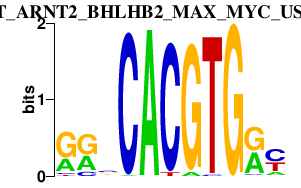
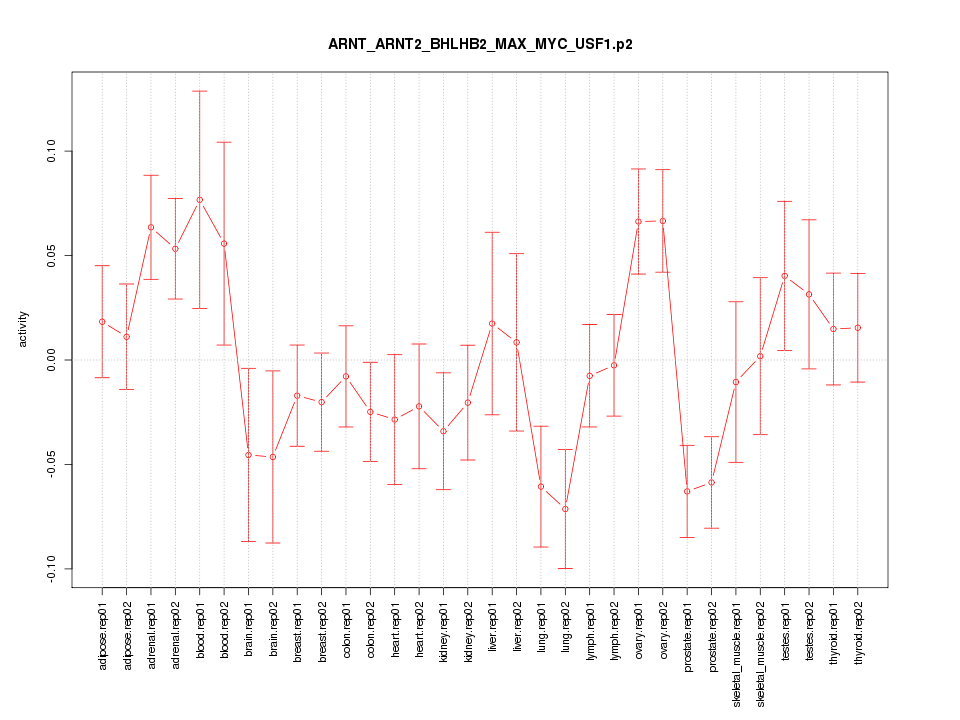
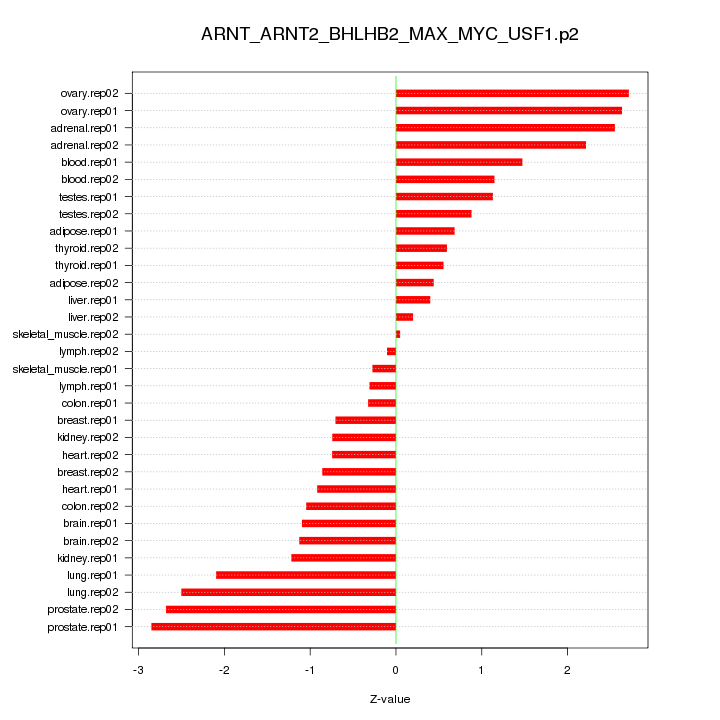
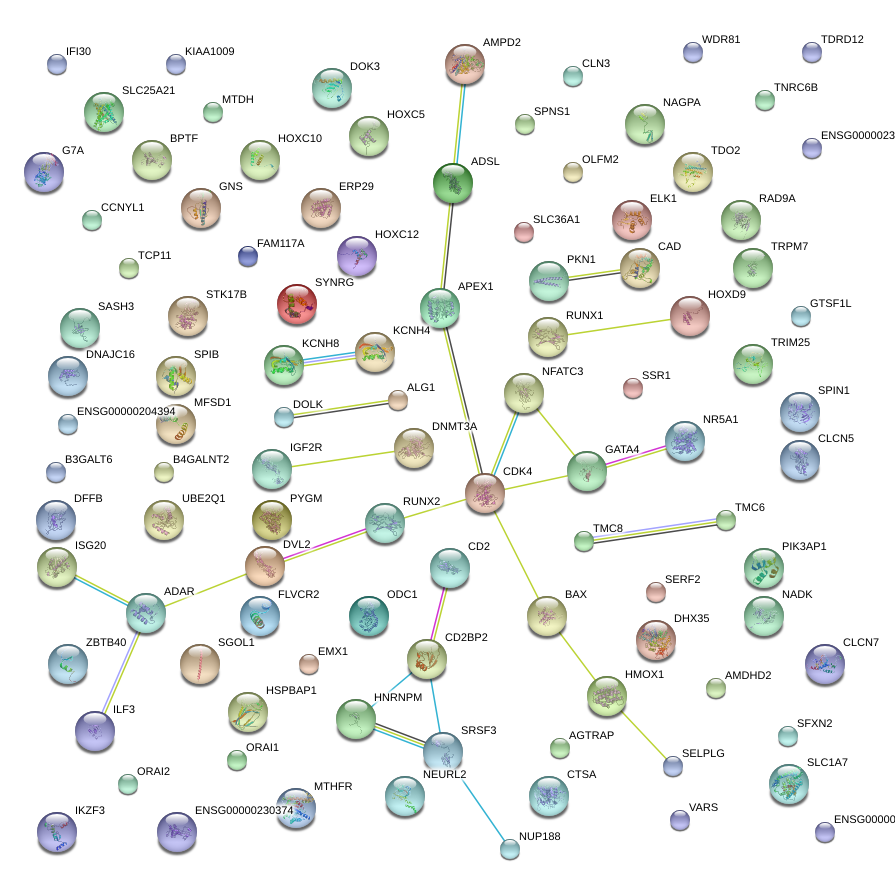
|
chr16_+_2570322
|
3.194
|
NM_001145815
NM_015944
|
AMDHD2
|
amidohydrolase domain containing 2
|
|
chr16_-_1525020
|
2.929
|
|
CLCN7
|
chloride channel 7
|
|
chr16_-_1524926
|
2.887
|
|
CLCN7
|
chloride channel 7
|
|
chr6_+_160390128
|
2.877
|
NM_000876
|
IGF2R
|
insulin-like growth factor 2 receptor
|
|
chr11_+_67159421
|
2.816
|
NM_004584
|
RAD9A
|
RAD9 homolog A (S. pombe)
|
|
chr20_-_44519629
|
2.753
|
NM_080749
|
NEURL2
|
neuralized homolog 2 (Drosophila)
|
|
chr16_-_1525006
|
2.723
|
|
CLCN7
|
chloride channel 7
|
|
chr9_-_127269689
|
2.415
|
NM_004959
|
NR5A1
|
nuclear receptor subfamily 5, group A, member 1
|
|
chr17_+_1628032
|
2.276
|
NM_001163809
|
WDR81
|
WD repeat domain 81
|
|
chr1_-_11866079
|
2.082
|
NM_005957
|
MTHFR
|
methylenetetrahydrofolate reductase (NAD(P)H)
|
|
chr17_-_56409868
|
1.999
|
|
|
|
|
chr9_-_131709783
|
1.944
|
NM_014908
|
DOLK
|
dolichol kinase
|
|
chr12_-_58146108
|
1.934
|
|
CDK4
|
cyclin-dependent kinase 4
|
|
chr12_+_54426831
|
1.904
|
NM_018953
|
HOXC5
|
homeobox C5
|
|
chr22_+_40573879
|
1.878
|
NM_001162501
NM_015088
|
TNRC6B
|
trinucleotide repeat containing 6B
|
|
chr3_-_122512526
|
1.871
|
NM_024610
|
HSPBAP1
|
HSPB (heat shock 27kDa) associated protein 1
|
|
chr19_+_50922194
|
1.861
|
NM_001243998
NM_001243999
NM_001244000
NM_003121
|
SPIB
|
Spi-B transcription factor (Spi-1/PU.1 related)
|
|
chr16_-_1525040
|
1.828
|
NM_001114331
NM_001287
|
CLCN7
|
chloride channel 7
|
|
chr17_-_7137722
|
1.821
|
NM_004422
|
DVL2
|
dishevelled, dsh homolog 2 (Drosophila)
|
|
chr1_+_117297056
|
1.816
|
NM_001767
|
CD2
|
CD2 molecule
|
|
chr7_+_102074002
|
1.794
|
|
ORAI2
|
ORAI calcium release-activated calcium modulator 2
|
|
chr12_-_58146025
|
1.785
|
|
CDK4
|
cyclin-dependent kinase 4
|
|
chr1_-_11865963
|
1.741
|
|
MTHFR
|
methylenetetrahydrofolate reductase (NAD(P)H)
|
|
chr17_+_1627833
|
1.705
|
NM_001163811
|
WDR81
|
WD repeat domain 81
|
|
chrX_+_128913881
|
1.689
|
NM_018990
|
SASH3
|
SAM and SH3 domain containing 3
|
|
chr3_+_158519911
|
1.673
|
|
MFSD1
|
major facilitator superfamily domain containing 1
|
|
chr12_-_58146110
|
1.620
|
|
CDK4
|
cyclin-dependent kinase 4
|
|
chr17_-_35969401
|
1.592
|
NM_001163544
NM_001163545
NM_001163546
NM_001163547
NM_007247
NM_080550
NM_198882
|
SYNRG
|
synergin, gamma
|
|
chrX_+_128913897
|
1.588
|
|
SASH3
|
SAM and SH3 domain containing 3
|
|
chr3_+_158519888
|
1.566
|
|
MFSD1
|
major facilitator superfamily domain containing 1
|
|
chr17_-_54991399
|
1.555
|
NM_005082
|
TRIM25
|
tripartite motif containing 25
|
|
chr19_+_10765106
|
1.547
|
|
ILF3
|
interleukin enhancer binding factor 3, 90kDa
|
|
chr20_+_44519965
|
1.542
|
|
CTSA
|
cathepsin A
|
|
chr19_+_33210678
|
1.536
|
NM_001110822
|
TDRD12
|
tudor domain containing 12
|
|
chr1_+_11796128
|
1.533
|
NM_001040194
NM_001040195
NM_001040196
NM_001040197
NM_020350
|
AGTRAP
|
angiotensin II receptor-associated protein
|
|
chr16_-_5083933
|
1.520
|
NM_016256
|
NAGPA
|
N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase
|
|
chr2_+_27440342
|
1.471
|
|
CAD
|
carbamoyl-phosphate synthetase 2, aspartate transcarbamylase, and dihydroorotase
|
|
chr8_+_11561710
|
1.471
|
NM_002052
|
GATA4
|
GATA binding protein 4
|
|
chr9_+_131709991
|
1.452
|
|
NUP188
|
nucleoporin 188kDa
|
|
chr8_+_98656329
|
1.449
|
NM_178812
|
MTDH
|
metadherin
|
|
chr15_-_50978921
|
1.441
|
|
TRPM7
|
transient receptor potential cation channel, subfamily M, member 7
|
|
chr19_+_49458151
|
1.440
|
|
BAX
|
BCL2-associated X protein
|
|
chr2_-_25474948
|
1.436
|
NM_153759
|
DNMT3A
|
DNA (cytosine-5-)-methyltransferase 3 alpha
|
|
chr10_+_104474297
|
1.417
|
NM_178858
|
SFXN2
|
sideroflexin 2
|
|
chr15_-_50978726
|
1.389
|
|
TRPM7
|
transient receptor potential cation channel, subfamily M, member 7
|
|
chr1_-_154600446
|
1.386
|
NM_001025107
|
ADAR
|
adenosine deaminase, RNA-specific
|
|
chr1_-_1710228
|
1.371
|
NM_001198993
|
NADK
|
NAD kinase
|
|
chr2_+_208576239
|
1.370
|
NM_001142300
NM_152523
|
CCNYL1
|
cyclin Y-like 1
|
|
chr5_-_176937337
|
1.367
|
NM_001144875
NM_001144876
|
DOK3
|
docking protein 3
|
|
chr20_+_37590969
|
1.362
|
NM_001190809
NM_021931
|
DHX35
|
DEAH (Asp-Glu-Ala-His) box polypeptide 35
|
|
chr9_+_131709970
|
1.356
|
NM_015354
|
NUP188
|
nucleoporin 188kDa
|
|
chr17_-_38020421
|
1.354
|
NM_012481
NM_183228
NM_183229
NM_183230
NM_183231
NM_183232
|
IKZF3
|
IKAROS family zinc finger 3 (Aiolos)
|
|
chr20_+_44519924
|
1.350
|
|
CTSA
|
cathepsin A
|
|
chr17_-_7137580
|
1.339
|
|
DVL2
|
dishevelled, dsh homolog 2 (Drosophila)
|
|
chr12_+_112451261
|
1.337
|
|
ERP29
|
endoplasmic reticulum protein 29
|
|
chr19_-_10047014
|
1.334
|
NM_058164
|
OLFM2
|
olfactomedin 2
|
|
chr2_-_10588451
|
1.330
|
NM_002539
|
ODC1
|
ornithine decarboxylase 1
|
|
chr16_-_5083913
|
1.317
|
|
NAGPA
|
N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase
|
|
chr2_-_197036301
|
1.316
|
NM_004226
|
STK17B
|
serine/threonine kinase 17b
|
|
chr20_+_44520021
|
1.311
|
|
CTSA
|
cathepsin A
|
|
chr15_+_89182038
|
1.308
|
NM_002201
|
ISG20
|
interferon stimulated exonuclease gene 20kDa
|
|
chr6_-_84937313
|
1.290
|
NM_014895
|
KIAA1009
|
KIAA1009
|
|
chr12_-_58146118
|
1.268
|
NM_000075
|
CDK4
|
cyclin-dependent kinase 4
|
|
chr1_+_11796211
|
1.266
|
|
AGTRAP
|
angiotensin II receptor-associated protein
|
|
chr1_-_154600383
|
1.266
|
|
ADAR
|
adenosine deaminase, RNA-specific
|
|
chr2_+_176987412
|
1.262
|
NM_014213
|
HOXD9
|
homeobox D9
|
|
chr3_-_20227618
|
1.255
|
NM_001012409
NM_001012410
NM_001012411
NM_001012412
NM_001012413
NM_138484
NM_001199251
NM_001199252
NM_001199253
NM_001199254
NM_001199255
NM_001199256
NM_001199257
|
SGOL1
|
shugoshin-like 1 (S. pombe)
|
|
chr12_+_112451229
|
1.250
|
|
|
|
|
chr20_-_42355566
|
1.245
|
NM_001008901
NM_176791
|
GTSF1L
|
gametocyte specific factor 1-like
|
|
chr3_+_158519714
|
1.244
|
NM_001167903
NM_022736
|
MFSD1
|
major facilitator superfamily domain containing 1
|
|
chr1_+_111742946
|
1.234
|
|
|
|
|
chr6_-_7313352
|
1.224
|
NM_003144
|
SSR1
|
signal sequence receptor, alpha
|
|
chr1_+_11796177
|
1.199
|
|
AGTRAP
|
angiotensin II receptor-associated protein
|
|
chr17_-_47841517
|
1.195
|
NM_030802
|
FAM117A
|
family with sequence similarity 117, member A
|
|
chr1_+_22778343
|
1.191
|
NM_001083621
NM_014870
|
ZBTB40
|
zinc finger and BTB domain containing 40
|
|
chr1_-_53608120
|
1.191
|
NM_006671
|
SLC1A7
|
solute carrier family 1 (glutamate transporter), member 7
