Project
Illumina Body Map 2: averaged replicates
Navigation
Downloads
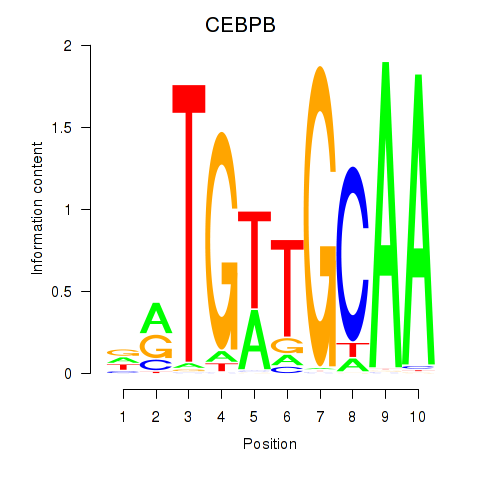
Results for CEBPB
Z-value: 2.61
Transcription factors associated with CEBPB
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
CEBPB
|
ENSG00000172216.4 | CCAAT enhancer binding protein beta |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| CEBPB | hg19_v2_chr20_+_48807351_48807384 | 0.31 | 8.5e-02 | Click! |
Activity profile of CEBPB motif
Sorted Z-values of CEBPB motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr20_+_56136136 | 9.15 |
ENST00000319441.4
ENST00000543666.1 |
PCK1
|
phosphoenolpyruvate carboxykinase 1 (soluble) |
| chr4_-_155533787 | 8.32 |
ENST00000407946.1
ENST00000405164.1 ENST00000336098.3 ENST00000393846.2 ENST00000404648.3 ENST00000443553.1 |
FGG
|
fibrinogen gamma chain |
| chr20_+_36974759 | 7.74 |
ENST00000217407.2
|
LBP
|
lipopolysaccharide binding protein |
| chr17_-_64216748 | 7.59 |
ENST00000585162.1
|
APOH
|
apolipoprotein H (beta-2-glycoprotein I) |
| chr5_-_141704566 | 4.97 |
ENST00000344120.4
ENST00000434127.2 |
SPRY4
|
sprouty homolog 4 (Drosophila) |
| chr3_+_186435065 | 4.84 |
ENST00000287611.2
ENST00000265023.4 |
KNG1
|
kininogen 1 |
| chr20_+_361890 | 4.82 |
ENST00000449710.1
ENST00000422053.2 |
TRIB3
|
tribbles pseudokinase 3 |
| chr2_+_211421262 | 4.76 |
ENST00000233072.5
|
CPS1
|
carbamoyl-phosphate synthase 1, mitochondrial |
| chr2_-_216300784 | 4.71 |
ENST00000421182.1
ENST00000432072.2 ENST00000323926.6 ENST00000336916.4 ENST00000357867.4 ENST00000359671.1 ENST00000346544.3 ENST00000345488.5 ENST00000357009.2 ENST00000446046.1 ENST00000356005.4 ENST00000443816.1 ENST00000426059.1 ENST00000354785.4 |
FN1
|
fibronectin 1 |
| chr9_-_34691201 | 4.55 |
ENST00000378800.3
ENST00000311925.2 |
CCL19
|
chemokine (C-C motif) ligand 19 |
| chr3_-_49722523 | 4.47 |
ENST00000448220.1
|
MST1
|
macrophage stimulating 1 (hepatocyte growth factor-like) |
| chr3_+_186435137 | 4.19 |
ENST00000447445.1
|
KNG1
|
kininogen 1 |
| chr6_+_31895287 | 4.12 |
ENST00000447952.2
|
C2
|
complement component 2 |
| chr6_+_31895254 | 4.11 |
ENST00000299367.5
ENST00000442278.2 |
C2
|
complement component 2 |
| chr16_+_72088376 | 4.04 |
ENST00000570083.1
ENST00000355906.5 ENST00000398131.2 ENST00000569639.1 ENST00000564499.1 ENST00000357763.4 ENST00000562526.1 ENST00000565574.1 ENST00000568417.2 ENST00000356967.5 |
HP
HPR
|
haptoglobin haptoglobin-related protein |
| chr20_+_361261 | 3.97 |
ENST00000217233.3
|
TRIB3
|
tribbles pseudokinase 3 |
| chr17_+_1665306 | 3.86 |
ENST00000571360.1
|
SERPINF1
|
serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 1 |
| chr6_+_31895467 | 3.85 |
ENST00000556679.1
ENST00000456570.1 |
CFB
CFB
|
complement factor B Complement factor B; Uncharacterized protein; cDNA FLJ55673, highly similar to Complement factor B |
| chr12_+_57849048 | 3.83 |
ENST00000266646.2
|
INHBE
|
inhibin, beta E |
| chr3_+_157154578 | 3.80 |
ENST00000295927.3
|
PTX3
|
pentraxin 3, long |
| chr3_+_12392971 | 3.70 |
ENST00000287820.6
|
PPARG
|
peroxisome proliferator-activated receptor gamma |
| chr12_-_56236711 | 3.62 |
ENST00000409200.3
|
MMP19
|
matrix metallopeptidase 19 |
| chr22_+_35776828 | 3.57 |
ENST00000216117.8
|
HMOX1
|
heme oxygenase (decycling) 1 |
| chr5_+_150400124 | 3.55 |
ENST00000388825.4
ENST00000521650.1 ENST00000517973.1 |
GPX3
|
glutathione peroxidase 3 (plasma) |
| chr6_+_31895480 | 3.47 |
ENST00000418949.2
ENST00000383177.3 ENST00000477310.1 |
C2
CFB
|
complement component 2 Complement factor B; Uncharacterized protein; cDNA FLJ55673, highly similar to Complement factor B |
| chr17_+_1665345 | 3.46 |
ENST00000576406.1
ENST00000571149.1 |
SERPINF1
|
serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 1 |
| chr19_-_11347173 | 3.46 |
ENST00000587656.1
|
DOCK6
|
dedicator of cytokinesis 6 |
| chr12_-_56236690 | 3.42 |
ENST00000322569.4
|
MMP19
|
matrix metallopeptidase 19 |
| chr16_-_28550348 | 3.34 |
ENST00000324873.6
|
NUPR1
|
nuclear protein, transcriptional regulator, 1 |
| chr12_-_56236734 | 3.33 |
ENST00000548629.1
|
MMP19
|
matrix metallopeptidase 19 |
| chr17_+_1665253 | 3.30 |
ENST00000254722.4
|
SERPINF1
|
serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 1 |
| chr1_+_212782012 | 3.26 |
ENST00000341491.4
ENST00000366985.1 |
ATF3
|
activating transcription factor 3 |
| chr16_-_28550320 | 3.19 |
ENST00000395641.2
|
NUPR1
|
nuclear protein, transcriptional regulator, 1 |
| chr8_+_22250334 | 2.95 |
ENST00000520832.1
|
SLC39A14
|
solute carrier family 39 (zinc transporter), member 14 |
| chr1_+_153232160 | 2.80 |
ENST00000368742.3
|
LOR
|
loricrin |
| chr20_+_9966728 | 2.79 |
ENST00000449270.1
|
RP5-839B4.8
|
RP5-839B4.8 |
| chr3_+_42897512 | 2.77 |
ENST00000493193.1
|
ACKR2
|
atypical chemokine receptor 2 |
| chr12_-_96390108 | 2.75 |
ENST00000538703.1
ENST00000261208.3 |
HAL
|
histidine ammonia-lyase |
| chr19_-_4540486 | 2.71 |
ENST00000306390.6
|
LRG1
|
leucine-rich alpha-2-glycoprotein 1 |
| chr11_+_47279155 | 2.70 |
ENST00000444396.1
ENST00000457932.1 ENST00000412937.1 |
NR1H3
|
nuclear receptor subfamily 1, group H, member 3 |
| chr11_+_47279248 | 2.62 |
ENST00000449369.1
|
NR1H3
|
nuclear receptor subfamily 1, group H, member 3 |
| chr21_-_44495964 | 2.61 |
ENST00000398168.1
ENST00000398165.3 |
CBS
|
cystathionine-beta-synthase |
| chr12_+_57623869 | 2.60 |
ENST00000414700.3
ENST00000557703.1 |
SHMT2
|
serine hydroxymethyltransferase 2 (mitochondrial) |
| chrX_-_55024967 | 2.56 |
ENST00000545676.1
|
PFKFB1
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 |
| chr1_-_203198790 | 2.49 |
ENST00000367229.1
ENST00000255427.3 ENST00000535569.1 |
CHIT1
|
chitinase 1 (chitotriosidase) |
| chr5_+_52083730 | 2.49 |
ENST00000282588.6
ENST00000274311.2 |
ITGA1
PELO
|
integrin, alpha 1 pelota homolog (Drosophila) |
| chr19_-_33360647 | 2.48 |
ENST00000590341.1
ENST00000587772.1 ENST00000023064.4 |
SLC7A9
|
solute carrier family 7 (amino acid transporter light chain, bo,+ system), member 9 |
| chr11_+_7506713 | 2.46 |
ENST00000329293.3
ENST00000534244.1 |
OLFML1
|
olfactomedin-like 1 |
| chr21_-_44495919 | 2.45 |
ENST00000398158.1
|
CBS
|
cystathionine-beta-synthase |
| chr12_+_57624119 | 2.45 |
ENST00000555773.1
ENST00000554975.1 ENST00000449049.3 ENST00000393827.4 |
SHMT2
|
serine hydroxymethyltransferase 2 (mitochondrial) |
| chr12_+_57623477 | 2.44 |
ENST00000557487.1
ENST00000555634.1 ENST00000556689.1 |
SHMT2
|
serine hydroxymethyltransferase 2 (mitochondrial) |
| chr11_+_47279504 | 2.44 |
ENST00000441012.2
ENST00000437276.1 ENST00000436029.1 ENST00000467728.1 ENST00000405853.3 |
NR1H3
|
nuclear receptor subfamily 1, group H, member 3 |
| chr12_+_57624085 | 2.41 |
ENST00000553474.1
|
SHMT2
|
serine hydroxymethyltransferase 2 (mitochondrial) |
| chrX_+_13587712 | 2.31 |
ENST00000361306.1
ENST00000380602.3 |
EGFL6
|
EGF-like-domain, multiple 6 |
| chr1_-_196577489 | 2.30 |
ENST00000609185.1
ENST00000451324.2 ENST00000367433.5 ENST00000367431.4 |
KCNT2
|
potassium channel, subfamily T, member 2 |
| chr12_+_57623907 | 2.28 |
ENST00000553529.1
ENST00000554310.1 |
SHMT2
|
serine hydroxymethyltransferase 2 (mitochondrial) |
| chr6_-_55739542 | 2.28 |
ENST00000446683.2
|
BMP5
|
bone morphogenetic protein 5 |
| chr9_-_95298254 | 2.24 |
ENST00000444490.2
|
ECM2
|
extracellular matrix protein 2, female organ and adipocyte specific |
| chr11_-_102595681 | 2.23 |
ENST00000236826.3
|
MMP8
|
matrix metallopeptidase 8 (neutrophil collagenase) |
| chr14_-_92413353 | 2.20 |
ENST00000556154.1
|
FBLN5
|
fibulin 5 |
| chr2_-_217559517 | 2.11 |
ENST00000449583.1
|
IGFBP5
|
insulin-like growth factor binding protein 5 |
| chr11_+_7506837 | 2.11 |
ENST00000528758.1
|
OLFML1
|
olfactomedin-like 1 |
| chr1_+_99127225 | 2.08 |
ENST00000370189.5
ENST00000529992.1 |
SNX7
|
sorting nexin 7 |
| chr9_+_131683174 | 2.05 |
ENST00000372592.3
ENST00000428610.1 |
PHYHD1
|
phytanoyl-CoA dioxygenase domain containing 1 |
| chr10_+_26727333 | 2.04 |
ENST00000356785.4
|
APBB1IP
|
amyloid beta (A4) precursor protein-binding, family B, member 1 interacting protein |
| chr19_-_36004543 | 2.00 |
ENST00000339686.3
ENST00000447113.2 ENST00000440396.1 |
DMKN
|
dermokine |
| chr1_+_28586006 | 1.99 |
ENST00000253063.3
|
SESN2
|
sestrin 2 |
| chr9_-_13175823 | 1.98 |
ENST00000545857.1
|
MPDZ
|
multiple PDZ domain protein |
| chr12_+_57624059 | 1.97 |
ENST00000557427.1
|
SHMT2
|
serine hydroxymethyltransferase 2 (mitochondrial) |
| chr5_+_38845960 | 1.93 |
ENST00000502536.1
|
OSMR
|
oncostatin M receptor |
| chr19_+_11350278 | 1.92 |
ENST00000252453.8
|
C19orf80
|
chromosome 19 open reading frame 80 |
| chr1_-_27952741 | 1.90 |
ENST00000399173.1
|
FGR
|
feline Gardner-Rasheed sarcoma viral oncogene homolog |
| chr11_+_7506584 | 1.90 |
ENST00000530135.1
|
OLFML1
|
olfactomedin-like 1 |
| chr7_+_114562616 | 1.89 |
ENST00000448022.1
|
MDFIC
|
MyoD family inhibitor domain containing |
| chr1_-_207143802 | 1.88 |
ENST00000324852.4
ENST00000400962.3 |
FCAMR
|
Fc receptor, IgA, IgM, high affinity |
| chr15_+_96869165 | 1.88 |
ENST00000421109.2
|
NR2F2
|
nuclear receptor subfamily 2, group F, member 2 |
| chr12_-_96390063 | 1.86 |
ENST00000541929.1
|
HAL
|
histidine ammonia-lyase |
| chr2_-_160654745 | 1.86 |
ENST00000259053.4
ENST00000429078.2 |
CD302
|
CD302 molecule |
| chr1_-_27953031 | 1.85 |
ENST00000374003.3
|
FGR
|
feline Gardner-Rasheed sarcoma viral oncogene homolog |
| chrX_-_32173579 | 1.76 |
ENST00000359836.1
ENST00000343523.2 ENST00000378707.3 ENST00000541735.1 ENST00000474231.1 |
DMD
|
dystrophin |
| chr10_+_26727125 | 1.74 |
ENST00000376236.4
|
APBB1IP
|
amyloid beta (A4) precursor protein-binding, family B, member 1 interacting protein |
| chr22_-_30901637 | 1.71 |
ENST00000381982.3
ENST00000255858.7 ENST00000540456.1 ENST00000392772.2 |
SEC14L4
|
SEC14-like 4 (S. cerevisiae) |
| chr5_+_150020214 | 1.71 |
ENST00000307662.4
|
SYNPO
|
synaptopodin |
| chr2_+_7118755 | 1.69 |
ENST00000433456.1
|
RNF144A
|
ring finger protein 144A |
| chr12_-_96389702 | 1.68 |
ENST00000552509.1
|
HAL
|
histidine ammonia-lyase |
| chr10_+_101542462 | 1.68 |
ENST00000370449.4
ENST00000370434.1 |
ABCC2
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 2 |
| chr1_-_27952617 | 1.65 |
ENST00000457296.1
|
FGR
|
feline Gardner-Rasheed sarcoma viral oncogene homolog |
| chr1_+_53480598 | 1.65 |
ENST00000430330.2
ENST00000408941.3 ENST00000478274.2 ENST00000484100.1 ENST00000435345.2 ENST00000488965.1 |
SCP2
|
sterol carrier protein 2 |
| chr1_+_53527854 | 1.64 |
ENST00000371500.3
ENST00000395871.2 ENST00000312553.5 |
PODN
|
podocan |
| chr6_+_151187074 | 1.63 |
ENST00000367308.4
|
MTHFD1L
|
methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 1-like |
| chr1_+_221051699 | 1.63 |
ENST00000366903.6
|
HLX
|
H2.0-like homeobox |
| chr5_+_38846101 | 1.59 |
ENST00000274276.3
|
OSMR
|
oncostatin M receptor |
| chr2_+_113885138 | 1.55 |
ENST00000409930.3
|
IL1RN
|
interleukin 1 receptor antagonist |
| chr5_-_150473127 | 1.50 |
ENST00000521001.1
|
TNIP1
|
TNFAIP3 interacting protein 1 |
| chrX_+_99839799 | 1.47 |
ENST00000373031.4
|
TNMD
|
tenomodulin |
| chr5_+_73109339 | 1.47 |
ENST00000296799.4
|
ARHGEF28
|
Rho guanine nucleotide exchange factor (GEF) 28 |
| chr7_+_114562172 | 1.46 |
ENST00000393486.1
ENST00000257724.3 |
MDFIC
|
MyoD family inhibitor domain containing |
| chrX_+_24072833 | 1.45 |
ENST00000253039.4
|
EIF2S3
|
eukaryotic translation initiation factor 2, subunit 3 gamma, 52kDa |
| chr22_-_30642728 | 1.43 |
ENST00000403987.3
|
LIF
|
leukemia inhibitory factor |
| chr1_+_162601774 | 1.40 |
ENST00000415555.1
|
DDR2
|
discoidin domain receptor tyrosine kinase 2 |
| chr12_+_10366223 | 1.40 |
ENST00000545290.1
|
GABARAPL1
|
GABA(A) receptor-associated protein like 1 |
| chr7_+_48211048 | 1.39 |
ENST00000435803.1
|
ABCA13
|
ATP-binding cassette, sub-family A (ABC1), member 13 |
| chr19_+_49259325 | 1.39 |
ENST00000222157.3
|
FGF21
|
fibroblast growth factor 21 |
| chr15_+_89182156 | 1.37 |
ENST00000379224.5
|
ISG20
|
interferon stimulated exonuclease gene 20kDa |
| chr14_-_92413727 | 1.37 |
ENST00000267620.10
|
FBLN5
|
fibulin 5 |
| chr15_+_89181974 | 1.35 |
ENST00000306072.5
|
ISG20
|
interferon stimulated exonuclease gene 20kDa |
| chr6_+_151187615 | 1.34 |
ENST00000441122.1
ENST00000423867.1 |
MTHFD1L
|
methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 1-like |
| chrX_-_106243451 | 1.34 |
ENST00000355610.4
ENST00000535534.1 |
MORC4
|
MORC family CW-type zinc finger 4 |
| chr15_+_89182178 | 1.33 |
ENST00000559876.1
|
ISG20
|
interferon stimulated exonuclease gene 20kDa |
| chr1_+_112016414 | 1.33 |
ENST00000343534.5
ENST00000369718.3 |
C1orf162
|
chromosome 1 open reading frame 162 |
| chr11_-_102595512 | 1.32 |
ENST00000438475.2
|
MMP8
|
matrix metallopeptidase 8 (neutrophil collagenase) |
| chr1_-_36945097 | 1.31 |
ENST00000331941.5
ENST00000418048.2 ENST00000338937.5 ENST00000440588.2 |
CSF3R
|
colony stimulating factor 3 receptor (granulocyte) |
| chr16_-_57809015 | 1.30 |
ENST00000540079.2
ENST00000569222.1 |
KIFC3
|
kinesin family member C3 |
| chr17_+_39846114 | 1.28 |
ENST00000586699.1
|
EIF1
|
eukaryotic translation initiation factor 1 |
| chr11_-_102668879 | 1.27 |
ENST00000315274.6
|
MMP1
|
matrix metallopeptidase 1 (interstitial collagenase) |
| chr11_-_64900791 | 1.26 |
ENST00000531018.1
|
SYVN1
|
synovial apoptosis inhibitor 1, synoviolin |
| chr4_+_77870856 | 1.24 |
ENST00000264893.6
ENST00000502584.1 ENST00000510641.1 |
SEPT11
|
septin 11 |
| chr11_-_119999539 | 1.24 |
ENST00000541857.1
|
TRIM29
|
tripartite motif containing 29 |
| chr6_-_134499037 | 1.23 |
ENST00000528577.1
|
SGK1
|
serum/glucocorticoid regulated kinase 1 |
| chr12_+_10124001 | 1.21 |
ENST00000396507.3
ENST00000304361.4 ENST00000434319.2 |
CLEC12A
|
C-type lectin domain family 12, member A |
| chr1_-_44497024 | 1.17 |
ENST00000372306.3
ENST00000372310.3 ENST00000475075.2 |
SLC6A9
|
solute carrier family 6 (neurotransmitter transporter, glycine), member 9 |
| chrX_-_106243294 | 1.17 |
ENST00000255495.7
|
MORC4
|
MORC family CW-type zinc finger 4 |
| chr4_+_77870960 | 1.11 |
ENST00000505788.1
ENST00000510515.1 ENST00000504637.1 |
SEPT11
|
septin 11 |
| chr6_-_32812420 | 1.11 |
ENST00000374881.2
|
PSMB8
|
proteasome (prosome, macropain) subunit, beta type, 8 |
| chr19_-_51869592 | 1.11 |
ENST00000596253.1
ENST00000309244.4 |
ETFB
|
electron-transfer-flavoprotein, beta polypeptide |
| chr18_+_21464737 | 1.07 |
ENST00000586751.1
|
LAMA3
|
laminin, alpha 3 |
| chr12_+_56390964 | 1.05 |
ENST00000356124.4
ENST00000266971.3 ENST00000394115.2 ENST00000547586.1 ENST00000552258.1 ENST00000548274.1 ENST00000546833.1 |
SUOX
|
sulfite oxidase |
| chr4_+_159727272 | 1.05 |
ENST00000379346.3
|
FNIP2
|
folliculin interacting protein 2 |
| chr11_-_119999611 | 1.04 |
ENST00000529044.1
|
TRIM29
|
tripartite motif containing 29 |
| chr17_-_38978847 | 1.02 |
ENST00000269576.5
|
KRT10
|
keratin 10 |
| chr6_+_126102292 | 1.02 |
ENST00000368357.3
|
NCOA7
|
nuclear receptor coactivator 7 |
| chr3_+_177159744 | 1.02 |
ENST00000439009.1
|
LINC00578
|
long intergenic non-protein coding RNA 578 |
| chr19_-_47288162 | 1.02 |
ENST00000594991.1
|
SLC1A5
|
solute carrier family 1 (neutral amino acid transporter), member 5 |
| chr17_+_53342311 | 1.00 |
ENST00000226067.5
|
HLF
|
hepatic leukemia factor |
| chr6_+_31939608 | 0.99 |
ENST00000375331.2
ENST00000375333.2 |
STK19
|
serine/threonine kinase 19 |
| chr5_-_172756506 | 0.97 |
ENST00000265087.4
|
STC2
|
stanniocalcin 2 |
| chr18_+_46065393 | 0.97 |
ENST00000256413.3
|
CTIF
|
CBP80/20-dependent translation initiation factor |
| chr16_+_56970567 | 0.97 |
ENST00000563911.1
|
HERPUD1
|
homocysteine-inducible, endoplasmic reticulum stress-inducible, ubiquitin-like domain member 1 |
| chr9_+_137218362 | 0.96 |
ENST00000481739.1
|
RXRA
|
retinoid X receptor, alpha |
| chr14_-_25045446 | 0.95 |
ENST00000216336.2
|
CTSG
|
cathepsin G |
| chr12_+_10366016 | 0.92 |
ENST00000546017.1
ENST00000535576.1 ENST00000539170.1 |
GABARAPL1
|
GABA(A) receptor-associated protein like 1 |
| chr3_+_127770455 | 0.92 |
ENST00000464451.1
|
SEC61A1
|
Sec61 alpha 1 subunit (S. cerevisiae) |
| chr3_+_187461442 | 0.92 |
ENST00000450760.1
|
RP11-211G3.2
|
RP11-211G3.2 |
| chr5_+_33441053 | 0.91 |
ENST00000541634.1
ENST00000455217.2 ENST00000414361.2 |
TARS
|
threonyl-tRNA synthetase |
| chr18_+_72168325 | 0.91 |
ENST00000582666.1
|
CNDP2
|
CNDP dipeptidase 2 (metallopeptidase M20 family) |
| chr1_-_204380919 | 0.91 |
ENST00000367188.4
|
PPP1R15B
|
protein phosphatase 1, regulatory subunit 15B |
| chr13_-_99667960 | 0.90 |
ENST00000448493.2
|
DOCK9
|
dedicator of cytokinesis 9 |
| chr9_+_2621798 | 0.89 |
ENST00000382100.3
|
VLDLR
|
very low density lipoprotein receptor |
| chr12_-_6451186 | 0.88 |
ENST00000540022.1
ENST00000536194.1 |
TNFRSF1A
|
tumor necrosis factor receptor superfamily, member 1A |
| chr8_-_21966893 | 0.87 |
ENST00000522405.1
ENST00000522379.1 ENST00000309188.6 ENST00000521807.2 |
NUDT18
|
nudix (nucleoside diphosphate linked moiety X)-type motif 18 |
| chr19_-_54824344 | 0.87 |
ENST00000346508.3
ENST00000446712.3 ENST00000432233.3 ENST00000301219.3 |
LILRA5
|
leukocyte immunoglobulin-like receptor, subfamily A (with TM domain), member 5 |
| chr12_-_12674032 | 0.87 |
ENST00000298573.4
|
DUSP16
|
dual specificity phosphatase 16 |
| chr17_+_65374075 | 0.86 |
ENST00000581322.1
|
PITPNC1
|
phosphatidylinositol transfer protein, cytoplasmic 1 |
| chr4_-_70725856 | 0.83 |
ENST00000226444.3
|
SULT1E1
|
sulfotransferase family 1E, estrogen-preferring, member 1 |
| chr16_-_4323015 | 0.83 |
ENST00000204517.6
|
TFAP4
|
transcription factor AP-4 (activating enhancer binding protein 4) |
| chr1_+_152784447 | 0.82 |
ENST00000360090.3
|
LCE1B
|
late cornified envelope 1B |
| chr10_+_45495898 | 0.82 |
ENST00000298299.3
|
ZNF22
|
zinc finger protein 22 |
| chrX_+_24073048 | 0.81 |
ENST00000423068.1
|
EIF2S3
|
eukaryotic translation initiation factor 2, subunit 3 gamma, 52kDa |
| chr10_+_81272287 | 0.81 |
ENST00000520547.2
|
EIF5AL1
|
eukaryotic translation initiation factor 5A-like 1 |
| chr5_+_33440802 | 0.80 |
ENST00000502553.1
ENST00000514259.1 ENST00000265112.3 |
TARS
|
threonyl-tRNA synthetase |
| chr11_+_118955583 | 0.80 |
ENST00000278715.3
ENST00000536813.1 ENST00000537841.1 ENST00000542729.1 ENST00000546302.1 ENST00000442944.2 ENST00000544387.1 ENST00000543090.1 |
HMBS
|
hydroxymethylbilane synthase |
| chr12_-_6451014 | 0.80 |
ENST00000366159.4
ENST00000539372.1 |
TNFRSF1A
|
tumor necrosis factor receptor superfamily, member 1A |
| chr1_-_36948879 | 0.78 |
ENST00000373106.1
ENST00000373104.1 ENST00000373103.1 |
CSF3R
|
colony stimulating factor 3 receptor (granulocyte) |
| chr4_+_129349188 | 0.78 |
ENST00000511497.1
|
RP11-420A23.1
|
RP11-420A23.1 |
| chr20_+_39969519 | 0.76 |
ENST00000373257.3
|
LPIN3
|
lipin 3 |
| chr12_-_6451235 | 0.76 |
ENST00000440083.2
ENST00000162749.2 |
TNFRSF1A
|
tumor necrosis factor receptor superfamily, member 1A |
| chr1_+_203595689 | 0.75 |
ENST00000357681.5
|
ATP2B4
|
ATPase, Ca++ transporting, plasma membrane 4 |
| chr10_-_21806759 | 0.75 |
ENST00000444772.3
|
SKIDA1
|
SKI/DACH domain containing 1 |
| chr17_+_29248918 | 0.73 |
ENST00000581548.1
ENST00000580525.1 |
ADAP2
|
ArfGAP with dual PH domains 2 |
| chr5_+_179105615 | 0.73 |
ENST00000514383.1
|
CANX
|
calnexin |
| chr19_+_49258775 | 0.72 |
ENST00000593756.1
|
FGF21
|
fibroblast growth factor 21 |
| chr5_-_137878887 | 0.72 |
ENST00000507939.1
ENST00000572514.1 ENST00000499810.2 ENST00000360541.5 |
ETF1
|
eukaryotic translation termination factor 1 |
| chr6_+_30687978 | 0.72 |
ENST00000327892.8
ENST00000435534.1 |
TUBB
|
tubulin, beta class I |
| chr18_+_29027696 | 0.71 |
ENST00000257189.4
|
DSG3
|
desmoglein 3 |
| chr2_+_216582766 | 0.70 |
ENST00000415479.1
|
AC093850.2
|
AC093850.2 |
| chr22_-_39640756 | 0.69 |
ENST00000331163.6
|
PDGFB
|
platelet-derived growth factor beta polypeptide |
| chr15_-_80263506 | 0.68 |
ENST00000335661.6
|
BCL2A1
|
BCL2-related protein A1 |
| chr4_+_159727222 | 0.66 |
ENST00000512986.1
|
FNIP2
|
folliculin interacting protein 2 |
| chr11_+_62649158 | 0.66 |
ENST00000539891.1
ENST00000536981.1 |
SLC3A2
|
solute carrier family 3 (amino acid transporter heavy chain), member 2 |
| chr19_-_47287990 | 0.65 |
ENST00000593713.1
ENST00000598022.1 ENST00000434726.2 |
SLC1A5
|
solute carrier family 1 (neutral amino acid transporter), member 5 |
| chr18_+_61420169 | 0.64 |
ENST00000425392.1
ENST00000336429.2 |
SERPINB7
|
serpin peptidase inhibitor, clade B (ovalbumin), member 7 |
| chr7_-_27224842 | 0.64 |
ENST00000517402.1
|
HOXA11
|
homeobox A11 |
| chr22_+_41258250 | 0.62 |
ENST00000544094.1
|
XPNPEP3
|
X-prolyl aminopeptidase (aminopeptidase P) 3, putative |
| chr14_+_75894714 | 0.61 |
ENST00000559060.1
|
JDP2
|
Jun dimerization protein 2 |
| chr1_+_203595903 | 0.60 |
ENST00000367218.3
ENST00000367219.3 ENST00000391954.2 |
ATP2B4
|
ATPase, Ca++ transporting, plasma membrane 4 |
| chr7_-_50860565 | 0.59 |
ENST00000403097.1
|
GRB10
|
growth factor receptor-bound protein 10 |
| chr17_+_29248953 | 0.58 |
ENST00000581285.1
|
ADAP2
|
ArfGAP with dual PH domains 2 |
| chr12_-_57914275 | 0.58 |
ENST00000547303.1
ENST00000552740.1 ENST00000547526.1 ENST00000551116.1 ENST00000346473.3 |
DDIT3
|
DNA-damage-inducible transcript 3 |
| chr16_+_89334512 | 0.56 |
ENST00000602042.1
|
AC137932.1
|
AC137932.1 |
| chrX_-_109683446 | 0.56 |
ENST00000372057.1
|
AMMECR1
|
Alport syndrome, mental retardation, midface hypoplasia and elliptocytosis chromosomal region gene 1 |
| chr3_-_126327398 | 0.55 |
ENST00000383572.2
|
TXNRD3NB
|
thioredoxin reductase 3 neighbor |
| chr6_+_32812568 | 0.54 |
ENST00000414474.1
|
PSMB9
|
proteasome (prosome, macropain) subunit, beta type, 9 |
| chr2_+_28618532 | 0.54 |
ENST00000545753.1
|
FOSL2
|
FOS-like antigen 2 |
| chr7_-_27224795 | 0.53 |
ENST00000006015.3
|
HOXA11
|
homeobox A11 |
| chrX_-_55208866 | 0.52 |
ENST00000545075.1
|
MTRNR2L10
|
MT-RNR2-like 10 |
| chr15_+_62359175 | 0.52 |
ENST00000355522.5
|
C2CD4A
|
C2 calcium-dependent domain containing 4A |
| chr5_-_35230434 | 0.51 |
ENST00000504500.1
|
PRLR
|
prolactin receptor |
| chr14_+_75894391 | 0.49 |
ENST00000419727.2
|
JDP2
|
Jun dimerization protein 2 |
| chr12_-_25102252 | 0.49 |
ENST00000261192.7
|
BCAT1
|
branched chain amino-acid transaminase 1, cytosolic |
| chr22_+_39052632 | 0.49 |
ENST00000411557.1
ENST00000396811.2 ENST00000216029.3 ENST00000416285.1 |
CBY1
|
chibby homolog 1 (Drosophila) |
Network of associatons between targets according to the STRING database.
First level regulatory network of CEBPB
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.6 | 7.7 | GO:0015920 | lipopolysaccharide transport(GO:0015920) |
| 2.3 | 11.3 | GO:1904640 | response to methionine(GO:1904640) |
| 1.6 | 4.8 | GO:0070409 | carbamoyl phosphate metabolic process(GO:0070408) carbamoyl phosphate biosynthetic process(GO:0070409) response to ammonia(GO:1903717) cellular response to ammonia(GO:1903718) |
| 1.6 | 4.7 | GO:1904237 | regulation of substrate-dependent cell migration, cell attachment to substrate(GO:1904235) positive regulation of substrate-dependent cell migration, cell attachment to substrate(GO:1904237) |
| 1.6 | 7.8 | GO:0090341 | negative regulation of secretion of lysosomal enzymes(GO:0090341) |
| 1.5 | 10.6 | GO:0071279 | cellular response to cobalt ion(GO:0071279) |
| 1.5 | 4.5 | GO:2000547 | regulation of dendritic cell dendrite assembly(GO:2000547) |
| 1.4 | 14.1 | GO:0019264 | glycine biosynthetic process from serine(GO:0019264) |
| 1.3 | 4.0 | GO:2000296 | negative regulation of hydrogen peroxide catabolic process(GO:2000296) |
| 1.3 | 5.1 | GO:0006535 | cysteine biosynthetic process from serine(GO:0006535) |
| 1.1 | 2.3 | GO:0071676 | negative regulation of mononuclear cell migration(GO:0071676) |
| 1.1 | 3.3 | GO:0061394 | regulation of transcription from RNA polymerase II promoter in response to arsenic-containing substance(GO:0061394) |
| 0.9 | 10.4 | GO:0001554 | luteolysis(GO:0001554) |
| 0.9 | 3.7 | GO:2000230 | response to metformin(GO:1901558) negative regulation of pancreatic stellate cell proliferation(GO:2000230) |
| 0.9 | 6.3 | GO:0019557 | histidine catabolic process to glutamate and formamide(GO:0019556) histidine catabolic process to glutamate and formate(GO:0019557) formamide metabolic process(GO:0043606) |
| 0.9 | 3.6 | GO:0006788 | heme oxidation(GO:0006788) negative regulation of mast cell cytokine production(GO:0032764) |
| 0.8 | 4.1 | GO:0000738 | DNA catabolic process, exonucleolytic(GO:0000738) |
| 0.7 | 2.1 | GO:1904204 | regulation of skeletal muscle hypertrophy(GO:1904204) |
| 0.7 | 11.7 | GO:2000427 | positive regulation of apoptotic cell clearance(GO:2000427) |
| 0.6 | 3.0 | GO:0009257 | 10-formyltetrahydrofolate biosynthetic process(GO:0009257) |
| 0.6 | 24.9 | GO:0072378 | blood coagulation, fibrin clot formation(GO:0072378) |
| 0.6 | 1.7 | GO:0050787 | antibiotic metabolic process(GO:0016999) detoxification of mercury ion(GO:0050787) |
| 0.5 | 3.8 | GO:0052199 | negative regulation of catalytic activity in other organism involved in symbiotic interaction(GO:0052199) |
| 0.5 | 1.6 | GO:0061537 | glycine secretion(GO:0061536) glycine secretion, neurotransmission(GO:0061537) |
| 0.5 | 3.6 | GO:0006982 | response to lipid hydroperoxide(GO:0006982) |
| 0.5 | 2.0 | GO:0090526 | regulation of gluconeogenesis involved in cellular glucose homeostasis(GO:0090526) |
| 0.5 | 1.4 | GO:0048861 | leukemia inhibitory factor signaling pathway(GO:0048861) |
| 0.5 | 1.9 | GO:0009956 | radial pattern formation(GO:0009956) |
| 0.4 | 1.3 | GO:2000282 | regulation of cellular amine catabolic process(GO:0033241) negative regulation of cellular amine catabolic process(GO:0033242) negative regulation of the force of heart contraction(GO:0098736) regulation of arginine catabolic process(GO:1900081) negative regulation of arginine catabolic process(GO:1900082) regulation of citrulline biosynthetic process(GO:1903248) negative regulation of citrulline biosynthetic process(GO:1903249) regulation of cellular amino acid biosynthetic process(GO:2000282) negative regulation of cellular amino acid biosynthetic process(GO:2000283) |
| 0.4 | 3.5 | GO:0038165 | oncostatin-M-mediated signaling pathway(GO:0038165) |
| 0.4 | 1.7 | GO:0006435 | threonyl-tRNA aminoacylation(GO:0006435) |
| 0.4 | 2.6 | GO:0033133 | positive regulation of glucokinase activity(GO:0033133) positive regulation of hexokinase activity(GO:1903301) |
| 0.4 | 1.7 | GO:0010585 | glutamine secretion(GO:0010585) L-glutamine import(GO:0036229) L-glutamine import into cell(GO:1903803) |
| 0.4 | 2.5 | GO:0044245 | polysaccharide digestion(GO:0044245) |
| 0.4 | 2.5 | GO:0070966 | nuclear-transcribed mRNA catabolic process, no-go decay(GO:0070966) |
| 0.3 | 1.0 | GO:0045994 | positive regulation of translational initiation by iron(GO:0045994) |
| 0.3 | 6.5 | GO:2000194 | regulation of female gonad development(GO:2000194) |
| 0.3 | 1.5 | GO:0085032 | modulation of signal transduction in other organism(GO:0044501) modulation by symbiont of host signal transduction pathway(GO:0052027) modulation of signal transduction in other organism involved in symbiotic interaction(GO:0052250) modulation by symbiont of host I-kappaB kinase/NF-kappaB cascade(GO:0085032) |
| 0.3 | 0.9 | GO:1901069 | guanosine-containing compound catabolic process(GO:1901069) |
| 0.3 | 2.0 | GO:1903575 | cornified envelope assembly(GO:1903575) |
| 0.3 | 0.8 | GO:0006711 | estrogen catabolic process(GO:0006711) |
| 0.3 | 3.6 | GO:0048251 | elastic fiber assembly(GO:0048251) |
| 0.3 | 2.4 | GO:0071550 | death-inducing signaling complex assembly(GO:0071550) |
| 0.3 | 7.0 | GO:0045717 | negative regulation of fatty acid biosynthetic process(GO:0045717) |
| 0.2 | 4.5 | GO:0045721 | negative regulation of gluconeogenesis(GO:0045721) |
| 0.2 | 1.6 | GO:0032380 | regulation of intracellular lipid transport(GO:0032377) regulation of intracellular sterol transport(GO:0032380) regulation of intracellular cholesterol transport(GO:0032383) |
| 0.2 | 5.4 | GO:0033008 | positive regulation of mast cell activation involved in immune response(GO:0033008) positive regulation of mast cell degranulation(GO:0043306) |
| 0.2 | 1.4 | GO:0098886 | modification of dendritic spine(GO:0098886) |
| 0.2 | 1.6 | GO:0045629 | negative regulation of T-helper 2 cell differentiation(GO:0045629) |
| 0.2 | 0.9 | GO:0045209 | MAPK phosphatase export from nucleus(GO:0045208) MAPK phosphatase export from nucleus, leptomycin B sensitive(GO:0045209) |
| 0.2 | 1.5 | GO:0035992 | tendon cell differentiation(GO:0035990) tendon formation(GO:0035992) |
| 0.2 | 1.7 | GO:0015811 | L-cystine transport(GO:0015811) |
| 0.2 | 3.8 | GO:0002291 | T cell activation via T cell receptor contact with antigen bound to MHC molecule on antigen presenting cell(GO:0002291) |
| 0.2 | 1.6 | GO:2000660 | negative regulation of interleukin-1-mediated signaling pathway(GO:2000660) |
| 0.2 | 1.1 | GO:0019418 | sulfide oxidation(GO:0019418) sulfide oxidation, using sulfide:quinone oxidoreductase(GO:0070221) |
| 0.2 | 3.9 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 0.2 | 0.7 | GO:0090678 | metanephric glomerular mesangial cell development(GO:0072255) reversible differentiation(GO:0090677) cell dedifferentiation involved in phenotypic switching(GO:0090678) positive regulation of phenotypic switching(GO:1900241) regulation of vascular smooth muscle cell dedifferentiation(GO:1905174) positive regulation of vascular smooth muscle cell dedifferentiation(GO:1905176) vascular smooth muscle cell dedifferentiation(GO:1990936) |
| 0.2 | 2.9 | GO:0015684 | ferrous iron transport(GO:0015684) ferrous iron transmembrane transport(GO:1903874) |
| 0.2 | 0.9 | GO:0039019 | pronephric nephron development(GO:0039019) |
| 0.1 | 0.6 | GO:0032792 | negative regulation of CREB transcription factor activity(GO:0032792) |
| 0.1 | 1.3 | GO:0009048 | dosage compensation by inactivation of X chromosome(GO:0009048) |
| 0.1 | 1.0 | GO:0070944 | neutrophil mediated cytotoxicity(GO:0070942) neutrophil mediated killing of symbiont cell(GO:0070943) neutrophil mediated killing of bacterium(GO:0070944) |
| 0.1 | 0.8 | GO:0009082 | branched-chain amino acid biosynthetic process(GO:0009082) leucine biosynthetic process(GO:0009098) valine biosynthetic process(GO:0009099) |
| 0.1 | 1.3 | GO:0045218 | zonula adherens maintenance(GO:0045218) |
| 0.1 | 0.6 | GO:0090362 | positive regulation of platelet-derived growth factor production(GO:0090362) |
| 0.1 | 0.9 | GO:0034436 | glycoprotein transport(GO:0034436) |
| 0.1 | 2.5 | GO:1902083 | negative regulation of peptidyl-cysteine S-nitrosylation(GO:1902083) |
| 0.1 | 0.3 | GO:0006419 | alanyl-tRNA aminoacylation(GO:0006419) |
| 0.1 | 1.5 | GO:0006449 | regulation of translational termination(GO:0006449) |
| 0.1 | 0.9 | GO:1903912 | negative regulation of PERK-mediated unfolded protein response(GO:1903898) negative regulation of endoplasmic reticulum stress-induced eIF2 alpha phosphorylation(GO:1903912) |
| 0.1 | 2.3 | GO:0006995 | cellular response to nitrogen starvation(GO:0006995) cellular response to nitrogen levels(GO:0043562) |
| 0.1 | 1.2 | GO:0070294 | renal sodium ion absorption(GO:0070294) |
| 0.1 | 0.7 | GO:0060356 | leucine import(GO:0060356) |
| 0.1 | 0.3 | GO:1904211 | membrane protein proteolysis involved in retrograde protein transport, ER to cytosol(GO:1904211) |
| 0.1 | 1.4 | GO:0090091 | positive regulation of extracellular matrix disassembly(GO:0090091) |
| 0.1 | 0.3 | GO:0060264 | respiratory burst involved in inflammatory response(GO:0002536) regulation of respiratory burst involved in inflammatory response(GO:0060264) |
| 0.1 | 0.3 | GO:0042144 | vacuole fusion, non-autophagic(GO:0042144) |
| 0.1 | 0.2 | GO:0006423 | cysteinyl-tRNA aminoacylation(GO:0006423) |
| 0.1 | 0.4 | GO:0060054 | positive regulation of epithelial cell proliferation involved in wound healing(GO:0060054) |
| 0.1 | 2.8 | GO:1902236 | negative regulation of endoplasmic reticulum stress-induced intrinsic apoptotic signaling pathway(GO:1902236) |
| 0.1 | 1.0 | GO:2001256 | regulation of store-operated calcium entry(GO:2001256) |
| 0.1 | 1.2 | GO:0060272 | embryonic skeletal joint morphogenesis(GO:0060272) |
| 0.1 | 3.8 | GO:0010862 | positive regulation of pathway-restricted SMAD protein phosphorylation(GO:0010862) |
| 0.1 | 0.3 | GO:0033031 | positive regulation of neutrophil apoptotic process(GO:0033031) |
| 0.1 | 3.4 | GO:0007257 | activation of JUN kinase activity(GO:0007257) |
| 0.1 | 0.4 | GO:0010728 | positive regulation of hydrogen peroxide metabolic process(GO:0010726) regulation of hydrogen peroxide biosynthetic process(GO:0010728) |
| 0.1 | 2.7 | GO:0030511 | positive regulation of transforming growth factor beta receptor signaling pathway(GO:0030511) positive regulation of cellular response to transforming growth factor beta stimulus(GO:1903846) |
| 0.1 | 0.5 | GO:0038161 | prolactin signaling pathway(GO:0038161) |
| 0.1 | 0.2 | GO:1903336 | negative regulation of vacuolar transport(GO:1903336) |
| 0.1 | 2.1 | GO:0097186 | amelogenesis(GO:0097186) |
| 0.1 | 4.7 | GO:0035987 | endodermal cell differentiation(GO:0035987) |
| 0.1 | 4.6 | GO:0018149 | peptide cross-linking(GO:0018149) |
| 0.1 | 1.1 | GO:0033539 | fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:0033539) |
| 0.1 | 0.4 | GO:0055129 | L-proline biosynthetic process(GO:0055129) |
| 0.1 | 0.8 | GO:0006782 | protoporphyrinogen IX biosynthetic process(GO:0006782) |
| 0.0 | 1.1 | GO:0031065 | positive regulation of histone deacetylation(GO:0031065) |
| 0.0 | 5.0 | GO:0070373 | negative regulation of ERK1 and ERK2 cascade(GO:0070373) |
| 0.0 | 0.8 | GO:0043922 | negative regulation by host of viral transcription(GO:0043922) |
| 0.0 | 0.3 | GO:0036093 | male germ cell proliferation(GO:0002176) germ cell proliferation(GO:0036093) |
| 0.0 | 0.2 | GO:0038112 | interleukin-8-mediated signaling pathway(GO:0038112) |
| 0.0 | 0.6 | GO:0030949 | positive regulation of vascular endothelial growth factor receptor signaling pathway(GO:0030949) |
| 0.0 | 0.2 | GO:0043353 | enucleate erythrocyte differentiation(GO:0043353) |
| 0.0 | 0.7 | GO:0034975 | protein folding in endoplasmic reticulum(GO:0034975) |
| 0.0 | 0.2 | GO:0032625 | interleukin-21 production(GO:0032625) interleukin-21 secretion(GO:0072619) |
| 0.0 | 1.3 | GO:0032461 | positive regulation of protein oligomerization(GO:0032461) |
| 0.0 | 1.9 | GO:0002250 | adaptive immune response(GO:0002250) |
| 0.0 | 1.0 | GO:0016032 | viral process(GO:0016032) |
| 0.0 | 0.8 | GO:0006646 | phosphatidylethanolamine biosynthetic process(GO:0006646) |
| 0.0 | 0.7 | GO:0008053 | mitochondrial fusion(GO:0008053) |
| 0.0 | 0.5 | GO:0003334 | keratinocyte development(GO:0003334) |
| 0.0 | 0.3 | GO:1903800 | positive regulation of production of miRNAs involved in gene silencing by miRNA(GO:1903800) |
| 0.0 | 4.6 | GO:0010811 | positive regulation of cell-substrate adhesion(GO:0010811) |
| 0.0 | 0.4 | GO:0031937 | positive regulation of chromatin silencing(GO:0031937) |
| 0.0 | 0.5 | GO:0031293 | membrane protein intracellular domain proteolysis(GO:0031293) |
| 0.0 | 0.6 | GO:0003094 | glomerular filtration(GO:0003094) renal filtration(GO:0097205) |
| 0.0 | 2.3 | GO:1900181 | negative regulation of protein localization to nucleus(GO:1900181) |
| 0.0 | 1.6 | GO:0002479 | antigen processing and presentation of exogenous peptide antigen via MHC class I, TAP-dependent(GO:0002479) |
| 0.0 | 0.4 | GO:0006750 | glutathione biosynthetic process(GO:0006750) |
| 0.0 | 1.2 | GO:0015914 | phospholipid transport(GO:0015914) |
| 0.0 | 0.2 | GO:0060052 | neurofilament cytoskeleton organization(GO:0060052) |
| 0.0 | 1.1 | GO:0046426 | negative regulation of JAK-STAT cascade(GO:0046426) negative regulation of STAT cascade(GO:1904893) |
| 0.0 | 0.5 | GO:0007420 | brain development(GO:0007420) |
| 0.0 | 2.4 | GO:0051291 | protein heterooligomerization(GO:0051291) |
| 0.0 | 1.7 | GO:0032436 | positive regulation of proteasomal ubiquitin-dependent protein catabolic process(GO:0032436) |
| 0.0 | 0.4 | GO:0046856 | phosphatidylinositol dephosphorylation(GO:0046856) |
| 0.0 | 0.5 | GO:0019221 | cytokine-mediated signaling pathway(GO:0019221) |
| 0.0 | 0.5 | GO:0016337 | single organismal cell-cell adhesion(GO:0016337) |
| 0.0 | 1.3 | GO:0007507 | heart development(GO:0007507) |
| 0.0 | 0.4 | GO:0016339 | calcium-dependent cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0016339) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 14.1 | GO:0070552 | BRISC complex(GO:0070552) |
| 1.3 | 3.8 | GO:1990622 | CHOP-ATF3 complex(GO:1990622) |
| 1.2 | 3.5 | GO:0005900 | oncostatin-M receptor complex(GO:0005900) |
| 1.0 | 4.0 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 1.0 | 13.4 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.6 | 2.5 | GO:0034665 | integrin alpha1-beta1 complex(GO:0034665) |
| 0.5 | 10.8 | GO:0043203 | axon hillock(GO:0043203) |
| 0.5 | 1.4 | GO:0097444 | spine apparatus(GO:0097444) |
| 0.4 | 7.6 | GO:0031089 | platelet dense granule lumen(GO:0031089) |
| 0.4 | 2.6 | GO:0005850 | eukaryotic translation initiation factor 2 complex(GO:0005850) |
| 0.4 | 3.6 | GO:0071953 | elastic fiber(GO:0071953) |
| 0.3 | 1.3 | GO:0000839 | Hrd1p ubiquitin ligase ERAD-L complex(GO:0000839) |
| 0.2 | 1.3 | GO:1990037 | Lewy body core(GO:1990037) |
| 0.2 | 2.1 | GO:0042567 | insulin-like growth factor ternary complex(GO:0042567) |
| 0.2 | 1.6 | GO:1990111 | spermatoproteasome complex(GO:1990111) |
| 0.1 | 2.0 | GO:1990316 | ATG1/ULK1 kinase complex(GO:1990316) |
| 0.1 | 1.7 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.1 | 12.5 | GO:1904724 | tertiary granule lumen(GO:1904724) |
| 0.1 | 1.4 | GO:0016013 | syntrophin complex(GO:0016013) |
| 0.1 | 3.8 | GO:0042101 | T cell receptor complex(GO:0042101) |
| 0.1 | 1.1 | GO:0005610 | laminin-5 complex(GO:0005610) |
| 0.1 | 9.7 | GO:0031093 | platelet alpha granule lumen(GO:0031093) |
| 0.1 | 4.8 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.1 | 2.0 | GO:0043220 | Schmidt-Lanterman incisure(GO:0043220) |
| 0.1 | 2.2 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.1 | 3.0 | GO:0032839 | dendrite cytoplasm(GO:0032839) |
| 0.1 | 5.5 | GO:0016235 | aggresome(GO:0016235) |
| 0.1 | 5.4 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.1 | 1.3 | GO:0005915 | zonula adherens(GO:0005915) |
| 0.1 | 0.7 | GO:0005641 | nuclear envelope lumen(GO:0005641) |
| 0.0 | 0.5 | GO:0070765 | gamma-secretase complex(GO:0070765) |
| 0.0 | 11.0 | GO:0090575 | RNA polymerase II transcription factor complex(GO:0090575) |
| 0.0 | 0.9 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.0 | 4.9 | GO:0005901 | caveola(GO:0005901) |
| 0.0 | 0.9 | GO:0034361 | very-low-density lipoprotein particle(GO:0034361) triglyceride-rich lipoprotein particle(GO:0034385) |
| 0.0 | 6.2 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.0 | 15.8 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.0 | 2.8 | GO:0031526 | brush border membrane(GO:0031526) |
| 0.0 | 5.7 | GO:0032993 | protein-DNA complex(GO:0032993) |
| 0.0 | 5.0 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.0 | 1.6 | GO:0005782 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.0 | 2.7 | GO:0001725 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.0 | 0.5 | GO:0031904 | endosome lumen(GO:0031904) |
| 0.0 | 0.3 | GO:0030897 | HOPS complex(GO:0030897) |
| 0.0 | 0.0 | GO:0002947 | tumor necrosis factor receptor superfamily complex(GO:0002947) |
| 0.0 | 0.3 | GO:0071141 | SMAD protein complex(GO:0071141) |
| 0.0 | 1.3 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.0 | 1.4 | GO:0035577 | azurophil granule membrane(GO:0035577) |
| 0.0 | 1.7 | GO:0070821 | tertiary granule membrane(GO:0070821) |
| 0.0 | 2.8 | GO:0055037 | recycling endosome(GO:0055037) |
| 0.0 | 1.9 | GO:0030175 | filopodium(GO:0030175) |
| 0.0 | 1.9 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 18.4 | GO:0005615 | extracellular space(GO:0005615) |
| 0.0 | 1.5 | GO:0042470 | melanosome(GO:0042470) pigment granule(GO:0048770) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.0 | 9.1 | GO:0004611 | phosphoenolpyruvate carboxykinase activity(GO:0004611) phosphoenolpyruvate carboxykinase (GTP) activity(GO:0004613) |
| 2.1 | 6.3 | GO:0004397 | histidine ammonia-lyase activity(GO:0004397) |
| 1.4 | 14.1 | GO:0004793 | glycine hydroxymethyltransferase activity(GO:0004372) threonine aldolase activity(GO:0004793) L-allo-threonine aldolase activity(GO:0008732) |
| 1.4 | 4.1 | GO:0008859 | exoribonuclease II activity(GO:0008859) |
| 1.3 | 5.1 | GO:0098809 | cystathionine beta-synthase activity(GO:0004122) oxidoreductase activity, acting on other nitrogenous compounds as donors, cytochrome as acceptor(GO:0016662) nitrite reductase (NO-forming) activity(GO:0050421) carbon monoxide binding(GO:0070025) nitrite reductase activity(GO:0098809) |
| 1.2 | 4.8 | GO:0004088 | carbamoyl-phosphate synthase (ammonia) activity(GO:0004087) carbamoyl-phosphate synthase (glutamine-hydrolyzing) activity(GO:0004088) |
| 1.1 | 4.5 | GO:0031735 | CCR10 chemokine receptor binding(GO:0031735) |
| 1.1 | 7.6 | GO:0060230 | lipoprotein lipase activator activity(GO:0060230) |
| 0.9 | 3.6 | GO:0004392 | heme oxygenase (decyclizing) activity(GO:0004392) |
| 0.9 | 7.8 | GO:0032810 | sterol response element binding(GO:0032810) |
| 0.9 | 7.7 | GO:0070891 | lipoteichoic acid binding(GO:0070891) |
| 0.8 | 5.4 | GO:0034988 | Fc-gamma receptor I complex binding(GO:0034988) |
| 0.7 | 4.0 | GO:0030492 | hemoglobin binding(GO:0030492) |
| 0.6 | 3.0 | GO:0004329 | formate-tetrahydrofolate ligase activity(GO:0004329) |
| 0.6 | 3.5 | GO:0004924 | oncostatin-M receptor activity(GO:0004924) |
| 0.5 | 1.6 | GO:0070538 | oleic acid binding(GO:0070538) |
| 0.5 | 4.7 | GO:0050692 | DBD domain binding(GO:0050692) |
| 0.5 | 1.6 | GO:0005150 | interleukin-1, Type I receptor binding(GO:0005150) |
| 0.5 | 2.6 | GO:0070095 | fructose-6-phosphate binding(GO:0070095) |
| 0.5 | 1.4 | GO:0005146 | leukemia inhibitory factor receptor binding(GO:0005146) |
| 0.4 | 2.2 | GO:0070052 | collagen V binding(GO:0070052) |
| 0.4 | 1.3 | GO:0036487 | nitric-oxide synthase inhibitor activity(GO:0036487) |
| 0.4 | 8.8 | GO:0055106 | ubiquitin-protein transferase regulator activity(GO:0055106) |
| 0.4 | 1.7 | GO:0004829 | threonine-tRNA ligase activity(GO:0004829) |
| 0.4 | 2.1 | GO:0051916 | granulocyte colony-stimulating factor binding(GO:0051916) |
| 0.4 | 1.6 | GO:0015375 | glycine:sodium symporter activity(GO:0015375) |
| 0.4 | 3.8 | GO:0001849 | complement component C1q binding(GO:0001849) |
| 0.3 | 2.4 | GO:0043120 | tumor necrosis factor binding(GO:0043120) |
| 0.3 | 2.5 | GO:0098639 | collagen binding involved in cell-matrix adhesion(GO:0098639) |
| 0.3 | 5.7 | GO:0030957 | Tat protein binding(GO:0030957) |
| 0.3 | 2.0 | GO:0070728 | leucine binding(GO:0070728) |
| 0.3 | 1.7 | GO:0015186 | L-glutamine transmembrane transporter activity(GO:0015186) |
| 0.2 | 7.4 | GO:0001848 | complement binding(GO:0001848) |
| 0.2 | 1.7 | GO:0042030 | ATPase inhibitor activity(GO:0042030) |
| 0.2 | 3.5 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.2 | 2.9 | GO:0015093 | ferrous iron transmembrane transporter activity(GO:0015093) |
| 0.2 | 0.9 | GO:0034189 | very-low-density lipoprotein particle binding(GO:0034189) |
| 0.2 | 0.9 | GO:0044715 | 8-oxo-dGDP phosphatase activity(GO:0044715) |
| 0.2 | 1.1 | GO:0030151 | molybdenum ion binding(GO:0030151) |
| 0.2 | 1.7 | GO:0015184 | L-cystine transmembrane transporter activity(GO:0015184) |
| 0.2 | 0.9 | GO:0102008 | cytosolic dipeptidase activity(GO:0102008) |
| 0.2 | 3.6 | GO:0008430 | selenium binding(GO:0008430) |
| 0.2 | 2.8 | GO:0019957 | C-C chemokine binding(GO:0019957) |
| 0.2 | 2.3 | GO:0070700 | BMP receptor binding(GO:0070700) |
| 0.2 | 2.1 | GO:0031995 | insulin-like growth factor II binding(GO:0031995) |
| 0.1 | 2.5 | GO:0004568 | chitinase activity(GO:0004568) |
| 0.1 | 0.8 | GO:0052655 | branched-chain-amino-acid transaminase activity(GO:0004084) L-leucine transaminase activity(GO:0052654) L-valine transaminase activity(GO:0052655) L-isoleucine transaminase activity(GO:0052656) |
| 0.1 | 0.8 | GO:0008195 | phosphatidate phosphatase activity(GO:0008195) |
| 0.1 | 0.8 | GO:0047894 | flavonol 3-sulfotransferase activity(GO:0047894) steroid sulfotransferase activity(GO:0050294) |
| 0.1 | 1.3 | GO:0043533 | inositol 1,3,4,5 tetrakisphosphate binding(GO:0043533) |
| 0.1 | 0.3 | GO:0004813 | alanine-tRNA ligase activity(GO:0004813) |
| 0.1 | 6.5 | GO:0005160 | transforming growth factor beta receptor binding(GO:0005160) |
| 0.1 | 1.4 | GO:0038062 | protein tyrosine kinase collagen receptor activity(GO:0038062) collagen receptor activity(GO:0038064) |
| 0.1 | 0.7 | GO:0003747 | translation release factor activity(GO:0003747) translation termination factor activity(GO:0008079) |
| 0.1 | 1.7 | GO:0043225 | anion transmembrane-transporting ATPase activity(GO:0043225) |
| 0.1 | 1.2 | GO:0008526 | phosphatidylcholine transporter activity(GO:0008525) phosphatidylinositol transporter activity(GO:0008526) |
| 0.1 | 15.2 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.1 | 0.5 | GO:0004925 | prolactin receptor activity(GO:0004925) |
| 0.1 | 11.3 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.1 | 9.0 | GO:0004869 | cysteine-type endopeptidase inhibitor activity(GO:0004869) |
| 0.1 | 4.7 | GO:0016504 | peptidase activator activity(GO:0016504) |
| 0.1 | 0.2 | GO:0004817 | cysteine-tRNA ligase activity(GO:0004817) |
| 0.1 | 0.4 | GO:0004735 | pyrroline-5-carboxylate reductase activity(GO:0004735) |
| 0.1 | 2.3 | GO:0015269 | calcium-activated potassium channel activity(GO:0015269) |
| 0.1 | 1.1 | GO:0050998 | nitric-oxide synthase binding(GO:0050998) |
| 0.1 | 1.2 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.1 | 2.1 | GO:0005104 | fibroblast growth factor receptor binding(GO:0005104) |
| 0.1 | 1.3 | GO:1904264 | ubiquitin protein ligase activity involved in ERAD pathway(GO:1904264) |
| 0.1 | 2.3 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 0.1 | 1.6 | GO:0004298 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 0.0 | 1.9 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.0 | 14.2 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 1.2 | GO:0017081 | chloride channel regulator activity(GO:0017081) |
| 0.0 | 0.2 | GO:0047066 | phospholipid-hydroperoxide glutathione peroxidase activity(GO:0047066) |
| 0.0 | 0.2 | GO:0070012 | oligopeptidase activity(GO:0070012) |
| 0.0 | 1.3 | GO:0008569 | ATP-dependent microtubule motor activity, minus-end-directed(GO:0008569) |
| 0.0 | 3.9 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.0 | 0.7 | GO:0016176 | superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 0.0 | 0.2 | GO:0019959 | interleukin-8 receptor activity(GO:0004918) interleukin-8 binding(GO:0019959) |
| 0.0 | 0.5 | GO:0052658 | inositol-1,4,5-trisphosphate 5-phosphatase activity(GO:0052658) |
| 0.0 | 0.7 | GO:0005432 | calcium:sodium antiporter activity(GO:0005432) |
| 0.0 | 0.3 | GO:0035500 | MH2 domain binding(GO:0035500) |
| 0.0 | 0.3 | GO:0019870 | potassium channel inhibitor activity(GO:0019870) |
| 0.0 | 0.7 | GO:0051400 | BH domain binding(GO:0051400) |
| 0.0 | 1.7 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.0 | 0.6 | GO:0008140 | cAMP response element binding protein binding(GO:0008140) |
| 0.0 | 0.7 | GO:0034185 | apolipoprotein binding(GO:0034185) |
| 0.0 | 5.9 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.7 | GO:0032794 | GTPase activating protein binding(GO:0032794) |
| 0.0 | 1.5 | GO:0051019 | mitogen-activated protein kinase binding(GO:0051019) |
| 0.0 | 0.8 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.0 | 1.6 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 3.3 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.0 | 0.4 | GO:0005328 | neurotransmitter:sodium symporter activity(GO:0005328) |
| 0.0 | 5.0 | GO:0030674 | protein binding, bridging(GO:0030674) |
| 0.0 | 1.0 | GO:0030374 | ligand-dependent nuclear receptor transcription coactivator activity(GO:0030374) |
| 0.0 | 2.0 | GO:0051213 | dioxygenase activity(GO:0051213) |
| 0.0 | 0.1 | GO:0008142 | oxysterol binding(GO:0008142) |
| 0.0 | 0.4 | GO:0015464 | acetylcholine receptor activity(GO:0015464) |
| 0.0 | 0.9 | GO:0043022 | ribosome binding(GO:0043022) |
| 0.0 | 0.0 | GO:0034353 | RNA pyrophosphohydrolase activity(GO:0034353) |
| 0.0 | 0.3 | GO:0016671 | oxidoreductase activity, acting on a sulfur group of donors, disulfide as acceptor(GO:0016671) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 17.4 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.2 | 11.1 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
| 0.2 | 12.4 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.2 | 4.7 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.1 | 9.9 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.1 | 7.9 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.1 | 9.1 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 2.5 | PID INTEGRIN CS PATHWAY | Integrin family cell surface interactions |
| 0.1 | 2.4 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 0.1 | 26.3 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.1 | 7.4 | PID IL6 7 PATHWAY | IL6-mediated signaling events |
| 0.0 | 1.9 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.0 | 3.0 | PID ATF2 PATHWAY | ATF-2 transcription factor network |
| 0.0 | 3.0 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.0 | 16.3 | NABA SECRETED FACTORS | Genes encoding secreted soluble factors |
| 0.0 | 0.9 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 0.0 | 1.9 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 1.6 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 0.5 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.0 | 1.6 | PID IL12 2PATHWAY | IL12-mediated signaling events |
| 0.0 | 1.2 | PID PI3KCI PATHWAY | Class I PI3K signaling events |
| 0.0 | 1.3 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.0 | 0.7 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.0 | 0.6 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.0 | 1.0 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.0 | 1.9 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.0 | 0.5 | PID PTP1B PATHWAY | Signaling events mediated by PTP1B |
| 0.0 | 0.7 | PID BCR 5PATHWAY | BCR signaling pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 15.6 | REACTOME INITIAL TRIGGERING OF COMPLEMENT | Genes involved in Initial triggering of complement |
| 0.5 | 8.8 | REACTOME NEGATIVE REGULATION OF THE PI3K AKT NETWORK | Genes involved in Negative regulation of the PI3K/AKT network |
| 0.5 | 9.1 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 0.4 | 16.8 | REACTOME GRB2 SOS PROVIDES LINKAGE TO MAPK SIGNALING FOR INTERGRINS | Genes involved in GRB2:SOS provides linkage to MAPK signaling for Intergrins |
| 0.4 | 9.0 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 0.3 | 3.8 | REACTOME GLYCOPROTEIN HORMONES | Genes involved in Glycoprotein hormones |
| 0.2 | 4.3 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.1 | 5.4 | REACTOME PLATELET SENSITIZATION BY LDL | Genes involved in Platelet sensitization by LDL |
| 0.1 | 6.8 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.1 | 4.4 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.1 | 5.1 | REACTOME ACTIVATION OF GENES BY ATF4 | Genes involved in Activation of Genes by ATF4 |
| 0.1 | 3.5 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.1 | 10.7 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.1 | 2.5 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.1 | 2.4 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.1 | 2.0 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.1 | 1.6 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 7ALPHA HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 7alpha-hydroxycholesterol |
| 0.1 | 5.2 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.0 | 0.9 | REACTOME ER PHAGOSOME PATHWAY | Genes involved in ER-Phagosome pathway |
| 0.0 | 1.6 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.0 | 11.9 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.0 | 2.3 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.0 | 0.7 | REACTOME CALNEXIN CALRETICULIN CYCLE | Genes involved in Calnexin/calreticulin cycle |
| 0.0 | 0.6 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.0 | 7.5 | REACTOME TOLL RECEPTOR CASCADES | Genes involved in Toll Receptor Cascades |
| 0.0 | 0.5 | REACTOME REGULATED PROTEOLYSIS OF P75NTR | Genes involved in Regulated proteolysis of p75NTR |
| 0.0 | 2.6 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.0 | 0.7 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.0 | 1.7 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.0 | 0.8 | REACTOME SYNTHESIS OF PE | Genes involved in Synthesis of PE |
| 0.0 | 0.8 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.0 | 1.2 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.0 | 4.4 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.0 | 0.5 | REACTOME PROLACTIN RECEPTOR SIGNALING | Genes involved in Prolactin receptor signaling |
| 0.0 | 1.4 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.0 | 1.3 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.0 | 1.1 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.0 | 0.9 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.0 | 0.4 | REACTOME AMYLOIDS | Genes involved in Amyloids |