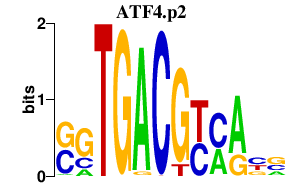
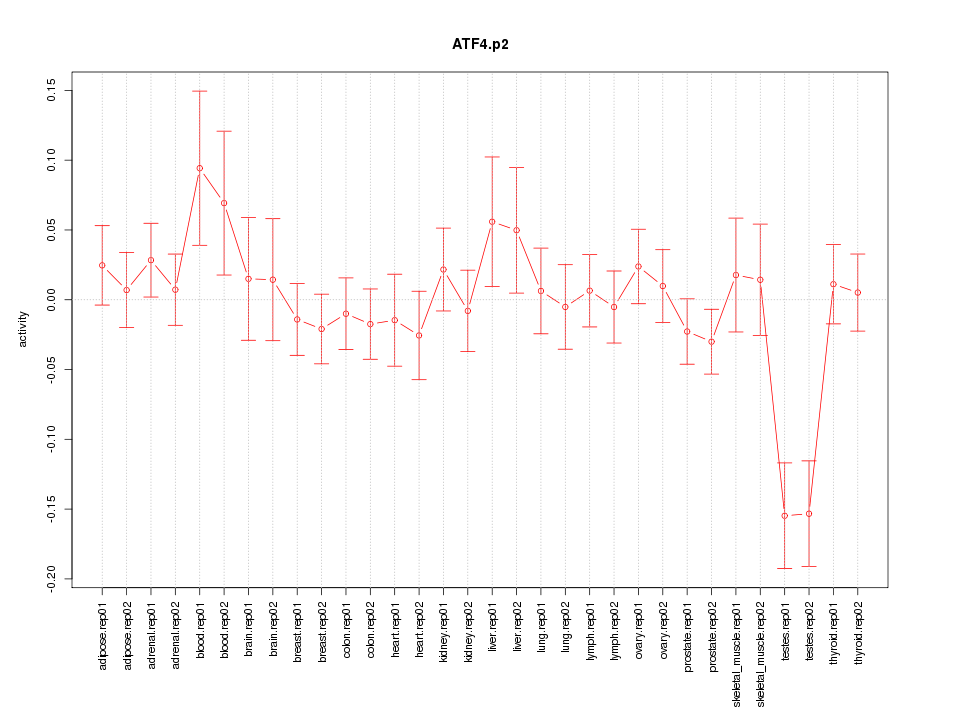
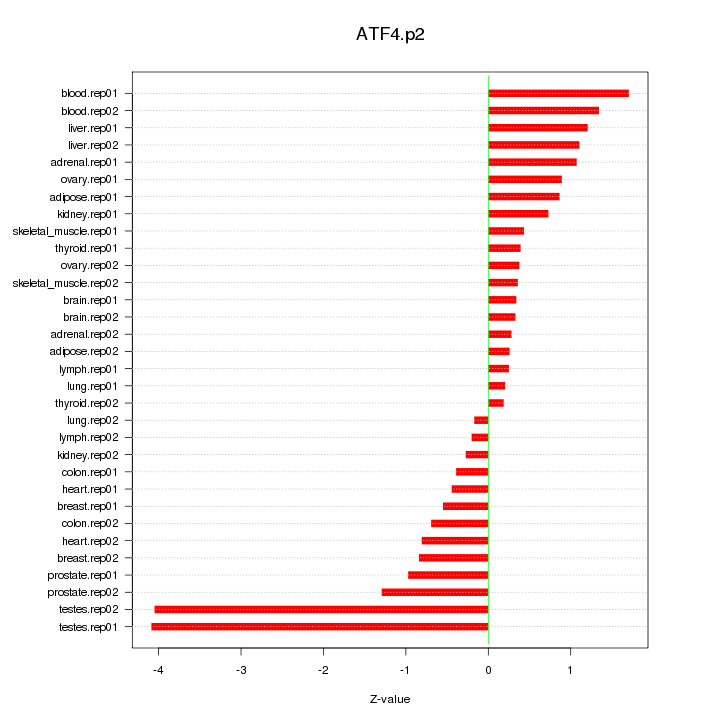
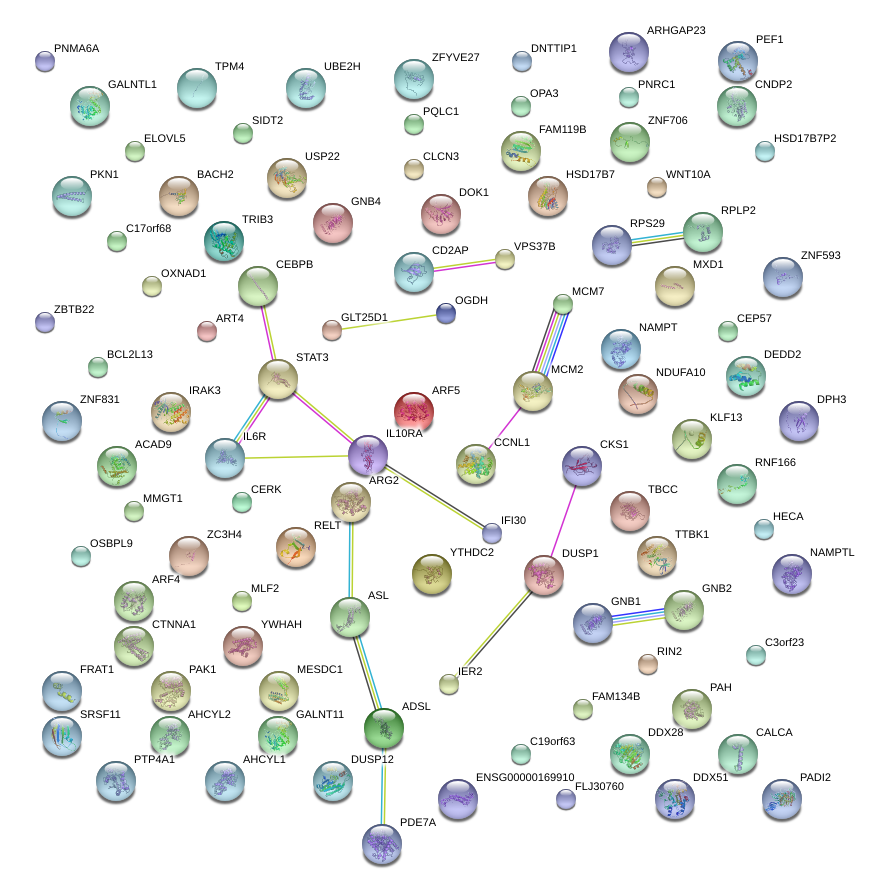
|
chr6_+_139456240
|
4.139
|
NM_016217
|
HECA
|
headcase homolog (Drosophila)
|
|
chr11_+_117857062
|
3.794
|
NM_001558
|
IL10RA
|
interleukin 10 receptor, alpha
|
|
chr11_-_77184549
|
2.410
|
NM_001128620
NM_002576
|
PAK1
|
p21 protein (Cdc42/Rac)-activated kinase 1
|
|
chr1_+_26496382
|
2.410
|
NM_015871
|
ZNF593
|
zinc finger protein 593
|
|
chr6_-_91006580
|
2.387
|
NM_001170794
|
BACH2
|
BTB and CNC homology 1, basic leucine zipper transcription factor 2
|
|
chr14_+_68086633
|
2.336
|
|
ARG2
|
arginase, type II
|
|
chr5_+_112849352
|
2.214
|
NM_022828
|
YTHDC2
|
YTH domain containing 2
|
|
chr6_-_91006460
|
2.164
|
NM_021813
|
BACH2
|
BTB and CNC homology 1, basic leucine zipper transcription factor 2
|
|
chr14_-_50329567
|
2.148
|
|
RN7SL2
|
RNA, 7SL, cytoplasmic 2
|
|
chr14_-_50053068
|
2.111
|
NM_001030001
NM_001032
|
RPS29
|
ribosomal protein S29
|
|
chr17_-_8151353
|
2.062
|
|
CTC1
|
CTS telomere maintenance complex component 1
|
|
chr7_-_105925329
|
2.035
|
|
NAMPT
|
nicotinamide phosphoribosyltransferase
|
|
chr11_+_117857123
|
1.941
|
|
IL10RA
|
interleukin 10 receptor, alpha
|
|
chr19_+_13263890
|
1.874
|
|
IER2
|
immediate early response 2
|
|
chr16_-_88772732
|
1.859
|
NM_001171815
|
RNF166
|
ring finger protein 166
|
|
chr19_+_18284578
|
1.858
|
NM_006332
|
IFI30
|
interferon, gamma-inducible protein 30
|
|
chr19_+_16187306
|
1.853
|
|
TPM4
|
tropomyosin 4
|
|
chr19_+_18284631
|
1.826
|
|
IFI30
|
interferon, gamma-inducible protein 30
|
|
chr18_+_72163582
|
1.810
|
|
CNDP2
|
CNDP dipeptidase 2 (metallopeptidase M20 family)
|
|
chr6_-_53213699
|
1.806
|
|
ELOVL5
|
ELOVL fatty acid elongase 5
|
|
chr17_-_8151373
|
1.715
|
NM_025099
|
CTC1
|
CTS telomere maintenance complex component 1
|
|
chr11_+_73087396
|
1.686
|
NM_152222
|
RELT
|
RELT tumor necrosis factor receptor
|
|
chr15_+_31619043
|
1.672
|
NM_015995
|
KLF13
|
Kruppel-like factor 13
|
|
chr18_+_72163335
|
1.669
|
NM_018235
|
CNDP2
|
CNDP dipeptidase 2 (metallopeptidase M20 family)
|
|
chrX_-_135055834
|
1.627
|
|
MMGT1
|
membrane magnesium transporter 1
|
|
chr12_-_123380682
|
1.619
|
NM_024667
|
VPS37B
|
vacuolar protein sorting 37 homolog B (S. cerevisiae)
|
|
chr7_-_105925410
|
1.509
|
NM_005746
|
NAMPT
|
nicotinamide phosphoribosyltransferase
|
|
chr19_+_16187315
|
1.475
|
|
TPM4
|
tropomyosin 4
|
|
chr11_+_809918
|
1.473
|
NM_001004
|
RPLP2
|
ribosomal protein, large, P2
|
|
chr12_-_123380608
|
1.472
|
|
|
|
|
chrX_+_152240726
|
1.457
|
NM_001170944
NM_001242319
NM_032882
|
PNMA6C
PNMA6D
PNMA6A
|
paraneoplastic antigen like 6C
paraneoplastic antigen like 6D
paraneoplastic antigen like 6A
|
|
chr19_-_42721802
|
1.453
|
NM_133328
|
DEDD2
|
death effector domain containing 2
|
|
chr8_-_66754276
|
1.439
|
|
PDE7A
|
phosphodiesterase 7A
|
|
chr1_+_154377847
|
1.436
|
|
IL6R
|
interleukin 6 receptor
|
|
chr12_-_132628841
|
1.431
|
NM_175066
|
DDX51
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 51
|
|
chr3_+_16306666
|
1.431
|
NM_138381
|
OXNAD1
|
oxidoreductase NAD-binding domain containing 1
|
|
chr7_+_44646197
|
1.344
|
|
OGDH
|
oxoglutarate (alpha-ketoglutarate) dehydrogenase (lipoamide)
|
|
chr7_+_44646189
|
1.341
|
|
OGDH
|
oxoglutarate (alpha-ketoglutarate) dehydrogenase (lipoamide)
|
|
chr7_+_65540821
|
1.323
|
NM_001024943
NM_001024944
NM_001024946
|
ASL
|
argininosuccinate lyase
|
|
chr16_-_68057098
|
1.319
|
|
DDX28
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 28
|
|
chr7_+_44646180
|
1.310
|
|
OGDH
|
oxoglutarate (alpha-ketoglutarate) dehydrogenase (lipoamide)
|
|
chr5_-_172198160
|
1.308
|
NM_004417
|
DUSP1
|
dual specificity phosphatase 1
|
|
chr6_-_42713837
|
1.305
|
NM_003192
|
TBCC
|
tubulin folding cofactor C
|
|
chr12_-_6862131
|
1.299
|
|
MLF2
|
myeloid leukemia factor 2
|
|
chr11_+_117049387
|
1.297
|
|
SIDT2
|
SID1 transmembrane family, member 2
|
|
chr16_-_68057120
|
1.289
|
|
DDX28
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 28
|
|
chr6_+_64281910
|
1.286
|
NM_003463
|
PTP4A1
|
protein tyrosine phosphatase type IVA, member 1
|
|
chr7_-_129592711
|
1.275
|
NM_001202498
|
UBE2H
|
ubiquitin-conjugating enzyme E2H
|
|
chr6_-_42713822
|
1.264
|
|
TBCC
|
tubulin folding cofactor C
|
|
chr19_+_16187266
|
1.264
|
|
TPM4
|
tropomyosin 4
|
|
chr7_+_44646168
|
1.262
|
|
OGDH
|
oxoglutarate (alpha-ketoglutarate) dehydrogenase (lipoamide)