|
|
chr1_-_154531083
|
1.189
|
NM_017582
|
UBE2Q1
|
ubiquitin-conjugating enzyme E2Q family member 1
|
|
chr16_+_68119211
|
1.176
|
NM_004555
NM_173163
NM_173165
|
NFATC3
|
nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 3
|
|
chr6_-_35109003
|
1.172
|
NM_001093728
NM_018679
|
TCP11
|
t-complex 11 homolog (mouse)
|
|
chr16_+_28986024
|
1.170
|
NM_001142448
NM_001142449
NM_001142450
NM_001142451
NM_032038
|
SPIN1
SPNS1
|
spindlin 1
spinster homolog 1 (Drosophila)
|
|
chrX_+_49832214
|
1.163
|
NM_000084
|
CLCN5
|
chloride channel 5
|
|
chr15_-_50978960
|
1.161
|
|
TRPM7
|
transient receptor potential cation channel, subfamily M, member 7
|
|
chr17_-_7137675
|
1.160
|
|
DVL2
|
dishevelled, dsh homolog 2 (Drosophila)
|
|
chr16_-_28503079
|
1.158
|
NM_000086
|
CLN3
|
ceroid-lipofuscinosis, neuronal 3
|
|
chr6_+_36562135
|
1.149
|
|
SRSF3
|
serine/arginine-rich splicing factor 3
|
|
chr20_+_44519590
|
1.149
|
NM_000308
NM_001127695
NM_001167594
|
CTSA
|
cathepsin A
|
|
chr19_+_18284631
|
1.147
|
|
IFI30
|
interferon, gamma-inducible protein 30
|
|
chr12_+_54348683
|
1.145
|
NM_173860
|
HOXC12
|
homeobox C12
|
|
chr16_+_5121809
|
1.130
|
NM_019109
|
ALG1
|
asparagine-linked glycosylation 1, beta-1,4-mannosyltransferase homolog (S. cerevisiae)
|
|
chr1_+_15853348
|
1.124
|
NM_015291
|
DNAJC16
|
DnaJ (Hsp40) homolog, subfamily C, member 16
|
|
chr17_+_76126835
|
1.096
|
NM_152468
|
TMC8
|
transmembrane channel-like 8
|
|
chr17_-_76124767
|
1.069
|
|
TMC6
|
transmembrane channel-like 6
|
|
chr22_+_35777087
|
1.067
|
|
HMOX1
|
heme oxygenase (decycling) 1
|
|
chr12_+_54378929
|
1.063
|
NM_017409
|
HOXC10
|
homeobox C10
|
|
chr21_-_36421586
|
1.062
|
NM_001754
|
RUNX1
|
runt-related transcription factor 1
|
|
chr12_-_65153182
|
1.060
|
|
GNS
|
glucosamine (N-acetyl)-6-sulfatase
|
|
chr19_+_8509875
|
1.052
|
|
HNRNPM
|
heterogeneous nuclear ribonucleoprotein M
|
|
chr17_+_65821625
|
1.049
|
NM_004459
NM_182641
|
BPTF
|
bromodomain PHD finger transcription factor
|
|
chr6_-_31763415
|
1.048
|
|
VARS
|
valyl-tRNA synthetase
|
|
chr5_+_150827138
|
1.046
|
NM_078483
|
SLC36A1
|
solute carrier family 36 (proton/amino acid symporter), member 1
|
|
chr1_+_1167622
|
1.045
|
NM_080605
|
B3GALT6
|
UDP-Gal:betaGal beta 1,3-galactosyltransferase polypeptide 6
|
|
chr15_+_44084517
|
1.042
|
NM_001199878
|
SERF2-C15ORF63
SERF2
|
SERF2-C15orf63 readthrough
small EDRK-rich factor 2
|
|
chr2_+_27440307
|
1.037
|
|
|
|
|
chr1_+_110162427
|
1.024
|
NM_139156
|
AMPD2
|
adenosine monophosphate deaminase 2
|
|
chrX_-_47509864
|
1.021
|
|
ELK1
|
ELK1, member of ETS oncogene family
|
|
chr21_-_36421630
|
1.016
|
|
RUNX1
|
runt-related transcription factor 1
|
|
chr10_-_98480174
|
1.014
|
NM_152309
|
PIK3AP1
|
phosphoinositide-3-kinase adaptor protein 1
|
|
chr14_+_76044939
|
1.013
|
NM_017791
|
FLVCR2
|
feline leukemia virus subgroup C cellular receptor family, member 2
|
|
chr12_-_65153189
|
1.013
|
NM_002076
|
GNS
|
glucosamine (N-acetyl)-6-sulfatase
|
|
chr2_-_197036262
|
1.009
|
|
STK17B
|
serine/threonine kinase 17b
|
|
chr17_-_47841468
|
1.008
|
|
FAM117A
|
family with sequence similarity 117, member A
|
|
chr12_-_109027654
|
1.001
|
NM_003006
|
SELPLG
|
selectin P ligand
|
|
chr19_+_14544216
|
0.996
|
|
PKN1
|
protein kinase N1
|
|
chr12_+_122064448
|
0.994
|
NM_032790
|
ORAI1
|
ORAI calcium release-activated calcium modulator 1
|
|
chr14_+_20923341
|
0.991
|
|
APEX1
|
APEX nuclease (multifunctional DNA repair enzyme) 1
|
|
chr14_-_106321626
|
0.990
|
|
|
|
|
chr2_-_85108202
|
0.989
|
|
C2orf89
|
chromosome 2 open reading frame 89
|
|
chr15_-_50978983
|
0.988
|
NM_017672
|
TRPM7
|
transient receptor potential cation channel, subfamily M, member 7
|
|
chr1_+_110162459
|
0.987
|
|
AMPD2
|
adenosine monophosphate deaminase 2
|
|
chr5_-_138725544
|
0.985
|
NM_016459
|
MZB1
|
marginal zone B and B1 cell-specific protein
|
|
chr19_-_15575333
|
0.984
|
NM_022904
|
RASAL3
|
RAS protein activator like 3
|
|
chr14_-_68283293
|
0.978
|
NM_015346
|
ZFYVE26
|
zinc finger, FYVE domain containing 26
|
|
chr19_+_8509859
|
0.978
|
|
HNRNPM
|
heterogeneous nuclear ribonucleoprotein M
|
|
chr20_+_44520203
|
0.975
|
|
CTSA
|
cathepsin A
|
|
chr14_-_20923266
|
0.974
|
NM_017807
|
OSGEP
|
O-sialoglycoprotein endopeptidase
|
|
chr17_+_78075360
|
0.966
|
|
GAA
|
glucosidase, alpha; acid
|
|
chr12_-_65153148
|
0.966
|
|
GNS
|
glucosamine (N-acetyl)-6-sulfatase
|
|
chr19_+_32836500
|
0.965
|
NM_001136156
NM_014910
|
ZNF507
|
zinc finger protein 507
|
|
chr3_+_158519877
|
0.964
|
|
MFSD1
|
major facilitator superfamily domain containing 1
|
|
chr12_-_109027615
|
0.964
|
|
SELPLG
|
selectin P ligand
|
|
chr12_-_65153055
|
0.959
|
|
GNS
|
glucosamine (N-acetyl)-6-sulfatase
|
|
chr10_-_27703178
|
0.950
|
NM_001034842
|
PTCHD3
|
patched domain containing 3
|
|
chr1_-_27226928
|
0.949
|
|
GPATCH3
|
G patch domain containing 3
|
|
chr19_+_1275510
|
0.946
|
NM_017914
|
C19orf24
|
chromosome 19 open reading frame 24
|
|
chr1_-_1709863
|
0.939
|
NM_023018
|
NADK
|
NAD kinase
|
|
chr12_-_110434177
|
0.938
|
|
GIT2
|
G protein-coupled receptor kinase interacting ArfGAP 2
|
|
chr19_+_1067293
|
0.931
|
|
HMHA1
|
histocompatibility (minor) HA-1
|
|
chr12_+_112451151
|
0.931
|
NM_001034025
NM_006817
|
ERP29
|
endoplasmic reticulum protein 29
|
|
chr2_+_27440233
|
0.929
|
NM_004341
|
CAD
|
carbamoyl-phosphate synthetase 2, aspartate transcarbamylase, and dihydroorotase
|
|
chr3_+_51428698
|
0.923
|
NM_013286
|
RBM15B
|
RNA binding motif protein 15B
|
|
chr14_+_20923289
|
0.923
|
NM_001244249
NM_001641
NM_080648
NM_080649
|
APEX1
|
APEX nuclease (multifunctional DNA repair enzyme) 1
|
|
chr7_+_102073953
|
0.918
|
NM_001126340
NM_032831
|
ORAI2
|
ORAI calcium release-activated calcium modulator 2
|
|
chr1_-_212873104
|
0.917
|
|
BATF3
|
basic leucine zipper transcription factor, ATF-like 3
|
|
chr15_+_44084493
|
0.912
|
|
SERF2
|
small EDRK-rich factor 2
|
|
chr1_+_153190059
|
0.911
|
NM_001195571
|
PRR9
|
proline rich 9
|
|
chr11_+_62538774
|
0.910
|
NM_006473
|
TAF6L
|
TAF6-like RNA polymerase II, p300/CBP-associated factor (PCAF)-associated factor, 65kDa
|
|
chr10_-_46090191
|
0.909
|
NM_001002265
NM_145021
|
MARCH8
|
membrane-associated ring finger (C3HC4) 8
|
|
chrX_+_100663205
|
0.908
|
|
HNRNPH2
|
heterogeneous nuclear ribonucleoprotein H2 (H')
|
|
chr1_+_17248425
|
0.908
|
NM_014675
|
CROCC
|
ciliary rootlet coiled-coil, rootletin
|
|
chr11_-_18343670
|
0.908
|
NM_007216
NM_181507
NM_181508
|
HPS5
|
Hermansky-Pudlak syndrome 5
|
|
chr11_-_71814371
|
0.902
|
NM_017907
