Project
PRJNA207719: Tissue specific transcriptome profiling
Navigation
Downloads
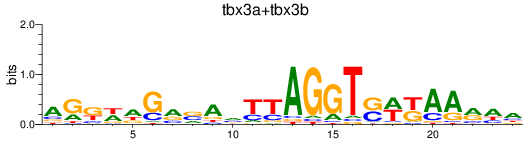
Results for tbx3a+tbx3b
Z-value: 1.26
Transcription factors associated with tbx3a+tbx3b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
tbx3a
|
ENSDARG00000002216 | T-box transcription factor 3a |
|
tbx3b
|
ENSDARG00000061509 | T-box transcription factor 3b |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| tbx3b | dr11_v1_chr5_+_26913120_26913120 | -0.94 | 1.7e-02 | Click! |
| tbx3a | dr11_v1_chr5_-_72324371_72324371 | -0.82 | 8.7e-02 | Click! |
Activity profile of tbx3a+tbx3b motif
Sorted Z-values of tbx3a+tbx3b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr25_-_13188678 | 1.55 |
ENSDART00000125754
|
si:ch211-147m6.1
|
si:ch211-147m6.1 |
| chr18_+_5547185 | 1.41 |
ENSDART00000193977
|
nnt2
|
nicotinamide nucleotide transhydrogenase 2 |
| chr13_-_37180815 | 1.28 |
ENSDART00000139907
|
si:dkeyp-77c8.1
|
si:dkeyp-77c8.1 |
| chr23_-_5683147 | 1.28 |
ENSDART00000102766
ENSDART00000067351 |
tnnt2a
|
troponin T type 2a (cardiac) |
| chr20_+_53577502 | 1.00 |
ENSDART00000126983
|
myh6
|
myosin, heavy chain 6, cardiac muscle, alpha |
| chr7_-_73752955 | 0.84 |
ENSDART00000171254
ENSDART00000009888 |
casq1b
|
calsequestrin 1b |
| chr3_-_33967767 | 0.84 |
ENSDART00000151493
ENSDART00000151160 |
ighv1-4
|
immunoglobulin heavy variable 1-4 |
| chr12_-_26415499 | 0.83 |
ENSDART00000185779
|
synpo2lb
|
synaptopodin 2-like b |
| chr13_-_51903150 | 0.83 |
ENSDART00000090644
|
mrpl2
|
mitochondrial ribosomal protein L2 |
| chr25_-_27819838 | 0.82 |
ENSDART00000067106
|
lmod2a
|
leiomodin 2 (cardiac) a |
| chr1_-_45177373 | 0.81 |
ENSDART00000143142
ENSDART00000034549 |
zgc:111983
|
zgc:111983 |
| chr2_-_37103622 | 0.67 |
ENSDART00000137849
|
zgc:101744
|
zgc:101744 |
| chr3_+_5297493 | 0.64 |
ENSDART00000138596
|
si:ch211-150d5.3
|
si:ch211-150d5.3 |
| chr21_-_27195256 | 0.57 |
ENSDART00000133152
ENSDART00000065401 |
zgc:110782
|
zgc:110782 |
| chr18_+_29145681 | 0.54 |
ENSDART00000089031
ENSDART00000193336 |
ppfibp2a
|
PTPRF interacting protein, binding protein 2a (liprin beta 2) |
| chr22_+_344763 | 0.51 |
ENSDART00000181934
|
CU914780.1
|
|
| chr22_+_9294336 | 0.51 |
ENSDART00000193694
|
si:ch211-250k18.7
|
si:ch211-250k18.7 |
| chr9_-_105135 | 0.51 |
ENSDART00000180126
|
FQ377903.3
|
|
| chr16_-_41990421 | 0.50 |
ENSDART00000055921
|
pycard
|
PYD and CARD domain containing |
| chr24_+_854877 | 0.50 |
ENSDART00000188408
|
piezo2a.2
|
piezo-type mechanosensitive ion channel component 2a, tandem duplicate 2 |
| chr2_-_21438492 | 0.47 |
ENSDART00000046098
|
plcd1b
|
phospholipase C, delta 1b |
| chr13_-_11984867 | 0.46 |
ENSDART00000157538
|
npm3
|
nucleophosmin/nucleoplasmin, 3 |
| chr14_-_14659023 | 0.46 |
ENSDART00000170355
ENSDART00000159888 ENSDART00000172241 |
nsdhl
|
NAD(P) dependent steroid dehydrogenase-like |
| chr17_-_50071748 | 0.45 |
ENSDART00000075188
|
zgc:113886
|
zgc:113886 |
| chr19_-_18127808 | 0.42 |
ENSDART00000108627
|
snx10a
|
sorting nexin 10a |
| chr5_-_38197080 | 0.41 |
ENSDART00000140708
|
si:ch211-284e13.9
|
si:ch211-284e13.9 |
| chr17_-_47090440 | 0.41 |
ENSDART00000163542
|
CABZ01056321.1
|
|
| chr22_-_5441893 | 0.40 |
ENSDART00000161421
|
zgc:194627
|
zgc:194627 |
| chr24_-_26479841 | 0.39 |
ENSDART00000079984
|
rpl22l1
|
ribosomal protein L22-like 1 |
| chr5_+_29803380 | 0.38 |
ENSDART00000005263
ENSDART00000137558 ENSDART00000146963 ENSDART00000134900 |
usf1l
|
upstream transcription factor 1, like |
| chr6_-_1514767 | 0.37 |
ENSDART00000067586
|
chchd6b
|
coiled-coil-helix-coiled-coil-helix domain containing 6b |
| chr7_+_65121743 | 0.36 |
ENSDART00000108471
|
ipo7
|
importin 7 |
| chr16_-_50918908 | 0.36 |
ENSDART00000108744
|
CU855821.1
|
|
| chr23_+_2421689 | 0.35 |
ENSDART00000180200
|
tcp1
|
t-complex 1 |
| chr1_+_56502706 | 0.34 |
ENSDART00000188665
|
CABZ01059403.1
|
|
| chr19_-_18127629 | 0.33 |
ENSDART00000187722
|
snx10a
|
sorting nexin 10a |
| chr10_-_15048781 | 0.33 |
ENSDART00000038401
ENSDART00000155674 |
si:ch211-95j8.2
|
si:ch211-95j8.2 |
| chr5_+_29803702 | 0.32 |
ENSDART00000147569
|
usf1l
|
upstream transcription factor 1, like |
| chr19_-_10810006 | 0.32 |
ENSDART00000151157
|
si:dkey-3n22.9
|
si:dkey-3n22.9 |
| chr4_+_14981854 | 0.32 |
ENSDART00000067046
|
cax1
|
cation/H+ exchanger protein 1 |
| chr7_-_3836635 | 0.32 |
ENSDART00000104350
|
si:dkey-88n24.8
|
si:dkey-88n24.8 |
| chr16_-_14587332 | 0.31 |
ENSDART00000012479
|
dscc1
|
DNA replication and sister chromatid cohesion 1 |
| chr24_-_31306724 | 0.31 |
ENSDART00000165399
|
acp5b
|
acid phosphatase 5b, tartrate resistant |
| chr18_-_21748966 | 0.30 |
ENSDART00000181910
|
pskh1
|
protein serine kinase H1 |
| chr9_+_50001746 | 0.30 |
ENSDART00000058892
|
slc38a11
|
solute carrier family 38, member 11 |
| chr2_+_27403300 | 0.29 |
ENSDART00000099180
|
elovl8a
|
ELOVL fatty acid elongase 8a |
| chr20_+_51464670 | 0.29 |
ENSDART00000150110
|
thbd
|
thrombomodulin |
| chr8_+_20918207 | 0.29 |
ENSDART00000144039
|
si:ch73-196i15.5
|
si:ch73-196i15.5 |
| chr6_-_55399214 | 0.28 |
ENSDART00000168367
|
ctsa
|
cathepsin A |
| chr4_+_121709 | 0.28 |
ENSDART00000186461
ENSDART00000172255 |
crebl2
|
cAMP responsive element binding protein-like 2 |
| chr13_+_25486608 | 0.28 |
ENSDART00000057689
|
bag3
|
BCL2 associated athanogene 3 |
| chr22_-_9649627 | 0.28 |
ENSDART00000164721
|
si:dkey-286j17.4
|
si:dkey-286j17.4 |
| chr7_+_2236317 | 0.28 |
ENSDART00000075859
|
zgc:172065
|
zgc:172065 |
| chr21_-_33478164 | 0.28 |
ENSDART00000191542
|
si:ch73-42p12.2
|
si:ch73-42p12.2 |
| chr12_-_26406323 | 0.28 |
ENSDART00000131896
|
myoz1b
|
myozenin 1b |
| chr21_+_7332963 | 0.27 |
ENSDART00000169834
|
AP3B1
|
adaptor related protein complex 3 beta 1 subunit |
| chr2_+_43895103 | 0.27 |
ENSDART00000007074
|
gbp3
|
guanylate binding protein 3 |
| chr22_-_9157364 | 0.27 |
ENSDART00000182762
|
si:ch211-213a13.5
|
si:ch211-213a13.5 |
| chr12_+_27141140 | 0.27 |
ENSDART00000136415
|
hoxb1b
|
homeobox B1b |
| chr8_-_25336589 | 0.26 |
ENSDART00000009682
|
atp5pb
|
ATP synthase peripheral stalk-membrane subunit b |
| chr20_-_26532167 | 0.26 |
ENSDART00000061914
|
mthfd1l
|
methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 1 like |
| chr8_+_23521974 | 0.26 |
ENSDART00000188130
ENSDART00000129378 |
sema3gb
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3Gb |
| chr25_-_35599887 | 0.26 |
ENSDART00000153827
|
clpxb
|
caseinolytic mitochondrial matrix peptidase chaperone subunit b |
| chr24_+_35947077 | 0.25 |
ENSDART00000173406
|
obscnb
|
obscurin, cytoskeletal calmodulin and titin-interacting RhoGEF b |
| chr15_-_41677689 | 0.24 |
ENSDART00000187063
|
spsb4b
|
splA/ryanodine receptor domain and SOCS box containing 4b |
| chr3_-_2072630 | 0.24 |
ENSDART00000189404
|
BX005442.3
|
|
| chr4_+_15899668 | 0.23 |
ENSDART00000101592
|
col7a1l
|
collagen type VII alpha 1-like |
| chr5_+_69856153 | 0.23 |
ENSDART00000124128
ENSDART00000073663 |
ugt2a6
|
UDP glucuronosyltransferase 2 family, polypeptide A6 |
| chr8_+_26818446 | 0.22 |
ENSDART00000134987
ENSDART00000138835 |
si:ch211-156j16.1
|
si:ch211-156j16.1 |
| chr4_-_5775507 | 0.22 |
ENSDART00000021753
|
ccnc
|
cyclin C |
| chr15_-_1765098 | 0.22 |
ENSDART00000149980
ENSDART00000093074 |
bud23
|
BUD23, rRNA methyltransferase and ribosome maturation factor |
| chr19_-_23674904 | 0.22 |
ENSDART00000187300
|
BX927234.2
|
|
| chr11_-_23219367 | 0.22 |
ENSDART00000003646
|
optc
|
opticin |
| chr25_-_15496485 | 0.21 |
ENSDART00000140245
|
si:dkeyp-67e1.6
|
si:dkeyp-67e1.6 |
| chr1_-_38813679 | 0.21 |
ENSDART00000148917
|
asb5b
|
ankyrin repeat and SOCS box containing 5b |
| chr12_+_18663154 | 0.21 |
ENSDART00000057918
|
si:ch211-147h1.4
|
si:ch211-147h1.4 |
| chr2_+_23482798 | 0.20 |
ENSDART00000187448
|
BX470197.1
|
|
| chr7_-_3836896 | 0.19 |
ENSDART00000136227
|
si:dkey-88n24.8
|
si:dkey-88n24.8 |
| chr1_-_2453174 | 0.19 |
ENSDART00000055779
ENSDART00000152555 |
ggact.2
ggact.3
|
gamma-glutamylamine cyclotransferase, tandem duplicate 2 gamma-glutamylamine cyclotransferase, tandem duplicate 3 |
| chr16_+_23331633 | 0.19 |
ENSDART00000187455
|
krtcap2
|
keratinocyte associated protein 2 |
| chr3_-_6191950 | 0.18 |
ENSDART00000161931
|
si:ch73-144l3.1
|
si:ch73-144l3.1 |
| chr23_-_19831739 | 0.18 |
ENSDART00000125066
|
haus7
|
HAUS augmin-like complex, subunit 7 |
| chr14_-_8903435 | 0.18 |
ENSDART00000160584
|
zgc:153681
|
zgc:153681 |
| chr9_-_25094181 | 0.18 |
ENSDART00000132160
|
rubcnl
|
RUN and cysteine rich domain containing beclin 1 interacting protein like |
| chr12_+_20506197 | 0.18 |
ENSDART00000153010
|
si:zfos-754c12.2
|
si:zfos-754c12.2 |
| chr3_-_36690348 | 0.17 |
ENSDART00000192513
|
myh11b
|
myosin, heavy chain 11b, smooth muscle |
| chr4_-_36792144 | 0.17 |
ENSDART00000159147
|
si:dkeyp-87d1.1
|
si:dkeyp-87d1.1 |
| chr8_+_7854130 | 0.17 |
ENSDART00000165575
|
cxxc1a
|
CXXC finger protein 1a |
| chr15_-_21678634 | 0.16 |
ENSDART00000061139
|
bco2b
|
beta-carotene oxygenase 2b |
| chr14_+_45645024 | 0.16 |
ENSDART00000168278
|
si:ch211-276i12.4
|
si:ch211-276i12.4 |
| chr24_+_12074340 | 0.16 |
ENSDART00000170997
ENSDART00000166227 |
ccr9b
|
chemokine (C-C motif) receptor 9b |
| chr12_+_20583552 | 0.16 |
ENSDART00000170035
|
arsg
|
arylsulfatase G |
| chr17_-_27200634 | 0.16 |
ENSDART00000185332
|
asap3
|
ArfGAP with SH3 domain, ankyrin repeat and PH domain 3 |
| chr25_-_15504559 | 0.16 |
ENSDART00000139294
|
BX323543.5
|
|
| chr20_-_34164278 | 0.16 |
ENSDART00000153128
|
hmcn1
|
hemicentin 1 |
| chr2_-_38992304 | 0.15 |
ENSDART00000114085
ENSDART00000146812 |
si:ch211-119o8.6
|
si:ch211-119o8.6 |
| chr19_+_22930177 | 0.15 |
ENSDART00000151737
|
si:ch211-244a23.1
|
si:ch211-244a23.1 |
| chr2_+_42005475 | 0.15 |
ENSDART00000056461
|
gbp2
|
guanylate binding protein 2 |
| chr17_-_27273296 | 0.14 |
ENSDART00000077087
|
id3
|
inhibitor of DNA binding 3 |
| chr25_-_15512819 | 0.14 |
ENSDART00000142684
|
si:dkeyp-67e1.2
|
si:dkeyp-67e1.2 |
| chr19_-_25464291 | 0.14 |
ENSDART00000112915
|
umad1
|
UBAP1-MVB12-associated (UMA) domain containing 1 |
| chr23_-_20345473 | 0.14 |
ENSDART00000140935
|
si:rp71-17i16.6
|
si:rp71-17i16.6 |
| chr3_+_37790351 | 0.14 |
ENSDART00000151506
|
si:dkey-260c8.8
|
si:dkey-260c8.8 |
| chr21_+_21611867 | 0.13 |
ENSDART00000189148
|
b9d2
|
B9 domain containing 2 |
| chr23_+_20687340 | 0.13 |
ENSDART00000143503
|
usp21
|
ubiquitin specific peptidase 21 |
| chr15_+_23523337 | 0.13 |
ENSDART00000182306
ENSDART00000185545 ENSDART00000152517 |
si:dkey-182i3.9
|
si:dkey-182i3.9 |
| chr8_-_46897734 | 0.13 |
ENSDART00000138125
|
hes2.2
|
hes family bHLH transcription factor 2, tandem duplicate 2 |
| chr17_+_8175998 | 0.12 |
ENSDART00000131200
|
myct1b
|
myc target 1b |
| chr7_-_3826794 | 0.12 |
ENSDART00000064229
ENSDART00000142138 |
si:dkey-88n24.10
|
si:dkey-88n24.10 |
| chr4_+_51542447 | 0.12 |
ENSDART00000163814
|
si:dkey-165e24.1
|
si:dkey-165e24.1 |
| chr4_+_62598975 | 0.12 |
ENSDART00000163548
|
si:ch211-79g12.2
|
si:ch211-79g12.2 |
| chr19_+_7424347 | 0.12 |
ENSDART00000004622
|
sf3b4
|
splicing factor 3b, subunit 4 |
| chr8_+_15251448 | 0.12 |
ENSDART00000063717
|
zgc:171480
|
zgc:171480 |
| chr14_+_3516567 | 0.12 |
ENSDART00000171153
ENSDART00000158416 |
fnta
|
farnesyltransferase, CAAX box, alpha |
| chr18_+_37272568 | 0.12 |
ENSDART00000132749
|
tmem123
|
transmembrane protein 123 |
| chr22_+_15323930 | 0.10 |
ENSDART00000142416
|
si:dkey-236e20.3
|
si:dkey-236e20.3 |
| chr4_-_75812937 | 0.10 |
ENSDART00000125096
|
si:ch211-203c5.3
|
si:ch211-203c5.3 |
| chr13_+_33462232 | 0.10 |
ENSDART00000177841
|
zgc:136302
|
zgc:136302 |
| chr19_+_9212031 | 0.10 |
ENSDART00000052930
|
ndufv1
|
NADH dehydrogenase (ubiquinone) flavoprotein 1 |
| chr7_+_22702225 | 0.09 |
ENSDART00000173672
|
si:dkey-165a24.9
|
si:dkey-165a24.9 |
| chr1_+_218524 | 0.09 |
ENSDART00000109529
|
tmco3
|
transmembrane and coiled-coil domains 3 |
| chr13_+_30163515 | 0.09 |
ENSDART00000040926
|
eif4ebp2
|
eukaryotic translation initiation factor 4E binding protein 2 |
| chr8_-_32894203 | 0.09 |
ENSDART00000147822
|
si:dkey-56i24.1
|
si:dkey-56i24.1 |
| chr12_-_26491464 | 0.09 |
ENSDART00000153361
|
si:dkey-287g12.6
|
si:dkey-287g12.6 |
| chr5_-_67783593 | 0.08 |
ENSDART00000010934
|
casr
|
calcium-sensing receptor |
| chr14_-_28052474 | 0.08 |
ENSDART00000172948
ENSDART00000135337 |
TSC22D3 (1 of many)
zgc:64189
|
si:ch211-220e11.3 zgc:64189 |
| chr4_-_49987980 | 0.08 |
ENSDART00000150428
|
si:dkey-156k2.4
|
si:dkey-156k2.4 |
| chr16_+_32029090 | 0.08 |
ENSDART00000041054
|
tmc4
|
transmembrane channel-like 4 |
| chr7_+_13756215 | 0.08 |
ENSDART00000172804
ENSDART00000018801 ENSDART00000115054 |
rasl12
|
RAS-like, family 12 |
| chr2_-_51611344 | 0.08 |
ENSDART00000158137
|
si:ch211-9d9.8
|
si:ch211-9d9.8 |
| chr18_+_32916351 | 0.07 |
ENSDART00000170387
ENSDART00000161553 |
olfcg2
|
olfactory receptor C family, g2 |
| chr16_-_47301376 | 0.07 |
ENSDART00000169697
|
mios
|
missing oocyte, meiosis regulator, homolog (Drosophila) |
| chr13_+_33304187 | 0.07 |
ENSDART00000075826
ENSDART00000145295 |
dcdc2b
|
doublecortin domain containing 2B |
| chr4_+_43700319 | 0.07 |
ENSDART00000141967
|
si:ch211-226o13.1
|
si:ch211-226o13.1 |
| chr15_-_5178899 | 0.06 |
ENSDART00000132148
|
or126-4
|
odorant receptor, family E, subfamily 126, member 4 |
| chr24_-_25184553 | 0.06 |
ENSDART00000166917
|
plcxd2
|
phosphatidylinositol-specific phospholipase C, X domain containing 2 |
| chr10_+_13000669 | 0.06 |
ENSDART00000158919
ENSDART00000172625 |
lpar1
|
lysophosphatidic acid receptor 1 |
| chr7_+_22702437 | 0.06 |
ENSDART00000182054
|
si:dkey-165a24.9
|
si:dkey-165a24.9 |
| chr7_+_17106160 | 0.05 |
ENSDART00000190048
ENSDART00000180004 ENSDART00000013409 |
prmt3
|
protein arginine methyltransferase 3 |
| chr7_+_13756374 | 0.05 |
ENSDART00000180808
|
rasl12
|
RAS-like, family 12 |
| chr6_-_40651944 | 0.05 |
ENSDART00000187423
|
ppil1
|
peptidylprolyl isomerase (cyclophilin)-like 1 |
| chr3_-_1187727 | 0.05 |
ENSDART00000180343
|
smdt1a
|
single-pass membrane protein with aspartate-rich tail 1a |
| chr2_-_37532772 | 0.04 |
ENSDART00000133951
|
arhgef18a
|
rho/rac guanine nucleotide exchange factor (GEF) 18a |
| chr19_+_18493782 | 0.04 |
ENSDART00000160992
|
fkbp10a
|
FK506 binding protein 10a |
| chr4_+_77908076 | 0.04 |
ENSDART00000168811
|
si:zfos-2131b9.2
|
si:zfos-2131b9.2 |
| chr21_-_20725853 | 0.04 |
ENSDART00000114502
|
si:ch211-22d5.2
|
si:ch211-22d5.2 |
| chr2_-_43745146 | 0.04 |
ENSDART00000056164
ENSDART00000098052 |
ftr96
|
finTRIM family, member 96 |
| chr14_+_23687678 | 0.03 |
ENSDART00000002469
|
hspa4b
|
heat shock protein 4b |
| chr1_+_25650917 | 0.02 |
ENSDART00000054235
|
plrg1
|
pleiotropic regulator 1 |
| chr10_+_26652859 | 0.02 |
ENSDART00000079174
|
htatsf1
|
HIV-1 Tat specific factor 1 |
| chr23_-_15090782 | 0.02 |
ENSDART00000133624
|
si:ch211-218g4.2
|
si:ch211-218g4.2 |
| chr17_-_53329704 | 0.02 |
ENSDART00000193895
|
exd1
|
exonuclease 3'-5' domain containing 1 |
| chr7_+_29163762 | 0.02 |
ENSDART00000173762
|
slc38a8b
|
solute carrier family 38, member 8b |
| chr2_+_20472150 | 0.01 |
ENSDART00000168537
|
agla
|
amylo-alpha-1, 6-glucosidase, 4-alpha-glucanotransferase a |
| chr15_-_38154616 | 0.01 |
ENSDART00000099392
|
irgq2
|
immunity-related GTPase family, q2 |
| chr18_-_40913294 | 0.01 |
ENSDART00000059196
ENSDART00000098878 |
polr2i
|
polymerase (RNA) II (DNA directed) polypeptide I |
| chr18_+_17538165 | 0.01 |
ENSDART00000147839
|
nup93
|
nucleoporin 93 |
| chr17_+_47090497 | 0.01 |
ENSDART00000169038
ENSDART00000159292 |
zgc:103755
|
zgc:103755 |
| chr4_+_53246788 | 0.01 |
ENSDART00000184708
ENSDART00000169256 |
si:dkey-250k10.1
|
si:dkey-250k10.1 |
| chr4_-_18775548 | 0.01 |
ENSDART00000141187
|
def6c
|
differentially expressed in FDCP 6c homolog |
| chr4_+_77681389 | 0.01 |
ENSDART00000099727
|
gbp4
|
guanylate binding protein 4 |
| chr5_-_10236599 | 0.00 |
ENSDART00000099834
|
si:ch73-42k18.1
|
si:ch73-42k18.1 |
| chr4_-_60518031 | 0.00 |
ENSDART00000157489
|
si:dkey-211i20.4
|
si:dkey-211i20.4 |
Network of associatons between targets according to the STRING database.
First level regulatory network of tbx3a+tbx3b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.4 | GO:0006740 | NADPH regeneration(GO:0006740) |
| 0.3 | 1.3 | GO:0003228 | atrial cardiac muscle tissue development(GO:0003228) |
| 0.3 | 0.8 | GO:0014809 | regulation of skeletal muscle contraction by calcium ion signaling(GO:0014722) regulation of skeletal muscle contraction by regulation of release of sequestered calcium ion(GO:0014809) |
| 0.2 | 1.0 | GO:0055014 | atrial cardiac muscle cell development(GO:0055014) |
| 0.1 | 0.7 | GO:0070986 | left/right axis specification(GO:0070986) |
| 0.1 | 0.3 | GO:0034086 | maintenance of sister chromatid cohesion(GO:0034086) maintenance of mitotic sister chromatid cohesion(GO:0034088) |
| 0.1 | 0.3 | GO:0010660 | negative regulation of muscle cell apoptotic process(GO:0010656) muscle cell apoptotic process(GO:0010657) striated muscle cell apoptotic process(GO:0010658) regulation of muscle cell apoptotic process(GO:0010660) regulation of striated muscle cell apoptotic process(GO:0010662) negative regulation of striated muscle cell apoptotic process(GO:0010664) |
| 0.1 | 0.8 | GO:0051694 | pointed-end actin filament capping(GO:0051694) |
| 0.1 | 0.3 | GO:0021570 | rhombomere 4 development(GO:0021570) rhombomere 4 morphogenesis(GO:0021661) |
| 0.1 | 0.3 | GO:0045600 | positive regulation of fat cell differentiation(GO:0045600) regulation of glucose import(GO:0046324) |
| 0.1 | 0.2 | GO:0042543 | protein N-linked glycosylation via arginine(GO:0042543) |
| 0.1 | 0.2 | GO:0036265 | RNA (guanine-N7)-methylation(GO:0036265) |
| 0.0 | 0.4 | GO:1900407 | regulation of cellular response to oxidative stress(GO:1900407) |
| 0.0 | 0.3 | GO:0071260 | cellular response to mechanical stimulus(GO:0071260) |
| 0.0 | 0.8 | GO:0032233 | positive regulation of actin filament bundle assembly(GO:0032233) |
| 0.0 | 0.1 | GO:0018343 | protein farnesylation(GO:0018343) |
| 0.0 | 0.3 | GO:0045453 | bone resorption(GO:0045453) |
| 0.0 | 0.8 | GO:0032543 | mitochondrial translation(GO:0032543) |
| 0.0 | 0.2 | GO:0016119 | carotene metabolic process(GO:0016119) carotene catabolic process(GO:0016121) terpene metabolic process(GO:0042214) terpene catabolic process(GO:0046247) |
| 0.0 | 0.8 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.0 | 0.3 | GO:0035999 | tetrahydrofolate interconversion(GO:0035999) |
| 0.0 | 0.5 | GO:0001878 | response to yeast(GO:0001878) |
| 0.0 | 0.5 | GO:0006919 | activation of cysteine-type endopeptidase activity involved in apoptotic process(GO:0006919) |
| 0.0 | 0.3 | GO:0019368 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.0 | 0.3 | GO:0048251 | elastic fiber assembly(GO:0048251) |
| 0.0 | 0.1 | GO:0005513 | detection of calcium ion(GO:0005513) |
| 0.0 | 0.4 | GO:0007007 | inner mitochondrial membrane organization(GO:0007007) |
| 0.0 | 0.3 | GO:0015986 | energy coupled proton transport, down electrochemical gradient(GO:0015985) ATP synthesis coupled proton transport(GO:0015986) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0061702 | inflammasome complex(GO:0061702) |
| 0.1 | 0.8 | GO:0033018 | sarcoplasmic reticulum lumen(GO:0033018) |
| 0.1 | 0.3 | GO:0031390 | Ctf18 RFC-like complex(GO:0031390) |
| 0.0 | 0.1 | GO:0005953 | CAAX-protein geranylgeranyltransferase complex(GO:0005953) |
| 0.0 | 2.1 | GO:0036379 | striated muscle thin filament(GO:0005865) myofilament(GO:0036379) |
| 0.0 | 0.4 | GO:0061617 | MICOS complex(GO:0061617) |
| 0.0 | 0.4 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.0 | 0.9 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.0 | 0.2 | GO:0070652 | HAUS complex(GO:0070652) |
| 0.0 | 0.8 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.0 | 0.3 | GO:0000276 | mitochondrial proton-transporting ATP synthase complex, coupling factor F(o)(GO:0000276) |
| 0.0 | 1.4 | GO:0030018 | Z disc(GO:0030018) |
| 0.0 | 0.2 | GO:0031430 | M band(GO:0031430) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.4 | GO:0016652 | NAD(P)+ transhydrogenase activity(GO:0008746) oxidoreductase activity, acting on NAD(P)H, NAD(P) as acceptor(GO:0016652) |
| 0.1 | 1.3 | GO:0030172 | troponin C binding(GO:0030172) troponin I binding(GO:0031013) |
| 0.1 | 0.3 | GO:0015369 | calcium:proton antiporter activity(GO:0015369) metal ion:proton antiporter activity(GO:0051139) |
| 0.1 | 0.8 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.1 | 0.3 | GO:0004185 | serine-type carboxypeptidase activity(GO:0004185) |
| 0.1 | 0.5 | GO:0003854 | 3-beta-hydroxy-delta5-steroid dehydrogenase activity(GO:0003854) |
| 0.0 | 0.1 | GO:0004662 | CAAX-protein geranylgeranyltransferase activity(GO:0004662) |
| 0.0 | 0.3 | GO:0003993 | acid phosphatase activity(GO:0003993) |
| 0.0 | 0.3 | GO:0004329 | formate-tetrahydrofolate ligase activity(GO:0004329) |
| 0.0 | 0.5 | GO:0097199 | cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:0097199) |
| 0.0 | 0.3 | GO:0033170 | DNA clamp loader activity(GO:0003689) protein-DNA loading ATPase activity(GO:0033170) |
| 0.0 | 0.3 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 0.0 | 0.3 | GO:0031433 | telethonin binding(GO:0031433) FATZ binding(GO:0051373) |
| 0.0 | 0.8 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.0 | 0.2 | GO:0003834 | beta-carotene 15,15'-monooxygenase activity(GO:0003834) carotenoid dioxygenase activity(GO:0010436) |
| 0.0 | 0.3 | GO:0102336 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.0 | 0.5 | GO:0004629 | phosphatidylinositol phospholipase C activity(GO:0004435) phospholipase C activity(GO:0004629) |
| 0.0 | 0.1 | GO:0008190 | eukaryotic initiation factor 4E binding(GO:0008190) |
| 0.0 | 0.2 | GO:0008649 | rRNA methyltransferase activity(GO:0008649) |
| 0.0 | 0.4 | GO:0008381 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 0.0 | 0.4 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.0 | 1.0 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.0 | 0.3 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.0 | 0.5 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 0.4 | REACTOME FORMATION OF TUBULIN FOLDING INTERMEDIATES BY CCT TRIC | Genes involved in Formation of tubulin folding intermediates by CCT/TriC |
| 0.0 | 0.2 | REACTOME THE ACTIVATION OF ARYLSULFATASES | Genes involved in The activation of arylsulfatases |
| 0.0 | 0.3 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |