Project
Illumina Body Map 2: averaged replicates
Navigation
Downloads
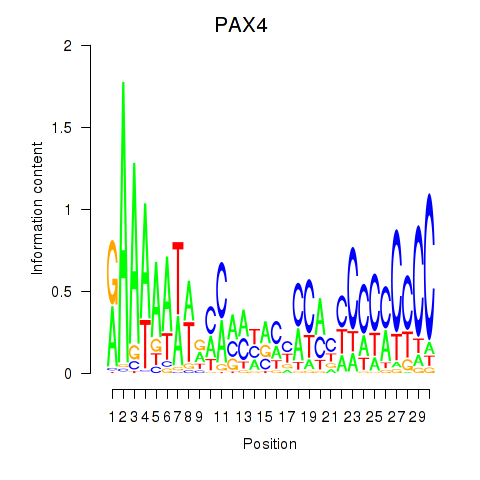
Results for PAX4
Z-value: 1.84
Transcription factors associated with PAX4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
PAX4
|
ENSG00000106331.10 | paired box 4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| PAX4 | hg19_v2_chr7_-_127255982_127255982 | -0.09 | 6.2e-01 | Click! |
Activity profile of PAX4 motif
Sorted Z-values of PAX4 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_+_26644441 | 6.90 |
ENST00000374213.2
|
CD52
|
CD52 molecule |
| chr1_-_153348067 | 5.65 |
ENST00000368737.3
|
S100A12
|
S100 calcium binding protein A12 |
| chr4_-_84030996 | 5.28 |
ENST00000411416.2
|
PLAC8
|
placenta-specific 8 |
| chr12_-_45269769 | 4.06 |
ENST00000548826.1
|
NELL2
|
NEL-like 2 (chicken) |
| chr5_+_147258266 | 4.03 |
ENST00000296694.4
|
SCGB3A2
|
secretoglobin, family 3A, member 2 |
| chr12_-_45269430 | 3.95 |
ENST00000395487.2
|
NELL2
|
NEL-like 2 (chicken) |
| chr1_-_25291475 | 3.81 |
ENST00000338888.3
ENST00000399916.1 |
RUNX3
|
runt-related transcription factor 3 |
| chr6_+_31553901 | 3.75 |
ENST00000418507.2
ENST00000438075.2 ENST00000376100.3 ENST00000376111.4 |
LST1
|
leukocyte specific transcript 1 |
| chr6_-_25042390 | 3.70 |
ENST00000606385.1
|
RP11-367G6.3
|
RP11-367G6.3 |
| chr12_-_45269251 | 3.67 |
ENST00000553120.1
|
NELL2
|
NEL-like 2 (chicken) |
| chr6_+_31553978 | 3.61 |
ENST00000376096.1
ENST00000376099.1 ENST00000376110.3 |
LST1
|
leukocyte specific transcript 1 |
| chr2_-_136875712 | 3.58 |
ENST00000241393.3
|
CXCR4
|
chemokine (C-X-C motif) receptor 4 |
| chr19_+_55105085 | 3.58 |
ENST00000251372.3
ENST00000453777.1 |
LILRA1
|
leukocyte immunoglobulin-like receptor, subfamily A (with TM domain), member 1 |
| chr7_+_26331541 | 3.57 |
ENST00000416246.1
ENST00000338523.4 ENST00000412416.1 |
SNX10
|
sorting nexin 10 |
| chr6_-_25042231 | 3.50 |
ENST00000510784.2
|
FAM65B
|
family with sequence similarity 65, member B |
| chr6_+_31554612 | 3.38 |
ENST00000211921.7
|
LST1
|
leukocyte specific transcript 1 |
| chr6_+_31554456 | 3.38 |
ENST00000339530.4
|
LST1
|
leukocyte specific transcript 1 |
| chr8_-_126963487 | 3.34 |
ENST00000518964.1
|
LINC00861
|
long intergenic non-protein coding RNA 861 |
| chr22_-_17700260 | 3.32 |
ENST00000399837.2
ENST00000543038.1 |
CECR1
|
cat eye syndrome chromosome region, candidate 1 |
| chr1_+_101702417 | 3.31 |
ENST00000305352.6
|
S1PR1
|
sphingosine-1-phosphate receptor 1 |
| chr7_+_26331678 | 3.26 |
ENST00000446848.2
|
SNX10
|
sorting nexin 10 |
| chr2_-_136873735 | 3.23 |
ENST00000409817.1
|
CXCR4
|
chemokine (C-X-C motif) receptor 4 |
| chr7_+_80267949 | 3.16 |
ENST00000482059.2
|
CD36
|
CD36 molecule (thrombospondin receptor) |
| chr7_+_80267973 | 3.13 |
ENST00000394788.3
ENST00000447544.2 |
CD36
|
CD36 molecule (thrombospondin receptor) |
| chr5_+_118668846 | 3.12 |
ENST00000513374.1
|
TNFAIP8
|
tumor necrosis factor, alpha-induced protein 8 |
| chr2_-_87088995 | 3.11 |
ENST00000393759.2
ENST00000349455.3 ENST00000331469.2 ENST00000431506.2 ENST00000393761.2 ENST00000390655.6 |
CD8B
|
CD8b molecule |
| chr22_+_23229960 | 3.10 |
ENST00000526893.1
ENST00000532223.2 ENST00000531372.1 |
IGLL5
|
immunoglobulin lambda-like polypeptide 5 |
| chr19_-_54784937 | 3.09 |
ENST00000434421.1
ENST00000314446.5 ENST00000391749.4 |
LILRB2
|
leukocyte immunoglobulin-like receptor, subfamily B (with TM and ITIM domains), member 2 |
| chr9_-_20622478 | 3.06 |
ENST00000355930.6
ENST00000380338.4 |
MLLT3
|
myeloid/lymphoid or mixed-lineage leukemia (trithorax homolog, Drosophila); translocated to, 3 |
| chr6_+_31554636 | 2.97 |
ENST00000433492.1
|
LST1
|
leukocyte specific transcript 1 |
| chr1_-_151032040 | 2.88 |
ENST00000540998.1
ENST00000357235.5 |
CDC42SE1
|
CDC42 small effector 1 |
| chr6_+_32407619 | 2.85 |
ENST00000395388.2
ENST00000374982.5 |
HLA-DRA
|
major histocompatibility complex, class II, DR alpha |
| chr11_-_124622134 | 2.78 |
ENST00000326621.5
|
VSIG2
|
V-set and immunoglobulin domain containing 2 |
| chr1_+_21880560 | 2.72 |
ENST00000425315.2
|
ALPL
|
alkaline phosphatase, liver/bone/kidney |
| chr19_+_55141861 | 2.64 |
ENST00000396327.3
ENST00000324602.7 ENST00000434867.2 |
LILRB1
|
leukocyte immunoglobulin-like receptor, subfamily B (with TM and ITIM domains), member 1 |
| chrX_-_49121165 | 2.63 |
ENST00000376207.4
ENST00000376199.2 |
FOXP3
|
forkhead box P3 |
| chr8_-_21771182 | 2.58 |
ENST00000523932.1
ENST00000544659.1 |
DOK2
|
docking protein 2, 56kDa |
| chr11_-_58980342 | 2.56 |
ENST00000361050.3
|
MPEG1
|
macrophage expressed 1 |
| chr16_-_28621353 | 2.43 |
ENST00000567512.1
|
SULT1A1
|
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 1 |
| chr19_+_10959043 | 2.41 |
ENST00000397820.4
|
C19orf38
|
chromosome 19 open reading frame 38 |
| chr3_+_27754397 | 2.38 |
ENST00000606069.1
|
RP11-222K16.2
|
RP11-222K16.2 |
| chr17_+_43299241 | 2.38 |
ENST00000328118.3
|
FMNL1
|
formin-like 1 |
| chr19_+_55141948 | 2.38 |
ENST00000396332.4
ENST00000427581.2 |
LILRB1
|
leukocyte immunoglobulin-like receptor, subfamily B (with TM and ITIM domains), member 1 |
| chr16_-_28621298 | 2.32 |
ENST00000566189.1
|
SULT1A1
|
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 1 |
| chr8_-_21771214 | 2.32 |
ENST00000276420.4
|
DOK2
|
docking protein 2, 56kDa |
| chr17_+_7253667 | 2.31 |
ENST00000570504.1
ENST00000574499.1 |
ACAP1
|
ArfGAP with coiled-coil, ankyrin repeat and PH domains 1 |
| chr7_-_23510086 | 2.28 |
ENST00000258729.3
|
IGF2BP3
|
insulin-like growth factor 2 mRNA binding protein 3 |