|
LAMTOR1
|
late endosomal/lysosomal adaptor, MAPK and MTOR activator 1
|
|
chr14_-_64761077
|
0.901
|
NM_001040275
NM_001437
|
ESR2
|
estrogen receptor 2 (ER beta)
|
|
chr12_+_54674531
|
0.895
|
|
HNRNPA1L2
HNRNPA1
|
heterogeneous nuclear ribonucleoprotein A1-like 2
heterogeneous nuclear ribonucleoprotein A1
|
|
chr10_-_25304865
|
0.894
|
NM_145010
|
ENKUR
|
enkurin, TRPC channel interacting protein
|
|
chr2_-_85108238
|
0.887
|
NM_001080824
|
C2orf89
|
chromosome 2 open reading frame 89
|
|
chr20_-_52210780
|
0.884
|
|
ZNF217
|
zinc finger protein 217
|
|
chr11_-_71814335
|
0.883
|
|
LAMTOR1
|
late endosomal/lysosomal adaptor, MAPK and MTOR activator 1
|
|
chr12_-_109027565
|
0.882
|
|
SELPLG
|
selectin P ligand
|
|
chr6_-_31763473
|
0.875
|
|
VARS
|
valyl-tRNA synthetase
|
|
chrX_-_47509993
|
0.873
|
NM_001114123
NM_005229
|
ELK1
|
ELK1, member of ETS oncogene family
|
|
chr10_+_46222647
|
0.868
|
NM_001169106
NM_001169107
NM_015262
|
FAM21C
|
family with sequence similarity 21, member C
|
|
chr1_-_111743261
|
0.867
|
NM_024901
|
DENND2D
|
DENN/MADD domain containing 2D
|
|
chr12_+_53645869
|
0.860
|
NM_032889
|
MFSD5
|
major facilitator superfamily domain containing 5
|
|
chr2_+_46926049
|
0.860
|
NM_014011
|
SOCS5
|
suppressor of cytokine signaling 5
|
|
chr12_+_132413776
|
0.854
|
NM_001002019
NM_025215
|
PUS1
|
pseudouridylate synthase 1
|
|
chr1_+_29213569
|
0.854
|
NM_001166005
NM_001166007
NM_004437
NM_203342
NM_203343
|
EPB41
|
erythrocyte membrane protein band 4.1 (elliptocytosis 1, RH-linked)
|
|
chrX_+_100663213
|
0.854
|
|
HNRNPH2
|
heterogeneous nuclear ribonucleoprotein H2 (H')
|
|
chr12_+_54332575
|
0.853
|
NM_017410
|
HOXC13
|
homeobox C13
|
|
chr15_-_31393906
|
0.848
|
NM_001252024
NM_001252030
NM_002420
|
TRPM1
|
transient receptor potential cation channel, subfamily M, member 1
|
|
chr14_-_47120940
|
0.847
|
NM_080746
|
RPL10L
|
ribosomal protein L10-like
|
|
chr12_+_54366909
|
0.846
|
NM_014212
|
HOXC11
|
homeobox C11
|
|
chr11_+_33037397
|
0.846
|
NM_001077242
|
DEPDC7
|
DEP domain containing 7
|
|
chr12_+_53645906
|
0.845
|
|
MFSD5
|
major facilitator superfamily domain containing 5
|
|
chr12_-_121341921
|
0.844
|
|
SPPL3
|
signal peptide peptidase-like 3
|
|
chr19_+_1071208
|
0.844
|
|
HMHA1
|
histocompatibility (minor) HA-1
|
|
chr20_-_33413368
|
0.844
|
NM_001242539
NM_014071
|
NCOA6
|
nuclear receptor coactivator 6
|
|
chr7_+_99070513
|
0.842
|
NM_001013258
NM_213603
|
ZNF789
|
zinc finger protein 789
|
|
chr4_-_164394885
|
0.840
|
NM_032136
|
TKTL2
|
transketolase-like 2
|
|
chr16_-_58718607
|
0.837
|
NM_018231
|
SLC38A7
|
solute carrier family 38, member 7
|
|
chr14_-_20922841
|
0.835
|
|
OSGEP
|
O-sialoglycoprotein endopeptidase
|
|
chr18_+_9708269
|
0.831
|
|
RAB31
|
RAB31, member RAS oncogene family
|
|
chr17_+_78075390
|
0.828
|
|
GAA
|
glucosidase, alpha; acid
|
|
chr7_+_99775521
|
0.825
|
NM_012447
|
STAG3
|
stromal antigen 3
|
|
chr7_+_2671582
|
0.814
|
NM_025250
|
TTYH3
|
tweety homolog 3 (Drosophila)
|
|
chr13_+_49684388
|
0.812
|
NM_014923
|
FNDC3A
|
fibronectin type III domain containing 3A
|
|
chr17_-_7590731
|
0.811
|
NM_000546
NM_001126112
NM_001126113
NM_001126114
|
TP53
|
tumor protein p53
|
|
chr19_+_1067114
|
0.809
|
NM_012292
|
HMHA1
|
histocompatibility (minor) HA-1
|
|
chr22_-_24181173
|
0.808
|
NM_001002862
NM_001135751
NM_198440
|
DERL3
|
Der1-like domain family, member 3
|
|
chr2_+_220042963
|
0.807
|
|
FAM134A
|
family with sequence similarity 134, member A
|
|
chr16_-_4466607
|
0.804
|
|
CORO7
|
coronin 7
|
|
chr1_+_29241002
|
0.803
|
NM_001166006
|
EPB41
|
erythrocyte membrane protein band 4.1 (elliptocytosis 1, RH-linked)
|
|
chr16_-_4466595
|
0.802
|
|
CORO7
|
coronin 7
|
|
chr20_+_43595176
|
0.801
|
|
STK4
|
serine/threonine kinase 4
|
|
chr8_+_99129488
|
0.799
|
NM_001145860
NM_001145861
|
POP1
|
processing of precursor 1, ribonuclease P/MRP subunit (S. cerevisiae)
|
|
chr14_-_20922969
|
0.799
|
|
OSGEP
|
O-sialoglycoprotein endopeptidase
|
|
chr16_-_88923233
|
0.798
|
|
GALNS
|
galactosamine (N-acetyl)-6-sulfate sulfatase
|
|
chr7_-_27135524
|
0.797
|
NM_005522
NM_153620
|
HOXA1
|
homeobox A1
|
|
chr17_+_49337892
|
0.796
|
NM_016001
|
UTP18
|
UTP18 small subunit (SSU) processome component homolog (yeast)
|
|
chr7_+_2671630
|
0.795
|
|
TTYH3
|
tweety homolog 3 (Drosophila)
|
|
chr17_+_42422490
|
0.794
|
NM_002087
|
GRN
|
granulin
|