|
|
chr10_+_99079017
|
1.254
|
NM_005479
|
FRAT1
|
frequently rearranged in advanced T-cell lymphomas
|
|
chr1_+_110527432
|
1.242
|
|
AHCYL1
|
adenosylhomocysteinase-like 1
|
|
chr1_-_17445865
|
1.236
|
NM_007365
|
PADI2
|
peptidyl arginine deiminase, type II
|
|
chr2_+_219745243
|
1.234
|
NM_025216
|
WNT10A
|
wingless-type MMTV integration site family, member 10A
|
|
chr11_+_809985
|
1.228
|
|
RPLP2
|
ribosomal protein, large, P2
|
|
chr5_+_138211084
|
1.222
|
|
CTNNA1
|
catenin (cadherin-associated protein), alpha 1, 102kDa
|
|
chr1_+_110527418
|
1.216
|
|
AHCYL1
|
adenosylhomocysteinase-like 1
|
|
chr7_-_129592794
|
1.215
|
NM_003344
NM_182697
|
UBE2H
|
ubiquitin-conjugating enzyme E2H
|
|
chr1_+_110527756
|
1.208
|
|
AHCYL1
|
adenosylhomocysteinase-like 1
|
|
chr7_+_44646220
|
1.208
|
|
OGDH
|
oxoglutarate (alpha-ketoglutarate) dehydrogenase (lipoamide)
|
|
chr7_-_45026204
|
1.206
|
|
SNHG15
|
small nucleolar RNA host gene 15 (non-protein coding)
|
|
chr20_+_57766074
|
1.199
|
NM_178457
|
ZNF831
|
zinc finger protein 831
|
|
chr6_-_53213586
|
1.175
|
|
ELOVL5
|
ELOVL fatty acid elongase 5
|
|
chr1_+_110527327
|
1.166
|
NM_001242673
NM_006621
|
AHCYL1
|
adenosylhomocysteinase-like 1
|
|
chr7_+_100273776
|
1.159
|
|
GNB2
|
guanine nucleotide binding protein (G protein), beta polypeptide 2
|
|
chr6_+_89791510
|
1.151
|
|
PNRC1
|
proline-rich nuclear receptor coactivator 1
|
|
chr12_-_123380628
|
1.148
|
|
VPS37B
|
vacuolar protein sorting 37 homolog B (S. cerevisiae)
|
|
chr3_-_16306283
|
1.146
|
NM_001047434
NM_206831
|
DPH3
|
DPH3, KTI11 homolog (S. cerevisiae)
|
|
chr11_-_14993790
|
1.139
|
NM_001033952
NM_001033953
NM_001741
|
CALCA
|
calcitonin-related polypeptide alpha
|
|
chr1_+_162760491
|
1.135
|
NM_016371
|
HSD17B7
|
hydroxysteroid (17-beta) dehydrogenase 7
|
|
chr4_+_170541690
|
1.128
|
|
CLCN3
|
chloride channel 3
|
|
chr19_-_47617008
|
1.124
|
NM_015168
|
ZC3H4
|
zinc finger CCCH-type containing 4
|
|
chr20_+_44420610
|
1.124
|
|
DNTTIP1
|
deoxynucleotidyltransferase, terminal, interacting protein 1
|
|
chr6_+_64282429
|
1.119
|
|
PTP4A1
|
protein tyrosine phosphatase type IVA, member 1
|
|
chr10_+_99496847
|
1.113
|
NM_001002262
NM_001174119
NM_001174120
NM_001174121
NM_001174122
NM_144588
|
ZFYVE27
|
zinc finger, FYVE domain containing 27
|
|
chr6_+_47445440
|
1.112
|
NM_012120
|
CD2AP
|
CD2-associated protein
|
|
chr4_+_170541728
|
1.091
|
|
CLCN3
|
chloride channel 3
|
|
chr1_-_32110464
|
1.087
|
|
PEF1
|
penta-EF-hand domain containing 1
|
|
chr19_+_50979788
|
1.086
|
|
C19orf63
|
chromosome 19 open reading frame 63
|
|
chr7_+_127228405
|
1.084
|
NM_001662
|
ARF5
|
ADP-ribosylation factor 5
|
|
chr18_-_77711603
|
1.078
|
|
PQLC1
|
PQ loop repeat containing 1
|
|
chr16_-_88772807
|
1.076
|
NM_178841
|
RNF166
|
ring finger protein 166
|
|
chr19_-_46088074
|
1.074
|
|
OPA3
|
optic atrophy 3 (autosomal recessive, with chorea and spastic paraplegia)
|
|
chr1_+_52082755
|
1.064
|
|
OSBPL9
|
oxysterol binding protein-like 9
|
|
chr22_+_32340435
|
1.061
|
NM_003405
|
YWHAH
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, eta polypeptide
|
|
chr7_+_151722826
|
1.054
|
|
GALNT11
|
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 11 (GalNAc-T11)
|
|
chr20_+_48807365
|
1.047
|
|
CEBPB
|
CCAAT/enhancer binding protein (C/EBP), beta
|
|
chr1_+_70671375
|
1.041
|
|
SRSF11
|
serine/arginine-rich splicing factor 11
|
|
chr3_-_156877882
|
1.027
|
|
CCNL1
|
cyclin L1
|
|
chr22_+_18121540
|
1.027
|
|
BCL2L13
|
BCL2-like 13 (apoptosis facilitator)
|
|
chr18_-_77711652
|
1.025
|
NM_001146343
NM_001146345
NM_025078
|
PQLC1
|
PQ loop repeat containing 1
|
|
chr12_+_66583029
|
1.019
|
|
IRAK3
|
interleukin-1 receptor-associated kinase 3
|
|
chr10_+_99496891
|
1.013
|
|
ZFYVE27
|
zinc finger, FYVE domain containing 27
|
|
chr6_+_64282449
|
1.006
|
|
PTP4A1
|
protein tyrosine phosphatase type IVA, member 1
|
|
chr7_+_65540937
|
0.998
|
|
ASL
|
argininosuccinate lyase
|
|
chr19_+_17666457
|
0.987
|
|
GLT25D1
|
glycosyltransferase 25 domain containing 1
|
|
chr2_-_240964717
|
0.984
|
|
NDUFA10
|
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 10, 42kDa
|
|
chr20_+_19915716
|
0.983
|
|
RIN2
|
Ras and Rab interactor 2
|
|
chr10_+_99496897
|
0.980
|
|
ZFYVE27
|
zinc finger, FYVE domain containing 27
|
|
chr5_-_16509106
|
0.978
|
NM_019000
|
FAM134B
|
family with sequence similarity 134, member B
|
|
chr19_+_50979772
|
0.974
|
|
C19orf63
|
chromosome 19 open reading frame 63
|
|
chr19_+_17666381
|
0.967
|
|
GLT25D1
|
glycosyltransferase 25 domain containing 1
|
|
chr22_-_47134044
|
0.964
|
|
CERK
|
ceramide kinase
|
|
chr17_-_40540576
|
0.951
|
|
STAT3
|
signal transducer and activator of transcription 3 (acute-phase response factor)
|
|
chr3_+_128598471
|
0.950
|
|
ACAD9
|
acyl-CoA dehydrogenase family, member 9
|
|
chr3_+_44380003
|
0.946
|
|
C3orf23
|
chromosome 3 open reading frame 23
|
|
chr15_+_81293286
|
0.945
|
NM_022566
|
MESDC1
|
mesoderm development candidate 1
|
|
chr12_-_103311053
|
0.943
|
|
PAH
|
phenylalanine hydroxylase
|
|
chr20_+_44420575
|
0.940
|
NM_052951
|
DNTTIP1
|
deoxynucleotidyltransferase, terminal, interacting protein 1
|
|
chr20_+_361675
|
0.939
|
|
TRIB3
|
tribbles homolog 3 (Drosophila)
|
|
chr3_+_128598459
|
0.933
|
|
ACAD9
|
acyl-CoA dehydrogenase family, member 9
|
|
chr17_-_20946088
|
0.927
|
NM_015276
|
USP22
|
ubiquitin specific peptidase 22
|
|
chr2_+_74781846
|
0.922
|
|
DOK1
|
docking protein 1, 62kDa (downstream of tyrosine kinase 1)
|
|
chr2_+_70142152
|
0.920
|
NM_001202513
NM_001202514
NM_002357
|
MXD1
|
MAX dimerization protein 1
|
|
chr20_+_44420606
|
0.918
|
|
DNTTIP1
|
deoxynucleotidyltransferase, terminal, interacting protein 1
|
|
chr6_+_43211221
|
0.918
|
NM_032538
|
TTBK1
|
tau tubulin kinase 1
|
|
chr7_+_151722774
|
0.917
|
NM_022087
|
GALNT11
|
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 11 (GalNAc-T11)
|
|
chr11_+_95523608
|
0.905
|
NM_001243776
NM_001243777
NM_014679
|
CEP57
|
centrosomal protein 57kDa
|
|
chr11_-_6633400
|
0.898
|
|
TAF10
|
TAF10 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 30kDa
|
|
chr15_-_41047421
|
0.895
|
|
FAM82A2
|
family with sequence similarity 82, member A2
|
|
chr1_-_150552005
|
0.887
|
|
MCL1
|
myeloid cell leukemia sequence 1 (BCL2-related)
|
|
chr1_+_110527701
|
0.882
|
|
AHCYL1
|
adenosylhomocysteinase-like 1
|
|
chr22_+_21271619
|
0.881
|
NM_005207
|
CRKL
|
v-crk sarcoma virus CT10 oncogene homolog (avian)-like
|
|
chr1_+_109289353
|
0.878
|
|
|
|
|
chr3_-_71632741
|
0.877
|
NM_001244808
NM_001012505
NM_001244810
NM_032682
|
FOXP1
|
forkhead box P1
|
|
chr20_+_48807460
|
0.869
|
|
CEBPB
|
CCAAT/enhancer binding protein (C/EBP), beta
|
|
chr22_+_21271721
|
0.867
|
|
CRKL
|
v-crk sarcoma virus CT10 oncogene homolog (avian)-like
|
|
chr14_+_68086655
|
0.863
|
|
ARG2
|
arginase, type II
|
|
chr19_+_18668652
|
0.861
|
|
KXD1
|
KxDL motif containing 1
|
|
chr3_+_128598497
|
0.861
|
|
ACAD9
|
acyl-CoA dehydrogenase family, member 9
|
|
chr12_+_66583004
|
0.860
|
|
IRAK3
|
interleukin-1 receptor-associated kinase 3
|
|
chr8_+_126442557
|
0.860
|
NM_025195
|
TRIB1
|
tribbles homolog 1 (Drosophila)
|
|
chr10_+_99496938
|
0.859
|
|
ZFYVE27
|
zinc finger, FYVE domain containing 27
|
|
chr9_-_70490145
|
0.850
|
|
CBWD5
|
COBW domain containing 5
|
|
chr3_+_44379923
|
0.845
|
NM_001029839
NM_173826
|
C3orf23
|
chromosome 3 open reading frame 23
|
|
chr20_-_17949384
|
0.844
|
NM_152227
|
SNX5
|
sorting nexin 5
|
|
chr13_-_52378230
|
0.841
|
NM_001031719
NM_024705
|
DHRS12
|
dehydrogenase/reductase (SDR family) member 12
|
|
chr3_+_141087298
|
0.841
|
|
ZBTB38
|
zinc finger and BTB domain containing 38
|
|
chr8_+_42128851
|
0.840
|
|
IKBKB
|
inhibitor of kappa light polypeptide gene enhancer in B-cells, kinase beta
|
|
chr3_+_128598440
|
0.837
|
|
ACAD9
|
acyl-CoA dehydrogenase family, member 9
|
|
chr6_-_32812409
|
0.836
|
|
PSMB8
|
proteasome (prosome, macropain) subunit, beta type, 8 (large multifunctional peptidase 7)
|
|
chr11_-_64889581
|
0.836
|
NM_001997
|
FAU
|
Finkel-Biskis-Reilly murine sarcoma virus (FBR-MuSV) ubiquitously expressed
|
|
chr19_+_1067536
|
0.836
|
|
HMHA1
|
histocompatibility (minor) HA-1
|
|
chr12_-_64616032
|
0.835
|
NM_152440
|
C12orf66
|
chromosome 12 open reading frame 66
|
|
chr4_-_8430185
|
0.833
|
|
ACOX3
|
acyl-CoA oxidase 3, pristanoyl
|
|
chr8_-_17104266
|
0.829
|
NM_013354
NM_054026
|
CNOT7
|
CCR4-NOT transcription complex, subunit 7
|
|
chr7_+_120590816
|
0.827
|
NM_019071
NM_198267
|
ING3
|
inhibitor of growth family, member 3
|
|
chr1_-_228290929
|
0.825
|
NM_024319
|
C1orf35
|
chromosome 1 open reading frame 35
|
|
chr14_+_55493837
|
0.824
|
NM_080867
NM_199421
|
SOCS4
|
suppressor of cytokine signaling 4
|
|
chr10_-_23003443
|
0.820
|
NM_005028
|
PIP4K2A
|
phosphatidylinositol-5-phosphate 4-kinase, type II, alpha
|
|
chr5_-_95297597
|
0.819
|
NM_012081
|
ELL2
|
elongation factor, RNA polymerase II, 2
|
|
chr11_+_60869929
|
0.818
|
NM_014207
|
CD5
|
CD5 molecule
|
|
chr9_+_132251323
|
0.816
|
|
LOC100506190
|
uncharacterized LOC100506190
|
|
chr15_-_41047448
|
0.815
|
NM_018145
|
FAM82A2
|
family with sequence similarity 82, member A2
|
|
chr11_-_67188614
|
0.814
|
|
PPP1CA
|
protein phosphatase 1, catalytic subunit, alpha isozyme
|
|
chr7_+_17338182
|
0.813
|
NM_001621
|
AHR
|
aryl hydrocarbon receptor
|
|
chr3_+_44379590
|
0.813
|
NM_001029840
|
C3orf23
|
chromosome 3 open reading frame 23
|
|
chr19_+_42388445
|
0.809
|
NM_199002
|
ARHGEF1
|
Rho guanine nucleotide exchange factor (GEF) 1
|
|
chr3_+_38206968
|
0.806
|
NM_005109
|
OXSR1
|
oxidative-stress responsive 1
|
|
chr2_-_202645653
|
0.800
|
|
ALS2
|
amyotrophic lateral sclerosis 2 (juvenile)
|
|
chr2_-_240964807
|
0.798
|
NM_004544
|
NDUFA10
|
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 10, 42kDa
|
|
chr9_-_178913
|
0.795
|
NM_001145355
NM_001145356
NM_018491
|
CBWD1
|
COBW domain containing 1
|
|
chr22_+_22550159
|
0.794
|
|
IGLV6-57
|
immunoglobulin lambda variable 6-57
|
|
chr12_+_69004708
|
0.789
|
|
RAP1B
|
RAP1B, member of RAS oncogene family
|
|
chr12_+_50135953
|
0.788
|
|
TMBIM6
|
transmembrane BAX inhibitor motif containing 6
|
|
chr1_+_154377668
|
0.787
|
NM_000565
NM_001206866
NM_181359
|
IL6R
|
interleukin 6 receptor
|
|
chr1_-_70671263
|
0.786
|
NM_017768
|
LRRC40
|
leucine rich repeat containing 40
|
|
chr17_-_56084656
|
0.784
|
NM_001078166
NM_006924
|
SRSF1
|
serine/arginine-rich splicing factor 1
|
|
chr10_+_64893003
|
0.782
|
NM_030759
|
NRBF2
|
nuclear receptor binding factor 2
|
|
chr19_+_50979656
|
0.781
|
NM_175063
NM_206538
|
C19orf63
|
chromosome 19 open reading frame 63
|
|
chr9_-_74979829
|
0.781
|
NM_001102420
NM_001102421
NM_006007
|
ZFAND5
|
zinc finger, AN1-type domain 5
|
|
chr11_+_62649160
|
0.778
|
|
SLC3A2
|
solute carrier family 3 (activators of dibasic and neutral amino acid transport), member 2
|
|
chr17_-_76778306
|
0.775
|
NM_004762
NM_017456
|
CYTH1
|
cytohesin 1
|
|
chr22_+_18121446
|
0.775
|
NM_015367
|
BCL2L13
|
BCL2-like 13 (apoptosis facilitator)
|
|
chr16_+_475605
|
0.771
|
|
RAB11FIP3
|
RAB11 family interacting protein 3 (class II)
|
|
chr19_+_45504686
|
0.769
|
NM_006509
|
RELB
|
v-rel reticuloendotheliosis viral oncogene homolog B
|
|
chr19_+_1941157
|
0.769
|
NM_001319
|
CSNK1G2
|
casein kinase 1, gamma 2
|
|
chr15_-_23034365
|
0.769
|
|
NIPA2
|
non imprinted in Prader-Willi/Angelman syndrome 2
|
|
chr3_-_33759743
|
0.768
|
|
CLASP2
|
cytoplasmic linker associated protein 2
|
|
chr7_-_994008
|
0.768
|
|
ADAP1
|
ArfGAP with dual PH domains 1
|
|
chr6_-_137540435
|
0.767
|
|
IFNGR1
|
interferon gamma receptor 1
|
|
chr19_+_18284619
|
0.764
|
|
IFI30
|
interferon, gamma-inducible protein 30
|
|
chr7_+_151722842
|
0.762
|
|
GALNT11
|
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 11 (GalNAc-T11)
|
|
chr20_+_30697512
|
0.762
|
|
TM9SF4
|
transmembrane 9 superfamily protein member 4
|
|
chr11_+_117049838
|
0.761
|
NM_001040455
|
SIDT2
|
SID1 transmembrane family, member 2
|
|
chr11_+_73087670
|
0.760
|
NM_032871
|
RELT
|
RELT tumor necrosis factor receptor
|
|
chr2_+_96068423
|
0.760
|
NM_016044
|
FAHD2A
|
fumarylacetoacetate hydrolase domain containing 2A
|
|
chr17_-_4890920
|
0.757
|
NM_001171166
NM_015099
|
CAMTA2
|
calmodulin binding transcription activator 2
|
|
chr8_+_42128834
|
0.757
|
|
IKBKB
|
inhibitor of kappa light polypeptide gene enhancer in B-cells, kinase beta
|
|
chr10_+_119000583
|
0.756
|
NM_003054
|
SLC18A2
|
solute carrier family 18 (vesicular monoamine), member 2
|
|
chr20_-_33865905
|
0.756
|
|
EDEM2
|
ER degradation enhancer, mannosidase alpha-like 2
|
|
chr2_+_61108702
|
0.753
|
|
REL
|
v-rel reticuloendotheliosis viral oncogene homolog (avian)
|
|
chr3_-_33759684
|
0.753
|
NM_015097
|
CLASP2
|
cytoplasmic linker associated protein 2
|
|
chr14_-_92588065
|
0.749
|
NM_004545
|
NDUFB1
|
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 1, 7kDa
|
|
chr5_-_32174355
|
0.749
|
|
GOLPH3
|
golgi phosphoprotein 3 (coat-protein)
|
|
chr16_+_11762235
|
0.748
|
NM_003498
|
SNN
|
stannin
|
|
chrX_-_152245957
|
0.748
|
NM_001242319
NM_032882
|
PNMA6D
PNMA6A
|
paraneoplastic antigen like 6D
paraneoplastic antigen like 6A
|
|
chr12_+_66582914
|
0.746
|
NM_001142523
NM_007199
|
IRAK3
|
interleukin-1 receptor-associated kinase 3
|
|
chr20_+_44637539
|
0.745
|
NM_004994
|
MMP9
|
matrix metallopeptidase 9 (gelatinase B, 92kDa gelatinase, 92kDa type IV collagenase)
|