
transmembrane protein 178 |
| chr17_-_43558494 | 0.05 |
ENSDART00000103830
|
nt5c1ab
|
5'-nucleotidase, cytosolic IAb |
| chr11_-_25734417 | 0.05 |
ENSDART00000103570
|
brpf3a
|
bromodomain and PHD finger containing, 3a |
| chr10_+_21650828 | 0.05 |
ENSDART00000160754
|
pcdh1g1
|
protocadherin 1 gamma 1 |
| chr5_+_30996322 | 0.05 |
ENSDART00000161765
|
ankfy1
|
ankyrin repeat and FYVE domain containing 1 |
| chr10_+_25204626 | 0.05 |
ENSDART00000024514
|
chordc1b
|
cysteine and histidine-rich domain (CHORD) containing 1b |
| chr5_-_35301800 | 0.05 |
ENSDART00000085142
|
map1b
|
microtubule-associated protein 1B |
| chr16_-_15988320 | 0.05 |
ENSDART00000160883
|
CABZ01060453.1
|
|
| chr1_+_26444986 | 0.05 |
ENSDART00000046376
|
g3bp2
|
GTPase activating protein (SH3 domain) binding protein 2 |
| chr21_-_41305748 | 0.05 |
ENSDART00000170457
|
nsg2
|
neuronal vesicle trafficking associated 2 |
| chr19_+_26718074 | 0.05 |
ENSDART00000134455
|
zgc:100906
|
zgc:100906 |
| chr1_+_10003193 | 0.05 |
ENSDART00000162675
|
trim2b
|
tripartite motif containing 2b |
| chr13_+_10023256 | 0.05 |
ENSDART00000110035
|
srbd1
|
S1 RNA binding domain 1 |
| chr5_-_24543526 | 0.04 |
ENSDART00000046384
|
trmt2a
|
tRNA methyltransferase 2 homolog A |
| chr4_+_28374628 | 0.04 |
ENSDART00000076037
|
alg10
|
asparagine-linked glycosylation 10 |
| chr23_-_33640959 | 0.04 |
ENSDART00000187599
ENSDART00000189475 |
csrnp2
|
cysteine-serine-rich nuclear protein 2 |
| chr23_+_4226341 | 0.04 |
ENSDART00000012445
|
zgc:113278
|
zgc:113278 |
| chr5_-_38342992 | 0.04 |
ENSDART00000140337
|
mink1
|
misshapen-like kinase 1 |
| chr13_+_13693722 | 0.04 |
ENSDART00000110509
|
si:ch211-194c3.5
|
si:ch211-194c3.5 |
| chr5_-_26118855 | 0.04 |
ENSDART00000009028
|
ela3l
|
elastase 3 like |
| chr16_+_36768674 | 0.04 |
ENSDART00000169208
ENSDART00000180470 |
si:ch73-215d9.1
|
si:ch73-215d9.1 |
| chr11_+_39672874 | 0.04 |
ENSDART00000046663
ENSDART00000157659 |
camta1b
|
calmodulin binding transcription activator 1b |
| chr5_+_66353750 | 0.04 |
ENSDART00000143410
|
si:ch211-261c8.5
|
si:ch211-261c8.5 |
| chr6_+_4229360 | 0.04 |
ENSDART00000191347
ENSDART00000130642 |
FO082877.1
|
|
| chr9_+_38883388 | 0.04 |
ENSDART00000135902
|
map2
|
microtubule-associated protein 2 |
| chr21_-_18648861 | 0.04 |
ENSDART00000112113
|
si:dkey-112m2.1
|
si:dkey-112m2.1 |
| chr18_-_8380090 | 0.04 |
ENSDART00000141581
ENSDART00000081143 |
sephs1
|
selenophosphate synthetase 1 |
| chr5_-_67911111 | 0.04 |
ENSDART00000051833
|
gsx1
|
GS homeobox 1 |
| chr2_+_26240631 | 0.04 |
ENSDART00000129895
|
palm1b
|
paralemmin 1b |
| chr5_-_26795438 | 0.04 |
ENSDART00000146124
|
si:ch211-102c2.7
|
si:ch211-102c2.7 |
| chr12_-_37734973 | 0.04 |
ENSDART00000140353
|
sdk2b
|
sidekick cell adhesion molecule 2b |
| chr9_+_29431763 | 0.04 |
ENSDART00000186095
ENSDART00000182640 |
uggt2
|
UDP-glucose glycoprotein glucosyltransferase 2 |
| chr18_-_2222128 | 0.04 |
ENSDART00000171402
|
pigb
|
phosphatidylinositol glycan anchor biosynthesis, class B |
| chr16_-_44878245 | 0.04 |
ENSDART00000154391
ENSDART00000154925 ENSDART00000154697 |
arhgap33
|
Rho GTPase activating protein 33 |
| chr20_+_50852356 | 0.04 |
ENSDART00000167517
ENSDART00000168396 |
gphnb
|
gephyrin b |
| chr4_-_16706776 | 0.04 |
ENSDART00000079461
|
dennd5b
|
DENN/MADD domain containing 5B |
| chr20_-_8304271 | 0.04 |
ENSDART00000179057
|
dab1a
|
Dab, reelin signal transducer, homolog 1a (Drosophila) |
| chr3_-_33175583 | 0.04 |
ENSDART00000126022
|
rarab
|
retinoic acid receptor, alpha b |
| chr8_+_34434345 | 0.04 |
ENSDART00000190246
ENSDART00000189447 ENSDART00000185557 ENSDART00000189230 |
zgc:174461
|
zgc:174461 |
| chr9_-_29985390 | 0.04 |
ENSDART00000134157
|
il1rapl1a
|
interleukin 1 receptor accessory protein-like 1a |
| chr5_+_69716458 | 0.03 |
ENSDART00000159594
|
mthfd2
|
methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 2, methenyltetrahydrofolate cyclohydrolase |
| chr12_+_26632448 | 0.03 |
ENSDART00000185762
|
arhgap12b
|
Rho GTPase activating protein 12b |
| chr18_-_26811959 | 0.03 |
ENSDART00000086250
|
znf592
|
zinc finger protein 592 |
| chr8_-_14050758 | 0.03 |
ENSDART00000133922
|
atp2b3a
|
ATPase plasma membrane Ca2+ transporting 3a |
| chr10_-_24659822 | 0.03 |
ENSDART00000171842
ENSDART00000163102 |
proser1
|
proline and serine rich 1 |
| chr10_-_7857494 | 0.03 |
ENSDART00000143215
|
inpp5ja
|
inositol polyphosphate-5-phosphatase Ja |
| chr8_+_28358161 | 0.03 |
ENSDART00000062682
|
adipor1b
|
adiponectin receptor 1b |
| chr12_-_15567104 | 0.03 |
ENSDART00000053003
|
hexim1
|
hexamethylene bis-acetamide inducible 1 |
| chr2_+_42112503 | 0.03 |
ENSDART00000136022
|
crha
|
corticotropin releasing hormone a |
| chr25_-_20378721 | 0.03 |
ENSDART00000181707
|
kctd15a
|
potassium channel tetramerization domain containing 15a |
| chr2_+_24177006 | 0.03 |
ENSDART00000132582
|
map4l
|
microtubule associated protein 4 like |
| chr20_-_26039841 | 0.03 |
ENSDART00000179929
|
si:dkey-12h9.6
|
si:dkey-12h9.6 |
| chr11_+_16153207 | 0.03 |
ENSDART00000192356
|
tada3l
|
transcriptional adaptor 3 (NGG1 homolog, yeast)-like |
| chr1_+_16397063 | 0.03 |
ENSDART00000159794
|
micu3a
|
mitochondrial calcium uptake family, member 3a |
| chr11_-_25733910 | 0.03 |
ENSDART00000171935
|
brpf3a
|
bromodomain and PHD finger containing, 3a |
| chr14_-_2187202 | 0.03 |
ENSDART00000160205
|
pcdh2ab12
|
protocadherin 2 alpha b 12 |
| chr10_-_41906609 | 0.03 |
ENSDART00000102530
|
kdm2bb
|
lysine (K)-specific demethylase 2Bb |
| chr10_+_38806719 | 0.03 |
ENSDART00000184416
|
dscama
|
Down syndrome cell adhesion molecule a |
| chr19_-_47456787 | 0.03 |
ENSDART00000168792
|
tfap2e
|
transcription factor AP-2 epsilon |
| chr10_+_42521943 | 0.03 |
ENSDART00000010420
ENSDART00000075303 |
actr1
|
ARP1 actin related protein 1, centractin |
| chr9_+_32978302 | 0.03 |
ENSDART00000007630
|
nhlh2
|
nescient helix loop helix 2 |
| chr19_-_6083761 | 0.03 |
ENSDART00000151185
ENSDART00000143941 |
gsk3aa
|
glycogen synthase kinase 3 alpha a |
| chr19_-_17864213 | 0.03 |
ENSDART00000151043
|
ints8
|
integrator complex subunit 8 |
| chr19_-_31402429 | 0.03 |
ENSDART00000137292
|
tmem106bb
|
transmembrane protein 106Bb |
| chr6_-_50704689 | 0.03 |
ENSDART00000074100
|
osgn1
|
oxidative stress induced growth inhibitor 1 |
| chr18_+_3579829 | 0.03 |
ENSDART00000158763
ENSDART00000182850 ENSDART00000162754 ENSDART00000178789 ENSDART00000172656 |
lrch3
|
leucine-rich repeats and calponin homology (CH) domain containing 3 |
| chr12_+_32323098 | 0.03 |
ENSDART00000188722
|
ANKFN1
|
si:ch211-277e21.2 |
| chr6_+_42820228 | 0.03 |
ENSDART00000185413
|
sema3fa
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3Fa |
| chr12_-_21834611 | 0.03 |
ENSDART00000179553
|
thrab
|
thyroid hormone receptor alpha b |
| chr10_-_41907213 | 0.03 |
ENSDART00000167004
|
kdm2bb
|
lysine (K)-specific demethylase 2Bb |
| chr16_+_23087326 | 0.03 |
ENSDART00000167518
ENSDART00000161087 |
efna3b
|
ephrin-A3b |
| chr6_+_39812475 | 0.03 |
ENSDART00000067063
|
c1ql4b
|
complement component 1, q subcomponent-like 4b |
| chr22_+_7439186 | 0.03 |
ENSDART00000190667
|
zgc:92041
|
zgc:92041 |
| chr19_+_23919096 | 0.03 |
ENSDART00000090200
|
snapin
|
SNAP-associated protein |
| chr6_-_15492030 | 0.03 |
ENSDART00000156141
ENSDART00000183992 |
st6gal2b
|
ST6 beta-galactosamide alpha-2,6-sialyltranferase 2b |
| chr4_+_14957360 | 0.03 |
ENSDART00000002770
ENSDART00000111882 ENSDART00000148292 |
tspan33a
|
tetraspanin 33a |
| chr11_-_30158191 | 0.03 |
ENSDART00000155278
ENSDART00000156121 |
scml2
|
Scm polycomb group protein like 2 |
| chr3_+_18807524 | 0.03 |
ENSDART00000055757
|
tnpo2
|
transportin 2 (importin 3, karyopherin beta 2b) |
| chr11_+_25539698 | 0.03 |
ENSDART00000035602
|
cxxc1b
|
CXXC finger protein 1b |
| chr23_-_24234371 | 0.03 |
ENSDART00000124539
|
ddost
|
dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit (non-catalytic) |
| chr20_+_44311448 | 0.03 |
ENSDART00000114660
|
opn8b
|
opsin 8, group member b |
| chr11_+_6881001 | 0.03 |
ENSDART00000170331
|
klhl26
|
kelch-like family member 26 |
| chr5_+_24543862 | 0.03 |
ENSDART00000029699
|
atp6v0a2b
|
ATPase H+ transporting V0 subunit a2b |
| chr22_+_3238474 | 0.03 |
ENSDART00000157954
|
si:ch1073-178p5.3
|
si:ch1073-178p5.3 |
| chr20_-_35012093 | 0.03 |
ENSDART00000062761
|
cnstb
|
consortin, connexin sorting protein b |
| chr7_-_40145097 | 0.03 |
ENSDART00000173634
|
wdr60
|
WD repeat domain 60 |
| chr5_-_37871526 | 0.03 |
ENSDART00000136450
|
arhgap35b
|
Rho GTPase activating protein 35b |
| chr15_+_16908085 | 0.03 |
ENSDART00000186870
|
ypel2b
|
yippee-like 2b |
| chr17_+_47090497 | 0.03 |
ENSDART00000169038
ENSDART00000159292 |
zgc:103755
|
zgc:103755 |
| chr5_-_51484156 | 0.03 |
ENSDART00000162064
|
CR388055.1
|
|
| chr2_+_9061885 | 0.03 |
ENSDART00000028906
|
pigk
|
phosphatidylinositol glycan anchor biosynthesis, class K |
| chr2_-_1569250 | 0.03 |
ENSDART00000167202
|
dab1b
|
Dab, reelin signal transducer, homolog 1b (Drosophila) |
| chr2_+_24177190 | 0.03 |
ENSDART00000099546
|
map4l
|
microtubule associated protein 4 like |
| chr1_-_36152131 | 0.03 |
ENSDART00000182113
ENSDART00000182904 |
znf827
|
zinc finger protein 827 |
| chr11_-_1409236 | 0.03 |
ENSDART00000121537
|
si:ch211-266k22.6
|
si:ch211-266k22.6 |
| chr5_-_24869213 | 0.03 |
ENSDART00000112287
ENSDART00000144635 |
gas2l1
|
growth arrest-specific 2 like 1 |
| chr2_-_32643738 | 0.02 |
ENSDART00000112452
|
si:dkeyp-73d8.9
|
si:dkeyp-73d8.9 |
| chr6_-_37744430 | 0.02 |
ENSDART00000150177
ENSDART00000149722 |
nipa2
|
non imprinted in Prader-Willi/Angelman syndrome 2 (human) |
| chr19_+_2279051 | 0.02 |
ENSDART00000182103
|
itgb8
|
integrin, beta 8 |
| chr19_+_32401278 | 0.02 |
ENSDART00000184353
|
atxn1a
|
ataxin 1a |
| chr15_-_14467394 | 0.02 |
ENSDART00000191944
|
numbl
|
numb homolog (Drosophila)-like |
| chr25_-_20691609 | 0.02 |
ENSDART00000186942
|
edc3
|
enhancer of mRNA decapping 3 homolog (S. cerevisiae) |
| chr1_-_22512063 | 0.02 |
ENSDART00000031546
ENSDART00000190987 |
chrna6
|
cholinergic receptor, nicotinic, alpha 6 |
| chr22_+_10440991 | 0.02 |
ENSDART00000064805
|
cenpp
|
centromere protein P |
| chr19_-_10915898 | 0.02 |
ENSDART00000163179
|
pip5k1aa
|
phosphatidylinositol-4-phosphate 5-kinase, type I, alpha, a |
| chr23_-_4225830 | 0.02 |
ENSDART00000170455
|
aar2
|
AAR2 splicing factor homolog (S. cerevisiae) |
| chr8_+_13368150 | 0.02 |
ENSDART00000114699
|
slc5a5
|
solute carrier family 5 (sodium/iodide cotransporter), member 5 |
| chr4_-_52165969 | 0.02 |
ENSDART00000171130
|
si:dkeyp-44b5.4
|
si:dkeyp-44b5.4 |
| chr16_-_24598042 | 0.02 |
ENSDART00000156407
ENSDART00000154072 |
si:dkey-56f14.4
|
si:dkey-56f14.4 |
| chr11_+_41243094 | 0.02 |
ENSDART00000191782
ENSDART00000169817 ENSDART00000162157 |
pax7a
|
paired box 7a |
| chr8_-_45867358 | 0.02 |
ENSDART00000132810
|
adam9
|
ADAM metallopeptidase domain 9 |
| chr1_-_14234076 | 0.02 |
ENSDART00000040049
|
camk2d2
|
calcium/calmodulin-dependent protein kinase (CaM kinase) II delta 2 |
| chr4_-_25859305 | 0.02 |
ENSDART00000159528
|
nr2c1
|
nuclear receptor subfamily 2, group C, member 1 |
| chr5_-_61431809 | 0.02 |
ENSDART00000082952
|
rcc1l
|
RCC1 like |
| chr22_-_23545307 | 0.02 |
ENSDART00000166915
|
crb1
|
crumbs family member 1, photoreceptor morphogenesis associated |
| chr3_-_12930217 | 0.02 |
ENSDART00000166322
|
pdgfab
|
platelet-derived growth factor alpha polypeptide b |
| chr10_+_24660017 | 0.02 |
ENSDART00000079597
|
vps36
|
vacuolar protein sorting 36 homolog (S. cerevisiae) |
| chr23_+_40460333 | 0.02 |
ENSDART00000184658
|
soga3b
|
SOGA family member 3b |
| chr2_-_22660232 | 0.02 |
ENSDART00000174742
|
thap4
|
THAP domain containing 4 |
| chr1_-_20928772 | 0.02 |
ENSDART00000078277
|
msmo1
|
methylsterol monooxygenase 1 |
| chr20_-_34663209 | 0.02 |
ENSDART00000132545
|
slc25a24
|
solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 24 |
| chr16_+_2899611 | 0.02 |
ENSDART00000149675
|
lars2
|
leucyl-tRNA synthetase 2, mitochondrial |
| chr20_+_50115335 | 0.02 |
ENSDART00000031139
|
slc24a4b
|
solute carrier family 24 (sodium/potassium/calcium exchanger), member 4b |
| chr24_-_21090447 | 0.02 |
ENSDART00000136507
ENSDART00000140786 ENSDART00000184841 |
qtrt2
|
queuine tRNA-ribosyltransferase accessory subunit 2 |
| chr9_-_1469083 | 0.02 |
ENSDART00000144096
|
rbm45
|
RNA binding motif protein 45 |
| chr12_-_18408566 | 0.02 |
ENSDART00000078767
ENSDART00000152748 |
tom1l2
|
target of myb1 like 2 membrane trafficking protein |
| chr11_-_891674 | 0.02 |
ENSDART00000172904
|
atg7
|
ATG7 autophagy related 7 homolog (S. cerevisiae) |
| chr11_+_26363435 | 0.02 |
ENSDART00000088800
|
ppcs
|
phosphopantothenoylcysteine synthetase |
| chr6_-_59271270 | 0.02 |
ENSDART00000126870
|
ndufa4l2b
|
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 4-like 2b |
| chr8_-_39822917 | 0.02 |
ENSDART00000067843
|
zgc:162025
|
zgc:162025 |
| chr24_+_26345609 | 0.02 |
ENSDART00000186844
|
lrrc34
|
leucine rich repeat containing 34 |
| chr17_-_39185336 | 0.02 |
ENSDART00000050534
|
crim1
|
cysteine rich transmembrane BMP regulator 1 (chordin-like) |
| chr20_+_9128829 | 0.02 |
ENSDART00000064144
ENSDART00000137450 |
bpnt1
|
bisphosphate nucleotidase 1 |
| chr9_+_32859967 | 0.02 |
ENSDART00000168992
|
si:dkey-145p14.5
|
si:dkey-145p14.5 |
| chr22_+_7462997 | 0.02 |
ENSDART00000106082
|
zgc:112368
|
zgc:112368 |
| chr16_+_54263921 | 0.02 |
ENSDART00000002856
|
drd2l
|
dopamine receptor D2 like |
| chr23_+_23022790 | 0.02 |
ENSDART00000189843
|
samd11
|
sterile alpha motif domain containing 11 |
| chr16_+_6957582 | 0.02 |
ENSDART00000111308
ENSDART00000187541 ENSDART00000149987 |
bbs9
|
Bardet-Biedl syndrome 9 |
| chr20_-_47731768 | 0.02 |
ENSDART00000031167
|
tfap2d
|
transcription factor AP-2 delta (activating enhancer binding protein 2 delta) |
| chr22_+_37902976 | 0.02 |
ENSDART00000169327
|
kcnh1a
|
potassium voltage-gated channel, subfamily H (eag-related), member 1a |
| chr23_-_17657348 | 0.02 |
ENSDART00000054736
|
bhlhe23
|
basic helix-loop-helix family, member e23 |
| chr20_-_47188966 | 0.02 |
ENSDART00000152965
|
si:dkeyp-104f11.9
|
si:dkeyp-104f11.9 |
| chr19_-_42588510 | 0.02 |
ENSDART00000102583
|
sytl1
|
synaptotagmin-like 1 |
| chr5_+_37069605 | 0.02 |
ENSDART00000161977
|
si:dkeyp-110c7.4
|
si:dkeyp-110c7.4 |
| chr6_-_31336317 | 0.02 |
ENSDART00000193927
|
dnajc6
|
DnaJ (Hsp40) homolog, subfamily C, member 6 |
| chr14_+_29200772 | 0.02 |
ENSDART00000166608
|
TENM2
|
si:dkey-34l15.2 |
| chr8_+_52530889 | 0.02 |
ENSDART00000127729
ENSDART00000170360 ENSDART00000162687 |
stambpb
|
STAM binding protein b |
| chr20_+_33519435 | 0.02 |
ENSDART00000061829
|
drc1
|
dynein regulatory complex subunit 1 homolog (Chlamydomonas) |
| chr19_-_18135724 | 0.02 |
ENSDART00000186609
|
cbx3a
|
chromobox homolog 3a (HP1 gamma homolog, Drosophila) |
| chr20_+_9128256 | 0.02 |
ENSDART00000163883
ENSDART00000183072 ENSDART00000187276 |
bpnt1
|
bisphosphate nucleotidase 1 |
| chr18_-_16392229 | 0.01 |
ENSDART00000189031
|
mgat4c
|
mgat4 family, member C |
| chr5_+_37729207 | 0.01 |
ENSDART00000184378
|
cdc42ep2
|
CDC42 effector protein (Rho GTPase binding) 2 |
| chr1_+_21309386 | 0.01 |
ENSDART00000054442
ENSDART00000145661 |
med28
|
mediator complex subunit 28 |
| chr13_+_24755049 | 0.01 |
ENSDART00000134102
|
col17a1b
|
collagen, type XVII, alpha 1b |
| chr11_-_16093018 | 0.01 |
ENSDART00000139309
ENSDART00000139819 |
SPATA1
|
si:dkey-205k8.5 |
| chr10_+_24660225 | 0.01 |
ENSDART00000190695
|
vps36
|
vacuolar protein sorting 36 homolog (S. cerevisiae) |
| chr24_-_41220538 | 0.01 |
ENSDART00000150207
|
acvr2ba
|
activin A receptor type 2Ba |
| chr2_+_47906240 | 0.01 |
ENSDART00000122206
|
ftr23
|
finTRIM family, member 23 |
| chr25_+_19485198 | 0.01 |
ENSDART00000156730
|
glsl
|
glutaminase like |
| chr5_-_57528943 | 0.01 |
ENSDART00000130320
|
pisd
|
phosphatidylserine decarboxylase |
| chr15_-_15357178 | 0.01 |
ENSDART00000106120
|
ywhag2
|
3-monooxygenase/tryptophan 5-monooxygenase activation protein, gamma polypeptide 2 |
| chr3_-_32873641 | 0.01 |
ENSDART00000075277
|
zgc:113090
|
zgc:113090 |
| chr5_+_3118231 | 0.01 |
ENSDART00000179809
|
supt4h1
|
SPT4 homolog, DSIF elongation factor subunit |
| chr5_+_33289057 | 0.01 |
ENSDART00000123210
|
med22
|
mediator complex subunit 22 |
| chr10_-_39321367 | 0.01 |
ENSDART00000129647
|
smtlb
|
somatolactin beta |
| chr11_+_13024002 | 0.01 |
ENSDART00000104113
|
btf3l4
|
basic transcription factor 3-like 4 |
| chr14_-_17576662 | 0.01 |
ENSDART00000193893
|
rnf4
|
ring finger protein 4 |
| chr7_+_56735195 | 0.01 |
ENSDART00000082830
|
KIAA0895L
|
KIAA0895 like |
| chr12_+_46696867 | 0.01 |
ENSDART00000152928
ENSDART00000153445 ENSDART00000123357 ENSDART00000152880 ENSDART00000123834 ENSDART00000174765 ENSDART00000189923 |
exoc7
|
exocyst complex component 7 |
| chr6_-_57539141 | 0.01 |
ENSDART00000156967
|
itcha
|
itchy E3 ubiquitin protein ligase a |
| chr9_+_29520696 | 0.01 |
ENSDART00000144430
|
fdx1
|
ferredoxin 1 |
| chr18_+_30028637 | 0.01 |
ENSDART00000139750
|
si:ch211-220f16.1
|
si:ch211-220f16.1 |
| chr3_-_8399774 | 0.01 |
ENSDART00000190238
ENSDART00000113140 ENSDART00000132949 |
rbfox3b
|
RNA binding fox-1 homolog 3b |
| chr5_+_66433287 | 0.01 |
ENSDART00000170757
|
kntc1
|
kinetochore associated 1 |
| chr21_+_26389391 | 0.01 |
ENSDART00000077197
|
tmsb
|
thymosin, beta |
| chr23_+_38953545 | 0.01 |
ENSDART00000184621
|
ATP9A
|
ATPase phospholipid transporting 9A (putative) |
| chr21_-_41025340 | 0.01 |
ENSDART00000148231
|
plac8l1
|
PLAC8-like 1 |
| chr16_-_43233509 | 0.01 |
ENSDART00000025877
|
cldn12
|
claudin 12 |
| chr17_-_24439672 | 0.01 |
ENSDART00000155020
|
wdpcp
|
WD repeat containing planar cell polarity effector |
| chr11_+_23760470 | 0.01 |
ENSDART00000175688
ENSDART00000121874 ENSDART00000086720 |
nfasca
|
neurofascin homolog (chicken) a |
| chr13_-_25198025 | 0.01 |
ENSDART00000159585
ENSDART00000144227 |
adka
|
adenosine kinase a |
Network of associatons between targets according to the STRING database.
First level regulatory network of sox11a+sox11b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0015824 | proline transport(GO:0015824) |
| 0.0 | 0.0 | GO:0045671 | negative regulation of osteoclast differentiation(GO:0045671) |
| 0.0 | 0.1 | GO:0003404 | optic vesicle morphogenesis(GO:0003404) |
| 0.0 | 0.0 | GO:0007529 | establishment of synaptic specificity at neuromuscular junction(GO:0007529) molybdenum incorporation into molybdenum-molybdopterin complex(GO:0018315) metal incorporation into metallo-molybdopterin complex(GO:0042040) glycine receptor clustering(GO:0072579) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0004771 | sterol esterase activity(GO:0004771) |
| 0.0 | 0.0 | GO:0061598 | molybdopterin adenylyltransferase activity(GO:0061598) molybdopterin molybdotransferase activity(GO:0061599) |
| 0.0 | 0.0 | GO:0050135 | NAD(P)+ nucleosidase activity(GO:0050135) |