Project
Illumina Body Map 2: averaged replicates
Navigation
Downloads
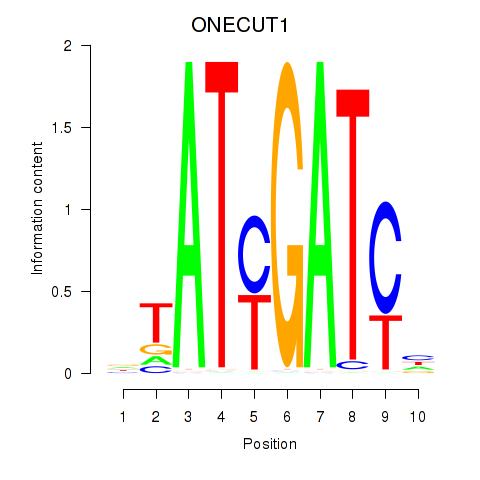
Results for ONECUT1
Z-value: 3.34
Transcription factors associated with ONECUT1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
ONECUT1
|
ENSG00000169856.7 | one cut homeobox 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| ONECUT1 | hg19_v2_chr15_-_53082178_53082209 | 0.45 | 1.1e-02 | Click! |
Activity profile of ONECUT1 motif
Sorted Z-values of ONECUT1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_+_57320437 | 18.62 |
ENST00000361249.3
|
C8A
|
complement component 8, alpha polypeptide |
| chr1_-_159684371 | 17.73 |
ENST00000255030.5
ENST00000437342.1 ENST00000368112.1 ENST00000368111.1 ENST00000368110.1 ENST00000343919.2 |
CRP
|
C-reactive protein, pentraxin-related |
| chr12_+_111471828 | 17.22 |
ENST00000261726.6
|
CUX2
|
cut-like homeobox 2 |
| chr1_+_159557607 | 17.11 |
ENST00000255040.2
|
APCS
|
amyloid P component, serum |
| chr13_-_46679144 | 15.96 |
ENST00000181383.4
|
CPB2
|
carboxypeptidase B2 (plasma) |
| chr9_-_116840728 | 15.41 |
ENST00000265132.3
|
AMBP
|
alpha-1-microglobulin/bikunin precursor |
| chr3_-_170744498 | 15.00 |
ENST00000382808.4
ENST00000314251.3 |
SLC2A2
|
solute carrier family 2 (facilitated glucose transporter), member 2 |
| chr13_-_46679185 | 14.52 |
ENST00000439329.3
|
CPB2
|
carboxypeptidase B2 (plasma) |
| chr17_-_26697304 | 13.88 |
ENST00000536498.1
|
VTN
|
vitronectin |
| chr3_+_186383741 | 13.70 |
ENST00000232003.4
|
HRG
|
histidine-rich glycoprotein |
| chr3_+_52811596 | 13.33 |
ENST00000542827.1
ENST00000273283.2 |
ITIH1
|
inter-alpha-trypsin inhibitor heavy chain 1 |
| chrX_+_70435044 | 10.93 |
ENST00000374029.1
ENST00000374022.3 ENST00000447581.1 |
GJB1
|
gap junction protein, beta 1, 32kDa |
| chr2_+_234959323 | 9.91 |
ENST00000373368.1
ENST00000168148.3 |
SPP2
|
secreted phosphoprotein 2, 24kDa |
| chr19_+_45449301 | 9.59 |
ENST00000591597.1
|
APOC2
|
apolipoprotein C-II |
| chr19_+_45449228 | 9.53 |
ENST00000252490.4
|
APOC2
|
apolipoprotein C-II |
| chr15_-_53097139 | 9.05 |
ENST00000560818.1
|
RP11-209K10.2
|
RP11-209K10.2 |
| chr17_-_64225508 | 8.95 |
ENST00000205948.6
|
APOH
|
apolipoprotein H (beta-2-glycoprotein I) |
| chr19_+_45449266 | 8.56 |
ENST00000592257.1
|
APOC2
|
apolipoprotein C-II |
| chr10_+_115312766 | 8.33 |
ENST00000351270.3
|
HABP2
|
hyaluronan binding protein 2 |
| chr16_+_27214802 | 8.29 |
ENST00000380948.2
ENST00000286096.4 |
KDM8
|
lysine (K)-specific demethylase 8 |
| chr1_-_9563433 | 8.28 |
ENST00000441033.1
|
RP13-392I16.1
|
RP13-392I16.1 |
| chr1_+_207262540 | 8.23 |
ENST00000452902.2
|
C4BPB
|
complement component 4 binding protein, beta |
| chr17_+_41052808 | 8.18 |
ENST00000592383.1
ENST00000253801.2 ENST00000585489.1 |
G6PC
|
glucose-6-phosphatase, catalytic subunit |
| chr1_+_207262578 | 8.14 |
ENST00000243611.5
ENST00000367076.3 |
C4BPB
|
complement component 4 binding protein, beta |
| chr12_+_56623827 | 8.09 |
ENST00000424625.1
ENST00000419753.1 ENST00000454355.2 ENST00000417965.1 ENST00000436633.1 |
SLC39A5
|
solute carrier family 39 (zinc transporter), member 5 |
| chr5_-_176836577 | 7.94 |
ENST00000253496.3
|
F12
|
coagulation factor XII (Hageman factor) |
| chr2_+_234959376 | 7.75 |
ENST00000425558.1
|
SPP2
|
secreted phosphoprotein 2, 24kDa |
| chr19_-_58864848 | 7.38 |
ENST00000263100.3
|
A1BG
|
alpha-1-B glycoprotein |
| chrX_+_122318006 | 7.29 |
ENST00000371266.1
ENST00000264357.5 |
GRIA3
|
glutamate receptor, ionotropic, AMPA 3 |
| chr12_+_56624436 | 7.06 |
ENST00000266980.4
ENST00000437277.1 |
SLC39A5
|
solute carrier family 39 (zinc transporter), member 5 |
| chr1_+_207262627 | 6.99 |
ENST00000391923.1
|
C4BPB
|
complement component 4 binding protein, beta |
| chr12_-_53297432 | 6.83 |
ENST00000546900.1
|
KRT8
|
keratin 8 |
| chr10_+_115312825 | 6.19 |
ENST00000537906.1
ENST00000541666.1 |
HABP2
|
hyaluronan binding protein 2 |
| chr11_+_63057412 | 6.07 |
ENST00000544661.1
|
SLC22A10
|
solute carrier family 22, member 10 |
| chr2_+_234600253 | 6.07 |
ENST00000373424.1
ENST00000441351.1 |
UGT1A6
|
UDP glucuronosyltransferase 1 family, polypeptide A6 |
| chr18_-_68004529 | 6.06 |
ENST00000578633.1
|
RP11-484N16.1
|
RP11-484N16.1 |
| chr20_-_22566089 | 6.02 |
ENST00000377115.4
|
FOXA2
|
forkhead box A2 |
| chr1_-_197036364 | 5.62 |
ENST00000367412.1
|
F13B
|
coagulation factor XIII, B polypeptide |
| chr1_-_151804314 | 5.49 |
ENST00000318247.6
|
RORC
|
RAR-related orphan receptor C |
| chr4_+_187187098 | 5.26 |
ENST00000403665.2
ENST00000264692.4 |
F11
|
coagulation factor XI |
| chr20_-_22559211 | 5.20 |
ENST00000564492.1
|
LINC00261
|
long intergenic non-protein coding RNA 261 |
| chr18_+_68002675 | 4.86 |
ENST00000584919.1
|
RP11-41O4.1
|
Uncharacterized protein |
| chr18_+_47087055 | 4.79 |
ENST00000577628.1
|
LIPG
|
lipase, endothelial |
| chr1_-_151804222 | 4.78 |
ENST00000392697.3
|
RORC
|
RAR-related orphan receptor C |
| chr13_-_114018400 | 4.72 |
ENST00000375430.4
ENST00000375431.4 |
GRTP1
|
growth hormone regulated TBC protein 1 |
| chrX_+_122318113 | 4.70 |
ENST00000371264.3
|
GRIA3
|
glutamate receptor, ionotropic, AMPA 3 |
| chr18_+_56113488 | 4.66 |
ENST00000590797.1
|
RP11-1151B14.3
|
RP11-1151B14.3 |
| chr2_+_223536428 | 4.60 |
ENST00000446656.3
|
MOGAT1
|
monoacylglycerol O-acyltransferase 1 |
| chr22_-_37505588 | 4.45 |
ENST00000406856.1
|
TMPRSS6
|