| chr17_+_7253635 | 2.28 |
ENST00000571471.1
|
ACAP1
|
ArfGAP with coiled-coil, ankyrin repeat and PH domains 1 |
| chrX_-_101410762 | 2.26 |
ENST00000543160.1
ENST00000333643.3 |
BEX5
|
brain expressed, X-linked 5 |
| chr12_-_121477039 | 2.26 |
ENST00000257570.5
|
OASL
|
2'-5'-oligoadenylate synthetase-like |
| chr20_-_52210368 | 2.22 |
ENST00000371471.2
|
ZNF217
|
zinc finger protein 217 |
| chr8_-_21771173 | 2.19 |
ENST00000518197.1
|
DOK2
|
docking protein 2, 56kDa |
| chr16_-_28621312 | 2.19 |
ENST00000314752.7
|
SULT1A1
|
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 1 |
| chrX_-_131623982 | 2.18 |
ENST00000370844.1
|
MBNL3
|
muscleblind-like splicing regulator 3 |
| chr11_-_124622083 | 2.03 |
ENST00000403470.1
|
VSIG2
|
V-set and immunoglobulin domain containing 2 |
| chrX_-_131623874 | 2.00 |
ENST00000436215.1
|
MBNL3
|
muscleblind-like splicing regulator 3 |
| chr18_-_3013307 | 1.99 |
ENST00000584294.1
|
LPIN2
|
lipin 2 |
| chr12_-_121476959 | 1.97 |
ENST00000339275.5
|
OASL
|
2'-5'-oligoadenylate synthetase-like |
| chr16_+_50730910 | 1.97 |
ENST00000300589.2
|
NOD2
|
nucleotide-binding oligomerization domain containing 2 |
| chr2_+_185463093 | 1.94 |
ENST00000302277.6
|
ZNF804A
|
zinc finger protein 804A |
| chr19_-_36391434 | 1.91 |
ENST00000396901.1
ENST00000585925.1 |
NFKBID
|
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, delta |
| chr14_-_50053081 | 1.90 |
ENST00000396020.3
ENST00000245458.6 |
RPS29
|
ribosomal protein S29 |
| chr9_-_20621834 | 1.89 |
ENST00000429426.2
|
MLLT3
|
myeloid/lymphoid or mixed-lineage leukemia (trithorax homolog, Drosophila); translocated to, 3 |
| chr13_+_28712614 | 1.88 |
ENST00000380958.3
|
PAN3
|
PAN3 poly(A) specific ribonuclease subunit homolog (S. cerevisiae) |
| chr3_-_44465475 | 1.80 |
ENST00000416124.1
|
LINC00694
|
long intergenic non-protein coding RNA 694 |
| chr6_+_32605134 | 1.75 |
ENST00000343139.5
ENST00000395363.1 ENST00000496318.1 |
HLA-DQA1
|
major histocompatibility complex, class II, DQ alpha 1 |
| chr19_-_55836697 | 1.73 |
ENST00000438693.1
ENST00000591570.1 |
TMEM150B
|
transmembrane protein 150B |
| chr6_+_15246501 | 1.67 |
ENST00000341776.2
|
JARID2
|
jumonji, AT rich interactive domain 2 |
| chr1_+_21877753 | 1.66 |
ENST00000374832.1
|
ALPL
|
alkaline phosphatase, liver/bone/kidney |
| chr11_+_1874200 | 1.64 |
ENST00000311604.3
|
LSP1
|
lymphocyte-specific protein 1 |
| chr19_-_55836669 | 1.61 |
ENST00000326652.4
|
TMEM150B
|
transmembrane protein 150B |
| chr8_-_101963482 | 1.59 |
ENST00000419477.2
|
YWHAZ
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta |
| chr3_-_71632894 | 1.59 |
ENST00000493089.1
|
FOXP1
|
forkhead box P1 |
| chr4_-_141074123 | 1.57 |
ENST00000502696.1
|
MAML3
|
mastermind-like 3 (Drosophila) |
| chr15_-_40401062 | 1.51 |
ENST00000354670.4
ENST00000559701.1 ENST00000557870.1 ENST00000558774.1 |
BMF
|
Bcl2 modifying factor |
| chr16_-_28608424 | 1.51 |
ENST00000335715.4
|
SULT1A2
|
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 2 |
| chr22_-_31688431 | 1.50 |
ENST00000402249.3
ENST00000443175.1 ENST00000215912.5 ENST00000441972.1 |
PIK3IP1
|
phosphoinositide-3-kinase interacting protein 1 |
| chr2_+_102972363 | 1.45 |
ENST00000409599.1
|
IL18R1
|
interleukin 18 receptor 1 |
| chr6_+_32605195 | 1.45 |
ENST00000374949.2
|
HLA-DQA1
|
major histocompatibility complex, class II, DQ alpha 1 |
| chr17_+_7254184 | 1.44 |
ENST00000575415.1
|
ACAP1
|
ArfGAP with coiled-coil, ankyrin repeat and PH domains 1 |
| chr22_-_31688381 | 1.43 |
ENST00000487265.2
|
PIK3IP1
|
phosphoinositide-3-kinase interacting protein 1 |
| chr9_+_134103496 | 1.42 |
ENST00000498010.1
ENST00000476004.1 ENST00000528406.1 |
NUP214
|
nucleoporin 214kDa |
| chr8_-_116680833 | 1.41 |
ENST00000220888.5
|
TRPS1
|
trichorhinophalangeal syndrome I |
| chr7_-_139763521 | 1.39 |
ENST00000263549.3
|
PARP12
|
poly (ADP-ribose) polymerase family, member 12 |
| chr1_+_12123414 | 1.36 |
ENST00000263932.2
|
TNFRSF8
|
tumor necrosis factor receptor superfamily, member 8 |
| chr6_+_133135580 | 1.35 |
ENST00000230050.3
|
RPS12
|
ribosomal protein S12 |
| chr7_+_26191809 | 1.34 |
ENST00000056233.3
|
NFE2L3
|
nuclear factor, erythroid 2-like 3 |
| chr6_-_90062543 | 1.34 |
ENST00000435041.2
|
UBE2J1
|
ubiquitin-conjugating enzyme E2, J1 |
| chr21_+_26934165 | 1.32 |
ENST00000456917.1
|
MIR155HG
|
MIR155 host gene (non-protein coding) |
| chr10_+_114135004 | 1.32 |
ENST00000393081.1
|
ACSL5
|
acyl-CoA synthetase long-chain family member 5 |
| chr8_-_623547 | 1.30 |
ENST00000522893.1
|
ERICH1
|
glutamate-rich 1 |
| chr8_-_101963677 | 1.29 |
ENST00000395956.3
ENST00000395953.2 |
YWHAZ
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta |
| chr7_+_69064566 | 1.28 |
ENST00000403018.2
|
AUTS2
|
autism susceptibility candidate 2 |
| chr17_+_72322346 | 1.27 |
ENST00000551294.1
ENST00000389916.4 |
KIF19
|
kinesin family member 19 |
| chr1_+_155146318 | 1.26 |
ENST00000368385.4
ENST00000545012.1 ENST00000392451.2 ENST00000368383.3 ENST00000368382.1 ENST00000334634.4 |
TRIM46
|
tripartite motif containing 46 |
| chr3_+_46283863 | 1.25 |
ENST00000545097.1
ENST00000541018.1 |
CCR3
|
chemokine (C-C motif) receptor 3 |
| chr1_+_161190160 | 1.25 |
ENST00000594609.1
|
AL590714.1
|
Uncharacterized protein |
| chr7_-_137686791 | 1.24 |
ENST00000452463.1
ENST00000330387.6 ENST00000456390.1 |
CREB3L2
|
cAMP responsive element binding protein 3-like 2 |
| chr16_+_69958887 | 1.24 |
ENST00000568684.1
|
WWP2
|
WW domain containing E3 ubiquitin protein ligase 2 |
| chr11_+_134144139 | 1.23 |
ENST00000389887.5
|
GLB1L3
|
galactosidase, beta 1-like 3 |
| chr4_-_113207048 | 1.22 |
ENST00000361717.3
|
TIFA
|
TRAF-interacting protein with forkhead-associated domain |
| chr19_-_10420459 | 1.19 |
ENST00000403352.1
ENST00000403903.3 |
ZGLP1
|
zinc finger, GATA-like protein 1 |
| chr18_-_53255379 | 1.17 |
ENST00000565908.2
|
TCF4
|
transcription factor 4 |
| chr11_-_93276582 | 1.17 |
ENST00000298966.2
|
SMCO4
|
single-pass membrane protein with coiled-coil domains 4 |
| chr7_-_38313235 | 1.13 |
ENST00000390338.2
|
TRGJP
|
T cell receptor gamma joining P |
| chr4_+_47487285 | 1.13 |
ENST00000273859.3
ENST00000504445.1 |
ATP10D
|
ATPase, class V, type 10D |
| chr1_+_12123463 | 1.11 |
ENST00000417814.2
|
TNFRSF8
|
tumor necrosis factor receptor superfamily, member 8 |
| chr15_-_60690932 | 1.08 |
ENST00000559818.1
|
ANXA2
|
annexin A2 |
| chr12_+_121570631 | 1.07 |
ENST00000546057.1
ENST00000377162.2 ENST00000328963.5 ENST00000535250.1 ENST00000541446.1 |
P2RX7
|
purinergic receptor P2X, ligand-gated ion channel, 7 |
| chr7_-_37488834 | 1.05 |
ENST00000310758.4
|
ELMO1
|
engulfment and cell motility 1 |
| chr2_+_99758161 | 1.05 |
ENST00000409684.1
|
C2ORF15
|
Uncharacterized protein C2orf15 |
| chr5_-_169407744 | 1.04 |
ENST00000377365.3
|
FAM196B
|
family with sequence similarity 196, member B |
| chr7_-_76955563 | 1.04 |
ENST00000441833.2
|
GSAP
|
gamma-secretase activating protein |
| chr18_-_53089723 | 1.04 |
ENST00000561992.1
ENST00000562512.2 |
TCF4
|
transcription factor 4 |
| chr11_+_20385327 | 1.03 |
ENST00000451739.2
ENST00000532505.1 |
HTATIP2
|
HIV-1 Tat interactive protein 2, 30kDa |
| chr7_+_69064300 | 1.03 |
ENST00000342771.4
|
AUTS2
|
autism susceptibility candidate 2 |
| chr9_-_123676827 | 1.02 |
ENST00000546084.1
|
TRAF1
|
TNF receptor-associated factor 1 |
| chr11_-_62368696 | 1.01 |
ENST00000527204.1
|
MTA2
|
metastasis associated 1 family, member 2 |
| chr1_-_85742773 | 1.01 |
ENST00000370580.1
|
BCL10
|
B-cell CLL/lymphoma 10 |
| chr7_-_37488777 | 1.00 |
ENST00000445322.1
ENST00000448602.1 |
ELMO1
|
engulfment and cell motility 1 |
| chrX_-_131547596 | 1.00 |
ENST00000538204.1
ENST00000370849.3 |
MBNL3
|
muscleblind-like splicing regulator 3 |
| chr18_-_60986613 | 0.99 |
ENST00000444484.1
|
BCL2
|
B-cell CLL/lymphoma 2 |
| chr11_+_46740730 | 0.99 |
ENST00000311907.5
ENST00000530231.1 ENST00000442468.1 |
F2
|
coagulation factor II (thrombin) |
| chr4_-_40516560 | 0.98 |
ENST00000513473.1
|
RBM47
|
RNA binding motif protein 47 |
| chr22_-_39151463 | 0.98 |
ENST00000405510.1
ENST00000433561.1 |
SUN2
|
Sad1 and UNC84 domain containing 2 |
| chr11_-_65374430 | 0.98 |
ENST00000532507.1
|
MAP3K11
|
mitogen-activated protein kinase kinase kinase 11 |
| chr17_-_77005801 | 0.95 |
ENST00000392446.5
|
CANT1
|
calcium activated nucleotidase 1 |
| chr2_+_12857043 | 0.92 |
ENST00000381465.2
|
TRIB2
|
tribbles pseudokinase 2 |
| chr6_-_32806483 | 0.92 |
ENST00000374899.4
|
TAP2
|
transporter 2, ATP-binding cassette, sub-family B (MDR/TAP) |
| chr1_-_152061537 | 0.92 |
ENST00000368806.1
|
TCHHL1
|
trichohyalin-like 1 |
| chr17_-_77005860 | 0.90 |
ENST00000591773.1
ENST00000588611.1 ENST00000586916.2 ENST00000592033.1 ENST00000588075.1 ENST00000302345.2 ENST00000591811.1 |
CANT1
|
calcium activated nucleotidase 1 |
| chr6_-_32095968 | 0.86 |
ENST00000375203.3
ENST00000375201.4 |
ATF6B
|
activating transcription factor 6 beta |
| chr2_-_191878162 | 0.86 |
ENST00000540176.1
|
STAT1
|
signal transducer and activator of transcription 1, 91kDa |
| chr3_+_46283916 | 0.86 |
ENST00000395940.2
|
CCR3
|
chemokine (C-C motif) receptor 3 |
| chr8_+_11666649 | 0.85 |
ENST00000528643.1
ENST00000525777.1 |
FDFT1
|
farnesyl-diphosphate farnesyltransferase 1 |
| chr17_+_72270380 | 0.83 |
ENST00000582036.1
ENST00000307504.5 |
DNAI2
|
dynein, axonemal, intermediate chain 2 |
| chr1_-_228594490 | 0.81 |
ENST00000366699.3
ENST00000284551.6 |
TRIM11
|
tripartite motif containing 11 |
| chr7_+_66386204 | 0.81 |
ENST00000341567.4
ENST00000607045.1 |
TMEM248
|
transmembrane protein 248 |
| chrX_-_131547625 | 0.80 |
ENST00000394311.2
|
MBNL3
|
muscleblind-like splicing regulator 3 |
| chr12_-_95945246 | 0.79 |
ENST00000258499.3
|
USP44
|
ubiquitin specific peptidase 44 |
| chr21_-_33651324 | 0.79 |
ENST00000290130.3
|
MIS18A
|
MIS18 kinetochore protein A |
| chr6_-_130031358 | 0.79 |
ENST00000368149.2
|
ARHGAP18
|
Rho GTPase activating protein 18 |
| chr3_+_107318157 | 0.78 |
ENST00000406780.1
|
BBX
|
bobby sox homolog (Drosophila) |
| chr18_+_72922710 | 0.78 |
ENST00000322038.5
|
TSHZ1
|
teashirt zinc finger homeobox 1 |
| chr11_+_20385231 | 0.77 |
ENST00000530266.1
ENST00000421577.2 ENST00000443524.2 ENST00000419348.2 |
HTATIP2
|
HIV-1 Tat interactive protein 2, 30kDa |
| chr12_-_109221160 | 0.77 |
ENST00000326470.5
|
SSH1
|
slingshot protein phosphatase 1 |
| chr16_-_28608364 | 0.77 |
ENST00000533150.1
|
SULT1A2
|
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 2 |
| chr2_-_21022818 | 0.76 |
ENST00000440866.2
ENST00000541941.1 ENST00000402479.2 ENST00000435420.2 ENST00000432947.1 ENST00000403006.2 ENST00000419825.2 ENST00000381090.3 ENST00000237822.3 ENST00000412261.1 |
C2orf43
|
chromosome 2 open reading frame 43 |
| chr5_-_54988448 | 0.76 |
ENST00000503817.1
ENST00000512595.1 |
SLC38A9
|
solute carrier family 38, member 9 |
| chr16_-_84273304 | 0.76 |
ENST00000308251.4
ENST00000568181.1 |
KCNG4
|
potassium voltage-gated channel, subfamily G, member 4 |
| chr17_-_77005813 | 0.76 |
ENST00000590370.1
ENST00000591625.1 |
CANT1
|
calcium activated nucleotidase 1 |
| chr3_-_197476560 | 0.75 |
ENST00000273582.5
|
KIAA0226
|
KIAA0226 |
| chr18_-_53255766 | 0.75 |
ENST00000566286.1
ENST00000564999.1 ENST00000566279.1 ENST00000354452.3 ENST00000356073.4 |
TCF4
|
transcription factor 4 |
| chr1_-_231376836 | 0.74 |
ENST00000451322.1
|
C1orf131
|
chromosome 1 open reading frame 131 |
| chr9_-_4859260 | 0.74 |
ENST00000599351.1
|
AL158147.2
|
HCG2011465; Uncharacterized protein |
| chr12_-_2986107 | 0.73 |
ENST00000359843.3
ENST00000342628.2 ENST00000361953.3 |
FOXM1
|
forkhead box M1 |
| chr2_-_208489275 | 0.72 |
ENST00000272839.3
ENST00000426075.1 |
METTL21A
|
methyltransferase like 21A |
| chr11_-_67211263 | 0.72 |
ENST00000393893.1
|
CORO1B
|
coronin, actin binding protein, 1B |
| chr8_-_144660771 | 0.72 |
ENST00000449291.2
|
NAPRT1
|
nicotinate phosphoribosyltransferase domain containing 1 |
| chr20_+_277737 | 0.72 |
ENST00000382352.3
|
ZCCHC3
|
zinc finger, CCHC domain containing 3 |
| chr14_-_50559361 | 0.71 |
ENST00000305273.1
|
C14orf183
|
chromosome 14 open reading frame 183 |
| chr6_-_32806506 | 0.71 |
ENST00000374897.2
ENST00000452392.2 |
TAP2
TAP2
|
transporter 2, ATP-binding cassette, sub-family B (MDR/TAP) Uncharacterized protein |
| chr4_+_109571800 | 0.70 |
ENST00000512478.2
|
OSTC
|
oligosaccharyltransferase complex subunit (non-catalytic) |
| chr12_-_99288536 | 0.70 |
ENST00000549797.1
ENST00000333732.7 ENST00000341752.7 |
ANKS1B
|
ankyrin repeat and sterile alpha motif domain containing 1B |
| chr1_-_231376867 | 0.70 |
ENST00000366649.2
ENST00000318906.2 ENST00000366651.3 |
C1orf131
|
chromosome 1 open reading frame 131 |
| chr14_+_104394770 | 0.69 |
ENST00000409874.4
ENST00000339063.5 |
TDRD9
|
tudor domain containing 9 |
| chr8_-_116681123 | 0.69 |
ENST00000519674.1
|
TRPS1
|
trichorhinophalangeal syndrome I |
| chr19_+_38779778 | 0.68 |
ENST00000590738.1
ENST00000587519.2 ENST00000591889.1 |
SPINT2
CTB-102L5.4
|
serine peptidase inhibitor, Kunitz type, 2 CTB-102L5.4 |
| chr3_-_142166904 | 0.68 |
ENST00000264951.4
|
XRN1
|
5'-3' exoribonuclease 1 |
| chr13_-_28194541 | 0.68 |
ENST00000316334.3
|
LNX2
|
ligand of numb-protein X 2 |
| chr1_-_151319283 | 0.68 |
ENST00000392746.3
|
RFX5
|
regulatory factor X, 5 (influences HLA class II expression) |
| chr14_+_76072096 | 0.68 |
ENST00000555058.1
|
FLVCR2
|
feline leukemia virus subgroup C cellular receptor family, member 2 |
| chr14_+_78227105 | 0.67 |
ENST00000439131.2
ENST00000355883.3 ENST00000557011.1 ENST00000556047.1 |
C14orf178
|
chromosome 14 open reading frame 178 |
| chr10_+_33271469 | 0.65 |
ENST00000414157.1
|
RP11-462L8.1
|
RP11-462L8.1 |
| chr11_-_62474803 | 0.64 |
ENST00000533982.1
ENST00000360796.5 |
BSCL2
|
Berardinelli-Seip congenital lipodystrophy 2 (seipin) |
| chr2_-_230787879 | 0.63 |
ENST00000435716.1
|
TRIP12
|
thyroid hormone receptor interactor 12 |
| chr1_-_1624083 | 0.63 |
ENST00000378662.1
ENST00000234800.6 |
SLC35E2B
|
solute carrier family 35, member E2B |
| chr18_+_3448455 | 0.62 |
ENST00000549780.1
|
TGIF1
|
TGFB-induced factor homeobox 1 |
| chr4_+_109571740 | 0.61 |
ENST00000361564.4
|
OSTC
|
oligosaccharyltransferase complex subunit (non-catalytic) |
| chr19_-_51308175 | 0.60 |
ENST00000345523.4
|
C19orf48
|
chromosome 19 open reading frame 48 |
| chr11_-_64545222 | 0.60 |
ENST00000433274.2
ENST00000432725.1 |
SF1
|
splicing factor 1 |
| chr10_-_27703269 | 0.59 |
ENST00000438700.3
|
PTCHD3
|
patched domain containing 3 |
| chr3_+_119298523 | 0.58 |
ENST00000357003.3
|
ADPRH
|
ADP-ribosylarginine hydrolase |
| chr17_+_73089382 | 0.57 |
ENST00000538213.2
ENST00000584118.1 |
SLC16A5
|
solute carrier family 16 (monocarboxylate transporter), member 5 |
| chr10_-_14920599 | 0.56 |
ENST00000609399.1
|
RP11-398C13.6
|
RP11-398C13.6 |
| chr5_+_150157444 | 0.55 |
ENST00000526627.1
|
SMIM3
|
small integral membrane protein 3 |
| chr21_+_43934633 | 0.54 |
ENST00000398343.2
|
SLC37A1
|
solute carrier family 37 (glucose-6-phosphate transporter), member 1 |
| chr5_-_130500922 | 0.53 |
ENST00000513012.1
ENST00000508488.1 ENST00000506908.1 ENST00000304043.5 |
HINT1
|
histidine triad nucleotide binding protein 1 |
| chr20_-_44993012 | 0.53 |
ENST00000372229.1
ENST00000372230.5 ENST00000543605.1 ENST00000243896.2 ENST00000317734.8 |
SLC35C2
|
solute carrier family 35 (GDP-fucose transporter), member C2 |
| chr18_-_60987220 | 0.52 |
ENST00000398117.1
|
BCL2
|
B-cell CLL/lymphoma 2 |
| chr1_-_155145721 | 0.52 |
ENST00000295682.4
|
KRTCAP2
|
keratinocyte associated protein 2 |
| chr18_+_29027696 | 0.51 |
ENST00000257189.4
|
DSG3
|
desmoglein 3 |
| chr10_-_70092671 | 0.51 |
ENST00000358769.2
ENST00000432941.1 ENST00000495025.2 |
PBLD
|
phenazine biosynthesis-like protein domain containing |
| chr4_+_95128996 | 0.50 |
ENST00000457823.2
|
SMARCAD1
|
SWI/SNF-related, matrix-associated actin-dependent regulator of chromatin, subfamily a, containing DEAD/H box 1 |
| chr4_-_176733897 | 0.48 |
ENST00000393658.2
|
GPM6A
|
glycoprotein M6A |
| chr18_-_5396271 | 0.47 |
ENST00000579951.1
|
EPB41L3
|
erythrocyte membrane protein band 4.1-like 3 |
| chr12_-_57146095 | 0.46 |
ENST00000550770.1
ENST00000338193.6 |
PRIM1
|
primase, DNA, polypeptide 1 (49kDa) |
| chr15_-_34502278 | 0.46 |
ENST00000559515.1
ENST00000256544.3 ENST00000560108.1 ENST00000559462.1 |
KATNBL1
|
katanin p80 subunit B-like 1 |
| chr6_-_113754604 | 0.45 |
ENST00000421737.1
|
RP1-124C6.1
|
RP1-124C6.1 |
| chr18_+_61254534 | 0.45 |
ENST00000269489.5
|
SERPINB13
|
serpin peptidase inhibitor, clade B (ovalbumin), member 13 |
Network of associatons between targets according to the STRING database.
First level regulatory network of PAX4
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.5 | 4.4 | GO:0071529 | cementum mineralization(GO:0071529) |
| 1.3 | 5.0 | GO:0002290 | gamma-delta T cell activation involved in immune response(GO:0002290) negative regulation of interferon-beta secretion(GO:0035548) regulation of gamma-delta T cell activation involved in immune response(GO:2001191) positive regulation of gamma-delta T cell activation involved in immune response(GO:2001193) |
| 1.1 | 3.3 | GO:0003245 | cardiac muscle tissue growth involved in heart morphogenesis(GO:0003245) |
| 0.9 | 2.6 | GO:0032831 | regulation of tolerance induction dependent upon immune response(GO:0002652) regulation of CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation(GO:0032829) positive regulation of CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation(GO:0032831) positive regulation of immature T cell proliferation in thymus(GO:0033092) |
| 0.8 | 2.5 | GO:0045553 | TRAIL biosynthetic process(GO:0045553) regulation of TRAIL biosynthetic process(GO:0045554) positive regulation of TRAIL biosynthetic process(GO:0045556) |
| 0.8 | 3.1 | GO:0002774 | Fc receptor mediated inhibitory signaling pathway(GO:0002774) |
| 0.8 | 6.8 | GO:0019064 | fusion of virus membrane with host plasma membrane(GO:0019064) membrane fusion involved in viral entry into host cell(GO:0039663) multi-organism membrane fusion(GO:0044800) |
| 0.7 | 2.9 | GO:0002503 | peptide antigen assembly with MHC class II protein complex(GO:0002503) |
| 0.6 | 11.7 | GO:0070050 | neuron cellular homeostasis(GO:0070050) |
| 0.6 | 2.9 | GO:0043553 | negative regulation of phosphatidylinositol 3-kinase activity(GO:0043553) |
| 0.6 | 2.3 | GO:0098582 | innate vocalization behavior(GO:0098582) |
| 0.5 | 1.6 | GO:0002476 | antigen processing and presentation of endogenous peptide antigen via MHC class Ib(GO:0002476) |
| 0.5 | 6.3 | GO:2000332 | blood microparticle formation(GO:0072564) regulation of blood microparticle formation(GO:2000332) positive regulation of blood microparticle formation(GO:2000334) |
| 0.5 | 2.0 | GO:0060585 | detection of peptidoglycan(GO:0032499) activation of MAPK activity involved in innate immune response(GO:0035419) regulation of prostaglandin-endoperoxide synthase activity(GO:0060584) positive regulation of prostaglandin-endoperoxide synthase activity(GO:0060585) positive regulation of protein K63-linked ubiquitination(GO:1902523) |
| 0.5 | 2.0 | GO:0046671 | negative regulation of cellular pH reduction(GO:0032848) CD8-positive, alpha-beta T cell lineage commitment(GO:0043375) negative regulation of retinal cell programmed cell death(GO:0046671) |
| 0.4 | 0.4 | GO:1901187 | regulation of ephrin receptor signaling pathway(GO:1901187) |
| 0.4 | 1.2 | GO:1902173 | negative regulation of keratinocyte apoptotic process(GO:1902173) |
| 0.4 | 1.1 | GO:0043132 | NAD transport(GO:0043132) |
| 0.3 | 1.0 | GO:0032765 | positive regulation of mast cell cytokine production(GO:0032765) |
| 0.3 | 9.2 | GO:0006068 | ethanol catabolic process(GO:0006068) |
| 0.3 | 0.9 | GO:0045081 | negative regulation of interleukin-10 biosynthetic process(GO:0045081) |
| 0.3 | 1.1 | GO:0052361 | multi-organism catabolic process(GO:0044035) development of symbiont involved in interaction with host(GO:0044115) modulation of development of symbiont involved in interaction with host(GO:0044145) negative regulation of development of symbiont involved in interaction with host(GO:0044147) metabolism of substance in other organism involved in symbiotic interaction(GO:0052214) catabolism of substance in other organism involved in symbiotic interaction(GO:0052227) metabolism of macromolecule in other organism involved in symbiotic interaction(GO:0052229) catabolism by host of symbiont macromolecule(GO:0052360) catabolism by organism of macromolecule in other organism involved in symbiotic interaction(GO:0052361) catabolism by host of symbiont protein(GO:0052362) catabolism by organism of protein in other organism involved in symbiotic interaction(GO:0052363) catabolism by host of substance in symbiont(GO:0052364) metabolism by host of symbiont macromolecule(GO:0052416) metabolism by host of symbiont protein(GO:0052417) metabolism by organism of protein in other organism involved in symbiotic interaction(GO:0052418) metabolism by host of substance in symbiont(GO:0052419) |
| 0.3 | 1.6 | GO:0032625 | interleukin-21 production(GO:0032625) interleukin-21 secretion(GO:0072619) |
| 0.3 | 0.8 | GO:0098976 | excitatory chemical synaptic transmission(GO:0098976) regulation of AMPA glutamate receptor clustering(GO:1904717) positive regulation of AMPA glutamate receptor clustering(GO:1904719) |
| 0.3 | 0.3 | GO:0021691 | cerebellar Purkinje cell layer maturation(GO:0021691) |
| 0.2 | 0.7 | GO:1902463 | protein localization to cell leading edge(GO:1902463) |
| 0.2 | 1.9 | GO:0070245 | positive regulation of thymocyte apoptotic process(GO:0070245) |
| 0.2 | 0.9 | GO:0060392 | negative regulation of SMAD protein import into nucleus(GO:0060392) |
| 0.2 | 5.3 | GO:0040015 | negative regulation of multicellular organism growth(GO:0040015) |
| 0.2 | 4.9 | GO:2000096 | positive regulation of Wnt signaling pathway, planar cell polarity pathway(GO:2000096) |
| 0.2 | 0.9 | GO:0046725 | negative regulation by virus of viral protein levels in host cell(GO:0046725) negative regulation of metanephric nephron tubule epithelial cell differentiation(GO:0072308) |
| 0.2 | 1.0 | GO:0071409 | cellular response to cycloheximide(GO:0071409) |
| 0.2 | 1.0 | GO:1900738 | cytolysis in other organism involved in symbiotic interaction(GO:0051801) positive regulation of phospholipase C-activating G-protein coupled receptor signaling pathway(GO:1900738) |
| 0.2 | 6.8 | GO:0071539 | protein localization to centrosome(GO:0071539) |
| 0.2 | 1.9 | GO:0010606 | positive regulation of cytoplasmic mRNA processing body assembly(GO:0010606) |
| 0.2 | 0.9 | GO:0008204 | ergosterol biosynthetic process(GO:0006696) ergosterol metabolic process(GO:0008204) |
| 0.2 | 6.9 | GO:0045730 | respiratory burst(GO:0045730) |
| 0.2 | 2.9 | GO:0090168 | Golgi reassembly(GO:0090168) |
| 0.2 | 0.8 | GO:0060023 | soft palate development(GO:0060023) |
| 0.1 | 17.4 | GO:0050672 | negative regulation of mononuclear cell proliferation(GO:0032945) negative regulation of lymphocyte proliferation(GO:0050672) |
| 0.1 | 6.0 | GO:0045662 | negative regulation of myoblast differentiation(GO:0045662) |
| 0.1 | 1.4 | GO:0035655 | interleukin-18-mediated signaling pathway(GO:0035655) cellular response to interleukin-18(GO:0071351) |
| 0.1 | 0.4 | GO:1901421 | positive regulation of response to alcohol(GO:1901421) |
| 0.1 | 3.8 | GO:0048935 | peripheral nervous system neuron differentiation(GO:0048934) peripheral nervous system neuron development(GO:0048935) |
| 0.1 | 5.8 | GO:0050832 | defense response to fungus(GO:0050832) |
| 0.1 | 1.6 | GO:0007221 | positive regulation of transcription of Notch receptor target(GO:0007221) |
| 0.1 | 1.3 | GO:0070862 | negative regulation of protein exit from endoplasmic reticulum(GO:0070862) negative regulation of retrograde protein transport, ER to cytosol(GO:1904153) |
| 0.1 | 1.0 | GO:0031022 | nuclear migration along microfilament(GO:0031022) |
| 0.1 | 0.4 | GO:0034721 | histone H3-K4 demethylation, trimethyl-H3-K4-specific(GO:0034721) |
| 0.1 | 1.7 | GO:0099612 | protein localization to axon(GO:0099612) |
| 0.1 | 0.7 | GO:1900383 | regulation of synaptic plasticity by receptor localization to synapse(GO:1900383) |
| 0.1 | 1.0 | GO:0007256 | activation of JNKK activity(GO:0007256) |
| 0.1 | 0.7 | GO:0060671 | epithelial cell differentiation involved in embryonic placenta development(GO:0060671) epithelial cell morphogenesis involved in placental branching(GO:0060672) |
| 0.1 | 1.7 | GO:0051574 | positive regulation of histone H3-K9 methylation(GO:0051574) |
| 0.1 | 0.8 | GO:0046598 | positive regulation of viral entry into host cell(GO:0046598) |
| 0.1 | 1.0 | GO:1902004 | positive regulation of beta-amyloid formation(GO:1902004) |
| 0.1 | 0.5 | GO:0006269 | DNA replication, synthesis of RNA primer(GO:0006269) |
| 0.1 | 1.0 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.1 | 5.2 | GO:0050690 | regulation of defense response to virus by virus(GO:0050690) |
| 0.1 | 0.6 | GO:0051725 | protein de-ADP-ribosylation(GO:0051725) |
| 0.1 | 0.9 | GO:1990440 | positive regulation of transcription from RNA polymerase II promoter in response to endoplasmic reticulum stress(GO:1990440) |
| 0.1 | 0.3 | GO:0072344 | rescue of stalled ribosome(GO:0072344) |
| 0.1 | 0.5 | GO:0009212 | dTTP biosynthetic process(GO:0006235) pyrimidine deoxyribonucleoside triphosphate biosynthetic process(GO:0009212) |
| 0.1 | 0.6 | GO:1901314 | negative regulation of histone ubiquitination(GO:0033183) regulation of histone H2A K63-linked ubiquitination(GO:1901314) negative regulation of histone H2A K63-linked ubiquitination(GO:1901315) |
| 0.1 | 2.0 | GO:0006646 | phosphatidylethanolamine biosynthetic process(GO:0006646) |
| 0.1 | 1.5 | GO:0032464 | positive regulation of protein homooligomerization(GO:0032464) |
| 0.1 | 0.7 | GO:0015712 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.1 | 0.6 | GO:0000389 | mRNA 3'-splice site recognition(GO:0000389) |
| 0.1 | 0.7 | GO:0018230 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 0.1 | 0.7 | GO:2000781 | positive regulation of double-strand break repair(GO:2000781) |
| 0.0 | 1.2 | GO:0051014 | actin filament severing(GO:0051014) |
| 0.0 | 1.2 | GO:0036158 | outer dynein arm assembly(GO:0036158) |
| 0.0 | 0.2 | GO:0018153 | isopeptide cross-linking via N6-(L-isoglutamyl)-L-lysine(GO:0018153) isopeptide cross-linking(GO:0018262) |
| 0.0 | 7.4 | GO:0060333 | interferon-gamma-mediated signaling pathway(GO:0060333) |
| 0.0 | 3.1 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.0 | 0.4 | GO:0002036 | regulation of L-glutamate transport(GO:0002036) |
| 0.0 | 0.2 | GO:0038098 | sequestering of BMP from receptor via BMP binding(GO:0038098) |
| 0.0 | 3.7 | GO:0006968 | cellular defense response(GO:0006968) |
| 0.0 | 0.7 | GO:0015886 | heme transport(GO:0015886) |
| 0.0 | 2.0 | GO:0006607 | NLS-bearing protein import into nucleus(GO:0006607) |
| 0.0 | 0.5 | GO:0036066 | protein O-linked fucosylation(GO:0036066) |
| 0.0 | 2.9 | GO:0034260 | negative regulation of GTPase activity(GO:0034260) |
| 0.0 | 0.2 | GO:0010446 | response to alkaline pH(GO:0010446) |
| 0.0 | 1.1 | GO:0045332 | phospholipid translocation(GO:0045332) |
| 0.0 | 0.8 | GO:0090266 | regulation of mitotic cell cycle spindle assembly checkpoint(GO:0090266) regulation of mitotic spindle checkpoint(GO:1903504) |
| 0.0 | 0.1 | GO:0019417 | sulfur oxidation(GO:0019417) |
| 0.0 | 1.1 | GO:0034389 | lipid particle organization(GO:0034389) |
| 0.0 | 1.3 | GO:0035338 | long-chain fatty-acyl-CoA biosynthetic process(GO:0035338) |
| 0.0 | 0.1 | GO:1903899 | positive regulation of PERK-mediated unfolded protein response(GO:1903899) |
| 0.0 | 1.2 | GO:0061641 | CENP-A containing nucleosome assembly(GO:0034080) CENP-A containing chromatin organization(GO:0061641) |
| 0.0 | 1.4 | GO:0006409 | tRNA export from nucleus(GO:0006409) tRNA-containing ribonucleoprotein complex export from nucleus(GO:0071431) |
| 0.0 | 1.6 | GO:0043044 | ATP-dependent chromatin remodeling(GO:0043044) |
| 0.0 | 0.7 | GO:0034587 | piRNA metabolic process(GO:0034587) |
| 0.0 | 1.2 | GO:0070534 | protein K63-linked ubiquitination(GO:0070534) |
| 0.0 | 2.6 | GO:0030166 | proteoglycan biosynthetic process(GO:0030166) |
| 0.0 | 2.3 | GO:0042035 | regulation of cytokine biosynthetic process(GO:0042035) |
| 0.0 | 0.1 | GO:0046901 | tetrahydrofolylpolyglutamate biosynthetic process(GO:0046901) |
| 0.0 | 1.3 | GO:0007019 | microtubule depolymerization(GO:0007019) |
| 0.0 | 0.1 | GO:0051684 | maintenance of Golgi location(GO:0051684) |
| 0.0 | 0.1 | GO:2000395 | regulation of ubiquitin-dependent endocytosis(GO:2000395) positive regulation of ubiquitin-dependent endocytosis(GO:2000397) |
| 0.0 | 0.1 | GO:0042791 | 5S class rRNA transcription from RNA polymerase III type 1 promoter(GO:0042791) tRNA transcription from RNA polymerase III promoter(GO:0042797) |
| 0.0 | 0.1 | GO:0048388 | endosomal lumen acidification(GO:0048388) |
| 0.0 | 0.1 | GO:0046121 | deoxyribonucleoside catabolic process(GO:0046121) |
| 0.0 | 0.5 | GO:0018195 | peptidyl-arginine modification(GO:0018195) |
| 0.0 | 2.6 | GO:0043154 | negative regulation of cysteine-type endopeptidase activity involved in apoptotic process(GO:0043154) |
| 0.0 | 0.1 | GO:0032468 | Golgi calcium ion homeostasis(GO:0032468) |
| 0.0 | 0.1 | GO:0042866 | pyruvate biosynthetic process(GO:0042866) |
| 0.0 | 0.4 | GO:0000462 | maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000462) |
| 0.0 | 0.1 | GO:1902527 | positive regulation of protein monoubiquitination(GO:1902527) |
| 0.0 | 0.7 | GO:0051491 | positive regulation of filopodium assembly(GO:0051491) |
| 0.0 | 0.2 | GO:0070935 | 3'-UTR-mediated mRNA stabilization(GO:0070935) |
| 0.0 | 0.2 | GO:0035635 | entry of bacterium into host cell(GO:0035635) |
| 0.0 | 0.2 | GO:0030321 | transepithelial chloride transport(GO:0030321) |
| 0.0 | 0.2 | GO:0046784 | viral mRNA export from host cell nucleus(GO:0046784) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 6.8 | GO:0031313 | extrinsic component of endosome membrane(GO:0031313) |
| 0.5 | 4.4 | GO:0065010 | extracellular membrane-bounded organelle(GO:0065010) |
| 0.5 | 1.9 | GO:0031251 | PAN complex(GO:0031251) |
| 0.3 | 1.4 | GO:1990876 | cytoplasmic side of nuclear pore(GO:1990876) |
| 0.3 | 6.0 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.2 | 1.6 | GO:0042825 | TAP complex(GO:0042825) |
| 0.2 | 1.1 | GO:1990667 | PCSK9-AnxA2 complex(GO:1990667) |
| 0.2 | 1.3 | GO:1990769 | proximal neuron projection(GO:1990769) |
| 0.2 | 1.0 | GO:0032449 | CBM complex(GO:0032449) |
| 0.1 | 4.0 | GO:0071682 | endocytic vesicle lumen(GO:0071682) |
| 0.1 | 4.2 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.1 | 1.2 | GO:0036157 | outer dynein arm(GO:0036157) |
| 0.1 | 1.8 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.1 | 3.1 | GO:0042101 | T cell receptor complex(GO:0042101) |
| 0.1 | 0.7 | GO:0071547 | piP-body(GO:0071547) |
| 0.1 | 5.4 | GO:0008023 | transcription elongation factor complex(GO:0008023) |
| 0.1 | 2.1 | GO:0032045 | guanyl-nucleotide exchange factor complex(GO:0032045) |
| 0.1 | 0.5 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 0.1 | 2.0 | GO:0031229 | integral component of nuclear inner membrane(GO:0005639) intrinsic component of nuclear inner membrane(GO:0031229) |
| 0.1 | 3.1 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.1 | 5.7 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.1 | 0.8 | GO:0071986 | Ragulator complex(GO:0071986) |
| 0.0 | 0.5 | GO:0005672 | transcription factor TFIIA complex(GO:0005672) |
| 0.0 | 1.7 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.0 | 3.3 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 0.1 | GO:0071062 | alphav-beta3 integrin-vitronectin complex(GO:0071062) |
| 0.0 | 5.3 | GO:0035578 | azurophil granule lumen(GO:0035578) |
| 0.0 | 3.3 | GO:1904724 | tertiary granule lumen(GO:1904724) |
| 0.0 | 0.1 | GO:1902912 | pyruvate kinase complex(GO:1902912) |
| 0.0 | 1.0 | GO:0016581 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.0 | 2.0 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.0 | 3.1 | GO:0101003 | ficolin-1-rich granule membrane(GO:0101003) |
| 0.0 | 0.6 | GO:0097225 | sperm midpiece(GO:0097225) |
| 0.0 | 2.5 | GO:0045335 | phagocytic vesicle(GO:0045335) |
| 0.0 | 0.4 | GO:0000812 | Swr1 complex(GO:0000812) |
| 0.0 | 2.3 | GO:0000118 | histone deacetylase complex(GO:0000118) |
| 0.0 | 0.5 | GO:0033116 | endoplasmic reticulum-Golgi intermediate compartment membrane(GO:0033116) |
| 0.0 | 5.7 | GO:0034774 | secretory granule lumen(GO:0034774) |
| 0.0 | 0.7 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.0 | 0.2 | GO:0005915 | zonula adherens(GO:0005915) |
| 0.0 | 4.9 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.0 | 0.5 | GO:0033270 | paranode region of axon(GO:0033270) |
| 0.0 | 0.3 | GO:0030126 | COPI vesicle coat(GO:0030126) |
| 0.0 | 0.1 | GO:0000127 | transcription factor TFIIIC complex(GO:0000127) |
| 0.0 | 0.6 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.0 | 0.2 | GO:0000445 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.0 | 0.5 | GO:0030057 | desmosome(GO:0030057) |
| 0.0 | 2.8 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.0 | 2.3 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.0 | 1.3 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.0 | 5.8 | GO:0005769 | early endosome(GO:0005769) |
| 0.0 | 5.8 | GO:0000790 | nuclear chromatin(GO:0000790) |
| 0.0 | 1.2 | GO:0005802 | trans-Golgi network(GO:0005802) |
| 0.0 | 2.8 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 11.9 | GO:0000139 | Golgi membrane(GO:0000139) |
| 0.0 | 0.4 | GO:0032040 | small-subunit processome(GO:0032040) |
| 0.0 | 1.2 | GO:0043202 | lysosomal lumen(GO:0043202) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.3 | 9.2 | GO:0047894 | flavonol 3-sulfotransferase activity(GO:0047894) |
| 1.3 | 5.0 | GO:0030107 | HLA-A specific inhibitory MHC class I receptor activity(GO:0030107) |
| 1.0 | 2.9 | GO:0036313 | phosphatidylinositol 3-kinase catalytic subunit binding(GO:0036313) |
| 0.8 | 3.1 | GO:0032396 | inhibitory MHC class I receptor activity(GO:0032396) |
| 0.6 | 6.3 | GO:0070892 | lipoteichoic acid receptor activity(GO:0070892) |
| 0.5 | 4.4 | GO:0004035 | alkaline phosphatase activity(GO:0004035) |
| 0.5 | 8.9 | GO:0019957 | C-C chemokine binding(GO:0019957) |
| 0.5 | 2.6 | GO:0004382 | guanosine-diphosphatase activity(GO:0004382) |
| 0.4 | 6.0 | GO:0032395 | MHC class II receptor activity(GO:0032395) |
| 0.4 | 4.2 | GO:0001730 | 2'-5'-oligoadenylate synthetase activity(GO:0001730) |
| 0.3 | 2.0 | GO:0008195 | phosphatidate phosphatase activity(GO:0008195) |
| 0.3 | 1.6 | GO:0015433 | peptide antigen-transporting ATPase activity(GO:0015433) |
| 0.2 | 1.0 | GO:0070053 | thrombospondin receptor activity(GO:0070053) |
| 0.2 | 5.6 | GO:0050786 | RAGE receptor binding(GO:0050786) |
| 0.2 | 1.0 | GO:0004706 | JUN kinase kinase kinase activity(GO:0004706) |
| 0.2 | 3.3 | GO:0038036 | sphingosine-1-phosphate receptor activity(GO:0038036) |
| 0.2 | 7.1 | GO:0005068 | transmembrane receptor protein tyrosine kinase adaptor activity(GO:0005068) |
| 0.2 | 0.9 | GO:0004310 | farnesyl-diphosphate farnesyltransferase activity(GO:0004310) squalene synthase activity(GO:0051996) |
| 0.2 | 2.6 | GO:0051525 | NFAT protein binding(GO:0051525) |
| 0.2 | 2.5 | GO:0048406 | nerve growth factor binding(GO:0048406) |
| 0.2 | 2.0 | GO:0051434 | BH3 domain binding(GO:0051434) |
| 0.2 | 0.6 | GO:0045131 | pre-mRNA branch point binding(GO:0045131) |
| 0.1 | 5.8 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.1 | 1.0 | GO:0004534 | 5'-3' exoribonuclease activity(GO:0004534) |
| 0.1 | 2.4 | GO:0001087 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.1 | 0.6 | GO:0003875 | ADP-ribosylarginine hydrolase activity(GO:0003875) |
| 0.1 | 0.5 | GO:0003896 | DNA primase activity(GO:0003896) |
| 0.1 | 2.4 | GO:0005522 | profilin binding(GO:0005522) |
| 0.1 | 0.4 | GO:0034647 | histone demethylase activity (H3-trimethyl-K4 specific)(GO:0034647) |
| 0.1 | 2.0 | GO:0042834 | peptidoglycan binding(GO:0042834) |
| 0.1 | 1.1 | GO:0019834 | phospholipase A2 inhibitor activity(GO:0019834) |
| 0.1 | 1.2 | GO:0004565 | beta-galactosidase activity(GO:0004565) |
| 0.1 | 3.1 | GO:0042288 | MHC class I protein binding(GO:0042288) |
| 0.1 | 1.1 | GO:0035381 | extracellular ATP-gated cation channel activity(GO:0004931) ATP-gated ion channel activity(GO:0035381) |
| 0.1 | 2.9 | GO:0005095 | GTPase inhibitor activity(GO:0005095) |
| 0.1 | 0.7 | GO:0015119 | hexose phosphate transmembrane transporter activity(GO:0015119) organophosphate:inorganic phosphate antiporter activity(GO:0015315) hexose-phosphate:inorganic phosphate antiporter activity(GO:0015526) glucose 6-phosphate:inorganic phosphate antiporter activity(GO:0061513) |
| 0.1 | 1.4 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.1 | 0.7 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 0.1 | 0.2 | GO:0004464 | leukotriene-C4 synthase activity(GO:0004464) |
| 0.1 | 1.3 | GO:0004579 | dolichyl-diphosphooligosaccharide-protein glycotransferase activity(GO:0004579) |
| 0.1 | 1.4 | GO:0004908 | interleukin-1 receptor activity(GO:0004908) |
| 0.1 | 1.9 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.1 | 1.0 | GO:0031996 | thioesterase binding(GO:0031996) |
| 0.1 | 2.1 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.1 | 0.4 | GO:0035402 | histone kinase activity (H3-T11 specific)(GO:0035402) |
| 0.1 | 0.2 | GO:0071633 | dihydroceramidase activity(GO:0071633) |
| 0.1 | 0.4 | GO:0034511 | U3 snoRNA binding(GO:0034511) |
| 0.1 | 0.4 | GO:0043141 | ATP-dependent 5'-3' DNA helicase activity(GO:0043141) |
| 0.1 | 2.1 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.1 | 3.1 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.1 | 2.5 | GO:0051059 | NF-kappaB binding(GO:0051059) |
| 0.0 | 3.1 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.0 | 2.3 | GO:0048027 | mRNA 5'-UTR binding(GO:0048027) |
| 0.0 | 1.3 | GO:0102391 | decanoate--CoA ligase activity(GO:0102391) |
| 0.0 | 0.9 | GO:0055106 | ubiquitin-protein transferase regulator activity(GO:0055106) |
| 0.0 | 0.3 | GO:0004045 | aminoacyl-tRNA hydrolase activity(GO:0004045) |
| 0.0 | 0.1 | GO:0051377 | mannose-ethanolamine phosphotransferase activity(GO:0051377) |
| 0.0 | 1.1 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.0 | 0.7 | GO:0015232 | heme transporter activity(GO:0015232) |
| 0.0 | 1.7 | GO:0032452 | histone demethylase activity(GO:0032452) |
| 0.0 | 1.0 | GO:0043495 | protein anchor(GO:0043495) |
| 0.0 | 0.1 | GO:0004139 | deoxyribose-phosphate aldolase activity(GO:0004139) |
| 0.0 | 1.4 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.0 | 0.2 | GO:0016403 | dimethylargininase activity(GO:0016403) |
| 0.0 | 1.3 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.0 | 1.3 | GO:0004869 | cysteine-type endopeptidase inhibitor activity(GO:0004869) |
| 0.0 | 2.0 | GO:0050681 | androgen receptor binding(GO:0050681) |
| 0.0 | 0.8 | GO:0005251 | delayed rectifier potassium channel activity(GO:0005251) |
| 0.0 | 0.1 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 0.0 | 5.0 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.0 | 1.0 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 0.0 | 0.1 | GO:0008841 | tetrahydrofolylpolyglutamate synthase activity(GO:0004326) dihydrofolate synthase activity(GO:0008841) |
| 0.0 | 0.1 | GO:0000026 | alpha-1,2-mannosyltransferase activity(GO:0000026) |
| 0.0 | 0.6 | GO:0046966 | thyroid hormone receptor binding(GO:0046966) |
| 0.0 | 1.1 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.0 | 0.9 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.0 | 3.6 | GO:0003823 | antigen binding(GO:0003823) |
| 0.0 | 1.1 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.0 | 0.7 | GO:0019706 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.0 | 0.2 | GO:0045029 | UDP-activated nucleotide receptor activity(GO:0045029) |
| 0.0 | 3.1 | GO:0001085 | RNA polymerase II transcription factor binding(GO:0001085) |
| 0.0 | 0.1 | GO:0072320 | volume-sensitive chloride channel activity(GO:0072320) |
| 0.0 | 0.4 | GO:0045505 | dynein intermediate chain binding(GO:0045505) |
| 0.0 | 0.2 | GO:0036122 | BMP binding(GO:0036122) |
| 0.0 | 1.3 | GO:0008270 | zinc ion binding(GO:0008270) |
| 0.0 | 1.0 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.0 | 0.8 | GO:0008138 | protein tyrosine/serine/threonine phosphatase activity(GO:0008138) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 10.1 | PID S1P S1P3 PATHWAY | S1P3 pathway |
| 0.1 | 2.6 | PID IL2 STAT5 PATHWAY | IL2 signaling events mediated by STAT5 |
| 0.1 | 3.0 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 0.1 | 4.3 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.1 | 6.0 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.1 | 4.5 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.1 | 7.9 | PID IL2 1PATHWAY | IL2-mediated signaling events |
| 0.1 | 2.5 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 0.1 | 1.0 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
| 0.1 | 3.1 | PID CD8 TCR DOWNSTREAM PATHWAY | Downstream signaling in naïve CD8+ T cells |
| 0.0 | 2.1 | PID IL8 CXCR2 PATHWAY | IL8- and CXCR2-mediated signaling events |
| 0.0 | 2.0 | PID NFKAPPAB CANONICAL PATHWAY | Canonical NF-kappaB pathway |
| 0.0 | 1.4 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.0 | 1.0 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.0 | 1.0 | PID IFNG PATHWAY | IFN-gamma pathway |
| 0.0 | 1.0 | PID FGF PATHWAY | FGF signaling pathway |
| 0.0 | 7.0 | NABA SECRETED FACTORS | Genes encoding secreted soluble factors |
| 0.0 | 0.4 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.0 | 0.8 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.0 | 0.4 | PID MYC PATHWAY | C-MYC pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 6.8 | REACTOME BINDING AND ENTRY OF HIV VIRION | Genes involved in Binding and entry of HIV virion |
| 0.3 | 9.2 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.2 | 5.6 | REACTOME ADVANCED GLYCOSYLATION ENDPRODUCT RECEPTOR SIGNALING | Genes involved in Advanced glycosylation endproduct receptor signaling |
| 0.1 | 3.2 | REACTOME TRANSLOCATION OF ZAP 70 TO IMMUNOLOGICAL SYNAPSE | Genes involved in Translocation of ZAP-70 to Immunological synapse |
| 0.1 | 3.5 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.1 | 2.0 | REACTOME NOTCH HLH TRANSCRIPTION PATHWAY | Genes involved in Notch-HLH transcription pathway |
| 0.1 | 1.0 | REACTOME DESTABILIZATION OF MRNA BY TRISTETRAPROLIN TTP | Genes involved in Destabilization of mRNA by Tristetraprolin (TTP) |
| 0.1 | 10.4 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.1 | 2.9 | REACTOME DESTABILIZATION OF MRNA BY KSRP | Genes involved in Destabilization of mRNA by KSRP |
| 0.1 | 2.0 | REACTOME SYNTHESIS OF PE | Genes involved in Synthesis of PE |
| 0.1 | 3.0 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.1 | 7.9 | REACTOME ANTIGEN PROCESSING CROSS PRESENTATION | Genes involved in Antigen processing-Cross presentation |
| 0.1 | 1.1 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.1 | 2.0 | REACTOME ACTIVATED TAK1 MEDIATES P38 MAPK ACTIVATION | Genes involved in activated TAK1 mediates p38 MAPK activation |
| 0.1 | 1.0 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.1 | 5.1 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.0 | 1.0 | REACTOME DOWNSTREAM TCR SIGNALING | Genes involved in Downstream TCR signaling |
| 0.0 | 3.3 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.0 | 2.1 | REACTOME THE ROLE OF NEF IN HIV1 REPLICATION AND DISEASE PATHOGENESIS | Genes involved in The role of Nef in HIV-1 replication and disease pathogenesis |
| 0.0 | 1.3 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.0 | 2.4 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 1.4 | REACTOME REGULATION OF MRNA STABILITY BY PROTEINS THAT BIND AU RICH ELEMENTS | Genes involved in Regulation of mRNA Stability by Proteins that Bind AU-rich Elements |
| 0.0 | 2.1 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.0 | 0.5 | REACTOME INHIBITION OF REPLICATION INITIATION OF DAMAGED DNA BY RB1 E2F1 | Genes involved in Inhibition of replication initiation of damaged DNA by RB1/E2F1 |
| 0.0 | 0.5 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.0 | 0.9 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 0.3 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.0 | 1.2 | REACTOME DEPOSITION OF NEW CENPA CONTAINING NUCLEOSOMES AT THE CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.0 | 3.3 | REACTOME G ALPHA I SIGNALLING EVENTS | Genes involved in G alpha (i) signalling events |
| 0.0 | 1.0 | REACTOME MEIOTIC SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.0 | 0.6 | REACTOME DOWNREGULATION OF SMAD2 3 SMAD4 TRANSCRIPTIONAL ACTIVITY | Genes involved in Downregulation of SMAD2/3:SMAD4 transcriptional activity |
| 0.0 | 0.2 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |