Project
Illumina Body Map 2: averaged replicates
Navigation
Downloads
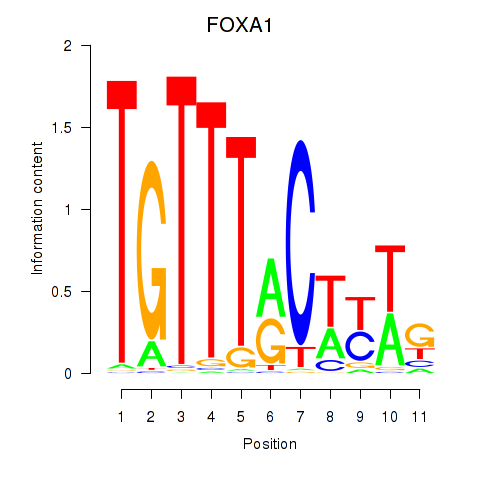
Results for FOXA1
Z-value: 2.67
Transcription factors associated with FOXA1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
FOXA1
|
ENSG00000129514.4 | forkhead box A1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| FOXA1 | hg19_v2_chr14_-_38064198_38064239 | 0.69 | 1.3e-05 | Click! |
Activity profile of FOXA1 motif
Sorted Z-values of FOXA1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr17_-_26695013 | 12.01 |
ENST00000555059.2
|
CTB-96E2.2
|
Homeobox protein SEBOX |
| chr17_-_26694979 | 11.83 |
ENST00000438614.1
|
VTN
|
vitronectin |
| chr1_-_57431679 | 9.77 |
ENST00000371237.4
ENST00000535057.1 ENST00000543257.1 |
C8B
|
complement component 8, beta polypeptide |
| chr14_-_94789663 | 9.39 |
ENST00000557225.1
ENST00000341584.3 |
SERPINA6
|
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 6 |
| chr4_-_155533787 | 9.12 |
ENST00000407946.1
ENST00000405164.1 ENST00000336098.3 ENST00000393846.2 ENST00000404648.3 ENST00000443553.1 |
FGG
|
fibrinogen gamma chain |
| chr17_-_39684550 | 7.63 |
ENST00000455635.1
ENST00000361566.3 |
KRT19
|
keratin 19 |
| chr14_-_38064198 | 6.54 |
ENST00000250448.2
|
FOXA1
|
forkhead box A1 |
| chr10_+_81370689 | 6.39 |
ENST00000372308.3
ENST00000398636.3 ENST00000428376.2 ENST00000372313.5 ENST00000419470.2 ENST00000429958.1 ENST00000439264.1 |
SFTPA1
|
surfactant protein A1 |
| chr4_-_72649763 | 6.19 |
ENST00000513476.1
|
GC
|
group-specific component (vitamin D binding protein) |
| chr17_-_46035187 | 6.09 |
ENST00000300557.2
|
PRR15L
|
proline rich 15-like |
| chr11_-_116663127 | 5.76 |
ENST00000433069.1
ENST00000542499.1 |
APOA5
|
apolipoprotein A-V |
| chr8_+_120079478 | 5.61 |
ENST00000332843.2
|
COLEC10
|
collectin sub-family member 10 (C-type lectin) |
| chr5_-_41213607 | 5.55 |
ENST00000337836.5
ENST00000433294.1 |
C6
|
complement component 6 |
| chr22_+_21128167 | 5.48 |
ENST00000215727.5
|
SERPIND1
|
serpin peptidase inhibitor, clade D (heparin cofactor), member 1 |
| chr11_+_62186498 | 5.48 |
ENST00000278282.2
|
SCGB1A1
|
secretoglobin, family 1A, member 1 (uteroglobin) |
| chr3_+_52811596 | 5.04 |
ENST00000542827.1
ENST00000273283.2 |
ITIH1
|
inter-alpha-trypsin inhibitor heavy chain 1 |
| chr4_+_187148556 | 4.96 |
ENST00000264690.6
ENST00000446598.2 ENST00000414291.1 ENST00000513864.1 |
KLKB1
|
kallikrein B, plasma (Fletcher factor) 1 |
| chr3_+_186330712 | 4.93 |
ENST00000411641.2
ENST00000273784.5 |
AHSG
|
alpha-2-HS-glycoprotein |
| chr5_-_41213123 | 4.92 |
ENST00000417809.1
|
C6
|
complement component 6 |
| chr15_+_69854027 | 4.87 |
ENST00000498938.2
|
RP11-279F6.1
|
RP11-279F6.1 |
| chr10_-_81320151 | 4.85 |
ENST00000372325.2
ENST00000372327.5 ENST00000417041.1 |
SFTPA2
|
surfactant protein A2 |
| chr10_+_51549498 | 4.82 |
ENST00000358559.2
ENST00000298239.6 |
MSMB
|
microseminoprotein, beta- |
| chr11_-_102401469 | 4.55 |
ENST00000260227.4
|
MMP7
|
matrix metallopeptidase 7 (matrilysin, uterine) |
| chr2_-_89161064 | 4.29 |
ENST00000390241.2
|
IGKJ2
|
immunoglobulin kappa joining 2 |
| chr20_-_7921090 | 4.28 |
ENST00000378789.3
|
HAO1
|
hydroxyacid oxidase (glycolate oxidase) 1 |
| chr12_-_53343602 | 4.28 |
ENST00000546897.1
ENST00000552551.1 |
KRT8
|
keratin 8 |
| chr2_+_7865923 | 4.25 |
ENST00000417930.1
|
AC092580.4
|
AC092580.4 |
| chr20_+_31595406 | 4.24 |
ENST00000170150.3
|
BPIFB2
|
BPI fold containing family B, member 2 |
| chr14_-_36988882 | 4.23 |
ENST00000498187.2
|
NKX2-1
|
NK2 homeobox 1 |
| chr3_+_137717571 | 3.88 |
ENST00000343735.4
|
CLDN18
|
claudin 18 |
| chr21_-_31588365 | 3.84 |
ENST00000399899.1
|
CLDN8
|
claudin 8 |
| chrX_+_135614293 | 3.81 |
ENST00000370634.3
|
VGLL1
|
vestigial like 1 (Drosophila) |
| chr1_-_27240455 | 3.80 |
ENST00000254227.3
|
NR0B2
|
nuclear receptor subfamily 0, group B, member 2 |
| chr16_-_86542455 | 3.78 |
ENST00000595886.1
ENST00000597578.1 ENST00000593604.1 |
FENDRR
|
FOXF1 adjacent non-coding developmental regulatory RNA |
| chr4_+_187187098 | 3.77 |
ENST00000403665.2
ENST00000264692.4 |
F11
|
coagulation factor XI |
| chr2_-_89160425 | 3.70 |
ENST00000390239.2
|
IGKJ4
|
immunoglobulin kappa joining 4 |
| chr12_-_53343633 | 3.66 |
ENST00000546826.1
|
KRT8
|
keratin 8 |
| chr2_-_69180083 | 3.65 |
ENST00000328895.4
|
GKN2
|
gastrokine 2 |
| chr3_+_137728842 | 3.60 |
ENST00000183605.5
|
CLDN18
|
claudin 18 |
| chr19_+_18496957 | 3.56 |
ENST00000252809.3
|
GDF15
|
growth differentiation factor 15 |
| chr3_-_148939598 | 3.52 |
ENST00000455472.3
|
CP
|
ceruloplasmin (ferroxidase) |
| chr12_-_53343560 | 3.48 |
ENST00000548998.1
|
KRT8
|
keratin 8 |
| chr21_-_31588338 | 3.48 |
ENST00000286809.1
|
CLDN8
|
claudin 8 |
| chr21_-_43735446 | 3.47 |
ENST00000398431.2
|
TFF3
|
trefoil factor 3 (intestinal) |
| chr1_+_200011711 | 3.40 |
ENST00000544748.1
|
NR5A2
|
nuclear receptor subfamily 5, group A, member 2 |
| chr2_-_165424973 | 3.30 |
ENST00000543549.1
|
GRB14
|
growth factor receptor-bound protein 14 |
| chr2_+_162101247 | 3.26 |
ENST00000439050.1
ENST00000436506.1 |
AC009299.3
|
AC009299.3 |
| chr11_-_124622134 | 3.25 |
ENST00000326621.5
|
VSIG2
|
V-set and immunoglobulin domain containing 2 |
| chr13_-_86373536 | 3.20 |
ENST00000400286.2
|
SLITRK6
|
SLIT and NTRK-like family, member 6 |
| chr2_-_89160770 | 3.19 |
ENST00000390240.2
|
IGKJ3
|
immunoglobulin kappa joining 3 |
| chr2_-_89160117 | 3.18 |
ENST00000390238.2
|
IGKJ5
|
immunoglobulin kappa joining 5 |
| chr12_-_120765565 | 3.18 |
ENST00000423423.3
ENST00000308366.4 |
PLA2G1B
|
phospholipase A2, group IB (pancreas) |
| chr12_-_91572278 | 3.15 |
ENST00000425043.1
ENST00000420120.2 ENST00000441303.2 ENST00000456569.2 |
DCN
|
decorin |
| chr7_+_134576151 | 3.11 |
ENST00000393118.2
|
CALD1
|
caldesmon 1 |
| chr2_-_21266935 | 3.11 |
ENST00000233242.1
|
APOB
|
apolipoprotein B |
| chr1_+_199996733 | 2.98 |
ENST00000236914.3
|
NR5A2
|
nuclear receptor subfamily 5, group A, member 2 |
| chr17_+_8924837 | 2.97 |
ENST00000173229.2
|
NTN1
|
netrin 1 |
| chr3_+_148447887 | 2.95 |
ENST00000475347.1
ENST00000474935.1 ENST00000461609.1 |
AGTR1
|
angiotensin II receptor, type 1 |
| chr3_-_148939835 | 2.94 |
ENST00000264613.6
|
CP
|
ceruloplasmin (ferroxidase) |
| chr6_+_135502501 | 2.93 |
ENST00000527615.1
ENST00000420123.2 ENST00000525369.1 ENST00000528774.1 ENST00000534121.1 ENST00000534044.1 ENST00000533624.1 |
MYB
|
v-myb avian myeloblastosis viral oncogene homolog |
| chr7_+_73242490 | 2.93 |
ENST00000431918.1
|
CLDN4
|
claudin 4 |
| chr1_+_199996702 | 2.91 |
ENST00000367362.3
|
NR5A2
|
nuclear receptor subfamily 5, group A, member 2 |
| chr15_+_96869165 | 2.91 |
ENST00000421109.2
|
NR2F2
|
nuclear receptor subfamily 2, group F, member 2 |
| chr11_-_124622083 | 2.82 |
ENST00000403470.1
|
VSIG2
|
V-set and immunoglobulin domain containing 2 |
| chr9_-_4859260 | 2.78 |
ENST00000599351.1
|
AL158147.2
|
HCG2011465; Uncharacterized protein |
| chr1_-_177939041 | 2.74 |
ENST00000308284.6
|
SEC16B
|
SEC16 homolog B (S. cerevisiae) |
| chr4_+_55095264 | 2.71 |
ENST00000257290.5
|
PDGFRA
|
platelet-derived growth factor receptor, alpha polypeptide |
| chr10_+_124320156 | 2.71 |
ENST00000338354.3
ENST00000344338.3 ENST00000330163.4 ENST00000368909.3 ENST00000368955.3 ENST00000368956.2 |
DMBT1
|
deleted in malignant brain tumors 1 |
| chr4_+_187187337 | 2.69 |
ENST00000492972.2
|
F11
|
coagulation factor XI |
| chr2_-_69180012 | 2.68 |
ENST00000481498.1
|
GKN2
|
gastrokine 2 |
| chr1_-_203320617 | 2.68 |
ENST00000354955.4
|
FMOD
|
fibromodulin |
| chr12_-_102872317 | 2.67 |
ENST00000424202.2
|
IGF1
|
insulin-like growth factor 1 (somatomedin C) |
| chr18_+_56113488 | 2.65 |
ENST00000590797.1
|
RP11-1151B14.3
|
RP11-1151B14.3 |
| chr6_+_161123270 | 2.65 |
ENST00000366924.2
ENST00000308192.9 ENST00000418964.1 |
PLG
|
plasminogen |
| chr20_+_31823792 | 2.63 |
ENST00000375413.4
ENST00000354297.4 ENST00000375422.2 |
BPIFA1
|
BPI fold containing family A, member 1 |
| chr14_-_70263979 | 2.63 |
ENST00000216540.4
|
SLC10A1
|
solute carrier family 10 (sodium/bile acid cotransporter), member 1 |
| chr9_+_27109133 | 2.63 |
ENST00000519097.1
ENST00000380036.4 |
TEK
|
TEK tyrosine kinase, endothelial |
| chr2_-_104496621 | 2.62 |
ENST00000455716.1
|
AC013727.1
|
AC013727.1 |
| chr7_-_81399438 | 2.62 |
ENST00000222390.5
|
HGF
|
hepatocyte growth factor (hepapoietin A; scatter factor) |
| chr3_+_186692745 | 2.62 |
ENST00000438590.1
|
ST6GAL1
|
ST6 beta-galactosamide alpha-2,6-sialyltranferase 1 |
| chr6_-_41715128 | 2.60 |
ENST00000356667.4
ENST00000373025.3 ENST00000425343.2 |
PGC
|
progastricsin (pepsinogen C) |
| chr4_-_70080449 | 2.51 |
ENST00000446444.1
|
UGT2B11
|
UDP glucuronosyltransferase 2 family, polypeptide B11 |
| chr17_-_48546324 | 2.49 |
ENST00000508540.1
|
CHAD
|
chondroadherin |
| chr1_-_111743285 | 2.46 |
ENST00000357640.4
|
DENND2D
|
DENN/MADD domain containing 2D |
| chr15_-_55541227 | 2.37 |
ENST00000566877.1
|
RAB27A
|
RAB27A, member RAS oncogene family |
| chr6_-_46922659 | 2.36 |
ENST00000265417.7
|
GPR116
|
G protein-coupled receptor 116 |
| chr18_+_66465475 | 2.32 |
ENST00000581520.1
|
CCDC102B
|
coiled-coil domain containing 102B |
| chr12_-_89746173 | 2.31 |
ENST00000308385.6
|
DUSP6
|
dual specificity phosphatase 6 |
| chrX_-_15402498 | 2.28 |
ENST00000297904.3
|
FIGF
|
c-fos induced growth factor (vascular endothelial growth factor D) |
| chr7_-_81399329 | 2.28 |
ENST00000453411.1
ENST00000444829.2 |
HGF
|
hepatocyte growth factor (hepapoietin A; scatter factor) |
| chr17_-_48546232 | 2.27 |
ENST00000258969.4
|
CHAD
|
chondroadherin |
| chr13_-_46716969 | 2.21 |
ENST00000435666.2
|
LCP1
|
lymphocyte cytosolic protein 1 (L-plastin) |
| chr21_+_17791648 | 2.17 |
ENST00000602892.1
ENST00000418813.2 ENST00000435697.1 |
LINC00478
|
long intergenic non-protein coding RNA 478 |
| chr8_+_126442563 | 2.16 |
ENST00000311922.3
|
TRIB1
|
tribbles pseudokinase 1 |
| chr14_-_36983034 | 2.15 |
ENST00000518529.2
|
SFTA3
|
surfactant associated 3 |
| chr7_-_81399355 | 2.13 |
ENST00000457544.2
|
HGF
|
hepatocyte growth factor (hepapoietin A; scatter factor) |
| chr15_+_71389281 | 2.11 |
ENST00000355327.3
|
THSD4
|
thrombospondin, type I, domain containing 4 |
| chr4_+_55095428 | 2.09 |
ENST00000508170.1
ENST00000512143.1 |