transmembrane protease, serine 6 |
| chr3_-_164914640 | 4.32 |
ENST00000241274.3
|
SLITRK3
|
SLIT and NTRK-like family, member 3 |
| chr22_-_37505449 | 4.27 |
ENST00000406725.1
|
TMPRSS6
|
transmembrane protease, serine 6 |
| chr2_+_108863651 | 4.19 |
ENST00000329106.2
ENST00000376700.1 |
SULT1C3
|
sulfotransferase family, cytosolic, 1C, member 3 |
| chr5_+_42756903 | 4.16 |
ENST00000361970.5
ENST00000388827.4 |
CCDC152
|
coiled-coil domain containing 152 |
| chr20_+_42984330 | 4.04 |
ENST00000316673.4
ENST00000609795.1 ENST00000457232.1 ENST00000609262.1 |
HNF4A
|
hepatocyte nuclear factor 4, alpha |
| chr10_-_94003003 | 3.91 |
ENST00000412050.4
|
CPEB3
|
cytoplasmic polyadenylation element binding protein 3 |
| chr14_+_74083548 | 3.82 |
ENST00000381139.1
|
ACOT6
|
acyl-CoA thioesterase 6 |
| chr10_-_5227096 | 3.81 |
ENST00000488756.1
ENST00000334314.3 |
AKR1CL1
|
aldo-keto reductase family 1, member C-like 1 |
| chr12_-_21487829 | 3.73 |
ENST00000445053.1
ENST00000452078.1 ENST00000458504.1 ENST00000422327.1 ENST00000421294.1 |
SLCO1A2
|
solute carrier organic anion transporter family, member 1A2 |
| chr7_-_96132835 | 3.65 |
ENST00000356686.1
|
C7orf76
|
chromosome 7 open reading frame 76 |
| chr18_+_47087390 | 3.45 |
ENST00000583083.1
|
LIPG
|
lipase, endothelial |
| chr2_-_85788652 | 3.37 |
ENST00000430215.3
|
GGCX
|
gamma-glutamyl carboxylase |
| chr3_-_24207039 | 3.25 |
ENST00000280696.5
|
THRB
|
thyroid hormone receptor, beta |
| chr17_-_17485731 | 3.22 |
ENST00000395783.1
|
PEMT
|
phosphatidylethanolamine N-methyltransferase |
| chr2_+_20650796 | 3.15 |
ENST00000448241.1
|
AC023137.2
|
AC023137.2 |
| chr17_+_53342311 | 3.15 |
ENST00000226067.5
|
HLF
|
hepatic leukemia factor |
| chrX_-_131623982 | 3.14 |
ENST00000370844.1
|
MBNL3
|
muscleblind-like splicing regulator 3 |
| chr3_+_159557637 | 3.12 |
ENST00000445224.2
|
SCHIP1
|
schwannomin interacting protein 1 |
| chr1_+_207262881 | 3.10 |
ENST00000451804.2
|
C4BPB
|
complement component 4 binding protein, beta |
| chr4_-_141348999 | 3.06 |
ENST00000325617.5
|
CLGN
|
calmegin |
| chr11_+_19799327 | 3.05 |
ENST00000540292.1
|
NAV2
|
neuron navigator 2 |
| chr7_-_121784285 | 2.99 |
ENST00000417368.2
|
AASS
|
aminoadipate-semialdehyde synthase |
| chr18_-_24765248 | 2.94 |
ENST00000580774.1
ENST00000284224.8 |
CHST9
|
carbohydrate (N-acetylgalactosamine 4-0) sulfotransferase 9 |
| chr1_-_177939348 | 2.89 |
ENST00000464631.2
|
SEC16B
|
SEC16 homolog B (S. cerevisiae) |
| chr1_-_8075693 | 2.86 |
ENST00000467067.1
|
ERRFI1
|
ERBB receptor feedback inhibitor 1 |
| chr4_-_175443484 | 2.83 |
ENST00000514584.1
ENST00000542498.1 ENST00000296521.7 ENST00000422112.2 ENST00000504433.1 |
HPGD
|
hydroxyprostaglandin dehydrogenase 15-(NAD) |
| chr4_+_159727222 | 2.77 |
ENST00000512986.1
|
FNIP2
|
folliculin interacting protein 2 |
| chr2_-_200715834 | 2.66 |
ENST00000420128.1
ENST00000416668.1 |
FTCDNL1
|
formiminotransferase cyclodeaminase N-terminal like |
| chr19_-_48016023 | 2.64 |
ENST00000598615.1
ENST00000597118.1 |
NAPA
|
N-ethylmaleimide-sensitive factor attachment protein, alpha |
| chr3_+_142315294 | 2.60 |
ENST00000464320.1
|
PLS1
|
plastin 1 |
| chr4_+_159727272 | 2.52 |
ENST00000379346.3
|
FNIP2
|
folliculin interacting protein 2 |
| chr2_-_197457335 | 2.50 |
ENST00000260983.3
|
HECW2
|
HECT, C2 and WW domain containing E3 ubiquitin protein ligase 2 |
| chr2_+_241375069 | 2.48 |
ENST00000264039.2
|
GPC1
|
glypican 1 |
| chr10_+_69869237 | 2.47 |
ENST00000373675.3
|
MYPN
|
myopalladin |
| chr12_+_21679220 | 2.46 |
ENST00000256969.2
|
C12orf39
|
chromosome 12 open reading frame 39 |
| chr11_+_57365150 | 2.44 |
ENST00000457869.1
ENST00000340687.6 ENST00000378323.4 ENST00000378324.2 ENST00000403558.1 |
SERPING1
|
serpin peptidase inhibitor, clade G (C1 inhibitor), member 1 |
| chr14_+_74058410 | 2.43 |
ENST00000326303.4
|
ACOT4
|
acyl-CoA thioesterase 4 |
| chr13_+_30002846 | 2.41 |
ENST00000542829.1
|
MTUS2
|
microtubule associated tumor suppressor candidate 2 |
| chr1_-_154167124 | 2.39 |
ENST00000515609.1
|
TPM3
|
tropomyosin 3 |
| chr3_+_142315225 | 2.39 |
ENST00000457734.2
ENST00000483373.1 ENST00000475296.1 ENST00000495744.1 ENST00000476044.1 ENST00000461644.1 |
PLS1
|
plastin 1 |
| chr8_-_121457608 | 2.25 |
ENST00000306185.3
|
MRPL13
|
mitochondrial ribosomal protein L13 |
| chr15_+_69854027 | 2.25 |
ENST00000498938.2
|
RP11-279F6.1
|
RP11-279F6.1 |
| chr1_-_209975494 | 2.22 |
ENST00000456314.1
|
IRF6
|
interferon regulatory factor 6 |
| chrX_-_33229636 | 2.18 |
ENST00000357033.4
|
DMD
|
dystrophin |
| chr10_+_28966271 | 2.17 |
ENST00000375533.3
|
BAMBI
|
BMP and activin membrane-bound inhibitor |
| chr15_-_86338100 | 2.16 |
ENST00000536947.1
|
KLHL25
|
kelch-like family member 25 |
| chr12_-_118490403 | 2.14 |
ENST00000535496.1
|
WSB2
|
WD repeat and SOCS box containing 2 |
| chr19_+_2785458 | 2.13 |
ENST00000307741.6
ENST00000585338.1 |
THOP1
|
thimet oligopeptidase 1 |
| chr7_+_99425633 | 2.06 |
ENST00000354829.2
ENST00000421837.2 ENST00000417625.1 ENST00000342499.4 ENST00000444905.1 ENST00000415413.1 ENST00000312017.5 ENST00000222382.5 |
CYP3A43
|
cytochrome P450, family 3, subfamily A, polypeptide 43 |
| chr7_-_121944491 | 2.06 |
ENST00000331178.4
ENST00000427185.2 ENST00000442488.2 |
FEZF1
|
FEZ family zinc finger 1 |
| chr3_+_148415571 | 2.02 |
ENST00000497524.1
ENST00000349243.3 ENST00000542281.1 ENST00000418473.2 ENST00000404754.2 |
AGTR1
|
angiotensin II receptor, type 1 |
| chr10_+_29135337 | 1.99 |
ENST00000375520.1
|
C10orf126
|
chromosome 10 open reading frame 126 |
| chr3_-_187694195 | 1.99 |
ENST00000446091.1
|
RP11-132N15.3
|
RP11-132N15.3 |
| chrX_+_10124977 | 1.98 |
ENST00000380833.4
|
CLCN4
|
chloride channel, voltage-sensitive 4 |
| chrM_+_8366 | 1.95 |
ENST00000361851.1
|
MT-ATP8
|
mitochondrially encoded ATP synthase 8 |
| chr18_-_31802056 | 1.93 |
ENST00000538587.1
|
NOL4
|
nucleolar protein 4 |
| chr4_+_185734773 | 1.90 |
ENST00000508020.1
|
RP11-701P16.2
|
Uncharacterized protein |
| chr4_-_17513604 | 1.86 |
ENST00000505710.1
|
QDPR
|
quinoid dihydropteridine reductase |
| chr12_-_56359802 | 1.82 |
ENST00000548803.1
ENST00000449260.2 ENST00000550447.1 ENST00000547137.1 ENST00000546543.1 ENST00000550464.1 |
PMEL
|
premelanosome protein |
| chr10_-_35104185 | 1.81 |
ENST00000374789.3
ENST00000374788.3 ENST00000346874.4 ENST00000374794.3 ENST00000350537.4 ENST00000374790.3 ENST00000374776.1 ENST00000374773.1 ENST00000545693.1 ENST00000545260.1 ENST00000340077.5 |
PARD3
|
par-3 family cell polarity regulator |
| chr2_-_86850949 | 1.81 |
ENST00000237455.4
|
RNF103
|
ring finger protein 103 |
| chr8_-_27468717 | 1.81 |
ENST00000520796.1
ENST00000520491.1 |
CLU
|
clusterin |
| chr13_-_30683005 | 1.81 |
ENST00000413591.1
ENST00000432770.1 |
LINC00365
|
long intergenic non-protein coding RNA 365 |
| chrX_-_131262048 | 1.80 |
ENST00000298542.4
|
FRMD7
|
FERM domain containing 7 |
| chr11_-_72385437 | 1.80 |
ENST00000418754.2
ENST00000542969.2 ENST00000334456.5 |
PDE2A
|
phosphodiesterase 2A, cGMP-stimulated |
| chr7_-_71912046 | 1.75 |
ENST00000395276.2
ENST00000431984.1 |
CALN1
|
calneuron 1 |
| chr1_+_10292308 | 1.71 |
ENST00000377081.1
|
KIF1B
|
kinesin family member 1B |
| chr16_-_73178346 | 1.71 |
ENST00000358463.2
|
C16orf47
|
chromosome 16 open reading frame 47 |
| chr3_+_32280159 | 1.69 |
ENST00000458535.2
ENST00000307526.3 |
CMTM8
|
CKLF-like MARVEL transmembrane domain containing 8 |
| chr15_-_86338134 | 1.68 |
ENST00000337975.5
|
KLHL25
|
kelch-like family member 25 |
| chr13_+_30002741 | 1.66 |
ENST00000380808.2
|
MTUS2
|
microtubule associated tumor suppressor candidate 2 |
| chr5_-_156362666 | 1.65 |
ENST00000406964.1
|
TIMD4
|
T-cell immunoglobulin and mucin domain containing 4 |
| chr8_+_22601 | 1.63 |
ENST00000522481.3
ENST00000518652.1 |
AC144568.2
|
Uncharacterized protein |
| chrM_+_8527 | 1.63 |
ENST00000361899.2
|
MT-ATP6
|
mitochondrially encoded ATP synthase 6 |
| chr12_-_118490217 | 1.61 |
ENST00000542304.1
|
WSB2
|
WD repeat and SOCS box containing 2 |
| chr3_+_40141502 | 1.60 |
ENST00000539167.1
|
MYRIP
|
myosin VIIA and Rab interacting protein |
| chr17_-_26733604 | 1.60 |
ENST00000584426.1
ENST00000584995.1 |
SLC46A1
|
solute carrier family 46 (folate transporter), member 1 |
| chr12_-_96336369 | 1.58 |
ENST00000546947.1
ENST00000546386.1 |
CCDC38
|
coiled-coil domain containing 38 |
| chr1_+_26496362 | 1.58 |
ENST00000374266.5
ENST00000270812.5 |
ZNF593
|
zinc finger protein 593 |
| chr9_-_75653627 | 1.58 |
ENST00000446946.1
|
ALDH1A1
|
aldehyde dehydrogenase 1 family, member A1 |
| chr1_-_165667545 | 1.57 |
ENST00000538148.1
|
ALDH9A1
|
aldehyde dehydrogenase 9 family, member A1 |
| chr8_-_27468842 | 1.55 |
ENST00000523500.1
|
CLU
|
clusterin |
| chr22_-_36220420 | 1.54 |
ENST00000473487.2
|
RBFOX2
|
RNA binding protein, fox-1 homolog (C. elegans) 2 |
| chr16_-_18573396 | 1.53 |
ENST00000543392.1
ENST00000381474.3 ENST00000330537.6 |
NOMO2
|
NODAL modulator 2 |
| chr15_+_31196080 | 1.53 |
ENST00000561607.1
ENST00000565466.1 |
FAN1
|
FANCD2/FANCI-associated nuclease 1 |
| chr9_+_130830451 | 1.52 |
ENST00000373068.2
ENST00000373069.5 |
SLC25A25
|
solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 25 |
| chrM_-_14670 | 1.49 |
ENST00000361681.2
|
MT-ND6
|
mitochondrially encoded NADH dehydrogenase 6 |
| chrM_+_14741 | 1.36 |
ENST00000361789.2
|
MT-CYB
|
mitochondrially encoded cytochrome b |
| chr1_-_162346657 | 1.33 |
ENST00000367935.5
|
C1orf111
|
chromosome 1 open reading frame 111 |
| chr1_+_156589198 | 1.30 |
ENST00000456112.1
|
HAPLN2
|
hyaluronan and proteoglycan link protein 2 |
| chr18_+_74207477 | 1.30 |
ENST00000532511.1
|
RP11-17M16.1
|
uncharacterized protein LOC400658 |
| chr5_+_139175380 | 1.29 |
ENST00000274710.3
|
PSD2
|
pleckstrin and Sec7 domain containing 2 |
| chr7_+_97736197 | 1.25 |
ENST00000297293.5
|
LMTK2
|
lemur tyrosine kinase 2 |
| chr3_+_127770455 | 1.25 |
ENST00000464451.1
|
SEC61A1
|
Sec61 alpha 1 subunit (S. cerevisiae) |
| chr15_-_74421477 | 1.25 |
ENST00000514871.1
|
RP11-247C2.2
|
HCG2004779; Uncharacterized protein |
| chr21_+_17961006 | 1.21 |
ENST00000602323.1
|
LINC00478
|
long intergenic non-protein coding RNA 478 |
| chr10_+_51576285 | 1.20 |
ENST00000443446.1
|
NCOA4
|
nuclear receptor coactivator 4 |
| chr1_-_165414414 | 1.20 |
ENST00000359842.5
|
RXRG
|
retinoid X receptor, gamma |
| chr1_-_25747283 | 1.18 |
ENST00000346452.4
ENST00000340849.4 ENST00000349438.4 ENST00000294413.7 ENST00000413854.1 ENST00000455194.1 ENST00000243186.6 ENST00000425135.1 |
RHCE
|
Rh blood group, CcEe antigens |
| chr11_+_85359002 | 1.16 |
ENST00000528105.1
ENST00000304511.2 |
TMEM126A
|
transmembrane protein 126A |
| chr19_+_13135386 | 1.15 |
ENST00000360105.4
ENST00000588228.1 ENST00000591028.1 |
NFIX
|
nuclear factor I/X (CCAAT-binding transcription factor) |
| chr18_+_39739223 | 1.13 |
ENST00000601948.1
|
LINC00907
|
long intergenic non-protein coding RNA 907 |
| chr11_+_134123389 | 1.12 |
ENST00000281182.4
ENST00000537423.1 ENST00000543332.1 ENST00000374752.4 |
ACAD8
|
acyl-CoA dehydrogenase family, member 8 |
| chr16_+_16326352 | 1.11 |
ENST00000399336.4
ENST00000263012.6 ENST00000538468.1 |
NOMO3
|
NODAL modulator 3 |
| chr3_+_177545563 | 1.07 |
ENST00000434309.1
|
RP11-91K9.1
|
RP11-91K9.1 |
| chr11_+_34999328 | 1.07 |
ENST00000526309.1
|
PDHX
|
pyruvate dehydrogenase complex, component X |
| chr1_-_63782888 | 1.05 |
ENST00000436475.2
|
LINC00466
|
long intergenic non-protein coding RNA 466 |
| chr22_+_51176624 | 1.04 |
ENST00000216139.5
ENST00000529621.1 |
ACR
|
acrosin |
| chr1_-_94586651 | 1.04 |
ENST00000535735.1
ENST00000370225.3 |
ABCA4
|
ATP-binding cassette, sub-family A (ABC1), member 4 |
| chr5_+_59726565 | 1.04 |
ENST00000412930.2
|
FKSG52
|
FKSG52 |
| chr15_-_99789736 | 1.02 |
ENST00000560235.1
ENST00000394132.2 ENST00000560860.1 ENST00000558078.1 ENST00000394136.1 ENST00000262074.4 ENST00000558613.1 ENST00000394130.1 ENST00000560772.1 |
TTC23
|
tetratricopeptide repeat domain 23 |
| chr18_-_31802282 | 1.02 |
ENST00000535475.1
|
NOL4
|
nucleolar protein 4 |
| chr10_+_114133773 | 1.01 |
ENST00000354655.4
|
ACSL5
|
acyl-CoA synthetase long-chain family member 5 |
| chr1_+_113933581 | 0.99 |
ENST00000307546.9
ENST00000369615.1 ENST00000369611.4 |
MAGI3
|
membrane associated guanylate kinase, WW and PDZ domain containing 3 |
| chr4_-_83483395 | 0.99 |
ENST00000515780.2
|
TMEM150C
|
transmembrane protein 150C |
| chr21_-_27107344 | 0.97 |
ENST00000457143.2
|
ATP5J
|
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit F6 |
| chr4_-_83483360 | 0.97 |
ENST00000449862.2
|
TMEM150C
|
transmembrane protein 150C |
| chr11_-_67407031 | 0.97 |
ENST00000335385.3
|
TBX10
|
T-box 10 |
| chr11_+_5905501 | 0.96 |
ENST00000316987.2
|
OR52E4
|
olfactory receptor, family 52, subfamily E, member 4 |
| chr19_-_6501778 | 0.95 |
ENST00000596291.1
|
TUBB4A
|
tubulin, beta 4A class IVa |
| chr13_+_111267866 | 0.93 |
ENST00000458711.2
ENST00000424185.2 ENST00000397191.4 ENST00000309957.2 |
CARKD
|
carbohydrate kinase domain containing |
| chr5_+_126984710 | 0.93 |
ENST00000379445.3
|
CTXN3
|
cortexin 3 |
| chr22_-_26961328 | 0.91 |
ENST00000398110.2
|
TPST2
|
tyrosylprotein sulfotransferase 2 |
| chr1_-_165668100 | 0.89 |
ENST00000354775.4
|
ALDH9A1
|
aldehyde dehydrogenase 9 family, member A1 |
| chr12_-_106477805 | 0.88 |
ENST00000553094.1
ENST00000549704.1 |
NUAK1
|
NUAK family, SNF1-like kinase, 1 |
| chr3_+_98699880 | 0.88 |
ENST00000473756.1
|
LINC00973
|
long intergenic non-protein coding RNA 973 |
| chr1_+_113933371 | 0.88 |
ENST00000369617.4
|
MAGI3
|
membrane associated guanylate kinase, WW and PDZ domain containing 3 |
| chr5_-_151784838 | 0.87 |
ENST00000255262.3
|
NMUR2
|
neuromedin U receptor 2 |
| chr21_-_27107198 | 0.86 |
ENST00000400094.1
|
ATP5J
|
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit F6 |
| chr1_+_50569575 | 0.85 |
ENST00000371827.1
|
ELAVL4
|
ELAV like neuron-specific RNA binding protein 4 |
| chr7_-_6523688 | 0.84 |
ENST00000490996.1
|
KDELR2
|
KDEL (Lys-Asp-Glu-Leu) endoplasmic reticulum protein retention receptor 2 |
| chr14_+_24407940 | 0.84 |
ENST00000354854.1
|
DHRS4-AS1
|
DHRS4-AS1 |
| chr19_+_11200038 | 0.84 |
ENST00000558518.1
ENST00000557933.1 ENST00000455727.2 ENST00000535915.1 ENST00000545707.1 ENST00000558013.1 |
LDLR
|
low density lipoprotein receptor |
| chr21_-_27107283 | 0.83 |
ENST00000284971.3
ENST00000400099.1 |
ATP5J
|
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit F6 |
| chr1_-_116383322 | 0.83 |
ENST00000429731.1
|
NHLH2
|
nescient helix loop helix 2 |
| chr10_+_70980051 | 0.82 |
ENST00000354624.5
ENST00000395086.2 |
HKDC1
|
hexokinase domain containing 1 |
| chr4_+_130692778 | 0.81 |
ENST00000513875.1
ENST00000508724.1 |
RP11-422J15.1
|
RP11-422J15.1 |
| chr3_+_101546827 | 0.81 |
ENST00000461724.1
ENST00000483180.1 ENST00000394054.2 |
NFKBIZ
|
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, zeta |
| chr13_+_52598827 | 0.80 |
ENST00000521776.2
|
UTP14C
|
UTP14, U3 small nucleolar ribonucleoprotein, homolog C (yeast) |
| chr12_-_10978957 | 0.79 |
ENST00000240619.2
|
TAS2R10
|
taste receptor, type 2, member 10 |
| chr1_-_202679535 | 0.79 |
ENST00000367268.4
|
SYT2
|
synaptotagmin II |
| chr11_-_126870683 | 0.79 |
ENST00000525704.2
|
KIRREL3
|
kin of IRRE like 3 (Drosophila) |
| chr6_+_108616243 | 0.77 |
ENST00000421954.1
|
LACE1
|
lactation elevated 1 |
| chr19_+_37998031 | 0.76 |
ENST00000586138.1
ENST00000588578.1 ENST00000587986.1 |
ZNF793
|
zinc finger protein 793 |
| chr1_-_116383738 | 0.74 |
ENST00000320238.3
|
NHLH2
|
nescient helix loop helix 2 |
| chr3_-_46735155 | 0.73 |
ENST00000318962.4
|
ALS2CL
|
ALS2 C-terminal like |
| chr20_+_22034809 | 0.73 |
ENST00000449427.1
|
RP11-125P18.1
|
RP11-125P18.1 |
| chr9_+_140135665 | 0.72 |
ENST00000340384.4
|
TUBB4B
|
tubulin, beta 4B class IVb |
| chr19_-_49243845 | 0.72 |
ENST00000222145.4
|
RASIP1
|
Ras interacting protein 1 |
| chr15_+_31196053 | 0.70 |
ENST00000561594.1
ENST00000362065.4 |
FAN1
|
FANCD2/FANCI-associated nuclease 1 |
| chrX_-_30877837 | 0.69 |
ENST00000378930.3
|
TAB3
|
TGF-beta activated kinase 1/MAP3K7 binding protein 3 |
Network of associatons between targets according to the STRING database.
First level regulatory network of ONECUT1
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.6 | 30.5 | GO:0003330 | regulation of extracellular matrix constituent secretion(GO:0003330) positive regulation of extracellular matrix constituent secretion(GO:0003331) |
| 5.5 | 27.7 | GO:0010902 | positive regulation of very-low-density lipoprotein particle remodeling(GO:0010902) positive regulation of phospholipid catabolic process(GO:0060697) |
| 3.4 | 13.7 | GO:2001027 | negative regulation of endothelial cell chemotaxis(GO:2001027) |
| 3.0 | 15.2 | GO:0034224 | cellular response to zinc ion starvation(GO:0034224) |
| 3.0 | 15.0 | GO:0070837 | dehydroascorbic acid transport(GO:0070837) |
| 2.6 | 26.5 | GO:0045959 | regulation of complement activation, classical pathway(GO:0030450) negative regulation of complement activation, classical pathway(GO:0045959) |
| 2.6 | 7.9 | GO:0002541 | activation of plasma proteins involved in acute inflammatory response(GO:0002541) |
| 2.4 | 17.1 | GO:0052199 | negative regulation of catalytic activity in other organism involved in symbiotic interaction(GO:0052199) |
| 2.1 | 8.2 | GO:0010983 | positive regulation of high-density lipoprotein particle clearance(GO:0010983) |
| 1.8 | 5.3 | GO:0051919 | positive regulation of fibrinolysis(GO:0051919) |
| 1.5 | 6.0 | GO:0045014 | carbon catabolite repression of transcription(GO:0045013) negative regulation of transcription by glucose(GO:0045014) |
| 1.3 | 3.9 | GO:1900365 | positive regulation of mRNA polyadenylation(GO:1900365) |
| 1.0 | 17.7 | GO:0032930 | positive regulation of superoxide anion generation(GO:0032930) |
| 1.0 | 8.9 | GO:0051918 | negative regulation of fibrinolysis(GO:0051918) |
| 1.0 | 15.4 | GO:0046149 | porphyrin-containing compound catabolic process(GO:0006787) tetrapyrrole catabolic process(GO:0033015) heme catabolic process(GO:0042167) pigment catabolic process(GO:0046149) |
| 0.9 | 0.9 | GO:0043006 | activation of phospholipase A2 activity by calcium-mediated signaling(GO:0043006) |
| 0.9 | 6.1 | GO:0052697 | flavonoid glucuronidation(GO:0052696) xenobiotic glucuronidation(GO:0052697) |
| 0.8 | 18.6 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 0.8 | 2.4 | GO:0001868 | regulation of complement activation, lectin pathway(GO:0001868) negative regulation of complement activation, lectin pathway(GO:0001869) |
| 0.8 | 13.9 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.8 | 4.6 | GO:0006651 | diacylglycerol biosynthetic process(GO:0006651) |
| 0.7 | 5.0 | GO:1902896 | terminal web assembly(GO:1902896) |
| 0.7 | 17.2 | GO:0007614 | short-term memory(GO:0007614) |
| 0.7 | 8.2 | GO:0015760 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.6 | 3.2 | GO:0008050 | female courtship behavior(GO:0008050) |
| 0.6 | 2.9 | GO:0019605 | butyrate metabolic process(GO:0019605) |
| 0.5 | 12.4 | GO:0015868 | purine ribonucleotide transport(GO:0015868) |
| 0.5 | 2.0 | GO:0086097 | phospholipase C-activating angiotensin-activated signaling pathway(GO:0086097) |
| 0.5 | 3.0 | GO:0033512 | L-lysine catabolic process to acetyl-CoA via saccharopine(GO:0033512) |
| 0.5 | 1.9 | GO:0051066 | dihydrobiopterin metabolic process(GO:0051066) |
| 0.5 | 4.6 | GO:2000189 | positive regulation of cholesterol homeostasis(GO:2000189) |
| 0.5 | 1.4 | GO:0002121 | inter-male aggressive behavior(GO:0002121) |
| 0.4 | 8.3 | GO:0070544 | histone H3-K36 demethylation(GO:0070544) |
| 0.4 | 1.7 | GO:1904647 | response to rotenone(GO:1904647) |
| 0.4 | 3.4 | GO:1902998 | regulation of neuronal signal transduction(GO:1902847) positive regulation of neurofibrillary tangle assembly(GO:1902998) |
| 0.4 | 6.8 | GO:0051599 | response to hydrostatic pressure(GO:0051599) |
| 0.4 | 8.7 | GO:0097264 | self proteolysis(GO:0097264) |
| 0.4 | 6.1 | GO:0015747 | urate transport(GO:0015747) |
| 0.4 | 2.6 | GO:0010807 | regulation of synaptic vesicle priming(GO:0010807) |
| 0.4 | 2.5 | GO:0030200 | heparan sulfate proteoglycan catabolic process(GO:0030200) |
| 0.3 | 2.1 | GO:0021797 | forebrain anterior/posterior pattern specification(GO:0021797) |
| 0.3 | 3.2 | GO:0050747 | positive regulation of lipoprotein metabolic process(GO:0050747) |
| 0.3 | 10.3 | GO:0036315 | cellular response to sterol(GO:0036315) |
| 0.3 | 2.9 | GO:0010616 | negative regulation of cardiac muscle adaptation(GO:0010616) negative regulation of cardiac muscle hypertrophy in response to stress(GO:1903243) |
| 0.3 | 3.1 | GO:0003025 | regulation of systemic arterial blood pressure by baroreceptor feedback(GO:0003025) |
| 0.3 | 0.9 | GO:0006478 | peptidyl-tyrosine sulfation(GO:0006478) |
| 0.3 | 0.6 | GO:0032289 | central nervous system myelin formation(GO:0032289) |
| 0.3 | 2.8 | GO:2001300 | lipoxin metabolic process(GO:2001300) |
| 0.3 | 1.7 | GO:0008065 | establishment of blood-nerve barrier(GO:0008065) |
| 0.3 | 0.8 | GO:0090118 | receptor-mediated endocytosis of low-density lipoprotein particle involved in cholesterol transport(GO:0090118) |
| 0.3 | 1.6 | GO:0051958 | methotrexate transport(GO:0051958) |
| 0.3 | 1.6 | GO:0061624 | fructose catabolic process(GO:0006001) fructose catabolic process to hydroxyacetone phosphate and glyceraldehyde-3-phosphate(GO:0061624) |
| 0.2 | 1.6 | GO:0033159 | negative regulation of protein import into nucleus, translocation(GO:0033159) |
| 0.2 | 0.7 | GO:0002188 | translation reinitiation(GO:0002188) |
| 0.2 | 1.2 | GO:0039019 | pronephric nephron development(GO:0039019) |
| 0.2 | 1.6 | GO:1903026 | negative regulation of RNA polymerase II regulatory region sequence-specific DNA binding(GO:1903026) |
| 0.2 | 0.6 | GO:0071418 | cellular response to amine stimulus(GO:0071418) |
| 0.2 | 2.5 | GO:0045329 | carnitine biosynthetic process(GO:0045329) |
| 0.2 | 3.4 | GO:0018214 | peptidyl-glutamic acid carboxylation(GO:0017187) protein carboxylation(GO:0018214) |
| 0.2 | 1.1 | GO:0006574 | valine catabolic process(GO:0006574) |
| 0.2 | 6.4 | GO:0042776 | mitochondrial ATP synthesis coupled proton transport(GO:0042776) |
| 0.2 | 12.1 | GO:0035235 | ionotropic glutamate receptor signaling pathway(GO:0035235) |
| 0.2 | 1.5 | GO:0010724 | regulation of definitive erythrocyte differentiation(GO:0010724) |
| 0.1 | 11.2 | GO:0030212 | hyaluronan metabolic process(GO:0030212) |
| 0.1 | 1.9 | GO:0014809 | regulation of skeletal muscle contraction by regulation of release of sequestered calcium ion(GO:0014809) |
| 0.1 | 2.2 | GO:0035413 | positive regulation of catenin import into nucleus(GO:0035413) |
| 0.1 | 2.9 | GO:0070863 | positive regulation of protein exit from endoplasmic reticulum(GO:0070863) |
| 0.1 | 1.9 | GO:0006122 | mitochondrial electron transport, ubiquinol to cytochrome c(GO:0006122) |
| 0.1 | 0.5 | GO:1903566 | positive regulation of protein localization to cilium(GO:1903566) |
| 0.1 | 16.9 | GO:0046849 | bone remodeling(GO:0046849) |
| 0.1 | 0.4 | GO:0071922 | establishment of sister chromatid cohesion(GO:0034085) cohesin loading(GO:0071921) regulation of cohesin loading(GO:0071922) |
| 0.1 | 1.7 | GO:0031580 | membrane raft polarization(GO:0001766) membrane raft distribution(GO:0031580) |
| 0.1 | 1.1 | GO:0061732 | mitochondrial acetyl-CoA biosynthetic process from pyruvate(GO:0061732) |
| 0.1 | 0.5 | GO:0070427 | nucleotide-binding oligomerization domain containing 1 signaling pathway(GO:0070427) |
| 0.1 | 2.5 | GO:0048739 | cardiac muscle fiber development(GO:0048739) |
| 0.1 | 1.6 | GO:0030050 | vesicle transport along actin filament(GO:0030050) |
| 0.1 | 3.7 | GO:0043252 | sodium-independent organic anion transport(GO:0043252) |
| 0.1 | 3.1 | GO:0045662 | negative regulation of myoblast differentiation(GO:0045662) |
| 0.1 | 0.9 | GO:0035507 | regulation of myosin-light-chain-phosphatase activity(GO:0035507) |
| 0.1 | 1.8 | GO:0031643 | positive regulation of myelination(GO:0031643) |
| 0.1 | 0.6 | GO:0086048 | membrane depolarization during bundle of His cell action potential(GO:0086048) |
| 0.1 | 1.6 | GO:0007617 | mating behavior(GO:0007617) multi-organism reproductive behavior(GO:0044705) |
| 0.1 | 0.8 | GO:0006621 | protein retention in ER lumen(GO:0006621) |
| 0.1 | 3.7 | GO:0007339 | binding of sperm to zona pellucida(GO:0007339) |
| 0.1 | 0.6 | GO:0031914 | negative regulation of synaptic plasticity(GO:0031914) |
| 0.1 | 2.2 | GO:0060644 | mammary gland epithelial cell differentiation(GO:0060644) |
| 0.1 | 1.2 | GO:0021554 | optic nerve development(GO:0021554) |
| 0.1 | 0.9 | GO:0060155 | platelet dense granule organization(GO:0060155) |
| 0.1 | 2.9 | GO:0030206 | chondroitin sulfate biosynthetic process(GO:0030206) |
| 0.1 | 1.8 | GO:1904380 | endoplasmic reticulum mannose trimming(GO:1904380) |
| 0.0 | 0.3 | GO:0043353 | enucleate erythrocyte differentiation(GO:0043353) |
| 0.0 | 0.3 | GO:0044806 | G-quadruplex DNA unwinding(GO:0044806) |
| 0.0 | 1.3 | GO:0032012 | regulation of ARF protein signal transduction(GO:0032012) |
| 0.0 | 1.8 | GO:0032438 | melanosome organization(GO:0032438) |
| 0.0 | 0.9 | GO:0010592 | positive regulation of lamellipodium assembly(GO:0010592) |
| 0.0 | 1.2 | GO:0006622 | protein targeting to lysosome(GO:0006622) |
| 0.0 | 0.4 | GO:0010839 | negative regulation of keratinocyte proliferation(GO:0010839) |
| 0.0 | 2.2 | GO:0033683 | nucleotide-excision repair, DNA incision(GO:0033683) |
| 0.0 | 0.4 | GO:0060012 | synaptic transmission, glycinergic(GO:0060012) |
| 0.0 | 1.0 | GO:0045332 | phospholipid translocation(GO:0045332) |
| 0.0 | 2.0 | GO:1903959 | regulation of anion transmembrane transport(GO:1903959) |
| 0.0 | 1.9 | GO:0046710 | GDP metabolic process(GO:0046710) |
| 0.0 | 0.6 | GO:0045792 | negative regulation of cell size(GO:0045792) |
| 0.0 | 6.6 | GO:0002576 | platelet degranulation(GO:0002576) |
| 0.0 | 0.9 | GO:0031115 | negative regulation of microtubule polymerization(GO:0031115) |
| 0.0 | 4.7 | GO:0090630 | activation of GTPase activity(GO:0090630) |
| 0.0 | 4.3 | GO:0006446 | regulation of translational initiation(GO:0006446) |
| 0.0 | 3.3 | GO:0051965 | positive regulation of synapse assembly(GO:0051965) |
| 0.0 | 0.3 | GO:0006553 | lysine metabolic process(GO:0006553) |
| 0.0 | 1.5 | GO:0031648 | protein destabilization(GO:0031648) |
| 0.0 | 4.7 | GO:0035383 | acyl-CoA metabolic process(GO:0006637) thioester metabolic process(GO:0035383) |
| 0.0 | 1.3 | GO:0033572 | transferrin transport(GO:0033572) |
| 0.0 | 1.1 | GO:1901522 | positive regulation of transcription from RNA polymerase II promoter involved in cellular response to chemical stimulus(GO:1901522) |
| 0.0 | 0.1 | GO:1902268 | negative regulation of polyamine transmembrane transport(GO:1902268) |
| 0.0 | 2.0 | GO:0033275 | muscle filament sliding(GO:0030049) actin-myosin filament sliding(GO:0033275) |
| 0.0 | 0.5 | GO:2001224 | positive regulation of neuron migration(GO:2001224) |
| 0.0 | 0.2 | GO:0003199 | endocardial cushion to mesenchymal transition involved in heart valve formation(GO:0003199) |
| 0.0 | 0.3 | GO:0048280 | vesicle fusion with Golgi apparatus(GO:0048280) |
| 0.0 | 1.0 | GO:0001580 | detection of chemical stimulus involved in sensory perception of bitter taste(GO:0001580) |
| 0.0 | 0.6 | GO:0005513 | detection of calcium ion(GO:0005513) |
| 0.0 | 1.5 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
| 0.0 | 0.8 | GO:0051156 | glucose 6-phosphate metabolic process(GO:0051156) |
| 0.0 | 1.9 | GO:0035914 | skeletal muscle cell differentiation(GO:0035914) |
| 0.0 | 0.5 | GO:0043457 | regulation of cellular respiration(GO:0043457) |
| 0.0 | 2.2 | GO:0070125 | mitochondrial translational elongation(GO:0070125) |
| 0.0 | 0.2 | GO:0071492 | cellular response to UV-A(GO:0071492) |
| 0.0 | 0.1 | GO:0042045 | epithelial fluid transport(GO:0042045) |
| 0.0 | 0.5 | GO:0061458 | reproductive system development(GO:0061458) |
| 0.0 | 0.8 | GO:0048488 | synaptic vesicle endocytosis(GO:0048488) |
| 0.0 | 0.7 | GO:0002755 | MyD88-dependent toll-like receptor signaling pathway(GO:0002755) |
| 0.0 | 0.7 | GO:0042572 | retinol metabolic process(GO:0042572) |
| 0.0 | 0.7 | GO:0043928 | exonucleolytic nuclear-transcribed mRNA catabolic process involved in deadenylation-dependent decay(GO:0043928) |
| 0.0 | 2.4 | GO:0008630 | intrinsic apoptotic signaling pathway in response to DNA damage(GO:0008630) |
| 0.0 | 6.3 | GO:0007596 | blood coagulation(GO:0007596) |
| 0.0 | 0.1 | GO:0015749 | hexose transport(GO:0008645) monosaccharide transport(GO:0015749) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.5 | 13.9 | GO:0071062 | rough endoplasmic reticulum lumen(GO:0048237) alphav-beta3 integrin-vitronectin complex(GO:0071062) |
| 3.1 | 27.7 | GO:0034363 | intermediate-density lipoprotein particle(GO:0034363) |
| 1.9 | 18.6 | GO:0005579 | membrane attack complex(GO:0005579) |
| 1.5 | 26.6 | GO:0031089 | platelet dense granule lumen(GO:0031089) |
| 0.6 | 5.0 | GO:1990357 | terminal web(GO:1990357) |
| 0.5 | 2.6 | GO:0070044 | synaptobrevin 2-SNAP-25-syntaxin-1a complex(GO:0070044) |
| 0.4 | 10.9 | GO:0005922 | connexon complex(GO:0005922) |
| 0.4 | 13.9 | GO:0036019 | endolysosome(GO:0036019) |
| 0.3 | 1.0 | GO:0043159 | acrosomal matrix(GO:0043159) |
| 0.3 | 6.2 | GO:0045263 | proton-transporting ATP synthase complex, coupling factor F(o)(GO:0045263) |
| 0.3 | 3.4 | GO:0034366 | spherical high-density lipoprotein particle(GO:0034366) |
| 0.2 | 12.0 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.2 | 51.8 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.2 | 2.8 | GO:0033269 | internode region of axon(GO:0033269) |
| 0.2 | 8.7 | GO:0090665 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.2 | 26.5 | GO:0044217 | other organism(GO:0044215) other organism cell(GO:0044216) other organism part(GO:0044217) |
| 0.1 | 1.9 | GO:0045275 | mitochondrial respiratory chain complex III(GO:0005750) respiratory chain complex III(GO:0045275) |
| 0.1 | 3.9 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.1 | 1.3 | GO:0045254 | pyruvate dehydrogenase complex(GO:0045254) |
| 0.1 | 2.4 | GO:0005862 | muscle thin filament tropomyosin(GO:0005862) |
| 0.1 | 0.6 | GO:0071817 | MMXD complex(GO:0071817) |
| 0.1 | 3.1 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.1 | 0.7 | GO:1990726 | Lsm1-7-Pat1 complex(GO:1990726) |
| 0.1 | 6.2 | GO:0031907 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.1 | 16.8 | GO:0005903 | brush border(GO:0005903) |
| 0.1 | 1.6 | GO:0031045 | dense core granule(GO:0031045) |
| 0.1 | 1.2 | GO:0044754 | autolysosome(GO:0044754) |
| 0.1 | 0.6 | GO:0044305 | calyx of Held(GO:0044305) |
| 0.1 | 0.8 | GO:0042584 | chromaffin granule membrane(GO:0042584) |
| 0.1 | 1.8 | GO:0032585 | multivesicular body membrane(GO:0032585) |
| 0.1 | 0.3 | GO:0017102 | methionyl glutamyl tRNA synthetase complex(GO:0017102) |
| 0.0 | 9.4 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.0 | 1.6 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.0 | 0.2 | GO:0038039 | G-protein coupled receptor heterodimeric complex(GO:0038039) |
| 0.0 | 3.8 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.0 | 2.7 | GO:1904115 | axon cytoplasm(GO:1904115) |
| 0.0 | 16.6 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.0 | 0.5 | GO:0033256 | I-kappaB/NF-kappaB complex(GO:0033256) |
| 0.0 | 0.6 | GO:1990454 | L-type voltage-gated calcium channel complex(GO:1990454) |
| 0.0 | 2.2 | GO:0045171 | intercellular bridge(GO:0045171) |
| 0.0 | 2.2 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.0 | 62.2 | GO:0005615 | extracellular space(GO:0005615) |
| 0.0 | 0.7 | GO:0044322 | endoplasmic reticulum quality control compartment(GO:0044322) |
| 0.0 | 0.2 | GO:0000275 | mitochondrial proton-transporting ATP synthase complex, catalytic core F(1)(GO:0000275) |
| 0.0 | 26.9 | GO:0000139 | Golgi membrane(GO:0000139) |
| 0.0 | 2.8 | GO:0012505 | endomembrane system(GO:0012505) |
| 0.0 | 0.8 | GO:0097381 | photoreceptor disc membrane(GO:0097381) |
| 0.0 | 0.7 | GO:0005795 | Golgi stack(GO:0005795) |
| 0.0 | 2.5 | GO:0072686 | mitotic spindle(GO:0072686) |
| 0.0 | 0.4 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.0 | 2.1 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) |
| 0.0 | 1.4 | GO:0043198 | dendritic shaft(GO:0043198) |
| 0.0 | 0.5 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.0 | 1.5 | GO:0070469 | respiratory chain(GO:0070469) |
| 0.0 | 1.3 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 0.5 | GO:0032040 | small-subunit processome(GO:0032040) |
| 0.0 | 0.3 | GO:0000974 | Prp19 complex(GO:0000974) |
| 0.0 | 1.8 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.0 | 1.7 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.0 | 2.4 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |
| 0.0 | 1.1 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.0 | 0.7 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.2 | 36.6 | GO:0060230 | lipoprotein lipase activator activity(GO:0060230) |
| 3.9 | 15.4 | GO:0019862 | IgA binding(GO:0019862) |
| 3.5 | 34.8 | GO:0001849 | complement component C1q binding(GO:0001849) |
| 3.0 | 15.0 | GO:0033300 | dehydroascorbic acid transporter activity(GO:0033300) |
| 2.0 | 8.2 | GO:0050309 | glucose-6-phosphatase activity(GO:0004346) sugar-terminal-phosphatase activity(GO:0050309) |
| 1.1 | 3.2 | GO:0080101 | phosphatidyl-N-methylethanolamine N-methyltransferase activity(GO:0000773) phosphatidylethanolamine N-methyltransferase activity(GO:0004608) phosphatidyl-N-dimethylethanolamine N-methyltransferase activity(GO:0080101) |
| 0.9 | 10.3 | GO:0008142 | oxysterol binding(GO:0008142) |
| 0.8 | 5.3 | GO:0042030 | ATPase inhibitor activity(GO:0042030) |
| 0.7 | 30.5 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.7 | 2.2 | GO:0070336 | flap-structured DNA binding(GO:0070336) |
| 0.7 | 2.8 | GO:0016404 | 15-hydroxyprostaglandin dehydrogenase (NAD+) activity(GO:0016404) |
| 0.7 | 2.0 | GO:0001596 | angiotensin type I receptor activity(GO:0001596) |
| 0.7 | 2.6 | GO:0005483 | soluble NSF attachment protein activity(GO:0005483) |
| 0.7 | 4.6 | GO:0004144 | diacylglycerol O-acyltransferase activity(GO:0004144) |
| 0.6 | 12.0 | GO:0004971 | AMPA glutamate receptor activity(GO:0004971) |
| 0.6 | 18.6 | GO:0001848 | complement binding(GO:0001848) |
| 0.6 | 1.9 | GO:0004155 | 6,7-dihydropteridine reductase activity(GO:0004155) |
| 0.5 | 1.6 | GO:0031177 | phosphopantetheine binding(GO:0031177) |
| 0.5 | 16.2 | GO:0043395 | heparan sulfate proteoglycan binding(GO:0043395) |
| 0.5 | 10.9 | GO:0005243 | gap junction channel activity(GO:0005243) |
| 0.5 | 13.4 | GO:0030247 | pattern binding(GO:0001871) polysaccharide binding(GO:0030247) |
| 0.4 | 8.3 | GO:0051864 | histone demethylase activity (H3-K36 specific)(GO:0051864) |
| 0.4 | 2.9 | GO:0047756 | chondroitin 4-sulfotransferase activity(GO:0047756) |
| 0.4 | 8.2 | GO:0008970 | phosphatidylcholine 1-acylhydrolase activity(GO:0008970) |
| 0.4 | 6.1 | GO:0015143 | urate transmembrane transporter activity(GO:0015143) salt transmembrane transporter activity(GO:1901702) |
| 0.3 | 11.3 | GO:0051787 | misfolded protein binding(GO:0051787) |
| 0.3 | 15.2 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.3 | 0.9 | GO:0008476 | protein-tyrosine sulfotransferase activity(GO:0008476) |
| 0.3 | 1.2 | GO:0004886 | 9-cis retinoic acid receptor activity(GO:0004886) |
| 0.3 | 1.6 | GO:0015350 | methotrexate transporter activity(GO:0015350) |
| 0.3 | 1.0 | GO:0004040 | amidase activity(GO:0004040) |
| 0.3 | 6.2 | GO:0047617 | acyl-CoA hydrolase activity(GO:0047617) |
| 0.2 | 0.7 | GO:0000253 | 3-keto sterol reductase activity(GO:0000253) |
| 0.2 | 1.8 | GO:0004118 | cGMP-stimulated cyclic-nucleotide phosphodiesterase activity(GO:0004118) |
| 0.2 | 2.5 | GO:0004028 | 3-chloroallyl aldehyde dehydrogenase activity(GO:0004028) |
| 0.2 | 3.2 | GO:0070324 | thyroid hormone binding(GO:0070324) |
| 0.2 | 0.8 | GO:0005046 | KDEL sequence binding(GO:0005046) |
| 0.2 | 0.6 | GO:0086057 | voltage-gated calcium channel activity involved in bundle of His cell action potential(GO:0086057) |
| 0.2 | 0.6 | GO:0004852 | uroporphyrinogen-III synthase activity(GO:0004852) |
| 0.2 | 2.2 | GO:0005114 | type II transforming growth factor beta receptor binding(GO:0005114) |
| 0.2 | 2.7 | GO:0005542 | folic acid binding(GO:0005542) |
| 0.2 | 1.6 | GO:0018479 | benzaldehyde dehydrogenase (NAD+) activity(GO:0018479) |
| 0.2 | 1.4 | GO:0016681 | ubiquinol-cytochrome-c reductase activity(GO:0008121) oxidoreductase activity, acting on diphenols and related substances as donors, cytochrome as acceptor(GO:0016681) |
| 0.2 | 3.9 | GO:0035613 | RNA stem-loop binding(GO:0035613) |
| 0.2 | 3.3 | GO:0070008 | serine-type exopeptidase activity(GO:0070008) |
| 0.1 | 4.2 | GO:0004062 | aryl sulfotransferase activity(GO:0004062) |
| 0.1 | 1.3 | GO:0034603 | pyruvate dehydrogenase activity(GO:0004738) pyruvate dehydrogenase [NAD(P)+] activity(GO:0034603) pyruvate dehydrogenase (NAD+) activity(GO:0034604) |
| 0.1 | 3.7 | GO:0015125 | bile acid transmembrane transporter activity(GO:0015125) |
| 0.1 | 6.1 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.1 | 0.8 | GO:0030229 | very-low-density lipoprotein particle receptor activity(GO:0030229) |
| 0.1 | 3.0 | GO:0016646 | oxidoreductase activity, acting on the CH-NH group of donors, NAD or NADP as acceptor(GO:0016646) |
| 0.1 | 1.5 | GO:0005347 | ATP transmembrane transporter activity(GO:0005347) ADP transmembrane transporter activity(GO:0015217) |
| 0.1 | 28.9 | GO:0004866 | endopeptidase inhibitor activity(GO:0004866) |
| 0.1 | 0.8 | GO:0004396 | glucokinase activity(GO:0004340) hexokinase activity(GO:0004396) fructokinase activity(GO:0008865) mannokinase activity(GO:0019158) |
| 0.1 | 1.7 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.1 | 1.9 | GO:0050998 | nitric-oxide synthase binding(GO:0050998) |
| 0.1 | 2.1 | GO:0070330 | aromatase activity(GO:0070330) |
| 0.1 | 17.8 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.1 | 8.5 | GO:0097110 | scaffold protein binding(GO:0097110) |
| 0.1 | 0.8 | GO:0043533 | inositol 1,3,4,5 tetrakisphosphate binding(GO:0043533) |
| 0.1 | 2.0 | GO:0005247 | voltage-gated chloride channel activity(GO:0005247) |
| 0.1 | 0.4 | GO:0016934 | extracellular-glycine-gated ion channel activity(GO:0016933) extracellular-glycine-gated chloride channel activity(GO:0016934) |
| 0.1 | 0.6 | GO:0043139 | 5'-3' DNA helicase activity(GO:0043139) |
| 0.1 | 19.3 | GO:0005539 | glycosaminoglycan binding(GO:0005539) |
| 0.1 | 0.5 | GO:0032810 | sterol response element binding(GO:0032810) |
| 0.1 | 10.2 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.1 | 2.7 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.1 | 0.3 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 0.0 | 4.0 | GO:0004879 | RNA polymerase II transcription factor activity, ligand-activated sequence-specific DNA binding(GO:0004879) transcription factor activity, direct ligand regulated sequence-specific DNA binding(GO:0098531) |
| 0.0 | 3.4 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.0 | 1.0 | GO:0033038 | bitter taste receptor activity(GO:0033038) |
| 0.0 | 1.1 | GO:0003995 | acyl-CoA dehydrogenase activity(GO:0003995) |
| 0.0 | 6.2 | GO:0015078 | hydrogen ion transmembrane transporter activity(GO:0015078) |
| 0.0 | 0.5 | GO:0004321 | fatty-acyl-CoA synthase activity(GO:0004321) |
| 0.0 | 1.9 | GO:0004385 | guanylate kinase activity(GO:0004385) |
| 0.0 | 1.0 | GO:0102391 | decanoate--CoA ligase activity(GO:0102391) |
| 0.0 | 0.9 | GO:0005229 | intracellular calcium activated chloride channel activity(GO:0005229) |
| 0.0 | 1.3 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.0 | 0.7 | GO:1904264 | ubiquitin protein ligase activity involved in ERAD pathway(GO:1904264) |
| 0.0 | 1.0 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.0 | 0.2 | GO:0004965 | G-protein coupled GABA receptor activity(GO:0004965) |
| 0.0 | 0.5 | GO:0043422 | protein kinase B binding(GO:0043422) |
| 0.0 | 1.8 | GO:0005547 | phosphatidylinositol-3,4,5-trisphosphate binding(GO:0005547) |
| 0.0 | 2.9 | GO:0017022 | myosin binding(GO:0017022) |
| 0.0 | 0.1 | GO:0008073 | ornithine decarboxylase inhibitor activity(GO:0008073) |
| 0.0 | 0.3 | GO:0097157 | pre-mRNA intronic binding(GO:0097157) |
| 0.0 | 1.2 | GO:0017091 | AU-rich element binding(GO:0017091) |
| 0.0 | 1.5 | GO:0050136 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.0 | 0.1 | GO:0017159 | pantetheine hydrolase activity(GO:0017159) |
| 0.0 | 7.5 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.0 | 0.9 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.0 | 0.7 | GO:0042288 | MHC class I protein binding(GO:0042288) |
| 0.0 | 1.2 | GO:0043022 | ribosome binding(GO:0043022) |
| 0.0 | 0.5 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 0.0 | 0.6 | GO:0019992 | diacylglycerol binding(GO:0019992) |
| 0.0 | 1.3 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.0 | 0.3 | GO:0016780 | phosphotransferase activity, for other substituted phosphate groups(GO:0016780) |
| 0.0 | 1.2 | GO:0050681 | androgen receptor binding(GO:0050681) |
| 0.0 | 0.5 | GO:0051059 | NF-kappaB binding(GO:0051059) |
| 0.0 | 3.3 | GO:0017137 | Rab GTPase binding(GO:0017137) |
| 0.0 | 1.1 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.0 | 0.1 | GO:0003828 | alpha-N-acetylneuraminate alpha-2,8-sialyltransferase activity(GO:0003828) |
| 0.0 | 0.4 | GO:0032041 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.0 | 0.2 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.0 | 0.3 | GO:0008301 | DNA binding, bending(GO:0008301) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 45.5 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.3 | 27.2 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.2 | 71.5 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.1 | 14.5 | PID IL6 7 PATHWAY | IL6-mediated signaling events |
| 0.1 | 7.4 | ST PHOSPHOINOSITIDE 3 KINASE PATHWAY | PI3K Pathway |
| 0.1 | 5.0 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.0 | 1.8 | SIG INSULIN RECEPTOR PATHWAY IN CARDIAC MYOCYTES | Genes related to the insulin receptor pathway |
| 0.0 | 7.0 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.0 | 2.5 | PID GLYPICAN 1PATHWAY | Glypican 1 network |
| 0.0 | 2.2 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.0 | 2.0 | PID ARF6 TRAFFICKING PATHWAY | Arf6 trafficking events |
| 0.0 | 2.2 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.0 | 0.5 | PID NFKAPPAB ATYPICAL PATHWAY | Atypical NF-kappaB pathway |
| 0.0 | 1.3 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 1.9 | PID LYSOPHOSPHOLIPID PATHWAY | LPA receptor mediated events |
| 0.0 | 1.0 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.0 | 0.4 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.0 | 0.6 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.3 | 17.7 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 1.0 | 45.1 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 1.0 | 27.7 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.7 | 15.6 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 0.6 | 25.1 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.4 | 15.2 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.4 | 10.9 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.3 | 3.4 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.3 | 5.6 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.3 | 12.0 | REACTOME UNBLOCKING OF NMDA RECEPTOR GLUTAMATE BINDING AND ACTIVATION | Genes involved in Unblocking of NMDA receptor, glutamate binding and activation |
| 0.2 | 3.7 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 0.2 | 6.1 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
| 0.1 | 15.2 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.1 | 12.3 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.1 | 8.2 | REACTOME GLUCOSE TRANSPORT | Genes involved in Glucose transport |
| 0.1 | 2.9 | REACTOME FORMATION OF ATP BY CHEMIOSMOTIC COUPLING | Genes involved in Formation of ATP by chemiosmotic coupling |
| 0.1 | 3.2 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.1 | 1.6 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.1 | 12.9 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.1 | 2.5 | REACTOME HS GAG DEGRADATION | Genes involved in HS-GAG degradation |
| 0.1 | 2.9 | REACTOME CHONDROITIN SULFATE BIOSYNTHESIS | Genes involved in Chondroitin sulfate biosynthesis |
| 0.1 | 10.2 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.1 | 0.7 | REACTOME IRAK2 MEDIATED ACTIVATION OF TAK1 COMPLEX UPON TLR7 8 OR 9 STIMULATION | Genes involved in IRAK2 mediated activation of TAK1 complex upon TLR7/8 or 9 stimulation |
| 0.1 | 1.8 | REACTOME TGF BETA RECEPTOR SIGNALING IN EMT EPITHELIAL TO MESENCHYMAL TRANSITION | Genes involved in TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) |
| 0.1 | 1.2 | REACTOME ER PHAGOSOME PATHWAY | Genes involved in ER-Phagosome pathway |
| 0.1 | 1.1 | REACTOME REGULATION OF PYRUVATE DEHYDROGENASE PDH COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |
| 0.1 | 3.9 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.0 | 2.4 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |
| 0.0 | 0.8 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.0 | 1.7 | REACTOME POST CHAPERONIN TUBULIN FOLDING PATHWAY | Genes involved in Post-chaperonin tubulin folding pathway |
| 0.0 | 0.5 | REACTOME NFKB IS ACTIVATED AND SIGNALS SURVIVAL | Genes involved in NF-kB is activated and signals survival |
| 0.0 | 1.1 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.0 | 0.7 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |
| 0.0 | 1.1 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.0 | 2.1 | REACTOME CYTOCHROME P450 ARRANGED BY SUBSTRATE TYPE | Genes involved in Cytochrome P450 - arranged by substrate type |
| 0.0 | 1.0 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.0 | 1.6 | REACTOME IRON UPTAKE AND TRANSPORT | Genes involved in Iron uptake and transport |
| 0.0 | 0.6 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.0 | 0.8 | REACTOME BOTULINUM NEUROTOXICITY | Genes involved in Botulinum neurotoxicity |
| 0.0 | 6.7 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.0 | 1.0 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.0 | 0.6 | REACTOME FORMATION OF INCISION COMPLEX IN GG NER | Genes involved in Formation of incision complex in GG-NER |
| 0.0 | 0.5 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.0 | 1.6 | REACTOME INTERFERON GAMMA SIGNALING | Genes involved in Interferon gamma signaling |
| 0.0 | 0.3 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.0 | 3.0 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |