Project
PRJNA207719: Tissue specific transcriptome profiling
Navigation
Downloads
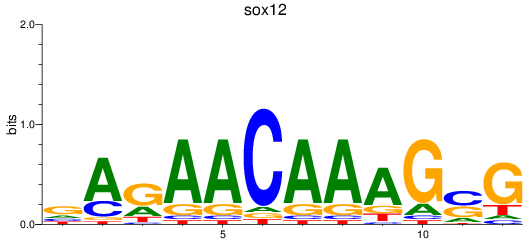
Results for sox12
Z-value: 1.11
Transcription factors associated with sox12
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
sox12
|
ENSDARG00000025847 | SRY-box transcription factor 12 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| sox12 | dr11_v1_chr11_-_24191928_24191928 | 0.75 | 1.5e-01 | Click! |
Activity profile of sox12 motif
Sorted Z-values of sox12 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr20_-_39265453 | 0.60 |
ENSDART00000141193
|
clu
|
clusterin |
| chr4_-_2525916 | 0.51 |
ENSDART00000134123
ENSDART00000132581 ENSDART00000019508 |
csrp2
|
cysteine and glycine-rich protein 2 |
| chr20_-_2355357 | 0.51 |
ENSDART00000085281
|
EPB41L2
|
si:ch73-18b11.1 |
| chr22_-_37349967 | 0.49 |
ENSDART00000104493
|
sox2
|
SRY (sex determining region Y)-box 2 |
| chr22_-_31517300 | 0.47 |
ENSDART00000164799
|
slc6a6b
|
solute carrier family 6 (neurotransmitter transporter), member 6b |
| chr14_-_8080416 | 0.45 |
ENSDART00000045109
|
zgc:92242
|
zgc:92242 |
| chr15_+_1148074 | 0.43 |
ENSDART00000152638
ENSDART00000152466 ENSDART00000188011 |
mlf1
|
myeloid leukemia factor 1 |
| chr2_-_44280061 | 0.43 |
ENSDART00000136818
|
mpz
|
myelin protein zero |
| chr19_-_4010263 | 0.40 |
ENSDART00000159605
ENSDART00000165541 |
map7d1b
|
MAP7 domain containing 1b |
| chr10_-_1718395 | 0.39 |
ENSDART00000137620
|
si:ch73-46j18.5
|
si:ch73-46j18.5 |
| chr18_-_14337450 | 0.39 |
ENSDART00000061435
|
hsbp1b
|
heat shock factor binding protein 1b |
| chr10_+_26834985 | 0.37 |
ENSDART00000147518
|
fam89b
|
family with sequence similarity 89, member B |
| chr5_-_38384755 | 0.36 |
ENSDART00000188573
ENSDART00000051233 |
mink1
|
misshapen-like kinase 1 |
| chr25_+_28825657 | 0.35 |
ENSDART00000153625
|
nfybb
|
nuclear transcription factor Y, beta b |
| chr7_-_38340674 | 0.34 |
ENSDART00000075782
|
slc7a10a
|
solute carrier family 7 (neutral amino acid transporter light chain, asc system), member 10a |
| chr11_+_24820542 | 0.34 |
ENSDART00000135443
|
kdm5ba
|
lysine (K)-specific demethylase 5Ba |
| chr17_+_37624704 | 0.34 |
ENSDART00000157001
|
tmem229b
|
transmembrane protein 229B |
| chr4_+_21129752 | 0.34 |
ENSDART00000169764
|
syt1a
|
synaptotagmin Ia |
| chr16_-_9897769 | 0.33 |
ENSDART00000187044
|
ncalda
|
neurocalcin delta a |
| chr25_+_25124684 | 0.32 |
ENSDART00000167542
|
ldha
|
lactate dehydrogenase A4 |
| chr24_+_20920563 | 0.32 |
ENSDART00000037224
|
cst14a.2
|
cystatin 14a, tandem duplicate 2 |
| chr14_-_33297287 | 0.32 |
ENSDART00000045555
ENSDART00000138294 ENSDART00000075056 |
rab41
|
RAB41, member RAS oncogene family |
| chr3_-_1146497 | 0.30 |
ENSDART00000149061
|
si:ch73-211l13.2
|
si:ch73-211l13.2 |
| chr8_+_42998944 | 0.30 |
ENSDART00000048819
|
rassf2a
|
Ras association (RalGDS/AF-6) domain family member 2a |
| chr22_+_21618121 | 0.30 |
ENSDART00000133939
|
tle2a
|
transducin like enhancer of split 2a |
| chr25_-_12788370 | 0.30 |
ENSDART00000158551
|
slc7a5
|
solute carrier family 7 (amino acid transporter light chain, L system), member 5 |
| chr6_-_13022166 | 0.29 |
ENSDART00000157139
|
tmbim1a
|
transmembrane BAX inhibitor motif containing 1a |
| chr7_+_25003313 | 0.29 |
ENSDART00000131935
|
naa40
|
N(alpha)-acetyltransferase 40, NatD catalytic subunit, homolog (S. cerevisiae) |
| chr2_+_24188502 | 0.28 |
ENSDART00000181955
ENSDART00000191706 ENSDART00000125909 |
map4l
|
microtubule associated protein 4 like |
| chr2_-_24348948 | 0.28 |
ENSDART00000136559
|
ano8a
|
anoctamin 8a |
| chr20_+_30445971 | 0.27 |
ENSDART00000153150
|
myt1la
|
myelin transcription factor 1-like, a |
| chr2_-_7696503 | 0.27 |
ENSDART00000169709
|
CABZ01055306.1
|
|
| chr8_-_2153147 | 0.27 |
ENSDART00000124093
|
si:dkeyp-117b11.1
|
si:dkeyp-117b11.1 |
| chr24_+_7637522 | 0.27 |
ENSDART00000082467
|
cavin1b
|
caveolae associated protein 1b |
| chr2_-_16217344 | 0.27 |
ENSDART00000152031
|
arhgef4
|
Rho guanine nucleotide exchange factor (GEF) 4 |
| chr12_+_5081759 | 0.27 |
ENSDART00000164178
|
prrt2
|
proline-rich transmembrane protein 2 |
| chr1_-_38361496 | 0.26 |
ENSDART00000015323
|
fbxo8
|
F-box protein 8 |
| chr23_-_7755373 | 0.26 |
ENSDART00000162868
|
PCMTD2 (1 of many)
|
protein-L-isoaspartate (D-aspartate) O-methyltransferase domain containing 2 |
| chr21_+_36774542 | 0.26 |
ENSDART00000130790
|
prr7
|
proline rich 7 (synaptic) |
| chr16_-_9897613 | 0.25 |
ENSDART00000104058
|
ncalda
|
neurocalcin delta a |
| chr8_+_26859639 | 0.24 |
ENSDART00000133440
|
prdm2a
|
PR domain containing 2, with ZNF domain a |
| chr5_+_26221011 | 0.24 |
ENSDART00000007587
|
gtf2h2
|
general transcription factor IIH, polypeptide 2 |
| chr7_+_69470442 | 0.23 |
ENSDART00000189593
|
gabarapb
|
GABA(A) receptor-associated protein b |
| chr23_+_9268083 | 0.23 |
ENSDART00000055054
|
acss2
|
acyl-CoA synthetase short chain family member 2 |
| chr10_-_32660716 | 0.23 |
ENSDART00000063544
ENSDART00000141013 |
atg101
|
autophagy related 101 |
| chr3_-_34027178 | 0.23 |
ENSDART00000170201
ENSDART00000151408 |
ighv1-4
ighv14-1
|
immunoglobulin heavy variable 1-4 immunoglobulin heavy variable 14-1 |
| chr14_+_48960078 | 0.22 |
ENSDART00000165514
|
mif
|
macrophage migration inhibitory factor |
| chr25_+_10485103 | 0.22 |
ENSDART00000104576
|
psmd13
|
proteasome 26S subunit, non-ATPase 13 |
| chr6_+_28877306 | 0.22 |
ENSDART00000065137
ENSDART00000123189 ENSDART00000065135 ENSDART00000181512 ENSDART00000130799 |
tp63
|
tumor protein p63 |
| chr1_-_44701313 | 0.22 |
ENSDART00000193926
|
si:dkey-28b4.8
|
si:dkey-28b4.8 |
| chr19_+_8481802 | 0.22 |
ENSDART00000050217
|
efna1a
|
ephrin-A1a |
| chr2_-_24348642 | 0.22 |
ENSDART00000181739
|
ano8a
|
anoctamin 8a |
| chr3_+_41922114 | 0.21 |
ENSDART00000138280
|
lfng
|
LFNG O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase |
| chr23_-_36449111 | 0.21 |
ENSDART00000110478
|
zgc:174906
|
zgc:174906 |
| chr10_-_25591194 | 0.21 |
ENSDART00000131640
|
tiam1a
|
T cell lymphoma invasion and metastasis 1a |
| chr6_+_27667359 | 0.20 |
ENSDART00000159624
ENSDART00000049177 |
rab6ba
|
RAB6B, member RAS oncogene family a |
| chr4_-_18939338 | 0.20 |
ENSDART00000132081
ENSDART00000042250 |
rap1b
|
RAP1B, member of RAS oncogene family |
| chr14_-_1955257 | 0.20 |
ENSDART00000193254
|
pcdh2g5
|
protocadherin 2 gamma 5 |
| chr2_+_36862473 | 0.20 |
ENSDART00000135624
|
si:dkey-193b15.8
|
si:dkey-193b15.8 |
| chr3_-_33437796 | 0.20 |
ENSDART00000075499
|
si:dkey-283b1.7
|
si:dkey-283b1.7 |
| chr12_+_10163585 | 0.19 |
ENSDART00000106191
|
psmc5
|
proteasome 26S subunit, ATPase 5 |
| chr4_-_68913650 | 0.19 |
ENSDART00000184297
|
si:dkey-264f17.5
|
si:dkey-264f17.5 |
| chr10_-_42131408 | 0.19 |
ENSDART00000076693
|
stambpa
|
STAM binding protein a |
| chr9_+_8929764 | 0.19 |
ENSDART00000102562
|
ankrd10b
|
ankyrin repeat domain 10b |
| chr5_-_38384289 | 0.19 |
ENSDART00000135260
|
mink1
|
misshapen-like kinase 1 |
| chr24_+_7637264 | 0.18 |
ENSDART00000124409
|
cavin1b
|
caveolae associated protein 1b |
| chr22_-_784110 | 0.18 |
ENSDART00000061775
|
rbbp5
|
retinoblastoma binding protein 5 |
| chr19_-_19339285 | 0.18 |
ENSDART00000158413
ENSDART00000170479 |
cspg5b
|
chondroitin sulfate proteoglycan 5b |
| chr5_-_37875636 | 0.18 |
ENSDART00000184674
|
arhgap35b
|
Rho GTPase activating protein 35b |
| chr9_-_31278048 | 0.18 |
ENSDART00000022204
|
zic5
|
zic family member 5 (odd-paired homolog, Drosophila) |
| chr23_+_19814371 | 0.18 |
ENSDART00000182897
|
emd
|
emerin (Emery-Dreifuss muscular dystrophy) |
| chr3_+_40284598 | 0.18 |
ENSDART00000009411
|
bud31
|
BUD31 homolog (S. cerevisiae) |
| chr20_+_4221978 | 0.18 |
ENSDART00000171898
|
ipcef1
|
interaction protein for cytohesin exchange factors 1 |
| chr19_+_20708460 | 0.17 |
ENSDART00000034124
|
rab5aa
|
RAB5A, member RAS oncogene family, a |
| chr15_+_37545855 | 0.17 |
ENSDART00000099456
|
psenen
|
presenilin enhancer gamma secretase subunit |
| chr14_-_48033073 | 0.17 |
ENSDART00000193115
ENSDART00000169300 ENSDART00000188036 ENSDART00000183432 ENSDART00000180973 |
rapgef2
|
Rap guanine nucleotide exchange factor (GEF) 2 |
| chr14_-_1958994 | 0.17 |
ENSDART00000161783
|
pcdh2g5
|
protocadherin 2 gamma 5 |
| chr1_-_26071535 | 0.17 |
ENSDART00000185187
|
pdcd4a
|
programmed cell death 4a |
| chr23_-_637347 | 0.17 |
ENSDART00000132175
|
l1camb
|
L1 cell adhesion molecule, paralog b |
| chr15_+_37546391 | 0.17 |
ENSDART00000186625
|
psenen
|
presenilin enhancer gamma secretase subunit |
| chr14_+_17376940 | 0.16 |
ENSDART00000054590
ENSDART00000010148 |
spon2b
|
spondin 2b, extracellular matrix protein |
| chr1_+_594736 | 0.16 |
ENSDART00000166731
|
jam2a
|
junctional adhesion molecule 2a |
| chr11_-_21267486 | 0.16 |
ENSDART00000128681
|
DYRK3
|
zgc:172180 |
| chr21_+_22845317 | 0.16 |
ENSDART00000065555
|
birc2
|
baculoviral IAP repeat containing 2 |
| chr3_-_16110100 | 0.16 |
ENSDART00000051807
|
lasp1
|
LIM and SH3 protein 1 |
| chr17_-_21057617 | 0.15 |
ENSDART00000148095
ENSDART00000048853 |
ube2d1a
|
ubiquitin-conjugating enzyme E2D 1a |
| chr16_+_25343934 | 0.15 |
ENSDART00000109710
|
zfat
|
zinc finger and AT hook domain containing |
| chr5_-_32363372 | 0.15 |
ENSDART00000098045
|
gas1b
|
growth arrest-specific 1b |
| chr6_-_55347616 | 0.15 |
ENSDART00000172081
|
pcif1
|
PDX1 C-terminal inhibiting factor 1 |
| chr3_-_16110351 | 0.15 |
ENSDART00000064838
|
lasp1
|
LIM and SH3 protein 1 |
| chr1_+_26071869 | 0.14 |
ENSDART00000059264
|
mxd4
|
MAX dimerization protein 4 |
| chr18_-_46063773 | 0.14 |
ENSDART00000078561
|
si:ch73-262h23.4
|
si:ch73-262h23.4 |
| chr16_-_16237844 | 0.14 |
ENSDART00000168747
ENSDART00000111912 |
rbm12b
|
RNA binding motif protein 12B |
| chr18_-_11595567 | 0.14 |
ENSDART00000098565
|
CRACR2A
|
calcium release activated channel regulator 2A |
| chr7_-_38861741 | 0.13 |
ENSDART00000173629
ENSDART00000037361 ENSDART00000173953 |
phf21aa
|
PHD finger protein 21Aa |
| chr17_+_25290136 | 0.13 |
ENSDART00000173295
|
kbtbd11
|
kelch repeat and BTB (POZ) domain containing 11 |
| chr16_+_12812472 | 0.13 |
ENSDART00000008535
|
u2af2a
|
U2 small nuclear RNA auxiliary factor 2a |
| chr10_+_21677058 | 0.13 |
ENSDART00000171499
ENSDART00000157516 |
pcdh1gb2
|
protocadherin 1 gamma b 2 |
| chr5_+_58455488 | 0.12 |
ENSDART00000038602
ENSDART00000127958 |
slc37a2
|
solute carrier family 37 (glucose-6-phosphate transporter), member 2 |
| chr8_+_16990120 | 0.12 |
ENSDART00000018934
|
pde4d
|
phosphodiesterase 4D, cAMP-specific |
| chr1_+_29798140 | 0.12 |
ENSDART00000019743
|
tm9sf2
|
transmembrane 9 superfamily member 2 |
| chr3_-_33175583 | 0.12 |
ENSDART00000126022
|
rarab
|
retinoic acid receptor, alpha b |
| chr3_-_60856157 | 0.12 |
ENSDART00000053502
|
CABZ01087513.1
|
|
| chr16_+_32082547 | 0.12 |
ENSDART00000190122
|
prpf31
|
PRP31 pre-mRNA processing factor 31 homolog (yeast) |
| chr10_+_43189325 | 0.12 |
ENSDART00000185584
|
vcanb
|
versican b |
| chr5_+_45138934 | 0.11 |
ENSDART00000041412
ENSDART00000136002 |
smarca2
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 |
| chr15_+_35691783 | 0.11 |
ENSDART00000183994
|
CT573366.1
|
|
| chr6_+_36942966 | 0.11 |
ENSDART00000028895
|
negr1
|
neuronal growth regulator 1 |
| chr17_-_7436766 | 0.11 |
ENSDART00000162002
|
grm1b
|
glutamate receptor, metabotropic 1b |
| chr6_+_18367388 | 0.11 |
ENSDART00000163394
|
dgke
|
diacylglycerol kinase, epsilon |
| chr17_+_45654724 | 0.11 |
ENSDART00000098952
|
zgc:162184
|
zgc:162184 |
| chr14_-_2004291 | 0.11 |
ENSDART00000114039
|
pcdh2g5
|
protocadherin 2 gamma 5 |
| chr17_-_26507289 | 0.11 |
ENSDART00000155616
|
ccser2a
|
coiled-coil serine-rich protein 2a |
| chr10_+_5268054 | 0.10 |
ENSDART00000114491
|
ror2
|
receptor tyrosine kinase-like orphan receptor 2 |
| chr24_+_36392040 | 0.10 |
ENSDART00000136057
|
arf2b
|
ADP-ribosylation factor 2b |
| chr8_+_9149436 | 0.10 |
ENSDART00000064090
|
pnck
|
pregnancy up-regulated nonubiquitous CaM kinase |
| chr7_+_24520518 | 0.10 |
ENSDART00000173604
|
btr09
|
bloodthirsty-related gene family, member 9 |
| chr14_-_2933185 | 0.10 |
ENSDART00000161677
ENSDART00000162446 ENSDART00000109378 |
si:dkey-201i24.6
|
si:dkey-201i24.6 |
| chr9_+_24920677 | 0.10 |
ENSDART00000037025
|
slc39a10
|
solute carrier family 39 (zinc transporter), member 10 |
| chr5_-_24231139 | 0.09 |
ENSDART00000143492
|
senp3a
|
SUMO1/sentrin/SMT3 specific peptidase 3a |
| chr7_+_49078890 | 0.09 |
ENSDART00000067857
|
b4galnt4b
|
beta-1,4-N-acetyl-galactosaminyl transferase 4b |
| chr8_-_16697615 | 0.09 |
ENSDART00000187929
|
rpe65b
|
retinal pigment epithelium-specific protein 65b |
| chr25_+_33063762 | 0.09 |
ENSDART00000189974
|
tln2b
|
talin 2b |
| chr4_-_13567387 | 0.09 |
ENSDART00000132971
ENSDART00000102010 |
mdm1
|
Mdm1 nuclear protein homolog (mouse) |
| chr21_+_31150438 | 0.09 |
ENSDART00000065366
|
st6gal1
|
ST6 beta-galactosamide alpha-2,6-sialyltranferase 1 |
| chr8_+_54135642 | 0.09 |
ENSDART00000170712
|
brpf1
|
bromodomain and PHD finger containing, 1 |
| chr8_-_43456025 | 0.09 |
ENSDART00000001092
ENSDART00000140618 |
ncor2
|
nuclear receptor corepressor 2 |
| chr7_-_28565230 | 0.09 |
ENSDART00000028887
|
tmem9b
|
TMEM9 domain family, member B |
| chr2_-_43130812 | 0.09 |
ENSDART00000039079
ENSDART00000056162 |
ftr13
|
finTRIM family, member 13 |
| chr10_+_10972795 | 0.09 |
ENSDART00000127331
|
cdc37l1
|
cell division cycle 37-like 1 |
| chr16_+_11242443 | 0.08 |
ENSDART00000024935
|
gsk3ab
|
glycogen synthase kinase 3 alpha b |
| chr4_-_4834617 | 0.07 |
ENSDART00000141539
|
coa6
|
cytochrome c oxidase assembly factor 6 |
| chr5_+_337215 | 0.07 |
ENSDART00000167982
|
rnf170
|
ring finger protein 170 |
| chr20_+_4222357 | 0.07 |
ENSDART00000188331
|
ipcef1
|
interaction protein for cytohesin exchange factors 1 |
| chr8_-_19313510 | 0.07 |
ENSDART00000164780
ENSDART00000137133 |
rgl1
|
ral guanine nucleotide dissociation stimulator-like 1 |
| chr19_-_38830582 | 0.07 |
ENSDART00000189966
ENSDART00000183055 |
adgrb2
|
adhesion G protein-coupled receptor B2 |
| chr1_-_52790724 | 0.07 |
ENSDART00000139577
ENSDART00000100937 |
patl1
|
protein associated with topoisomerase II homolog 1 (yeast) |
| chr7_-_30217000 | 0.07 |
ENSDART00000173646
|
znf710b
|
zinc finger protein 710b |
| chr6_+_20647155 | 0.07 |
ENSDART00000193477
|
slc19a1
|
solute carrier family 19 (folate transporter), member 1 |
| chr20_-_29051696 | 0.07 |
ENSDART00000140350
|
thbs1b
|
thrombospondin 1b |
| chr21_+_30549512 | 0.06 |
ENSDART00000132831
|
rab38c
|
RAB38c, member of RAS oncogene family |
| chr22_-_16494406 | 0.06 |
ENSDART00000062727
|
stx6
|
syntaxin 6 |
| chr10_-_35321625 | 0.06 |
ENSDART00000131268
|
rhbdd2
|
rhomboid domain containing 2 |
| chr18_+_22138924 | 0.06 |
ENSDART00000183961
|
ripor1
|
RHO family interacting cell polarization regulator 1 |
| chr16_+_32082359 | 0.06 |
ENSDART00000140794
ENSDART00000137029 |
prpf31
|
PRP31 pre-mRNA processing factor 31 homolog (yeast) |
| chr18_+_6479963 | 0.06 |
ENSDART00000092752
ENSDART00000136333 |
wash1
|
WAS protein family homolog 1 |
| chr6_+_11829867 | 0.05 |
ENSDART00000151044
|
baz2ba
|
bromodomain adjacent to zinc finger domain, 2Ba |
| chr13_+_9559461 | 0.05 |
ENSDART00000047740
|
wdr32
|
WD repeat domain 32 |
| chr23_+_30707837 | 0.05 |
ENSDART00000016096
|
dnajc11a
|
DnaJ (Hsp40) homolog, subfamily C, member 11a |
| chr12_+_4160804 | 0.05 |
ENSDART00000152515
|
itgam
|
integrin, alpha M (complement component 3 receptor 3 subunit) |
| chr11_+_37768298 | 0.05 |
ENSDART00000166886
|
sox13
|
SRY (sex determining region Y)-box 13 |
| chr23_+_21544227 | 0.04 |
ENSDART00000140253
|
arhgef10lb
|
Rho guanine nucleotide exchange factor (GEF) 10-like b |
| chr25_-_31171948 | 0.03 |
ENSDART00000184275
|
FQ976913.1
|
|
| chr1_+_47331644 | 0.03 |
ENSDART00000053284
|
bcl9
|
B cell CLL/lymphoma 9 |
| chr15_-_35930070 | 0.03 |
ENSDART00000076229
|
irs1
|
insulin receptor substrate 1 |
| chr4_+_15954293 | 0.03 |
ENSDART00000132695
|
si:dkey-117n7.4
|
si:dkey-117n7.4 |
| chr14_-_2036604 | 0.03 |
ENSDART00000192446
|
BX005294.2
|
|
| chr25_+_22017182 | 0.03 |
ENSDART00000156517
|
si:dkey-217l24.1
|
si:dkey-217l24.1 |
| chr1_+_46509176 | 0.03 |
ENSDART00000166028
|
mcf2la
|
mcf.2 cell line derived transforming sequence-like a |
| chr1_+_45839927 | 0.03 |
ENSDART00000148086
ENSDART00000180413 ENSDART00000048191 ENSDART00000179047 |
map2k7
|
mitogen-activated protein kinase kinase 7 |
| chr14_-_2318590 | 0.02 |
ENSDART00000192735
|
pcdh2ab8
|
protocadherin 2 alpha b 8 |
| chr7_+_24729558 | 0.02 |
ENSDART00000111542
ENSDART00000170100 |
shroom4
|
shroom family member 4 |
| chr12_+_28995942 | 0.02 |
ENSDART00000076334
|
valopb
|
vertebrate ancient long opsin b |
| chr9_+_537297 | 0.02 |
ENSDART00000165149
|
CU984600.3
|
|
| chr8_-_52229462 | 0.02 |
ENSDART00000185949
ENSDART00000051825 |
tcf7l1b
|
transcription factor 7 like 1b |
| chr17_-_23412705 | 0.01 |
ENSDART00000126995
|
si:ch211-149k12.3
|
si:ch211-149k12.3 |
| chr15_+_26933196 | 0.01 |
ENSDART00000023842
|
ppm1da
|
protein phosphatase, Mg2+/Mn2+ dependent, 1Da |
| chr1_+_33668236 | 0.01 |
ENSDART00000122316
ENSDART00000102184 |
arl13b
|
ADP-ribosylation factor-like 13b |
| chr19_+_20724347 | 0.01 |
ENSDART00000090757
|
kat2b
|
K(lysine) acetyltransferase 2B |
| chr13_-_33317323 | 0.01 |
ENSDART00000110295
ENSDART00000144848 ENSDART00000136701 |
tmem234
|
transmembrane protein 234 |
| chr5_+_45139196 | 0.00 |
ENSDART00000113738
|
smarca2
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 |
| chr3_-_40284744 | 0.00 |
ENSDART00000018626
|
pdap1b
|
pdgfa associated protein 1b |
Network of associatons between targets according to the STRING database.
First level regulatory network of sox12
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0042940 | D-amino acid transport(GO:0042940) D-alanine transport(GO:0042941) D-serine transport(GO:0042942) |
| 0.1 | 0.5 | GO:0021982 | pineal gland development(GO:0021982) |
| 0.1 | 0.2 | GO:0048169 | regulation of long-term neuronal synaptic plasticity(GO:0048169) |
| 0.1 | 0.5 | GO:0006361 | transcription initiation from RNA polymerase I promoter(GO:0006361) |
| 0.0 | 0.1 | GO:0015712 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.0 | 0.2 | GO:0060547 | negative regulation of necroptotic process(GO:0060546) negative regulation of necrotic cell death(GO:0060547) |
| 0.0 | 0.2 | GO:2000301 | negative regulation of neurotransmitter secretion(GO:0046929) response to cAMP(GO:0051591) cellular response to cAMP(GO:0071320) negative regulation of synaptic vesicle transport(GO:1902804) negative regulation of synaptic vesicle exocytosis(GO:2000301) |
| 0.0 | 0.2 | GO:0006083 | acetate metabolic process(GO:0006083) acetyl-CoA biosynthetic process from acetate(GO:0019427) |
| 0.0 | 0.3 | GO:0034205 | Notch receptor processing(GO:0007220) beta-amyloid formation(GO:0034205) amyloid precursor protein catabolic process(GO:0042987) |
| 0.0 | 0.2 | GO:0055016 | hypochord development(GO:0055016) |
| 0.0 | 0.2 | GO:0045899 | regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0045898) positive regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0045899) |
| 0.0 | 0.1 | GO:0050861 | positive regulation of B cell receptor signaling pathway(GO:0050861) |
| 0.0 | 0.2 | GO:0000244 | spliceosomal tri-snRNP complex assembly(GO:0000244) |
| 0.0 | 0.2 | GO:0048730 | epidermis morphogenesis(GO:0048730) |
| 0.0 | 0.2 | GO:0033278 | cell proliferation in midbrain(GO:0033278) |
| 0.0 | 0.1 | GO:0051958 | folic acid transport(GO:0015884) drug transport(GO:0015893) methotrexate transport(GO:0051958) reduced folate transmembrane transport(GO:0098838) |
| 0.0 | 0.1 | GO:0031507 | heterochromatin assembly(GO:0031507) |
| 0.0 | 0.2 | GO:0099550 | trans-synaptic signalling, modulating synaptic transmission(GO:0099550) |
| 0.0 | 0.3 | GO:0006474 | N-terminal protein amino acid acetylation(GO:0006474) |
| 0.0 | 0.3 | GO:1901222 | regulation of NIK/NF-kappaB signaling(GO:1901222) |
| 0.0 | 0.5 | GO:0006891 | intra-Golgi vesicle-mediated transport(GO:0006891) |
| 0.0 | 0.3 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.0 | 0.2 | GO:0043248 | proteasome assembly(GO:0043248) |
| 0.0 | 0.1 | GO:0046600 | negative regulation of centriole replication(GO:0046600) |
| 0.0 | 0.5 | GO:0072348 | sulfur compound transport(GO:0072348) |
| 0.0 | 0.2 | GO:0006995 | cellular response to nitrogen starvation(GO:0006995) cellular response to nitrogen levels(GO:0043562) |
| 0.0 | 0.4 | GO:0014029 | neural crest formation(GO:0014029) |
| 0.0 | 0.1 | GO:0001957 | intramembranous ossification(GO:0001957) direct ossification(GO:0036072) |
| 0.0 | 0.1 | GO:0003139 | secondary heart field specification(GO:0003139) |
| 0.0 | 0.1 | GO:0000290 | deadenylation-dependent decapping of nuclear-transcribed mRNA(GO:0000290) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.9 | GO:0042583 | chromaffin granule(GO:0042583) |
| 0.0 | 0.2 | GO:0098842 | postsynaptic early endosome(GO:0098842) |
| 0.0 | 0.3 | GO:0070765 | gamma-secretase complex(GO:0070765) |
| 0.0 | 0.2 | GO:0000439 | core TFIIH complex(GO:0000439) |
| 0.0 | 0.3 | GO:0016602 | CCAAT-binding factor complex(GO:0016602) |
| 0.0 | 0.2 | GO:0031597 | cytosolic proteasome complex(GO:0031597) |
| 0.0 | 0.4 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.0 | 0.2 | GO:0008541 | proteasome regulatory particle, lid subcomplex(GO:0008541) |
| 0.0 | 0.2 | GO:0071005 | U2-type precatalytic spliceosome(GO:0071005) |
| 0.0 | 0.1 | GO:0089701 | U2AF(GO:0089701) |
| 0.0 | 0.2 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.0 | 0.1 | GO:0071203 | WASH complex(GO:0071203) |
| 0.0 | 0.5 | GO:0005901 | caveola(GO:0005901) plasma membrane raft(GO:0044853) |
| 0.0 | 0.2 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0042134 | rRNA primary transcript binding(GO:0042134) |
| 0.1 | 0.3 | GO:0043998 | H2A histone acetyltransferase activity(GO:0043998) |
| 0.1 | 0.2 | GO:0033829 | O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase activity(GO:0033829) |
| 0.0 | 0.2 | GO:0050218 | propionate-CoA ligase activity(GO:0050218) |
| 0.0 | 0.3 | GO:0004459 | L-lactate dehydrogenase activity(GO:0004459) |
| 0.0 | 0.1 | GO:0061513 | hexose phosphate transmembrane transporter activity(GO:0015119) organophosphate:inorganic phosphate antiporter activity(GO:0015315) hexose-phosphate:inorganic phosphate antiporter activity(GO:0015526) glucose 6-phosphate:inorganic phosphate antiporter activity(GO:0061513) |
| 0.0 | 0.1 | GO:0016422 | mRNA (2'-O-methyladenosine-N6-)-methyltransferase activity(GO:0016422) |
| 0.0 | 0.3 | GO:0034647 | histone demethylase activity (H3-trimethyl-K4 specific)(GO:0034647) |
| 0.0 | 0.6 | GO:0051787 | misfolded protein binding(GO:0051787) |
| 0.0 | 0.1 | GO:0015350 | reduced folate carrier activity(GO:0008518) methotrexate transporter activity(GO:0015350) |
| 0.0 | 0.5 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.0 | 0.1 | GO:0033842 | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N-acetylgalactosaminyltransferase activity(GO:0033842) |
| 0.0 | 0.1 | GO:0099530 | G-protein coupled receptor activity involved in regulation of postsynaptic membrane potential(GO:0099530) |
| 0.0 | 0.2 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.0 | 0.1 | GO:0030628 | pre-mRNA 3'-splice site binding(GO:0030628) |
| 0.0 | 0.2 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) cysteine-type endopeptidase regulator activity involved in apoptotic process(GO:0043028) |
| 0.0 | 0.5 | GO:1901682 | sulfur compound transmembrane transporter activity(GO:1901682) |
| 0.0 | 0.2 | GO:0017025 | TBP-class protein binding(GO:0017025) |
| 0.0 | 0.2 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.0 | 0.3 | GO:0004869 | cysteine-type endopeptidase inhibitor activity(GO:0004869) |
| 0.0 | 0.2 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.0 | 0.1 | GO:0003835 | beta-galactoside alpha-2,6-sialyltransferase activity(GO:0003835) |
| 0.0 | 0.2 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.0 | 0.2 | GO:0046875 | ephrin receptor binding(GO:0046875) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.0 | 0.5 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.0 | 1.0 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.0 | 0.2 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.0 | 0.4 | PID ATF2 PATHWAY | ATF-2 transcription factor network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.0 | 0.3 | REACTOME SIGNALING BY NOTCH4 | Genes involved in Signaling by NOTCH4 |
| 0.0 | 0.2 | REACTOME RAP1 SIGNALLING | Genes involved in Rap1 signalling |
| 0.0 | 0.3 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |
| 0.0 | 0.3 | REACTOME PYRUVATE METABOLISM | Genes involved in Pyruvate metabolism |
| 0.0 | 0.2 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.0 | 0.2 | REACTOME RNA POL I TRANSCRIPTION TERMINATION | Genes involved in RNA Polymerase I Transcription Termination |
| 0.0 | 0.1 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.0 | 0.4 | REACTOME CROSS PRESENTATION OF SOLUBLE EXOGENOUS ANTIGENS ENDOSOMES | Genes involved in Cross-presentation of soluble exogenous antigens (endosomes) |