PDGFRA
|
platelet-derived growth factor receptor, alpha polypeptide |
| chr10_+_124320195 | 2.05 |
ENST00000359586.6
|
DMBT1
|
deleted in malignant brain tumors 1 |
| chr12_+_103981044 | 2.04 |
ENST00000388887.2
|
STAB2
|
stabilin 2 |
| chr2_-_21266816 | 2.02 |
ENST00000399256.4
|
APOB
|
apolipoprotein B |
| chr6_-_90025011 | 2.00 |
ENST00000402938.3
|
GABRR2
|
gamma-aminobutyric acid (GABA) A receptor, rho 2 |
| chr10_-_81708854 | 2.00 |
ENST00000372292.3
|
SFTPD
|
surfactant protein D |
| chr6_-_90024967 | 1.97 |
ENST00000602399.1
|
GABRR2
|
gamma-aminobutyric acid (GABA) A receptor, rho 2 |
| chr12_+_27849378 | 1.96 |
ENST00000310791.2
|
REP15
|
RAB15 effector protein |
| chr21_+_17791838 | 1.94 |
ENST00000453910.1
|
LINC00478
|
long intergenic non-protein coding RNA 478 |
| chr21_+_17443521 | 1.93 |
ENST00000456342.1
|
LINC00478
|
long intergenic non-protein coding RNA 478 |
| chr12_-_71551652 | 1.92 |
ENST00000546561.1
|
TSPAN8
|
tetraspanin 8 |
| chr19_-_49118067 | 1.91 |
ENST00000593772.1
|
FAM83E
|
family with sequence similarity 83, member E |
| chr7_+_134576317 | 1.90 |
ENST00000424922.1
ENST00000495522.1 |
CALD1
|
caldesmon 1 |
| chr21_+_17792672 | 1.90 |
ENST00000602620.1
|
LINC00478
|
long intergenic non-protein coding RNA 478 |
| chr3_+_52245458 | 1.87 |
ENST00000459884.1
|
ALAS1
|
aminolevulinate, delta-, synthase 1 |
| chr3_-_145878954 | 1.86 |
ENST00000282903.5
ENST00000360060.3 |
PLOD2
|
procollagen-lysine, 2-oxoglutarate 5-dioxygenase 2 |
| chr3_+_142315225 | 1.86 |
ENST00000457734.2
ENST00000483373.1 ENST00000475296.1 ENST00000495744.1 ENST00000476044.1 ENST00000461644.1 |
PLS1
|
plastin 1 |
| chr1_+_223101757 | 1.86 |
ENST00000284476.6
|
DISP1
|
dispatched homolog 1 (Drosophila) |
| chr9_+_136287444 | 1.80 |
ENST00000355699.2
ENST00000356589.2 ENST00000371911.3 |
ADAMTS13
|
ADAM metallopeptidase with thrombospondin type 1 motif, 13 |
| chr7_+_80231466 | 1.78 |
ENST00000309881.7
ENST00000534394.1 |
CD36
|
CD36 molecule (thrombospondin receptor) |
| chr12_-_89746264 | 1.76 |
ENST00000548755.1
|
DUSP6
|
dual specificity phosphatase 6 |
| chr2_+_88047606 | 1.74 |
ENST00000359481.4
|
PLGLB2
|
plasminogen-like B2 |
| chr7_-_81399411 | 1.74 |
ENST00000423064.2
|
HGF
|
hepatocyte growth factor (hepapoietin A; scatter factor) |
| chr4_-_111119804 | 1.74 |
ENST00000394607.3
ENST00000302274.3 |
ELOVL6
|
ELOVL fatty acid elongase 6 |
| chr12_-_71551868 | 1.72 |
ENST00000247829.3
|
TSPAN8
|
tetraspanin 8 |
| chr7_-_81399678 | 1.72 |
ENST00000412881.1
|
HGF
|
hepatocyte growth factor (hepapoietin A; scatter factor) |
| chr21_-_40033618 | 1.72 |
ENST00000417133.2
ENST00000398910.1 ENST00000442448.1 |
ERG
|
v-ets avian erythroblastosis virus E26 oncogene homolog |
| chr9_-_95186739 | 1.70 |
ENST00000375550.4
|
OMD
|
osteomodulin |
| chr7_-_81399287 | 1.70 |
ENST00000354224.6
|
HGF
|
hepatocyte growth factor (hepapoietin A; scatter factor) |
| chr21_-_43771226 | 1.70 |
ENST00000291526.4
|
TFF2
|
trefoil factor 2 |
| chr13_+_78109804 | 1.68 |
ENST00000535157.1
|
SCEL
|
sciellin |
| chr16_-_68034452 | 1.68 |
ENST00000575510.1
|
DPEP2
|
dipeptidase 2 |
| chr19_+_18492973 | 1.68 |
ENST00000595973.2
|
GDF15
|
growth differentiation factor 15 |
| chr3_+_174577070 | 1.67 |
ENST00000454872.1
|
NAALADL2
|
N-acetylated alpha-linked acidic dipeptidase-like 2 |
| chr4_-_74486109 | 1.66 |
ENST00000395777.2
|
RASSF6
|
Ras association (RalGDS/AF-6) domain family member 6 |
| chr6_+_10528560 | 1.65 |
ENST00000379597.3
|
GCNT2
|
glucosaminyl (N-acetyl) transferase 2, I-branching enzyme (I blood group) |
| chr11_-_108464465 | 1.64 |
ENST00000525344.1
|
EXPH5
|
exophilin 5 |
| chr11_-_108464321 | 1.64 |
ENST00000265843.4
|
EXPH5
|
exophilin 5 |
| chr6_-_161085291 | 1.61 |
ENST00000316300.5
|
LPA
|
lipoprotein, Lp(a) |
| chr5_-_121659052 | 1.59 |
ENST00000512105.1
|
CTD-2544H17.1
|
CTD-2544H17.1 |
| chr18_+_68002675 | 1.58 |
ENST00000584919.1
|
RP11-41O4.1
|
Uncharacterized protein |
| chr1_-_95391315 | 1.57 |
ENST00000545882.1
ENST00000415017.1 |
CNN3
|
calponin 3, acidic |
| chr2_-_182545603 | 1.55 |
ENST00000295108.3
|
NEUROD1
|
neuronal differentiation 1 |
| chr14_+_37126765 | 1.54 |
ENST00000402703.2
|
PAX9
|
paired box 9 |
| chr17_+_72427477 | 1.53 |
ENST00000342648.5
ENST00000481232.1 |
GPRC5C
|
G protein-coupled receptor, family C, group 5, member C |
| chr2_+_171640291 | 1.52 |
ENST00000409885.1
|
ERICH2
|
glutamate-rich 2 |
| chr18_+_66465302 | 1.51 |
ENST00000360242.5
ENST00000358653.5 |
CCDC102B
|
coiled-coil domain containing 102B |
| chr13_+_78109884 | 1.46 |
ENST00000377246.3
ENST00000349847.3 |
SCEL
|
sciellin |
| chr5_-_148758839 | 1.45 |
ENST00000261796.3
|
IL17B
|
interleukin 17B |
| chr14_+_38065052 | 1.44 |
ENST00000556845.1
|
TTC6
|
tetratricopeptide repeat domain 6 |
| chr1_-_216596738 | 1.41 |
ENST00000307340.3
ENST00000366943.2 ENST00000366942.3 |
USH2A
|
Usher syndrome 2A (autosomal recessive, mild) |
| chr3_-_185538849 | 1.39 |
ENST00000421047.2
|
IGF2BP2
|
insulin-like growth factor 2 mRNA binding protein 2 |
| chr15_+_59910132 | 1.38 |
ENST00000559200.1
|
GCNT3
|
glucosaminyl (N-acetyl) transferase 3, mucin type |
| chr1_-_178840157 | 1.37 |
ENST00000367629.1
ENST00000234816.2 |
ANGPTL1
|
angiopoietin-like 1 |
| chr7_+_116165754 | 1.37 |
ENST00000405348.1
|
CAV1
|
caveolin 1, caveolae protein, 22kDa |
| chr7_-_81399744 | 1.36 |
ENST00000421558.1
|
HGF
|
hepatocyte growth factor (hepapoietin A; scatter factor) |
| chr2_+_71162995 | 1.35 |
ENST00000234396.4
|
ATP6V1B1
|
ATPase, H+ transporting, lysosomal 56/58kDa, V1 subunit B1 |
| chr13_+_76334795 | 1.32 |
ENST00000526202.1
ENST00000465261.2 |
LMO7
|
LIM domain 7 |
| chr10_+_102891048 | 1.30 |
ENST00000467928.2
|
TLX1
|
T-cell leukemia homeobox 1 |
| chr21_+_17443434 | 1.29 |
ENST00000400178.2
|
LINC00478
|
long intergenic non-protein coding RNA 478 |
| chr6_-_87804815 | 1.27 |
ENST00000369582.2
|
CGA
|
glycoprotein hormones, alpha polypeptide |
| chr2_+_71163051 | 1.27 |
ENST00000412314.1
|
ATP6V1B1
|
ATPase, H+ transporting, lysosomal 56/58kDa, V1 subunit B1 |
| chr18_-_52626622 | 1.24 |
ENST00000591504.1
|
CCDC68
|
coiled-coil domain containing 68 |
| chr2_-_158345462 | 1.23 |
ENST00000439355.1
ENST00000540637.1 |
CYTIP
|
cytohesin 1 interacting protein |
| chr13_+_76334567 | 1.21 |
ENST00000321797.8
|
LMO7
|
LIM domain 7 |
| chr3_+_142315294 | 1.19 |
ENST00000464320.1
|
PLS1
|
plastin 1 |
| chr3_+_160394940 | 1.18 |
ENST00000320767.2
|
ARL14
|
ADP-ribosylation factor-like 14 |
| chr12_+_121416437 | 1.18 |
ENST00000402929.1
ENST00000535955.1 ENST00000538626.1 ENST00000543427.1 |
HNF1A
|
HNF1 homeobox A |
| chr12_+_121416340 | 1.17 |
ENST00000257555.6
ENST00000400024.2 |
HNF1A
|
HNF1 homeobox A |
| chr6_-_108145499 | 1.13 |
ENST00000369020.3
ENST00000369022.2 |
SCML4
|
sex comb on midleg-like 4 (Drosophila) |
| chr7_-_107443652 | 1.13 |
ENST00000340010.5
ENST00000422236.2 ENST00000453332.1 |
SLC26A3
|
solute carrier family 26 (anion exchanger), member 3 |
| chr14_-_104408032 | 1.12 |
ENST00000455348.2
|
RD3L
|
retinal degeneration 3-like |
| chr11_+_844406 | 1.10 |
ENST00000397404.1
|
TSPAN4
|
tetraspanin 4 |
| chr12_+_121416489 | 1.08 |
ENST00000541395.1
ENST00000544413.1 |
HNF1A
|
HNF1 homeobox A |
| chr6_+_135502466 | 1.08 |
ENST00000367814.4
|
MYB
|
v-myb avian myeloblastosis viral oncogene homolog |
| chr2_-_158345341 | 1.08 |
ENST00000435117.1
|
CYTIP
|
cytohesin 1 interacting protein |
| chr8_-_133772794 | 1.06 |
ENST00000519187.1
ENST00000523829.1 ENST00000356838.3 ENST00000377901.4 ENST00000519304.1 |
TMEM71
|
transmembrane protein 71 |
| chr10_-_90751038 | 1.04 |
ENST00000458159.1
ENST00000415557.1 ENST00000458208.1 |
ACTA2
|
actin, alpha 2, smooth muscle, aorta |
| chr10_-_21806759 | 1.03 |
ENST00000444772.3
|
SKIDA1
|
SKI/DACH domain containing 1 |
| chrX_+_37865804 | 1.00 |
ENST00000297875.2
ENST00000357972.5 |
SYTL5
|
synaptotagmin-like 5 |
| chr17_-_17485731 | 1.00 |
ENST00000395783.1
|
PEMT
|
phosphatidylethanolamine N-methyltransferase |
| chr17_-_39140549 | 1.00 |
ENST00000377755.4
|
KRT40
|
keratin 40 |
| chr3_+_72201910 | 0.99 |
ENST00000469178.1
ENST00000485404.1 |
LINC00870
|
long intergenic non-protein coding RNA 870 |
| chr16_-_73178346 | 0.99 |
ENST00000358463.2
|
C16orf47
|
chromosome 16 open reading frame 47 |
| chr4_+_75174180 | 0.98 |
ENST00000413830.1
|
EPGN
|
epithelial mitogen |
| chr4_-_74486347 | 0.96 |
ENST00000342081.3
|
RASSF6
|
Ras association (RalGDS/AF-6) domain family member 6 |
| chr3_+_121902511 | 0.96 |
ENST00000490131.1
|
CASR
|
calcium-sensing receptor |
| chr4_+_68424434 | 0.94 |
ENST00000265404.2
ENST00000396225.1 |
STAP1
|
signal transducing adaptor family member 1 |
| chr10_-_23528745 | 0.91 |
ENST00000376501.5
|
C10orf115
|
chromosome 10 open reading frame 115 |
| chr7_+_134430212 | 0.91 |
ENST00000436461.2
|
CALD1
|
caldesmon 1 |
| chr7_-_92855762 | 0.91 |
ENST00000453812.2
ENST00000394468.2 |
HEPACAM2
|
HEPACAM family member 2 |
| chr19_-_48894104 | 0.90 |
ENST00000597017.1
|
KDELR1
|
KDEL (Lys-Asp-Glu-Leu) endoplasmic reticulum protein retention receptor 1 |
| chr17_+_72426891 | 0.90 |
ENST00000392627.1
|
GPRC5C
|
G protein-coupled receptor, family C, group 5, member C |
| chr7_-_77325545 | 0.85 |
ENST00000447009.1
ENST00000416650.1 ENST00000440088.1 ENST00000430801.1 ENST00000398043.2 |
RSBN1L-AS1
|
RSBN1L antisense RNA 1 |
| chr5_+_179921430 | 0.85 |
ENST00000393356.1
|
CNOT6
|
CCR4-NOT transcription complex, subunit 6 |
| chr6_+_139135648 | 0.83 |
ENST00000541398.1
|
ECT2L
|
epithelial cell transforming sequence 2 oncogene-like |
| chr2_+_204801471 | 0.83 |
ENST00000316386.6
ENST00000435193.1 |
ICOS
|
inducible T-cell co-stimulator |
| chr1_+_94798754 | 0.81 |
ENST00000418242.1
|
RP11-148B18.3
|
RP11-148B18.3 |
| chr4_+_165675197 | 0.81 |
ENST00000515485.1
|
RP11-294O2.2
|
RP11-294O2.2 |
| chr4_-_186456766 | 0.80 |
ENST00000284771.6
|
PDLIM3
|
PDZ and LIM domain 3 |
| chr8_-_133772870 | 0.78 |
ENST00000522334.1
ENST00000519016.1 |
TMEM71
|
transmembrane protein 71 |
| chr2_-_192711968 | 0.77 |
ENST00000304141.4
|
SDPR
|
serum deprivation response |
| chr6_+_112375275 | 0.76 |
ENST00000368666.2
ENST00000604763.1 ENST00000230529.5 |
WISP3
|
WNT1 inducible signaling pathway protein 3 |
Network of associatons between targets according to the STRING database.
First level regulatory network of FOXA1
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.8 | 14.1 | GO:0051919 | positive regulation of fibrinolysis(GO:0051919) |
| 2.2 | 6.5 | GO:0060738 | epithelial-mesenchymal signaling involved in prostate gland development(GO:0060738) |
| 2.1 | 10.5 | GO:0001970 | positive regulation of activation of membrane attack complex(GO:0001970) |
| 1.5 | 4.5 | GO:1904897 | regulation of hepatic stellate cell proliferation(GO:1904897) positive regulation of hepatic stellate cell proliferation(GO:1904899) hepatic stellate cell proliferation(GO:1990922) |
| 1.2 | 4.8 | GO:1900154 | regulation of bone trabecula formation(GO:1900154) negative regulation of bone trabecula formation(GO:1900155) |
| 1.2 | 5.8 | GO:0010902 | positive regulation of very-low-density lipoprotein particle remodeling(GO:0010902) |
| 1.1 | 3.4 | GO:0035623 | renal glucose absorption(GO:0035623) |
| 1.1 | 9.5 | GO:0061113 | pancreas morphogenesis(GO:0061113) |
| 1.0 | 13.5 | GO:0060665 | regulation of branching involved in salivary gland morphogenesis by mesenchymal-epithelial signaling(GO:0060665) |
| 0.9 | 5.5 | GO:0010193 | response to ozone(GO:0010193) |
| 0.9 | 7.0 | GO:0009441 | glycolate metabolic process(GO:0009441) |
| 0.8 | 4.1 | GO:0042663 | regulation of endodermal cell fate specification(GO:0042663) |
| 0.8 | 5.5 | GO:0008218 | bioluminescence(GO:0008218) |
| 0.7 | 7.5 | GO:0071847 | TNFSF11-mediated signaling pathway(GO:0071847) |
| 0.7 | 2.9 | GO:0086097 | phospholipase C-activating angiotensin-activated signaling pathway(GO:0086097) |
| 0.7 | 2.9 | GO:0060849 | radial pattern formation(GO:0009956) regulation of transcription involved in lymphatic endothelial cell fate commitment(GO:0060849) |
| 0.7 | 2.2 | GO:0045645 | regulation of eosinophil differentiation(GO:0045643) positive regulation of eosinophil differentiation(GO:0045645) |
| 0.7 | 4.8 | GO:0035790 | platelet-derived growth factor receptor-alpha signaling pathway(GO:0035790) |
| 0.7 | 11.4 | GO:0051599 | response to hydrostatic pressure(GO:0051599) |
| 0.7 | 2.6 | GO:1900191 | biofilm formation(GO:0042710) single-species biofilm formation(GO:0044010) single-species biofilm formation in or on host organism(GO:0044407) regulation of single-species biofilm formation(GO:1900190) negative regulation of single-species biofilm formation(GO:1900191) regulation of single-species biofilm formation in or on host organism(GO:1900228) negative regulation of single-species biofilm formation in or on host organism(GO:1900229) |
| 0.6 | 5.1 | GO:0006642 | triglyceride mobilization(GO:0006642) |
| 0.6 | 3.2 | GO:0044240 | multicellular organism lipid catabolic process(GO:0044240) |
| 0.6 | 11.8 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.5 | 2.7 | GO:0007181 | transforming growth factor beta receptor complex assembly(GO:0007181) |
| 0.5 | 3.2 | GO:0060005 | vestibular reflex(GO:0060005) |
| 0.5 | 1.5 | GO:0060730 | regulation of intestinal epithelial structure maintenance(GO:0060730) |
| 0.5 | 6.8 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 0.5 | 1.9 | GO:0046947 | hydroxylysine metabolic process(GO:0046946) hydroxylysine biosynthetic process(GO:0046947) |
| 0.5 | 1.4 | GO:1903609 | negative regulation of peptidyl-tyrosine autophosphorylation(GO:1900085) negative regulation of inward rectifier potassium channel activity(GO:1903609) |
| 0.4 | 3.0 | GO:1902896 | terminal web assembly(GO:1902896) |
| 0.4 | 2.6 | GO:0002760 | positive regulation of antimicrobial peptide production(GO:0002225) positive regulation of antimicrobial humoral response(GO:0002760) positive regulation of antibacterial peptide production(GO:0002803) |
| 0.4 | 9.8 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 0.4 | 9.1 | GO:0031639 | plasminogen activation(GO:0031639) |
| 0.4 | 6.3 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 0.4 | 3.0 | GO:0033564 | anterior/posterior axon guidance(GO:0033564) |
| 0.4 | 6.4 | GO:0008228 | opsonization(GO:0008228) |
| 0.4 | 4.2 | GO:0021759 | globus pallidus development(GO:0021759) |
| 0.3 | 1.4 | GO:0002426 | immunoglobulin production in mucosal tissue(GO:0002426) |
| 0.3 | 2.4 | GO:1903435 | positive regulation of constitutive secretory pathway(GO:1903435) |
| 0.3 | 2.6 | GO:0048014 | Tie signaling pathway(GO:0048014) |
| 0.3 | 1.3 | GO:1903691 | positive regulation of wound healing, spreading of epidermal cells(GO:1903691) |
| 0.3 | 0.9 | GO:1903980 | negative regulation of macrophage colony-stimulating factor signaling pathway(GO:1902227) negative regulation of response to macrophage colony-stimulating factor(GO:1903970) negative regulation of cellular response to macrophage colony-stimulating factor stimulus(GO:1903973) positive regulation of microglial cell activation(GO:1903980) |
| 0.3 | 10.2 | GO:0016338 | calcium-independent cell-cell adhesion via plasma membrane cell-adhesion molecules(GO:0016338) |
| 0.3 | 1.4 | GO:0048496 | maintenance of organ identity(GO:0048496) |
| 0.3 | 0.8 | GO:1902396 | protein localization to bicellular tight junction(GO:1902396) |
| 0.2 | 1.7 | GO:0003199 | endocardial cushion to mesenchymal transition involved in heart valve formation(GO:0003199) |
| 0.2 | 9.4 | GO:0008211 | glucocorticoid metabolic process(GO:0008211) |
| 0.2 | 0.8 | GO:0002304 | gamma-delta intraepithelial T cell differentiation(GO:0002304) CD8-positive, gamma-delta intraepithelial T cell differentiation(GO:0002305) |
| 0.2 | 3.1 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 0.2 | 0.4 | GO:0061030 | epithelial cell differentiation involved in mammary gland alveolus development(GO:0061030) |
| 0.2 | 1.6 | GO:0036438 | maintenance of lens transparency(GO:0036438) |
| 0.2 | 0.6 | GO:0010966 | regulation of phosphate transport(GO:0010966) |
| 0.2 | 6.5 | GO:0006825 | copper ion transport(GO:0006825) |
| 0.2 | 6.2 | GO:0042359 | vitamin D metabolic process(GO:0042359) |
| 0.2 | 4.6 | GO:0044849 | estrous cycle(GO:0044849) |
| 0.2 | 4.9 | GO:0030502 | negative regulation of bone mineralization(GO:0030502) |
| 0.2 | 0.7 | GO:0050883 | negative regulation of sodium:proton antiporter activity(GO:0032416) musculoskeletal movement, spinal reflex action(GO:0050883) |
| 0.2 | 1.0 | GO:0090131 | mesenchyme migration(GO:0090131) |
| 0.2 | 3.3 | GO:0071985 | multivesicular body sorting pathway(GO:0071985) |
| 0.2 | 1.8 | GO:0035864 | response to potassium ion(GO:0035864) |
| 0.2 | 2.1 | GO:0048251 | elastic fiber assembly(GO:0048251) |
| 0.2 | 1.8 | GO:0070543 | response to linoleic acid(GO:0070543) blood microparticle formation(GO:0072564) regulation of blood microparticle formation(GO:2000332) positive regulation of blood microparticle formation(GO:2000334) |
| 0.2 | 7.7 | GO:0060706 | cell differentiation involved in embryonic placenta development(GO:0060706) |
| 0.1 | 4.4 | GO:1901741 | positive regulation of myoblast fusion(GO:1901741) |
| 0.1 | 1.7 | GO:0034626 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.1 | 2.9 | GO:0070863 | positive regulation of protein exit from endoplasmic reticulum(GO:0070863) |
| 0.1 | 0.4 | GO:0002276 | basophil activation involved in immune response(GO:0002276) |
| 0.1 | 1.9 | GO:0006782 | protoporphyrinogen IX biosynthetic process(GO:0006782) |
| 0.1 | 7.8 | GO:0030212 | hyaluronan metabolic process(GO:0030212) |
| 0.1 | 0.2 | GO:1904017 | cellular response to Thyroglobulin triiodothyronine(GO:1904017) |
| 0.1 | 1.0 | GO:0050747 | positive regulation of lipoprotein metabolic process(GO:0050747) |
| 0.1 | 2.6 | GO:0006054 | N-acetylneuraminate metabolic process(GO:0006054) |
| 0.1 | 2.2 | GO:0071803 | positive regulation of podosome assembly(GO:0071803) |
| 0.1 | 0.3 | GO:0034392 | negative regulation of smooth muscle cell apoptotic process(GO:0034392) |
| 0.1 | 0.4 | GO:0035711 | plasmacytoid dendritic cell activation(GO:0002270) regulation of restriction endodeoxyribonuclease activity(GO:0032072) T-helper 1 cell activation(GO:0035711) |
| 0.1 | 1.7 | GO:0042340 | keratan sulfate catabolic process(GO:0042340) |
| 0.1 | 1.6 | GO:0034374 | low-density lipoprotein particle remodeling(GO:0034374) |
| 0.1 | 0.5 | GO:2001045 | negative regulation of integrin-mediated signaling pathway(GO:2001045) |
| 0.1 | 1.9 | GO:0060539 | diaphragm development(GO:0060539) |
| 0.1 | 1.0 | GO:0010606 | positive regulation of cytoplasmic mRNA processing body assembly(GO:0010606) |
| 0.1 | 0.5 | GO:0050916 | sensory perception of sweet taste(GO:0050916) |
| 0.1 | 4.6 | GO:0033572 | transferrin transport(GO:0033572) |
| 0.1 | 0.7 | GO:0072733 | response to staurosporine(GO:0072733) cellular response to staurosporine(GO:0072734) |
| 0.1 | 1.6 | GO:0032780 | negative regulation of ATPase activity(GO:0032780) |
| 0.1 | 0.2 | GO:0045084 | positive regulation of interleukin-12 biosynthetic process(GO:0045084) |
| 0.1 | 0.4 | GO:0045349 | interferon-alpha biosynthetic process(GO:0045349) regulation of interferon-alpha biosynthetic process(GO:0045354) |
| 0.1 | 1.4 | GO:1900017 | positive regulation of cytokine production involved in inflammatory response(GO:1900017) |
| 0.1 | 1.1 | GO:0045852 | pH elevation(GO:0045852) intracellular pH elevation(GO:0051454) |
| 0.1 | 0.8 | GO:0001672 | regulation of chromatin assembly or disassembly(GO:0001672) |
| 0.1 | 0.9 | GO:0006621 | protein retention in ER lumen(GO:0006621) |
| 0.1 | 0.3 | GO:0006391 | transcription initiation from mitochondrial promoter(GO:0006391) |
| 0.1 | 0.8 | GO:0060174 | limb bud formation(GO:0060174) |
| 0.1 | 0.6 | GO:0097646 | calcitonin family receptor signaling pathway(GO:0097646) amylin receptor signaling pathway(GO:0097647) |
| 0.1 | 2.2 | GO:0060325 | face morphogenesis(GO:0060325) |
| 0.1 | 0.6 | GO:0060019 | radial glial cell differentiation(GO:0060019) |
| 0.1 | 3.6 | GO:1900047 | negative regulation of blood coagulation(GO:0030195) negative regulation of hemostasis(GO:1900047) |
| 0.1 | 3.3 | GO:0046627 | negative regulation of insulin receptor signaling pathway(GO:0046627) |
| 0.1 | 0.7 | GO:0061088 | regulation of sequestering of zinc ion(GO:0061088) |
| 0.1 | 0.4 | GO:0070857 | regulation of bile acid biosynthetic process(GO:0070857) |
| 0.0 | 4.8 | GO:0019730 | antimicrobial humoral response(GO:0019730) |
| 0.0 | 2.4 | GO:0015721 | bile acid and bile salt transport(GO:0015721) |
| 0.0 | 0.2 | GO:1904383 | response to sodium phosphate(GO:1904383) response to insulin-like growth factor stimulus(GO:1990418) |
| 0.0 | 1.5 | GO:0007635 | chemosensory behavior(GO:0007635) |
| 0.0 | 4.8 | GO:0007585 | respiratory gaseous exchange(GO:0007585) |
| 0.0 | 0.5 | GO:0008343 | adult feeding behavior(GO:0008343) |
| 0.0 | 0.6 | GO:0006828 | manganese ion transport(GO:0006828) |
| 0.0 | 0.5 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.0 | 0.7 | GO:0060628 | regulation of ER to Golgi vesicle-mediated transport(GO:0060628) |
| 0.0 | 0.4 | GO:0019509 | L-methionine biosynthetic process from methylthioadenosine(GO:0019509) |
| 0.0 | 3.8 | GO:0032024 | positive regulation of insulin secretion(GO:0032024) |
| 0.0 | 0.8 | GO:0097094 | craniofacial suture morphogenesis(GO:0097094) |
| 0.0 | 1.4 | GO:0008210 | estrogen metabolic process(GO:0008210) |
| 0.0 | 0.6 | GO:0002517 | T cell tolerance induction(GO:0002517) |
| 0.0 | 0.6 | GO:2000096 | positive regulation of Wnt signaling pathway, planar cell polarity pathway(GO:2000096) |
| 0.0 | 0.5 | GO:0045741 | positive regulation of epidermal growth factor-activated receptor activity(GO:0045741) |
| 0.0 | 0.6 | GO:0003215 | cardiac right ventricle morphogenesis(GO:0003215) |
| 0.0 | 1.3 | GO:0016486 | peptide hormone processing(GO:0016486) |
| 0.0 | 0.4 | GO:0060347 | heart trabecula formation(GO:0060347) |
| 0.0 | 0.1 | GO:0060023 | soft palate development(GO:0060023) |
| 0.0 | 0.3 | GO:0019441 | tryptophan catabolic process to kynurenine(GO:0019441) |
| 0.0 | 0.2 | GO:0061002 | negative regulation of dendritic spine morphogenesis(GO:0061002) |
| 0.0 | 0.4 | GO:0016254 | preassembly of GPI anchor in ER membrane(GO:0016254) |
| 0.0 | 2.6 | GO:0007586 | digestion(GO:0007586) |
| 0.0 | 0.4 | GO:0002089 | lens morphogenesis in camera-type eye(GO:0002089) |
| 0.0 | 1.4 | GO:0042035 | regulation of cytokine biosynthetic process(GO:0042035) |
| 0.0 | 0.1 | GO:0035826 | rubidium ion transport(GO:0035826) regulation of rubidium ion transport(GO:2000680) |
| 0.0 | 1.0 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.0 | 11.8 | GO:0071062 | rough endoplasmic reticulum lumen(GO:0048237) alphav-beta3 integrin-vitronectin complex(GO:0071062) |
| 2.0 | 20.2 | GO:0005579 | membrane attack complex(GO:0005579) |
| 1.3 | 10.7 | GO:1990357 | terminal web(GO:1990357) |
| 1.3 | 1.3 | GO:0005796 | Golgi lumen(GO:0005796) |
| 0.7 | 9.1 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.6 | 5.1 | GO:0034363 | intermediate-density lipoprotein particle(GO:0034363) |
| 0.6 | 21.7 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.5 | 15.5 | GO:0042599 | lamellar body(GO:0042599) |
| 0.5 | 1.4 | GO:0002139 | stereocilia coupling link(GO:0002139) |
| 0.4 | 5.9 | GO:0030478 | actin cap(GO:0030478) |
| 0.4 | 2.7 | GO:0035867 | alphav-beta3 integrin-IGF-1-IGF1R complex(GO:0035867) |
| 0.4 | 3.0 | GO:0044279 | other organism cell membrane(GO:0044218) other organism membrane(GO:0044279) |
| 0.3 | 5.8 | GO:0042627 | chylomicron(GO:0042627) |
| 0.2 | 3.9 | GO:0042589 | zymogen granule membrane(GO:0042589) |
| 0.2 | 3.1 | GO:0098647 | collagen type VI trimer(GO:0005589) collagen beaded filament(GO:0098647) |
| 0.2 | 1.0 | GO:0030485 | smooth muscle contractile fiber(GO:0030485) |
| 0.2 | 2.6 | GO:0033180 | proton-transporting V-type ATPase, V1 domain(GO:0033180) |
| 0.1 | 2.4 | GO:0033093 | Weibel-Palade body(GO:0033093) |
| 0.1 | 16.7 | GO:0031093 | platelet alpha granule lumen(GO:0031093) |
| 0.1 | 13.1 | GO:0005902 | microvillus(GO:0005902) |
| 0.1 | 18.0 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.1 | 1.4 | GO:0034098 | VCP-NPL4-UFD1 AAA ATPase complex(GO:0034098) |
| 0.1 | 1.1 | GO:0016342 | catenin complex(GO:0016342) |
| 0.1 | 6.1 | GO:0045178 | basal part of cell(GO:0045178) |
| 0.1 | 1.5 | GO:0043205 | microfibril(GO:0001527) fibril(GO:0043205) |
| 0.1 | 4.3 | GO:0031907 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.1 | 6.6 | GO:0005791 | rough endoplasmic reticulum(GO:0005791) |
| 0.1 | 1.8 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.1 | 6.9 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.1 | 1.1 | GO:0097225 | sperm midpiece(GO:0097225) |
| 0.0 | 1.6 | GO:0034358 | plasma lipoprotein particle(GO:0034358) lipoprotein particle(GO:1990777) |
| 0.0 | 0.9 | GO:0030663 | COPI-coated vesicle membrane(GO:0030663) |
| 0.0 | 3.1 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.0 | 0.6 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.0 | 8.8 | GO:0090575 | RNA polymerase II transcription factor complex(GO:0090575) |
| 0.0 | 16.8 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.0 | 1.0 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.0 | 0.4 | GO:0000506 | glycosylphosphatidylinositol-N-acetylglucosaminyltransferase (GPI-GnT) complex(GO:0000506) |
| 0.0 | 2.2 | GO:0031526 | brush border membrane(GO:0031526) |
| 0.0 | 4.3 | GO:0005923 | bicellular tight junction(GO:0005923) |
| 0.0 | 4.5 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.0 | 37.6 | GO:0005615 | extracellular space(GO:0005615) |
| 0.0 | 0.7 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.0 | 2.6 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.0 | 0.5 | GO:0031095 | platelet dense tubular network membrane(GO:0031095) |
| 0.0 | 0.5 | GO:0005922 | connexon complex(GO:0005922) |
| 0.0 | 1.6 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.0 | 4.4 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.0 | 0.6 | GO:0005801 | cis-Golgi network(GO:0005801) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.1 | 6.2 | GO:1902271 | D3 vitamins binding(GO:1902271) |
| 1.4 | 4.3 | GO:0052854 | (S)-2-hydroxy-acid oxidase activity(GO:0003973) very-long-chain-(S)-2-hydroxy-acid oxidase activity(GO:0052852) long-chain-(S)-2-hydroxy-long-chain-acid oxidase activity(GO:0052853) medium-chain-(S)-2-hydroxy-acid oxidase activity(GO:0052854) |
| 1.4 | 10.9 | GO:0035473 | lipase binding(GO:0035473) |
| 1.2 | 4.8 | GO:0005018 | platelet-derived growth factor alpha-receptor activity(GO:0005018) |
| 1.0 | 2.9 | GO:0001596 | angiotensin type I receptor activity(GO:0001596) |
| 0.9 | 2.6 | GO:1904854 | proteasome core complex binding(GO:1904854) |
| 0.6 | 1.9 | GO:0003870 | 5-aminolevulinate synthase activity(GO:0003870) N-succinyltransferase activity(GO:0016749) |
| 0.5 | 6.5 | GO:0016724 | ferroxidase activity(GO:0004322) oxidoreductase activity, oxidizing metal ions, oxygen as acceptor(GO:0016724) |
| 0.5 | 5.5 | GO:0019834 | phospholipase A2 inhibitor activity(GO:0019834) |
| 0.5 | 1.9 | GO:0033823 | procollagen-lysine 5-dioxygenase activity(GO:0008475) procollagen glucosyltransferase activity(GO:0033823) |
| 0.5 | 1.4 | GO:0047225 | acetylgalactosaminyl-O-glycosyl-glycoprotein beta-1,6-N-acetylglucosaminyltransferase activity(GO:0047225) |
| 0.5 | 1.4 | GO:0070320 | inward rectifier potassium channel inhibitor activity(GO:0070320) |
| 0.4 | 2.6 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.3 | 10.1 | GO:0030247 | pattern binding(GO:0001871) polysaccharide binding(GO:0030247) |
| 0.3 | 1.0 | GO:0080101 | phosphatidyl-N-methylethanolamine N-methyltransferase activity(GO:0000773) phosphatidylethanolamine N-methyltransferase activity(GO:0004608) phosphatidyl-N-dimethylethanolamine N-methyltransferase activity(GO:0080101) |
| 0.3 | 18.5 | GO:0042056 | chemoattractant activity(GO:0042056) |
| 0.3 | 4.5 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 0.3 | 1.6 | GO:0008109 | N-acetyllactosaminide beta-1,6-N-acetylglucosaminyltransferase activity(GO:0008109) |
| 0.3 | 3.2 | GO:0047498 | calcium-dependent phospholipase A2 activity(GO:0047498) |
| 0.3 | 2.6 | GO:0003835 | beta-galactoside alpha-2,6-sialyltransferase activity(GO:0003835) |
| 0.2 | 0.9 | GO:0005046 | KDEL sequence binding(GO:0005046) |
| 0.2 | 3.8 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.2 | 4.5 | GO:0001161 | intronic transcription regulatory region sequence-specific DNA binding(GO:0001161) intronic transcription regulatory region DNA binding(GO:0044213) |
| 0.2 | 6.5 | GO:0070008 | serine-type exopeptidase activity(GO:0070008) |
| 0.2 | 3.8 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.2 | 6.5 | GO:0038187 | signaling pattern recognition receptor activity(GO:0008329) pattern recognition receptor activity(GO:0038187) |
| 0.2 | 0.7 | GO:0001855 | complement component C4b binding(GO:0001855) |
| 0.2 | 0.5 | GO:0050692 | DBD domain binding(GO:0050692) |
| 0.1 | 1.7 | GO:0009922 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.1 | 0.4 | GO:0010309 | acireductone dioxygenase [iron(II)-requiring] activity(GO:0010309) |
| 0.1 | 0.5 | GO:0000822 | inositol hexakisphosphate binding(GO:0000822) inositol 1,4,5-trisphosphate-sensitive calcium-release channel activity(GO:0005220) |
| 0.1 | 0.5 | GO:0004967 | glucagon receptor activity(GO:0004967) |
| 0.1 | 0.9 | GO:0005157 | macrophage colony-stimulating factor receptor binding(GO:0005157) |
| 0.1 | 18.4 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.1 | 2.2 | GO:0055106 | ubiquitin-protein transferase regulator activity(GO:0055106) |
| 0.1 | 3.5 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.1 | 0.7 | GO:0031748 | D1 dopamine receptor binding(GO:0031748) |
| 0.1 | 11.9 | GO:0098531 | RNA polymerase II transcription factor activity, ligand-activated sequence-specific DNA binding(GO:0004879) transcription factor activity, direct ligand regulated sequence-specific DNA binding(GO:0098531) |
| 0.1 | 1.9 | GO:0015197 | peptide transporter activity(GO:0015197) |
| 0.1 | 0.4 | GO:0035651 | AP-3 adaptor complex binding(GO:0035651) |
| 0.1 | 2.0 | GO:0030169 | low-density lipoprotein particle binding(GO:0030169) |
| 0.1 | 0.3 | GO:0031859 | platelet activating factor receptor binding(GO:0031859) |
| 0.1 | 0.8 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 0.1 | 10.5 | GO:0097110 | scaffold protein binding(GO:0097110) |
| 0.1 | 0.7 | GO:0004563 | beta-N-acetylhexosaminidase activity(GO:0004563) |
| 0.1 | 2.6 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) |
| 0.1 | 0.2 | GO:0045518 | interleukin-22 receptor binding(GO:0045518) |
| 0.1 | 0.3 | GO:0033754 | indoleamine 2,3-dioxygenase activity(GO:0033754) |
| 0.1 | 2.2 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.1 | 2.4 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.1 | 1.6 | GO:0034185 | apolipoprotein binding(GO:0034185) |
| 0.1 | 12.8 | GO:0019210 | kinase inhibitor activity(GO:0019210) |
| 0.1 | 8.3 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.1 | 2.7 | GO:0005159 | insulin-like growth factor receptor binding(GO:0005159) |
| 0.1 | 1.1 | GO:0019531 | oxalate transmembrane transporter activity(GO:0019531) |
| 0.1 | 1.6 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 0.0 | 2.5 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.0 | 0.3 | GO:0000179 | rRNA (adenine-N6,N6-)-dimethyltransferase activity(GO:0000179) |
| 0.0 | 1.1 | GO:0045295 | gamma-catenin binding(GO:0045295) |
| 0.0 | 1.0 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.0 | 0.4 | GO:0017176 | phosphatidylinositol N-acetylglucosaminyltransferase activity(GO:0017176) |
| 0.0 | 6.2 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 11.5 | GO:0030674 | protein binding, bridging(GO:0030674) |
| 0.0 | 2.7 | GO:0005160 | transforming growth factor beta receptor binding(GO:0005160) |
| 0.0 | 0.6 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.0 | 1.4 | GO:0005520 | insulin-like growth factor binding(GO:0005520) |
| 0.0 | 1.4 | GO:0048027 | mRNA 5'-UTR binding(GO:0048027) |
| 0.0 | 6.2 | GO:0005319 | lipid transporter activity(GO:0005319) |
| 0.0 | 0.8 | GO:0042974 | retinoic acid receptor binding(GO:0042974) |
| 0.0 | 3.2 | GO:0004714 | transmembrane receptor protein tyrosine kinase activity(GO:0004714) |
| 0.0 | 0.5 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.0 | 0.6 | GO:0070410 | co-SMAD binding(GO:0070410) |
| 0.0 | 0.1 | GO:0005174 | CD40 receptor binding(GO:0005174) |
| 0.0 | 2.2 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.0 | 0.2 | GO:0019788 | NEDD8 transferase activity(GO:0019788) |
| 0.0 | 1.0 | GO:0004435 | phosphatidylinositol phospholipase C activity(GO:0004435) |
| 0.0 | 1.4 | GO:0017022 | myosin binding(GO:0017022) |
| 0.0 | 0.7 | GO:0097200 | cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:0097200) |
| 0.0 | 0.2 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.0 | 1.5 | GO:0001102 | RNA polymerase II activating transcription factor binding(GO:0001102) |
| 0.0 | 19.6 | GO:0005198 | structural molecule activity(GO:0005198) |
| 0.0 | 0.5 | GO:0001530 | lipopolysaccharide binding(GO:0001530) |
| 0.0 | 4.8 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 2.5 | GO:0016820 | hydrolase activity, acting on acid anhydrides, catalyzing transmembrane movement of substances(GO:0016820) |
| 0.0 | 7.7 | GO:0001077 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
| 0.0 | 0.5 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.0 | 2.0 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.5 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.0 | 4.4 | GO:0030246 | carbohydrate binding(GO:0030246) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 44.7 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.3 | 25.3 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 0.2 | 17.5 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.2 | 4.0 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.2 | 11.2 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.1 | 6.2 | PID PDGFRA PATHWAY | PDGFR-alpha signaling pathway |
| 0.1 | 9.5 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 2.3 | PID LYMPH ANGIOGENESIS PATHWAY | VEGFR3 signaling in lymphatic endothelium |
| 0.1 | 8.9 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.1 | 9.9 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.1 | 4.4 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.1 | 5.3 | PID SYNDECAN 1 PATHWAY | Syndecan-1-mediated signaling events |
| 0.1 | 24.0 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.1 | 2.7 | PID IGF1 PATHWAY | IGF1 pathway |
| 0.1 | 2.9 | PID IL2 PI3K PATHWAY | IL2 signaling events mediated by PI3K |
| 0.1 | 4.7 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.0 | 1.5 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
| 0.0 | 1.4 | PID ECADHERIN KERATINOCYTE PATHWAY | E-cadherin signaling in keratinocytes |
| 0.0 | 0.9 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.0 | 0.3 | PID NFKAPPAB ATYPICAL PATHWAY | Atypical NF-kappaB pathway |
| 0.0 | 2.0 | PID BMP PATHWAY | BMP receptor signaling |
| 0.0 | 0.5 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.0 | 6.1 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 0.7 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 0.0 | 0.5 | SIG PIP3 SIGNALING IN B LYMPHOCYTES | Genes related to PIP3 signaling in B lymphocytes |
| 0.0 | 0.9 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
| 0.0 | 3.7 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 0.4 | PID WNT CANONICAL PATHWAY | Canonical Wnt signaling pathway |
| 0.0 | 0.7 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.0 | 0.4 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.0 | 0.4 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.0 | 0.6 | PID PTP1B PATHWAY | Signaling events mediated by PTP1B |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 9.1 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.4 | 11.4 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 0.4 | 13.5 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.4 | 17.7 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.4 | 20.9 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.4 | 10.9 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.2 | 5.1 | REACTOME KERATAN SULFATE DEGRADATION | Genes involved in Keratan sulfate degradation |
| 0.2 | 3.6 | REACTOME ROLE OF SECOND MESSENGERS IN NETRIN1 SIGNALING | Genes involved in Role of second messengers in netrin-1 signaling |
| 0.2 | 12.7 | REACTOME REGULATION OF BETA CELL DEVELOPMENT | Genes involved in Regulation of beta-cell development |
| 0.2 | 4.1 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.2 | 5.9 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.2 | 7.2 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.2 | 3.2 | REACTOME ACYL CHAIN REMODELLING OF PS | Genes involved in Acyl chain remodelling of PS |
| 0.1 | 9.1 | REACTOME IRON UPTAKE AND TRANSPORT | Genes involved in Iron uptake and transport |
| 0.1 | 2.4 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 0.1 | 6.3 | REACTOME STEROID HORMONES | Genes involved in Steroid hormones |
| 0.1 | 3.1 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.1 | 2.0 | REACTOME HYALURONAN UPTAKE AND DEGRADATION | Genes involved in Hyaluronan uptake and degradation |
| 0.1 | 1.6 | REACTOME LIPOPROTEIN METABOLISM | Genes involved in Lipoprotein metabolism |
| 0.1 | 2.3 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.1 | 7.1 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.1 | 13.1 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.1 | 1.9 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.1 | 2.5 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
| 0.1 | 2.6 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.1 | 0.5 | REACTOME ELEVATION OF CYTOSOLIC CA2 LEVELS | Genes involved in Elevation of cytosolic Ca2+ levels |
| 0.1 | 2.2 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.0 | 1.7 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.0 | 1.4 | REACTOME ENOS ACTIVATION AND REGULATION | Genes involved in eNOS activation and regulation |
| 0.0 | 0.4 | REACTOME ENDOGENOUS STEROLS | Genes involved in Endogenous sterols |
| 0.0 | 4.7 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.0 | 3.8 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 0.5 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.0 | 0.3 | REACTOME THROMBIN SIGNALLING THROUGH PROTEINASE ACTIVATED RECEPTORS PARS | Genes involved in Thrombin signalling through proteinase activated receptors (PARs) |
| 0.0 | 1.9 | REACTOME EXTRACELLULAR MATRIX ORGANIZATION | Genes involved in Extracellular matrix organization |
| 0.0 | 0.8 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.0 | 0.5 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.0 | 1.1 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.0 | 1.0 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.0 | 0.4 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 0.0 | 0.4 | REACTOME SYNTHESIS OF PIPS AT THE EARLY ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the early endosome membrane |
| 0.0 | 0.7 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.0 | 0.4 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.0 | 4.6 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.0 | 0.4 | REACTOME ADVANCED GLYCOSYLATION ENDPRODUCT RECEPTOR SIGNALING | Genes involved in Advanced glycosylation endproduct receptor signaling |
| 0.0 | 0.7 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.0 | 0.4 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.0 | 0.6 | REACTOME DOWNREGULATION OF SMAD2 3 SMAD4 TRANSCRIPTIONAL ACTIVITY | Genes involved in Downregulation of SMAD2/3:SMAD4 transcriptional activity |
| 0.0 | 0.3 | REACTOME GLUCAGON TYPE LIGAND RECEPTORS | Genes involved in Glucagon-type ligand receptors |
| 0.0 | 1.1 | REACTOME DOWNSTREAM SIGNAL TRANSDUCTION | Genes involved in Downstream signal transduction |
| 0.0 | 0.8 | REACTOME COSTIMULATION BY THE CD28 FAMILY | Genes involved in Costimulation by the CD28 family |
| 0.0 | 2.6 | REACTOME G ALPHA Q SIGNALLING EVENTS | Genes involved in G alpha (q) signalling events |