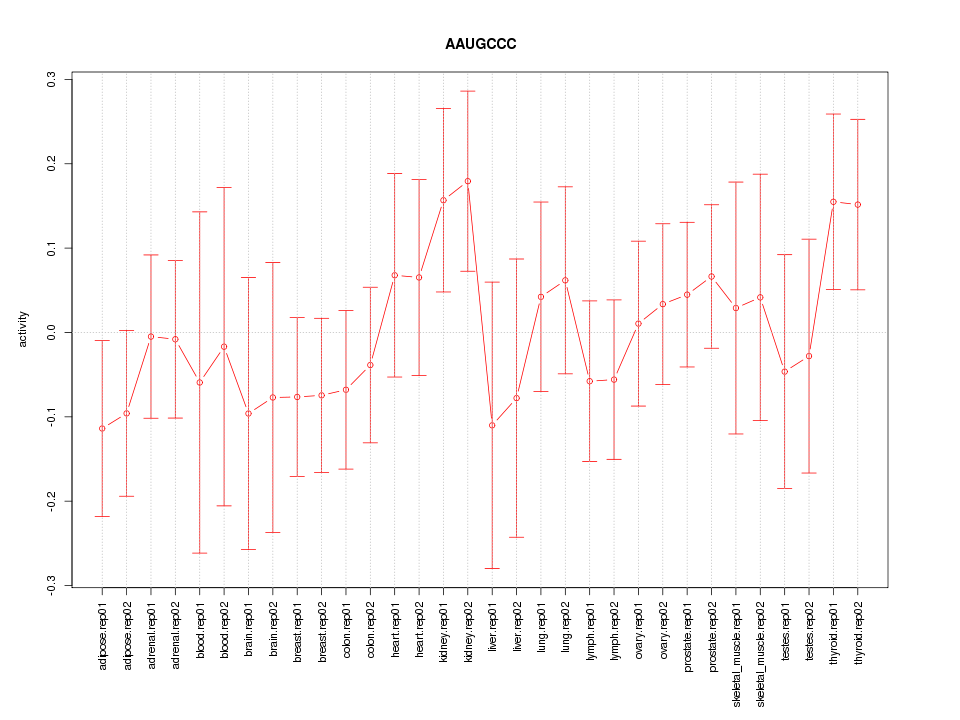
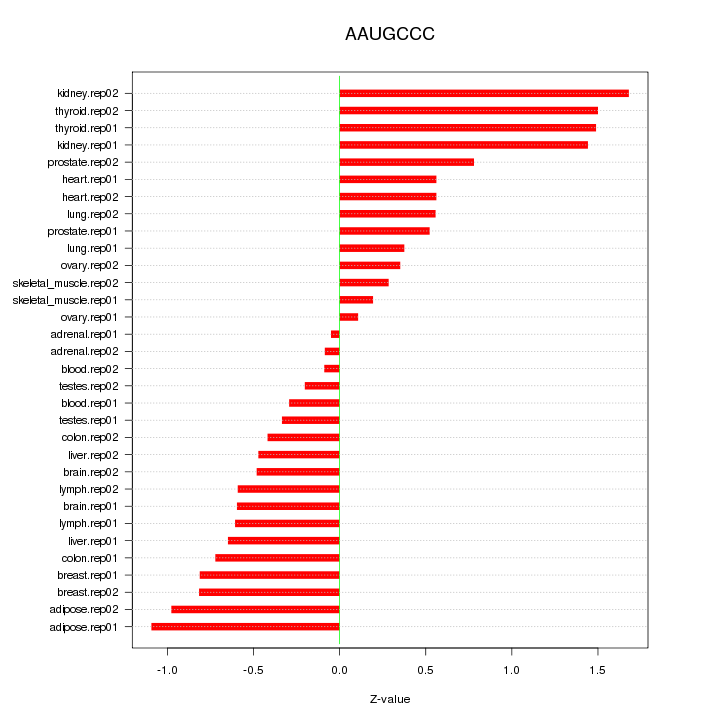
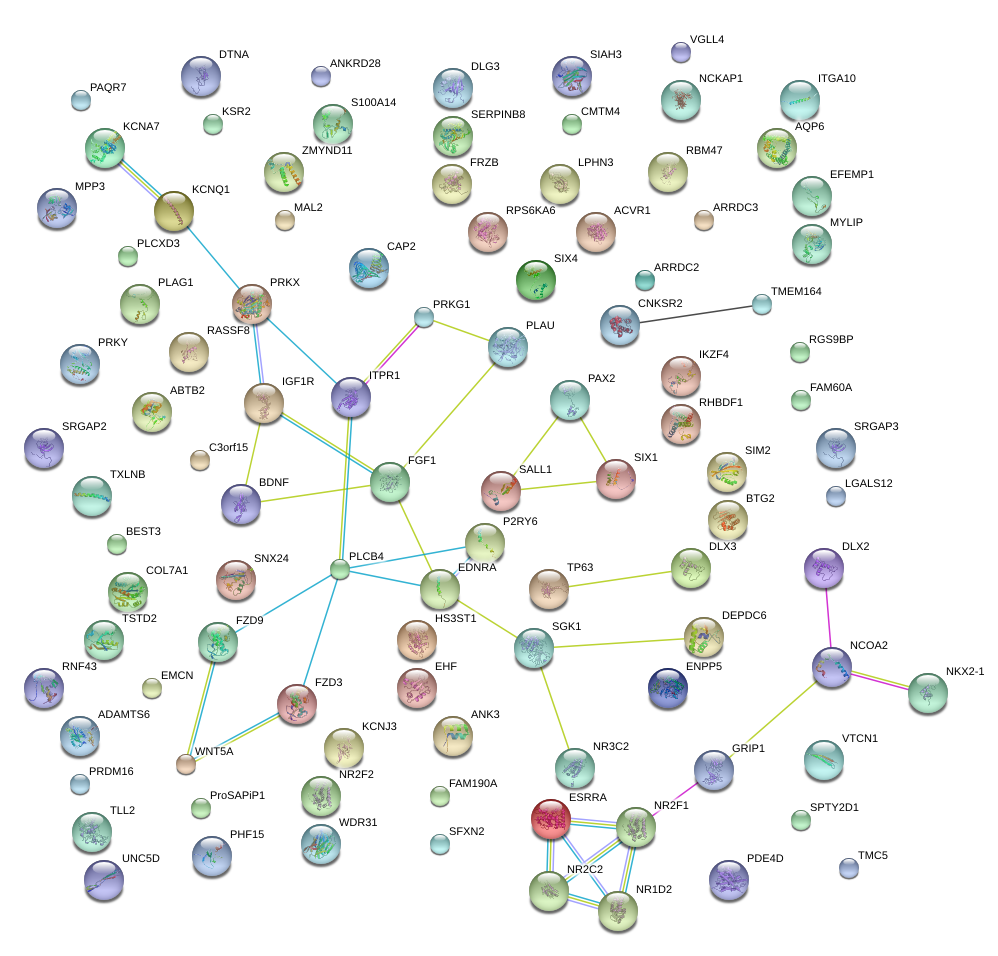
|
chr11_+_34642587
|
1.783
|
NM_001206615
NM_012153
|
EHF
|
ets homologous factor
|
|
chr11_+_34654010
|
1.640
|
NM_001206616
|
EHF
|
ets homologous factor
|
|
chr2_+_155555092
|
1.217
|
NM_002239
|
KCNJ3
|
potassium inwardly-rectifying channel, subfamily J, member 3
|
|
chr14_-_36989333
|
1.147
|
NM_001079668
|
NKX2-1
|
NK2 homeobox 1
|
|
chr1_+_2985730
|
1.079
|
NM_022114
NM_199454
|
PRDM16
|
PR domain containing 16
|
|
chr14_-_36988902
|
1.036
|
NM_003317
|
NKX2-1
|
NK2 homeobox 1
|
|
chr1_-_153588789
|
0.901
|
NM_020672
|
S100A14
|
S100 calcium binding protein A14
|
|
chr10_+_102505337
|
0.828
|
NM_000278
NM_003987
NM_003988
NM_003989
NM_003990
|
PAX2
|
paired box 2
|
|
chr11_+_2466171
|
0.819
|
NM_000218
|
KCNQ1
|
potassium voltage-gated channel, KQT-like subfamily, member 1
|
|
chr19_-_49576120
|
0.735
|
NM_031886
|
KCNA7
|
potassium voltage-gated channel, shaker-related subfamily, member 7
|
|
chrX_-_83442906
|
0.722
|
NM_014496
|
RPS6KA6
|
ribosomal protein S6 kinase, 90kDa, polypeptide 6
|
|
chr16_-_51185150
|
0.721
|
NM_002968
|
SALL1
|
sal-like 1 (Drosophila)
|
|
chr14_-_61190458
|
0.716
|
NM_017420
|
SIX4
|
SIX homeobox 4
|
|
chr6_-_46138584
|
0.687
|
NM_021572
|
ENPP5
|
ectonucleotide pyrophosphatase/phosphodiesterase 5 (putative)
|
|
chr4_-_101439071
|
0.660
|
NM_001159694
NM_016242
|
EMCN
|
endomucin
|
|
chr6_+_17393610
|
0.658
|
NM_006366
|
CAP2
|
CAP, adenylate cyclase-associated protein, 2 (yeast)
|
|
chr16_-_51184507
|
0.647
|
NM_001127892
|
SALL1
|
sal-like 1 (Drosophila)
|
|
chr10_+_52834233
|
0.640
|
NM_006258
|
PRKG1
|
protein kinase, cGMP-dependent, type I
|
|
chr2_-_158732341
|
0.600
|
NM_001111067
|
ACVR1
|
activin A receptor, type I
|
|
chr10_+_52750904
|
0.590
|
NM_001098512
|
PRKG1
|
protein kinase, cGMP-dependent, type I
|
|
chr6_-_134498973
|
0.588
|
NM_001143677
|
SGK1
|
serum/glucocorticoid regulated kinase 1
|
|
chr10_-_62332666
|
0.585
|
NM_001204404
|
ANK3
|
ankyrin 3, node of Ranvier (ankyrin G)
|
|
chr13_-_46425756
|
0.581
|
NM_198849
|
SIAH3
|
seven in absentia homolog 3 (Drosophila)
|
|
chr11_-_27742224
|
0.564
|
NM_001143805
NM_001143806
NM_170732
|
BDNF
|
brain-derived neurotrophic factor
|
|
chr10_-_61900659
|
0.521
|
NM_001149
|
ANK3
|
ankyrin 3, node of Ranvier (ankyrin G)
|
|
chr11_-_27721975
|
0.506
|
NM_001143813
NM_001143814
NM_001709
|
BDNF
|
brain-derived neurotrophic factor
|
|
chr5_-_142077508
|
0.504
|
NM_001144934
NM_001144935
|
FGF1
|
fibroblast growth factor 1 (acidic)
|
|
chr12_-_70093064
|
0.496
|
NM_032735
|
BEST3
|
bestrophin 3
|
|
chr5_-_141993691
|
0.493
|
NM_033137
|
FGF1
|
fibroblast growth factor 1 (acidic)
|
|
chr12_-_70083055
|
0.492
|
NM_152439
|
BEST3
|
bestrophin 3
|
|
chr9_-_116102533
|
0.492
|
NM_001012361
NM_145241
|
WDR31
|
WD repeat domain 31
|
|
chrX_-_3631654
|
0.489
|
NM_005044
|
PRKX
|
protein kinase, X-linked
|
|
chr12_+_50366619
|
0.480
|
NM_001652
|
AQP6
|
aquaporin 6, kidney specific
|
|
chr10_-_62493282
|
0.468
|
NM_001204403
|
ANK3
|
ankyrin 3, node of Ranvier (ankyrin G)
|
|
chr2_-_183731279
|
0.463
|
NM_001463
|
FRZB
|
frizzled-related protein
|
|
chr12_+_26205490
|
0.461
|
NM_001164746
|
RASSF8
|
Ras association (RalGDS/AF-6) domain family (N-terminal) member 8
|
|
chr10_-_98273444
|
0.461
|
NM_012465
|
TLL2
|
tolloid-like 2
|
|
chr3_-_55521330
|
0.460
|
NM_003392
|
WNT5A
|
wingless-type MMTV integration site family, member 5A
|
|
chr11_-_34379526
|
0.460
|
NM_145804
|
ABTB2
|
ankyrin repeat and BTB (POZ) domain containing 2
|
|
chr12_+_26126687
|
0.454
|
NM_001164747
|
RASSF8
|
Ras association (RalGDS/AF-6) domain family (N-terminal) member 8
|
|
chr6_-_134496006
|
0.429
|
NM_005627
|
SGK1
|
serum/glucocorticoid regulated kinase 1
|
|
chr11_+_64073040
|
0.428
|
NM_004451
|
ESRRA
|
estrogen-related receptor alpha
|
|
chr2_-_56151273
|
0.413
|
NM_001039348
NM_001039349
|
EFEMP1
|
EGF containing fibulin-like extracellular matrix protein 1
|
|
chr11_-_27722599
|
0.394
|
NM_001143808
NM_001143809
NM_001143810
NM_001143811
|
BDNF
|
brain-derived neurotrophic factor
|
|
chr12_-_118406027
|
0.389
|
NM_173598
|
KSR2
|
kinase suppressor of ras 2
|
|
chr16_+_19422056
|
0.389
|
NM_001105248
NM_001105249
|
TMC5
|
transmembrane channel-like 5
|
|
chr20_+_9198036
|
0.386
|
NM_182797
|
PLCB4
|
phospholipase C, beta 4
|
|
chr10_+_75670861
|
0.386
|
NM_001145031
NM_002658
|
PLAU
|
plasminogen activator, urokinase
|
|
chr20_-_3149070
|
0.383
|
NM_014731
|
ProSAPiP1
|
ProSAPiP1 protein
|
|
chr12_-_67072881
|
0.370
|
NM_001178074
NM_021150
|
GRIP1
|
glutamate receptor interacting protein 1
|
|
chr11_-_27721179
|
0.370
|
NM_170734
|
BDNF
|
brain-derived neurotrophic factor
|
|
chr3_+_189349215
|
0.367
|
NM_001114978
NM_001114979
NM_003722
|
TP63
|
tumor protein p63
|
|
chr5_-_90679115
|
0.366
|
NM_020801
|
ARRDC3
|
arrestin domain containing 3
|
|
chr5_+_133861685
|
0.365
|
NM_015288
|
PHF15
|
PHD finger protein 15
|
|
chr4_+_91048683
|
0.364
|
NM_001145065
|
FAM190A
|
family with sequence similarity 190, member A
|
|
chr5_-_58571916
|
0.363
|
NM_001197220
|
PDE4D
|
phosphodiesterase 4D, cAMP-specific
|
|
chr20_+_9049660
|
0.358
|
NM_001172646
|
PLCB4
|
phospholipase C, beta 4
|
|
chr8_+_35092963
|
0.349
|
NM_080872
|
UNC5D
|
unc-5 homolog D (C. elegans)
|
|
chr4_-_149363498
|
0.342
|
NM_000901
NM_001166104
|
NR3C2
|
nuclear receptor subfamily 3, group C, member 2
|
|
chr16_-_122573
|
0.337
|
NM_022450
|
RHBDF1
|
rhomboid 5 homolog 1 (Drosophila)
|
|
chr3_+_189507448
|
0.337
|
NM_001114980
NM_001114981
NM_001114982
|
TP63
|
tumor protein p63
|
|
chr1_+_145524989
|
0.334
|
NM_003637
|
ITGA10
|
integrin, alpha 10
|
|
chr8_+_120220554
|
0.334
|
NM_052886
|
MAL2
|
mal, T-cell differentiation protein 2 (gene/pseudogene)
|
|
chr5_-_142000899
|
0.331
|
NM_001144892
|
FGF1
|
fibroblast growth factor 1 (acidic)
|
|
chr15_+_99192696
|
0.330
|
NM_000875
|
IGF1R
|
insulin-like growth factor 1 receptor
|
|
chr6_-_139613114
|
0.330
|
NM_153235
|
TXLNB
|
taxilin beta
|
|
chr9_-_100395961
|
0.328
|
NM_139246
|
TSTD2
|
thiosulfate sulfurtransferase (rhodanese)-like domain containing 2
|
|
chr6_-_134639195
|
0.324
|
NM_001143676
|
SGK1
|
serum/glucocorticoid regulated kinase 1
|
|
chr21_+_38071990
|
0.318
|
NM_005069
NM_009586
|
SIM2
|
single-minded homolog 2 (Drosophila)
|
|
chr5_-_142065991
|
0.316
|
NM_033136
|
FGF1
|
fibroblast growth factor 1 (acidic)
|
|
chr4_+_148402068
|
0.315
|
NM_001166055
NM_001957
|
EDNRA
|
endothelin receptor type A
|
|
chr2_-_158731622
|
0.313
|
NM_001105
|
ACVR1
|
activin A receptor, type I
|
|
chrX_+_109246338
|
0.312
|
NM_032227
|
TMEM164
|
transmembrane protein 164
|
|
chrX_+_69664704
|
0.312
|
NM_021120
|
DLG3
|
discs, large homolog 3 (Drosophila)
|
|
chr5_-_59189620
|
0.305
|
NM_001104631
|
PDE4D
|
phosphodiesterase 4D, cAMP-specific
|
|
chr5_-_142065586
|
0.299
|
NM_000800
|
FGF1
|
fibroblast growth factor 1 (acidic)
|
|
chr5_-_64777703
|
0.299
|
NM_197941
|
ADAMTS6
|
ADAM metallopeptidase with thrombospondin type 1 motif, 6
|
|
chr5_-_59064437
|
0.293
|
NM_001197218
|
PDE4D
|
phosphodiesterase 4D, cAMP-specific
|
|
chr20_+_9288446
|
0.290
|
NM_000933
|
PLCB4
|
phospholipase C, beta 4
|
|
chr3_-_48632592
|
0.286
|
NM_000094
|
COL7A1
|
collagen, type VII, alpha 1
|
|
chr5_-_58882274
|
0.285
|
NM_006203
|
PDE4D
|
phosphodiesterase 4D, cAMP-specific
|
|
chr10_+_181410
|
0.285
|
NM_001202464
|
ZMYND11
|
zinc finger, MYND-type containing 11
|
|
chrX_+_109245851
|
0.284
|
NM_017698
|
TMEM164
|
transmembrane protein 164
|
|
chr19_+_18111923
|
0.281
|
NM_001025604
|
ARRDC2
|
arrestin domain containing 2
|
|
chr6_-_134497069
|
0.279
|
NM_001143678
|
SGK1
|
serum/glucocorticoid regulated kinase 1
|
|
chr12_-_31479120
|
0.278
|
NM_001135811
NM_021238
NM_001135812
|
FAM60A
|
family with sequence similarity 60, member A
|
|
chr11_-_27723061
|
0.278
|
NM_170733
|
BDNF
|
brain-derived neurotrophic factor
|
|
chr3_+_23986735
|
0.272
|
NM_005126
|
NR1D2
|
nuclear receptor subfamily 1, group D, member 2
|
|
chr11_+_72983246
|
0.271
|
NM_004154
|
P2RY6
|
pyrimidinergic receptor P2Y, G-protein coupled, 6
|
|
chr6_+_16129285
|
0.268
|
NM_013262
|
MYLIP
|
myosin regulatory light chain interacting protein
|
|
chr1_-_117753476
|
0.266
|
NM_001253850
NM_024626
|
VTCN1
|
V-set domain containing T cell activation inhibitor 1
|
|
chr11_-_27741192
|
0.265
|
NM_001143807
|
BDNF
|
brain-derived neurotrophic factor
|
|
chr14_-_61115906
|
0.263
|
NM_005982
|
SIX1
|
SIX homeobox 1
|
|
chr5_-_58334936
|
0.261
|
NM_001197221
NM_001197222
|
PDE4D
|
phosphodiesterase 4D, cAMP-specific
|
|
chr3_-_15839674
|
0.260
|
NM_001195098
|
ANKRD28
|
ankyrin repeat domain 28
|
|
chr4_-_11430502
|
0.258
|
NM_005114
|
HS3ST1
|
heparan sulfate (glucosamine) 3-O-sulfotransferase 1
|
|
chr2_-_183903225
|
0.258
|
NM_013436
NM_205842
|
NCKAP1
|
NCK-associated protein 1
|
|
chr8_+_120885899
|
0.258
|
NM_022783
|
DEPTOR
|
DEP domain containing MTOR-interacting protein
|
|
chr10_+_104474297
|
0.255
|
NM_178858
|
SFXN2
|
sideroflexin 2
|
|
chr15_+_96873845
|
0.254
|
NM_021005
|
NR2F2
|
nuclear receptor subfamily 2, group F, member 2
|
|
chr18_+_32397925
|
0.252
|
NM_001198943
|
DTNA
|
dystrobrevin, alpha
|
|
chr8_+_28351694
|
0.249
|
NM_017412
NM_145866
|
FZD3
|
frizzled family receptor 3
|
|
chrX_+_69672110
|
0.249
|
NM_020730
|
DLG3
|
discs, large homolog 3 (Drosophila)
|
|
chr4_-_40517913
|
0.247
|
NM_019027
|
RBM47
|
RNA binding motif protein 47
|
|
chr17_-_48072566
|
0.246
|
NM_005220
|
DLX3
|
distal-less homeobox 3
|
|
chrX_+_69674914
|
0.246
|
NM_001166278
|
DLG3
|
discs, large homolog 3 (Drosophila)
|
|
chr3_+_23987514
|
0.245
|
NM_001145425
|
NR1D2
|
nuclear receptor subfamily 1, group D, member 2
|
|
chr10_-_62149487
|
0.245
|
NM_020987
|
ANK3
|
ankyrin 3, node of Ranvier (ankyrin G)
|
|
chr17_-_56494930
|
0.244
|
NM_017763
|
RNF43
|
ring finger protein 43
|
|
chr5_-_58652671
|
0.240
|
NM_001197219
|
PDE4D
|
phosphodiesterase 4D, cAMP-specific
|
|
chr3_+_4535030
|
0.236
|
NM_001099952
NM_001168272
NM_002222
|
ITPR1
|
inositol 1,4,5-trisphosphate receptor, type 1
|
|
chr15_+_96875793
|
0.235
|
NM_001145156
|
NR2F2
|
nuclear receptor subfamily 2, group F, member 2
|
|
chr16_+_19467771
|
0.234
|
NM_024780
|
TMC5
|
transmembrane channel-like 5
|
|
chr1_+_203274646
|
0.233
|
NM_006763
|
BTG2
|
BTG family, member 2
|
|
chr3_-_9291251
|
0.231
|
NM_001033117
NM_014850
|
SRGAP3
|
SLIT-ROBO Rho GTPase activating protein 3
|
|
chr3_+_14989076
|
0.231
|
NM_003298
|
NR2C2
|
nuclear receptor subfamily 2, group C, member 2
|
|
chr3_-_11685395
|
0.229
|
NM_001128219
|
VGLL4
|
vestigial like 4 (Drosophila)
|
|
chr5_+_122181159
|
0.229
|
NM_014035
|
SNX24
|
sorting nexin 24
|
|
chr3_+_119421865
|
0.229
|
NM_033364
|
C3orf15
|
chromosome 3 open reading frame 15
|
|
chr19_+_33166312
|
0.223
|
NM_207391
|
RGS9BP
|
regulator of G protein signaling 9 binding protein
|
|
chr5_-_58295758
|
0.222
|
NM_001197223
|
PDE4D
|
phosphodiesterase 4D, cAMP-specific
|
|
chr12_+_56414688
|
0.221
|
NM_022465
|
IKZF4
|
IKAROS family zinc finger 4 (Eos)
|
|
chr19_+_18118921
|
0.221
|
NM_015683
|
ARRDC2
|
arrestin domain containing 2
|
|
chr8_-_57123858
|
0.219
|
NM_001114634
NM_001114635
NM_002655
|
PLAG1
|
pleiomorphic adenoma gene 1
|
|
chr18_+_32398225
|
0.218
|
NM_001198942
NM_001198944
NM_032980
NM_032981
|
DTNA
|
dystrobrevin, alpha
|
|
chr12_+_26111951
|
0.218
|
NM_001164748
NM_007211
|
RASSF8
|
Ras association (RalGDS/AF-6) domain family (N-terminal) member 8
|
|
chr11_-_27681195
|
0.212
|
NM_001143816
NM_170735
|
BDNF
|
brain-derived neurotrophic factor
|
|
chr1_-_26197743
|
0.207
|
NM_178422
|
PAQR7
|
progestin and adipoQ receptor family member VII
|
|
chr5_-_59783885
|
0.206
|
NM_001165899
|
PDE4D
|
phosphodiesterase 4D, cAMP-specific
|
|
chr18_+_32335939
|
0.206
|
NM_001390
NM_001391
|
DTNA
|
dystrobrevin, alpha
|
|
chr3_-_15798057
|
0.205
|
NM_001195099
|
ANKRD28
|
ankyrin repeat domain 28
|
|
chr16_-_66730466
|
0.203
|
NM_178818
NM_181521
|
CMTM4
|
CKLF-like MARVEL transmembrane domain containing 4
|
|
chr5_-_41510627
|
0.202
|
NM_001005473
|
PLCXD3
|
phosphatidylinositol-specific phospholipase C, X domain containing 3
|
|
chr11_-_18655962
|
0.197
|
NM_194285
|
SPTY2D1
|
SPT2, Suppressor of Ty, domain containing 1 (S. cerevisiae)
|
|
chr4_+_88928797
|
0.196
|
NM_000297
|
PKD2
|
polycystic kidney disease 2 (autosomal dominant)
|
|
chr4_+_41992492
|
0.195
|
NM_006345
|
SLC30A9
|
solute carrier family 30 (zinc transporter), member 9
|
|
chr3_+_14444083
|
0.193
|
NM_001134367
NM_001134368
NM_003043
|
SLC6A6
|
solute carrier family 6 (neurotransmitter transporter, taurine), member 6
|
|
chr9_+_116638544
|
0.193
|
NM_133374
|
ZNF618
|
zinc finger protein 618
|
|
chr2_-_153573858
|
0.192
|
NM_017892
|
PRPF40A
|
PRP40 pre-mRNA processing factor 40 homolog A (S. cerevisiae)
|
|
chr11_-_27722446
|
0.191
|
NM_001143812
|
BDNF
|
brain-derived neurotrophic factor
|
|
chr12_+_56137063
|
0.191
|
NM_005811
|
GDF11
|
growth differentiation factor 11
|
|
chr12_+_74931550
|
0.190
|
NM_001136262
|
ATXN7L3B
|
ataxin 7-like 3B
|
|
chr16_+_67063035
|
0.189
|
NM_001755
NM_022845
|
CBFB
|
core-binding factor, beta subunit
|
|
chr7_+_101460881
|
0.189
|
NM_181552
|
CUX1
|
cut-like homeobox 1
|
|
chr11_-_16497917
|
0.188
|
NM_033326
|
SOX6
|
SRY (sex determining region Y)-box 6
|
|
chr16_+_53088791
|
0.188
|
NM_025134
|
CHD9
|
chromodomain helicase DNA binding protein 9
|
|
chr20_+_45523247
|
0.185
|
NM_005244
NM_172110
|
EYA2
|
eyes absent homolog 2 (Drosophila)
|
|
chr1_+_157963062
|
0.184
|
NM_018240
|
KIRREL
|
kin of IRRE like (Drosophila)
|
|
chr3_+_2933898
|
0.182
|
NM_001206956
NM_175613
|
CNTN4
|
contactin 4
|
|
chr5_-_39074494
|
0.182
|
NM_152756
|
RICTOR
|
RPTOR independent companion of MTOR, complex 2
|
|
chr18_+_32173281
|
0.180
|
NM_001128175
NM_001198938
NM_001198939
NM_001198940
|
DTNA
|
dystrobrevin, alpha
|
|
chr3_+_113616317
|
0.180
|
NM_001172105
|
GRAMD1C
|
GRAM domain containing 1C
|
|
chr3_-_113465101
|
0.179
|
NM_025146
|
NAA50
|
N(alpha)-acetyltransferase 50, NatE catalytic subunit
|
|
chr13_+_30002776
|
0.178
|
NM_015233
|
MTUS2
|
microtubule associated tumor suppressor candidate 2
|
|
chr3_-_11623829
|
0.177
|
NM_001128220
|
VGLL4
|
vestigial like 4 (Drosophila)
|
|
chr3_-_11761946
|
0.176
|
NM_014667
|
VGLL4
|
vestigial like 4 (Drosophila)
|
|
chr9_-_14398981
|
0.175
|
NM_001190738
|
NFIB
|
nuclear factor I/B
|
|
chr3_-_15900928
|
0.175
|
NM_015199
|
ANKRD28
|
ankyrin repeat domain 28
|
|
chr1_+_236958539
|
0.173
|
NM_000254
|
MTR
|
5-methyltetrahydrofolate-homocysteine methyltransferase
|
|
chr12_-_49182718
|
0.173
|
NM_020983
|
ADCY6
|
adenylate cyclase 6
|
|
chr12_+_2162388
|
0.172
|
NM_000719
NM_001129827
NM_001129829
NM_001129830
NM_001129831
NM_001129832
NM_001129833
NM_001129834
NM_001129835
NM_001129836
NM_001129837
NM_001129838
NM_001129839
NM_001129840
NM_001129841
NM_001129842
NM_001129843
NM_001129844
NM_001129846
NM_001167623
NM_001167624
NM_001167625
NM_199460
|
CACNA1C
|
calcium channel, voltage-dependent, L type, alpha 1C subunit
|
|
chr7_-_150675236
|
0.170
|
NM_000238
NM_172056
|
KCNH2
|
potassium voltage-gated channel, subfamily H (eag-related), member 2
|
|
chr1_-_100715388
|
0.170
|
NM_001918
|
DBT
|
dihydrolipoamide branched chain transacylase E2
|
|
chr10_-_101190201
|
0.169
|
NM_002079
|
GOT1
|
glutamic-oxaloacetic transaminase 1, soluble (aspartate aminotransferase 1)
|
|
chr6_+_46620631
|
0.169
|
NM_001204051
NM_001204052
NM_004277
|
SLC25A27
|
solute carrier family 25, member 27
|
|
chr20_+_33292117
|
0.168
|
NM_021202
|
TP53INP2
|
tumor protein p53 inducible nuclear protein 2
|
|
chr3_+_113557679
|
0.168
|
NM_017577
|
GRAMD1C
|
GRAM domain containing 1C
|
|
chr20_-_3388219
|
0.166
|
NM_001009984
|
C20orf194
|
chromosome 20 open reading frame 194
|
|
chr4_-_74124456
|
0.165
|
NM_032217
NM_198889
|
ANKRD17
|
ankyrin repeat domain 17
|
|
chr10_+_180404
|
0.165
|
NM_006624
NM_001202465
NM_001202466
NM_212479
|
ZMYND11
|
zinc finger, MYND-type containing 11
|
|
chr3_-_125094003
|
0.162
|
NM_021964
|
ZNF148
|
zinc finger protein 148
|
|
chr18_+_7567313
|
0.162
|
NM_001105244
NM_002845
|
PTPRM
|
protein tyrosine phosphatase, receptor type, M
|
|
chr10_-_7829906
|
0.160
|
NM_012311
|
KIN
|
KIN, antigenic determinant of recA protein homolog (mouse)
|
|
chr22_-_41842853
|
0.160
|
NM_016272
|
TOB2
|
transducer of ERBB2, 2
|
|
chr4_-_40631846
|
0.159
|
NM_001098634
|
RBM47
|
RNA binding motif protein 47
|
|
chr3_+_171757413
|
0.159
|
NM_022763
|
FNDC3B
|
fibronectin type III domain containing 3B
|
|
chr1_-_155904186
|
0.157
|
NM_014949
|
KIAA0907
|
KIAA0907
|
|
chr2_+_46524448
|
0.155
|
NM_001430
|
EPAS1
|
endothelial PAS domain protein 1
|
|
chr10_-_103815868
|
0.154
|
NM_024541
|
C10orf76
|
chromosome 10 open reading frame 76
|
|
chr7_-_27205148
|
0.152
|
NM_152739
|
HOXA9
|
homeobox A9
|
|
chr16_-_67840464
|
0.151
|
NM_020850
|
RANBP10
|
RAN binding protein 10
|
|
chr11_+_111895537
|
0.148
|
NM_001931
|
DLAT
|
dihydrolipoamide S-acetyltransferase
|
|
chr12_-_49177876
|
0.147
|
NM_015270
|
ADCY6
|
adenylate cyclase 6
|
|
chr17_-_17399494
|
0.146
|
NM_001199989
NM_016084
|
RASD1
|
RAS, dexamethasone-induced 1
|
|
chr17_-_44270144
|
0.146
|
NM_001193466
|
KIAA1267
|
KIAA1267
|
|
chr12_+_64238511
|
0.146
|
NM_020762
|
SRGAP1
|
SLIT-ROBO Rho GTPase activating protein 1
|
|
chr2_+_85980600
|
0.146
|
NM_032827
|
ATOH8
|
atonal homolog 8 (Drosophila)
|
|
chr11_+_63655986
|
0.142
|
NM_017490
|
MARK2
|
MAP/microtubule affinity-regulating kinase 2
|
|
chr8_+_21946636
|
0.142
|
NM_022749
|
FAM160B2
|
family with sequence similarity 160, member B2
|
|
chr15_+_96876568
|
0.137
|
NM_001145157
|
NR2F2
|
nuclear receptor subfamily 2, group F, member 2
|
|
chr6_+_43138919
|
0.136
|
NM_003131
|
SRF
|
serum response factor (c-fos serum response element-binding transcription factor)
|
|
chr3_-_47823386
|
0.133
|
NM_003074
|
SMARCC1
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily c, member 1
|
|
chr6_+_36644236
|
0.129
|
NM_001220777
NM_078467
|
CDKN1A
|
cyclin-dependent kinase inhibitor 1A (p21, Cip1)
|
|
chr3_+_2280512
|
0.128
|
NM_001206955
|
CNTN4
|
contactin 4
|
|
chr11_-_16430425
|
0.124
|
NM_001145811
|
SOX6
|
SRY (sex determining region Y)-box 6
|
|
chr14_+_55518313
|
0.124
|
NM_144578
|
MAPK1IP1L
|
mitogen-activated protein kinase 1 interacting protein 1-like
|
|
chr10_-_103874661
|
0.122
|
NM_003893
|
LDB1
|
LIM domain binding 1
|
|
chr13_+_29598746
|
0.119
|
NM_001033602
|
MTUS2
|
microtubule associated tumor suppressor candidate 2
|
|
chr1_-_28415071
|
0.118
|
NM_001990
|
EYA3
|
eyes absent homolog 3 (Drosophila)
|
|
chr1_-_91487769
|
0.118
|
NM_032186
NM_201269
|
ZNF644
|
zinc finger protein 644
|