Project
GSE53960: rat RNA-Seq transcriptomic Bodymap
Navigation
Downloads
Results for Glis1
Z-value: 0.11
Transcription factors associated with Glis1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Glis1
|
ENSRNOG00000000138 | GLIS family zinc finger 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Glis1 | rn6_v1_chr5_+_127160495_127160495 | -0.06 | 2.5e-01 | Click! |
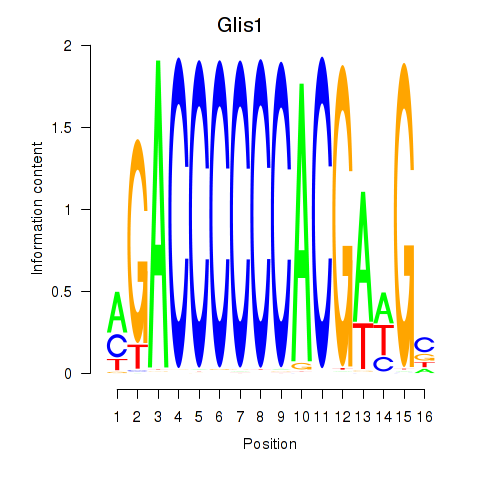
Activity profile of Glis1 motif
Sorted Z-values of Glis1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr13_+_36378356 | 6.27 |
ENSRNOT00000071305
|
C1ql2
|
complement C1q like 2 |
| chr5_-_151396507 | 0.65 |
ENSRNOT00000012710
|
Gpr3
|
G protein-coupled receptor 3 |
| chr5_-_136762986 | 0.54 |
ENSRNOT00000026852
|
Artn
|
artemin |
| chr10_-_10831782 | 0.48 |
ENSRNOT00000004314
|
Nudt16l1
|
nudix hydrolase 16 like 1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of Glis1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.5 | GO:0061146 | Peyer's patch morphogenesis(GO:0061146) |
| 0.0 | 0.7 | GO:0040020 | regulation of meiotic nuclear division(GO:0040020) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 6.3 | GO:0005581 | collagen trimer(GO:0005581) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | GO:0050072 | m7G(5')pppN diphosphatase activity(GO:0050072) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 6.3 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 0.5 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |