Project
Illumina Body Map 2: averaged replicates
Navigation
Downloads
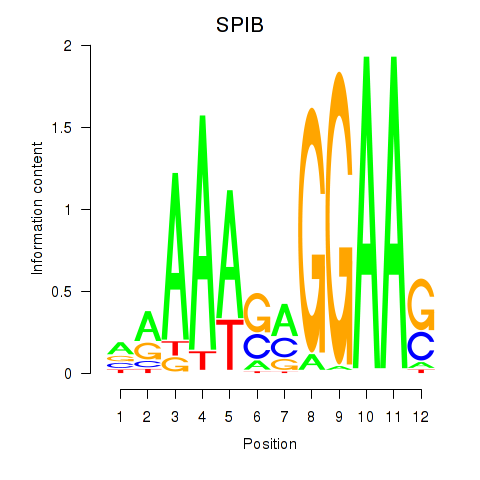
Results for SPIB
Z-value: 6.88
Transcription factors associated with SPIB
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
SPIB
|
ENSG00000269404.2 | Spi-B transcription factor |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| SPIB | hg19_v2_chr19_+_50922187_50922215 | 0.70 | 8.3e-06 | Click! |
Activity profile of SPIB motif
Sorted Z-values of SPIB motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr17_-_72709050 | 38.38 |
ENST00000583937.1
ENST00000301573.9 ENST00000326165.6 ENST00000469092.1 |
CD300LF
|
CD300 molecule-like family member f |
| chr1_-_160681593 | 28.61 |
ENST00000368045.3
ENST00000368046.3 |
CD48
|
CD48 molecule |
| chrX_-_47489244 | 23.03 |
ENST00000469388.1
ENST00000396992.3 ENST00000377005.2 |
CFP
|
complement factor properdin |
| chr22_+_37309662 | 20.08 |
ENST00000403662.3
ENST00000262825.5 |
CSF2RB
|
colony stimulating factor 2 receptor, beta, low-affinity (granulocyte-macrophage) |
| chr3_-_151047327 | 19.40 |
ENST00000325602.5
|
P2RY13
|
purinergic receptor P2Y, G-protein coupled, 13 |
| chr1_+_161676739 | 19.38 |
ENST00000236938.6
ENST00000367959.2 ENST00000546024.1 ENST00000540521.1 ENST00000367949.2 ENST00000350710.3 ENST00000540926.1 |
FCRLA
|
Fc receptor-like A |
| chr19_+_42381337 | 19.30 |
ENST00000597454.1
ENST00000444740.2 |
CD79A
|
CD79a molecule, immunoglobulin-associated alpha |
| chr12_+_10124001 | 18.79 |
ENST00000396507.3
ENST00000304361.4 ENST00000434319.2 |
CLEC12A
|
C-type lectin domain family 12, member A |
| chr17_-_72619783 | 18.76 |
ENST00000328630.3
|
CD300E
|
CD300e molecule |
| chrX_+_78200829 | 18.47 |
ENST00000544091.1
|
P2RY10
|
purinergic receptor P2Y, G-protein coupled, 10 |
| chrX_+_78200913 | 18.43 |
ENST00000171757.2
|
P2RY10
|
purinergic receptor P2Y, G-protein coupled, 10 |
| chr5_-_66492562 | 18.38 |
ENST00000256447.4
|
CD180
|
CD180 molecule |
| chr12_+_10124110 | 17.86 |
ENST00000350667.4
|
CLEC12A
|
C-type lectin domain family 12, member A |
| chr7_-_36764004 | 17.75 |
ENST00000431169.1
|
AOAH
|
acyloxyacyl hydrolase (neutrophil) |
| chr17_-_72619869 | 17.67 |
ENST00000392619.1
ENST00000426295.2 |
CD300E
|
CD300e molecule |
| chr1_+_209941827 | 17.58 |
ENST00000367023.1
|
TRAF3IP3
|
TRAF3 interacting protein 3 |
| chr7_+_74188309 | 17.57 |
ENST00000289473.4
ENST00000433458.1 |
NCF1
|
neutrophil cytosolic factor 1 |
| chr16_+_30194916 | 17.49 |
ENST00000570045.1
ENST00000565497.1 ENST00000570244.1 |
CORO1A
|
coronin, actin binding protein, 1A |
| chr9_-_71161505 | 17.43 |
ENST00000446290.1
|
RP11-274B18.2
|
RP11-274B18.2 |
| chr12_+_8276433 | 17.33 |
ENST00000345999.3
ENST00000352620.3 ENST00000360500.3 |
CLEC4A
|
C-type lectin domain family 4, member A |
| chr6_-_159466136 | 16.90 |
ENST00000367066.3
ENST00000326965.6 |
TAGAP
|
T-cell activation RhoGTPase activating protein |
| chr7_-_36764142 | 16.86 |
ENST00000258749.5
ENST00000535891.1 |
AOAH
|
acyloxyacyl hydrolase (neutrophil) |
| chr12_-_55378470 | 16.67 |
ENST00000524668.1
ENST00000533607.1 |
TESPA1
|
thymocyte expressed, positive selection associated 1 |
| chr12_-_55378452 | 16.46 |
ENST00000449076.1
|
TESPA1
|
thymocyte expressed, positive selection associated 1 |
| chr5_+_169064245 | 16.34 |
ENST00000256935.8
|
DOCK2
|
dedicator of cytokinesis 2 |
| chr1_+_161677034 | 16.04 |
ENST00000349527.4
ENST00000309691.6 ENST00000294796.4 ENST00000367953.3 ENST00000367950.1 |
FCRLA
|
Fc receptor-like A |
| chr7_-_36764062 | 15.90 |
ENST00000435386.1
|
AOAH
|
acyloxyacyl hydrolase (neutrophil) |
| chr6_-_159466042 | 15.61 |
ENST00000338313.5
|
TAGAP
|
T-cell activation RhoGTPase activating protein |
| chr5_+_57787254 | 15.48 |
ENST00000502276.1
ENST00000396776.2 ENST00000511930.1 |
GAPT
|
GRB2-binding adaptor protein, transmembrane |
| chr19_-_54850417 | 15.24 |
ENST00000291759.4
|
LILRA4
|
leukocyte immunoglobulin-like receptor, subfamily A (with TM domain), member 4 |
| chr1_+_112016414 | 15.23 |
ENST00000343534.5
ENST00000369718.3 |
C1orf162
|
chromosome 1 open reading frame 162 |
| chr1_-_160616804 | 15.01 |
ENST00000538290.1
|
SLAMF1
|
signaling lymphocytic activation molecule family member 1 |
| chr7_+_50344289 | 14.80 |
ENST00000413698.1
ENST00000359197.5 ENST00000331340.3 ENST00000357364.4 ENST00000343574.5 ENST00000349824.4 ENST00000346667.4 ENST00000440768.2 |
IKZF1
|
IKAROS family zinc finger 1 (Ikaros) |
| chr17_-_8868991 | 14.73 |
ENST00000447110.1
|
PIK3R5
|
phosphoinositide-3-kinase, regulatory subunit 5 |
| chr6_-_24936170 | 14.72 |
ENST00000538035.1
|
FAM65B
|
family with sequence similarity 65, member B |
| chr7_+_139528952 | 14.54 |
ENST00000416849.2
ENST00000436047.2 ENST00000414508.2 ENST00000448866.1 |
TBXAS1
|
thromboxane A synthase 1 (platelet) |
| chr10_+_49917751 | 14.50 |
ENST00000325239.5
ENST00000413659.2 |
WDFY4
|
WDFY family member 4 |
| chr17_+_72667239 | 14.30 |
ENST00000402449.4
|
RAB37
|
RAB37, member RAS oncogene family |
| chr12_+_8276224 | 14.17 |
ENST00000229332.5
|
CLEC4A
|
C-type lectin domain family 4, member A |
| chr17_-_43487780 | 14.12 |
ENST00000532038.1
ENST00000528677.1 |
ARHGAP27
|
Rho GTPase activating protein 27 |
| chr12_-_15114492 | 14.10 |
ENST00000541546.1
|
ARHGDIB
|
Rho GDP dissociation inhibitor (GDI) beta |
| chr3_+_121774202 | 13.96 |
ENST00000469710.1
ENST00000493101.1 ENST00000330540.2 ENST00000264468.5 |
CD86
|
CD86 molecule |
| chr1_+_161676983 | 13.92 |
ENST00000367957.2
|
FCRLA
|
Fc receptor-like A |
| chr5_+_54398463 | 13.80 |
ENST00000274306.6
|
GZMA
|
granzyme A (granzyme 1, cytotoxic T-lymphocyte-associated serine esterase 3) |
| chr17_-_5138099 | 13.70 |
ENST00000571800.1
ENST00000574081.1 ENST00000399600.4 ENST00000574297.1 |
SCIMP
|
SLP adaptor and CSK interacting membrane protein |
| chr16_-_50715239 | 13.63 |
ENST00000330943.4
ENST00000300590.3 |
SNX20
|
sorting nexin 20 |
| chrX_-_77582980 | 13.61 |
ENST00000373304.3
|
CYSLTR1
|
cysteinyl leukotriene receptor 1 |
| chr9_-_35618364 | 13.52 |
ENST00000378431.1
ENST00000378430.3 ENST00000259633.4 |
CD72
|
CD72 molecule |
| chr6_+_391739 | 13.44 |
ENST00000380956.4
|
IRF4
|
interferon regulatory factor 4 |
| chr6_-_133079022 | 13.32 |
ENST00000525289.1
ENST00000326499.6 |
VNN2
|
vanin 2 |
| chr1_+_26644441 | 13.32 |
ENST00000374213.2
|
CD52
|
CD52 molecule |
| chr7_+_139529040 | 13.32 |
ENST00000455353.1
ENST00000458722.1 ENST00000411653.1 |
TBXAS1
|
thromboxane A synthase 1 (platelet) |
| chr1_-_31230650 | 13.24 |
ENST00000294507.3
|
LAPTM5
|
lysosomal protein transmembrane 5 |
| chr19_+_16254488 | 13.19 |
ENST00000588246.1
ENST00000593031.1 |
HSH2D
|
hematopoietic SH2 domain containing |
| chr14_+_88471468 | 13.18 |
ENST00000267549.3
|
GPR65
|
G protein-coupled receptor 65 |
| chr22_+_37318082 | 13.07 |
ENST00000406230.1
|
CSF2RB
|
colony stimulating factor 2 receptor, beta, low-affinity (granulocyte-macrophage) |
| chr2_+_68592305 | 13.02 |
ENST00000234313.7
|
PLEK
|
pleckstrin |
| chr19_-_3786253 | 12.98 |
ENST00000585778.1
|
MATK
|
megakaryocyte-associated tyrosine kinase |
| chr6_+_31554779 | 12.63 |
ENST00000376090.2
|
LST1
|
leukocyte specific transcript 1 |
| chr9_+_215158 | 12.42 |
ENST00000479404.1
|
DOCK8
|
dedicator of cytokinesis 8 |
| chr9_+_214842 | 12.34 |
ENST00000453981.1
ENST00000432829.2 |
DOCK8
|
dedicator of cytokinesis 8 |
| chr19_-_3786354 | 12.28 |
ENST00000395040.2
ENST00000310132.6 |
MATK
|
megakaryocyte-associated tyrosine kinase |
| chr7_+_139529085 | 12.09 |
ENST00000539806.1
|
TBXAS1
|
thromboxane A synthase 1 (platelet) |
| chr6_-_41254403 | 12.05 |
ENST00000589614.1
ENST00000334475.6 ENST00000591620.1 ENST00000244709.4 |
TREM1
|
triggering receptor expressed on myeloid cells 1 |
| chr12_-_9885888 | 11.95 |
ENST00000327839.3
|
CLECL1
|
C-type lectin-like 1 |
| chr3_+_121796697 | 11.94 |
ENST00000482356.1
ENST00000393627.2 |
CD86
|
CD86 molecule |
| chr12_-_15114191 | 11.94 |
ENST00000541380.1
|
ARHGDIB
|
Rho GDP dissociation inhibitor (GDI) beta |
| chr17_-_29641104 | 11.81 |
ENST00000577894.1
ENST00000330927.4 |
EVI2B
|
ecotropic viral integration site 2B |
| chr11_-_58980342 | 11.77 |
ENST00000361050.3
|
MPEG1
|
macrophage expressed 1 |
| chr17_-_29641084 | 11.75 |
ENST00000544462.1
|
EVI2B
|
ecotropic viral integration site 2B |
| chr17_-_39093672 | 11.70 |
ENST00000209718.3
ENST00000436344.3 ENST00000485751.1 |
KRT23
|
keratin 23 (histone deacetylase inducible) |
| chr19_+_6772710 | 11.58 |
ENST00000304076.2
ENST00000602142.1 ENST00000596764.1 |
VAV1
|
vav 1 guanine nucleotide exchange factor |
| chr20_-_62710832 | 11.46 |
ENST00000395042.1
|
RGS19
|
regulator of G-protein signaling 19 |
| chr3_+_46395219 | 11.42 |
ENST00000445132.2
ENST00000292301.4 |
CCR2
|
chemokine (C-C motif) receptor 2 |
| chr13_+_108922228 | 11.42 |
ENST00000542136.1
|
TNFSF13B
|
tumor necrosis factor (ligand) superfamily, member 13b |
| chr5_+_35856951 | 11.32 |
ENST00000303115.3
ENST00000343305.4 ENST00000506850.1 ENST00000511982.1 |
IL7R
|
interleukin 7 receptor |
| chr1_+_161475208 | 11.31 |
ENST00000367972.4
ENST00000271450.6 |
FCGR2A
|
Fc fragment of IgG, low affinity IIa, receptor (CD32) |
| chr14_+_22217447 | 11.30 |
ENST00000390427.3
|
TRAV5
|
T cell receptor alpha variable 5 |
| chr16_+_57576584 | 11.22 |
ENST00000340339.4
|
GPR114
|
G protein-coupled receptor 114 |
| chr8_-_131028782 | 11.20 |
ENST00000519020.1
|
FAM49B
|
family with sequence similarity 49, member B |
| chr12_+_54891495 | 10.90 |
ENST00000293373.6
|
NCKAP1L
|
NCK-associated protein 1-like |
| chr12_-_15114658 | 10.84 |
ENST00000542276.1
|
ARHGDIB
|
Rho GDP dissociation inhibitor (GDI) beta |
| chr6_+_36973406 | 10.84 |
ENST00000274963.8
|
FGD2
|
FYVE, RhoGEF and PH domain containing 2 |
| chr16_+_3096638 | 10.79 |
ENST00000336577.4
|
MMP25
|
matrix metallopeptidase 25 |
| chr22_-_37640277 | 10.78 |
ENST00000401529.3
ENST00000249071.6 |
RAC2
|
ras-related C3 botulinum toxin substrate 2 (rho family, small GTP binding protein Rac2) |
| chr20_-_62711259 | 10.76 |
ENST00000332298.5
|
RGS19
|
regulator of G-protein signaling 19 |
| chr12_-_15114603 | 10.72 |
ENST00000228945.4
|
ARHGDIB
|
Rho GDP dissociation inhibitor (GDI) beta |
| chr19_-_54746600 | 10.69 |
ENST00000245621.5
ENST00000270464.5 ENST00000419410.2 ENST00000391735.3 ENST00000396365.2 ENST00000440558.2 ENST00000407860.2 |
LILRA6
LILRB3
|
leukocyte immunoglobulin-like receptor, subfamily A (with TM domain), member 6 leukocyte immunoglobulin-like receptor, subfamily B (with TM and ITIM domains), member 3 |
| chrX_-_30595959 | 10.62 |
ENST00000378962.3
|
CXorf21
|
chromosome X open reading frame 21 |
| chr13_+_31309645 | 10.61 |
ENST00000380490.3
|
ALOX5AP
|
arachidonate 5-lipoxygenase-activating protein |
| chr19_+_48828788 | 10.41 |
ENST00000594198.1
ENST00000597279.1 ENST00000593437.1 |
EMP3
|
epithelial membrane protein 3 |
| chr7_+_150211918 | 10.40 |
ENST00000313543.4
|
GIMAP7
|
GTPase, IMAP family member 7 |
| chr7_-_115799942 | 10.36 |
ENST00000484212.1
|
TFEC
|
transcription factor EC |
| chr11_+_65408273 | 10.24 |
ENST00000394227.3
|
SIPA1
|
signal-induced proliferation-associated 1 |
| chr12_+_7060508 | 10.21 |
ENST00000541698.1
ENST00000542462.1 |
PTPN6
|
protein tyrosine phosphatase, non-receptor type 6 |
| chr4_+_68424434 | 10.14 |
ENST00000265404.2
ENST00000396225.1 |
STAP1
|
signal transducing adaptor family member 1 |
| chr12_+_8276495 | 10.12 |
ENST00000546339.1
|
CLEC4A
|
C-type lectin domain family 4, member A |
| chr5_+_118668846 | 10.07 |
ENST00000513374.1
|
TNFAIP8
|
tumor necrosis factor, alpha-induced protein 8 |
| chr2_+_113885138 | 10.01 |
ENST00000409930.3
|
IL1RN
|
interleukin 1 receptor antagonist |
| chr22_-_37640456 | 9.89 |
ENST00000405484.1
ENST00000441619.1 ENST00000406508.1 |
RAC2
|
ras-related C3 botulinum toxin substrate 2 (rho family, small GTP binding protein Rac2) |
| chr3_+_32433363 | 9.88 |
ENST00000465248.1
|
CMTM7
|
CKLF-like MARVEL transmembrane domain containing 7 |
| chr12_-_9885702 | 9.84 |
ENST00000542530.1
|
CLECL1
|
C-type lectin-like 1 |
| chr3_+_46283863 | 9.74 |
ENST00000545097.1
ENST00000541018.1 |
CCR3
|
chemokine (C-C motif) receptor 3 |
| chr3_+_46395579 | 9.73 |
ENST00000421659.1
|
CCR2
|
chemokine (C-C motif) receptor 2 |
| chr10_+_26727333 | 9.72 |
ENST00000356785.4
|
APBB1IP
|
amyloid beta (A4) precursor protein-binding, family B, member 1 interacting protein |
| chr1_+_117544366 | 9.58 |
ENST00000256652.4
ENST00000369470.1 |
CD101
|
CD101 molecule |
| chr11_-_64512273 | 9.53 |
ENST00000377497.3
ENST00000377487.1 ENST00000430645.1 |
RASGRP2
|
RAS guanyl releasing protein 2 (calcium and DAG-regulated) |
| chr12_+_7060414 | 9.52 |
ENST00000538715.1
|
PTPN6
|
protein tyrosine phosphatase, non-receptor type 6 |
| chr6_+_31582961 | 9.49 |
ENST00000376059.3
ENST00000337917.7 |
AIF1
|
allograft inflammatory factor 1 |
| chr19_+_7413835 | 9.43 |
ENST00000576789.1
|
CTB-133G6.1
|
CTB-133G6.1 |
| chr11_-_3859089 | 9.38 |
ENST00000396979.1
|
RHOG
|
ras homolog family member G |
| chr21_-_46340807 | 9.34 |
ENST00000397846.3
ENST00000524251.1 ENST00000522688.1 |
ITGB2
|
integrin, beta 2 (complement component 3 receptor 3 and 4 subunit) |
| chr13_+_108921977 | 9.33 |
ENST00000430559.1
ENST00000375887.4 |
TNFSF13B
|
tumor necrosis factor (ligand) superfamily, member 13b |
| chr19_+_48828582 | 9.31 |
ENST00000270221.6
ENST00000596315.1 |
EMP3
|
epithelial membrane protein 3 |
| chr17_-_33885117 | 9.30 |
ENST00000415846.3
|
SLFN14
|
schlafen family member 14 |
| chr15_+_77287426 | 9.26 |
ENST00000558012.1
ENST00000267939.5 ENST00000379595.3 |
PSTPIP1
|
proline-serine-threonine phosphatase interacting protein 1 |
| chr19_-_54804173 | 9.20 |
ENST00000391744.3
ENST00000251390.3 |
LILRA3
|
leukocyte immunoglobulin-like receptor, subfamily A (without TM domain), member 3 |
| chr1_+_40840320 | 9.16 |
ENST00000372708.1
|
SMAP2
|
small ArfGAP2 |
| chr11_-_64512803 | 9.15 |
ENST00000377489.1
ENST00000354024.3 |
RASGRP2
|
RAS guanyl releasing protein 2 (calcium and DAG-regulated) |
| chr19_+_50919056 | 9.10 |
ENST00000599632.1
|
CTD-2545M3.6
|
CTD-2545M3.6 |
| chr10_-_98480243 | 9.01 |
ENST00000339364.5
|
PIK3AP1
|
phosphoinositide-3-kinase adaptor protein 1 |
| chr20_+_30640004 | 8.96 |
ENST00000520553.1
ENST00000518730.1 ENST00000375852.2 |
HCK
|
hemopoietic cell kinase |
| chr17_-_18950310 | 8.96 |
ENST00000573099.1
|
GRAP
|
GRB2-related adaptor protein |
| chr17_-_73389854 | 8.95 |
ENST00000578961.1
ENST00000392564.1 ENST00000582582.1 |
GRB2
|
growth factor receptor-bound protein 2 |
| chr9_+_93564039 | 8.95 |
ENST00000375754.4
ENST00000375751.4 |
SYK
|
spleen tyrosine kinase |
| chr12_+_7060432 | 8.93 |
ENST00000318974.9
ENST00000456013.1 |
PTPN6
|
protein tyrosine phosphatase, non-receptor type 6 |
| chr19_+_35940486 | 8.93 |
ENST00000246549.2
|
FFAR2
|
free fatty acid receptor 2 |
| chr11_-_47400062 | 8.89 |
ENST00000533030.1
|
SPI1
|
spleen focus forming virus (SFFV) proviral integration oncogene |
| chr14_-_23288930 | 8.86 |
ENST00000554517.1
ENST00000285850.7 ENST00000397529.2 ENST00000555702.1 |
SLC7A7
|
solute carrier family 7 (amino acid transporter light chain, y+L system), member 7 |
| chr19_-_52035044 | 8.80 |
ENST00000359982.4
ENST00000436458.1 ENST00000425629.3 ENST00000391797.3 ENST00000343300.4 |
SIGLEC6
|
sialic acid binding Ig-like lectin 6 |
| chr1_+_159272111 | 8.76 |
ENST00000368114.1
|
FCER1A
|
Fc fragment of IgE, high affinity I, receptor for; alpha polypeptide |
| chr11_+_60145967 | 8.76 |
ENST00000534016.1
|
MS4A7
|
membrane-spanning 4-domains, subfamily A, member 7 |
| chr19_-_13213954 | 8.71 |
ENST00000590974.1
|
LYL1
|
lymphoblastic leukemia derived sequence 1 |
| chr19_-_10446449 | 8.68 |
ENST00000592439.1
|
ICAM3
|
intercellular adhesion molecule 3 |
| chr17_-_73389737 | 8.68 |
ENST00000392563.1
|
GRB2
|
growth factor receptor-bound protein 2 |
| chr11_-_47399942 | 8.67 |
ENST00000227163.4
|
SPI1
|
spleen focus forming virus (SFFV) proviral integration oncogene |
| chr21_-_32716556 | 8.64 |
ENST00000455508.1
|
TIAM1
|
T-cell lymphoma invasion and metastasis 1 |
| chr9_+_93564191 | 8.63 |
ENST00000375747.1
|
SYK
|
spleen tyrosine kinase |
| chr16_-_50715196 | 8.55 |
ENST00000423026.2
|
SNX20
|
sorting nexin 20 |
| chr19_+_49838653 | 8.54 |
ENST00000598095.1
ENST00000426897.2 ENST00000323906.4 ENST00000535669.2 ENST00000597602.1 ENST00000595660.1 |
CD37
|
CD37 molecule |
| chr11_-_64512469 | 8.53 |
ENST00000377485.1
|
RASGRP2
|
RAS guanyl releasing protein 2 (calcium and DAG-regulated) |
| chr21_-_46340884 | 8.48 |
ENST00000302347.5
ENST00000517819.1 |
ITGB2
|
integrin, beta 2 (complement component 3 receptor 3 and 4 subunit) |
| chr2_+_143886877 | 8.47 |
ENST00000295095.6
|
ARHGAP15
|
Rho GTPase activating protein 15 |
| chr5_-_39270725 | 8.47 |
ENST00000512138.1
ENST00000512982.1 ENST00000540520.1 |
FYB
|
FYN binding protein |
| chr6_+_31553901 | 8.35 |
ENST00000418507.2
ENST00000438075.2 ENST00000376100.3 ENST00000376111.4 |
LST1
|
leukocyte specific transcript 1 |
| chr19_-_13213662 | 8.32 |
ENST00000264824.4
|
LYL1
|
lymphoblastic leukemia derived sequence 1 |
| chr2_-_145188137 | 8.29 |
ENST00000440875.1
|
ZEB2
|
zinc finger E-box binding homeobox 2 |
| chr2_+_233924548 | 8.29 |
ENST00000422935.1
|
INPP5D
|
inositol polyphosphate-5-phosphatase, 145kDa |
| chr1_+_149754227 | 8.27 |
ENST00000444948.1
ENST00000369168.4 |
FCGR1A
|
Fc fragment of IgG, high affinity Ia, receptor (CD64) |
| chr6_+_31553978 | 8.26 |
ENST00000376096.1
ENST00000376099.1 ENST00000376110.3 |
LST1
|
leukocyte specific transcript 1 |
| chr19_+_35939154 | 8.25 |
ENST00000599180.2
|
FFAR2
|
free fatty acid receptor 2 |
| chr9_-_130541017 | 8.25 |
ENST00000314830.8
|
SH2D3C
|
SH2 domain containing 3C |
| chr6_+_31554612 | 8.25 |
ENST00000211921.7
|
LST1
|
leukocyte specific transcript 1 |
| chr2_+_113931513 | 8.16 |
ENST00000245796.6
ENST00000441564.3 |
PSD4
|
pleckstrin and Sec7 domain containing 4 |
| chr20_-_35274548 | 8.14 |
ENST00000262866.4
|
SLA2
|
Src-like-adaptor 2 |
| chr11_-_47400032 | 8.13 |
ENST00000533968.1
|
SPI1
|
spleen focus forming virus (SFFV) proviral integration oncogene |
| chr16_+_31213206 | 8.12 |
ENST00000561916.2
|
C16orf98
|
chromosome 16 open reading frame 98 |
| chr1_+_15736359 | 8.07 |
ENST00000375980.4
|
EFHD2
|
EF-hand domain family, member D2 |
| chr12_+_25205666 | 8.04 |
ENST00000547044.1
|
LRMP
|
lymphoid-restricted membrane protein |
| chr6_+_31554826 | 8.04 |
ENST00000376089.2
ENST00000396112.2 |
LST1
|
leukocyte specific transcript 1 |
| chr11_-_47400078 | 8.01 |
ENST00000378538.3
|
SPI1
|
spleen focus forming virus (SFFV) proviral integration oncogene |
| chr1_+_151129103 | 8.01 |
ENST00000368910.3
|
TNFAIP8L2
|
tumor necrosis factor, alpha-induced protein 8-like 2 |
| chr15_+_77287715 | 8.01 |
ENST00000559161.1
|
PSTPIP1
|
proline-serine-threonine phosphatase interacting protein 1 |
| chr2_-_242089677 | 7.96 |
ENST00000405260.1
|
PASK
|
PAS domain containing serine/threonine kinase |
| chr17_-_43487741 | 7.95 |
ENST00000455881.1
|
ARHGAP27
|
Rho GTPase activating protein 27 |
| chr8_-_101734308 | 7.86 |
ENST00000519004.1
ENST00000519363.1 ENST00000520142.1 |
PABPC1
|
poly(A) binding protein, cytoplasmic 1 |
| chr10_+_129785536 | 7.82 |
ENST00000419012.2
|
PTPRE
|
protein tyrosine phosphatase, receptor type, E |
| chr22_-_37880543 | 7.78 |
ENST00000442496.1
|
MFNG
|
MFNG O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase |
| chr6_+_31554456 | 7.78 |
ENST00000339530.4
|
LST1
|
leukocyte specific transcript 1 |
| chr1_-_120935894 | 7.66 |
ENST00000369383.4
ENST00000369384.4 |
FCGR1B
|
Fc fragment of IgG, high affinity Ib, receptor (CD64) |
| chr16_+_12059050 | 7.63 |
ENST00000396495.3
|
TNFRSF17
|
tumor necrosis factor receptor superfamily, member 17 |
| chr21_-_46340770 | 7.53 |
ENST00000397854.3
|
ITGB2
|
integrin, beta 2 (complement component 3 receptor 3 and 4 subunit) |
| chr3_+_46283916 | 7.44 |
ENST00000395940.2
|
CCR3
|
chemokine (C-C motif) receptor 3 |
| chr12_+_25205628 | 7.43 |
ENST00000554942.1
|
LRMP
|
lymphoid-restricted membrane protein |
| chr6_+_31554636 | 7.41 |
ENST00000433492.1
|
LST1
|
leukocyte specific transcript 1 |
| chr11_+_60145948 | 7.33 |
ENST00000300184.3
ENST00000358246.1 |
MS4A7
|
membrane-spanning 4-domains, subfamily A, member 7 |
| chr17_+_8869157 | 7.19 |
ENST00000585297.1
|
CTB-41I6.1
|
CTB-41I6.1 |
| chr5_-_156536126 | 7.15 |
ENST00000522593.1
|
HAVCR2
|
hepatitis A virus cellular receptor 2 |
| chr1_+_167599532 | 7.14 |
ENST00000537350.1
|
RCSD1
|
RCSD domain containing 1 |
| chr10_+_129785574 | 7.11 |
ENST00000430713.2
ENST00000471218.1 |
PTPRE
|
protein tyrosine phosphatase, receptor type, E |
| chr6_-_154677900 | 6.98 |
ENST00000265198.4
ENST00000520261.1 |
IPCEF1
|
interaction protein for cytohesin exchange factors 1 |
| chr15_+_91427642 | 6.97 |
ENST00000328850.3
ENST00000414248.2 |
FES
|
feline sarcoma oncogene |
| chr3_-_3151664 | 6.92 |
ENST00000256452.3
ENST00000311981.8 ENST00000430514.2 ENST00000456302.1 |
IL5RA
|
interleukin 5 receptor, alpha |
| chr9_+_93589734 | 6.92 |
ENST00000375746.1
|
SYK
|
spleen tyrosine kinase |
| chr8_-_101734170 | 6.89 |
ENST00000522387.1
ENST00000518196.1 |
PABPC1
|
poly(A) binding protein, cytoplasmic 1 |
| chr14_-_98444369 | 6.86 |
ENST00000554822.1
|
C14orf64
|
chromosome 14 open reading frame 64 |
| chr20_+_62711482 | 6.84 |
ENST00000336866.2
ENST00000355631.4 |
OPRL1
|
opiate receptor-like 1 |
| chr14_-_98444438 | 6.82 |
ENST00000512901.2
|
C14orf64
|
chromosome 14 open reading frame 64 |
| chr14_-_23588816 | 6.80 |
ENST00000206513.5
|
CEBPE
|
CCAAT/enhancer binding protein (C/EBP), epsilon |
| chr1_+_206643787 | 6.79 |
ENST00000367120.3
|
IKBKE
|
inhibitor of kappa light polypeptide gene enhancer in B-cells, kinase epsilon |
| chr6_+_130339710 | 6.66 |
ENST00000526087.1
ENST00000533560.1 ENST00000361794.2 |
L3MBTL3
|
l(3)mbt-like 3 (Drosophila) |
| chr11_+_64107663 | 6.65 |
ENST00000356786.5
|
CCDC88B
|
coiled-coil domain containing 88B |
| chr7_+_106505912 | 6.63 |
ENST00000359195.3
|
PIK3CG
|
phosphatidylinositol-4,5-bisphosphate 3-kinase, catalytic subunit gamma |
| chr17_+_19030782 | 6.63 |
ENST00000344415.4
ENST00000577213.1 |
GRAPL
|
GRB2-related adaptor protein-like |
| chr11_+_60145997 | 6.61 |
ENST00000530614.1
ENST00000530027.1 ENST00000530234.2 ENST00000528215.1 ENST00000531787.1 |
MS4A7
MS4A14
|
membrane-spanning 4-domains, subfamily A, member 7 membrane-spanning 4-domains, subfamily A, member 14 |
| chr12_+_15699286 | 6.58 |
ENST00000442921.2
ENST00000542557.1 ENST00000445537.2 ENST00000544244.1 |
PTPRO
|
protein tyrosine phosphatase, receptor type, O |
| chr2_-_231084659 | 6.53 |
ENST00000258381.6
ENST00000358662.4 ENST00000455674.1 ENST00000392048.3 |
SP110
|
SP110 nuclear body protein |
| chr14_+_105391147 | 6.52 |
ENST00000540372.1
ENST00000392593.4 |
PLD4
|
phospholipase D family, member 4 |
| chr2_-_231084617 | 6.49 |
ENST00000409815.2
|
SP110
|
SP110 nuclear body protein |
| chr15_+_91427691 | 6.48 |
ENST00000559355.1
ENST00000394302.1 |
FES
|
feline sarcoma oncogene |
Network of associatons between targets according to the STRING database.
First level regulatory network of SPIB
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 13.1 | 39.3 | GO:0045404 | positive regulation of interleukin-4 biosynthetic process(GO:0045404) |
| 12.8 | 38.4 | GO:0034125 | negative regulation of MyD88-dependent toll-like receptor signaling pathway(GO:0034125) regulation of interleukin-4-mediated signaling pathway(GO:1902214) |
| 10.6 | 21.2 | GO:1902567 | negative regulation of eosinophil activation(GO:1902567) |
| 10.5 | 31.4 | GO:0002277 | myeloid dendritic cell activation involved in immune response(GO:0002277) |
| 10.1 | 10.1 | GO:1903973 | negative regulation of macrophage colony-stimulating factor signaling pathway(GO:1902227) negative regulation of response to macrophage colony-stimulating factor(GO:1903970) negative regulation of cellular response to macrophage colony-stimulating factor stimulus(GO:1903973) |
| 8.3 | 33.3 | GO:0045399 | positive regulation of interleukin-3 production(GO:0032752) interleukin-3 biosynthetic process(GO:0042223) regulation of interleukin-3 biosynthetic process(GO:0045399) positive regulation of interleukin-3 biosynthetic process(GO:0045401) |
| 8.3 | 24.8 | GO:0061485 | memory T cell proliferation(GO:0061485) |
| 8.0 | 40.1 | GO:0038043 | interleukin-5-mediated signaling pathway(GO:0038043) |
| 6.8 | 47.6 | GO:0071461 | cellular response to redox state(GO:0071461) |
| 6.7 | 33.7 | GO:0044027 | hypermethylation of CpG island(GO:0044027) |
| 6.0 | 42.3 | GO:0002322 | B cell proliferation involved in immune response(GO:0002322) |
| 5.7 | 17.2 | GO:0002232 | leukocyte chemotaxis involved in inflammatory response(GO:0002232) |
| 4.9 | 9.9 | GO:0002337 | B-1a B cell differentiation(GO:0002337) |
| 4.8 | 28.7 | GO:0033277 | abortive mitotic cell cycle(GO:0033277) |
| 4.7 | 33.1 | GO:0010387 | COP9 signalosome assembly(GO:0010387) |
| 4.1 | 20.7 | GO:0002636 | positive regulation of germinal center formation(GO:0002636) B cell costimulation(GO:0031296) |
| 4.1 | 12.2 | GO:0002434 | immune complex clearance(GO:0002434) immune complex clearance by monocytes and macrophages(GO:0002436) negative regulation of dendritic cell antigen processing and presentation(GO:0002605) regulation of cytotoxic T cell degranulation(GO:0043317) negative regulation of cytotoxic T cell degranulation(GO:0043318) regulation of immune complex clearance by monocytes and macrophages(GO:0090264) |
| 4.0 | 19.9 | GO:0045659 | regulation of neutrophil differentiation(GO:0045658) negative regulation of neutrophil differentiation(GO:0045659) |
| 3.6 | 10.9 | GO:0045588 | positive regulation of gamma-delta T cell differentiation(GO:0045588) |
| 3.6 | 7.1 | GO:0032672 | regulation of interleukin-3 production(GO:0032672) |
| 3.5 | 17.5 | GO:0032796 | uropod organization(GO:0032796) |
| 3.5 | 17.3 | GO:0042539 | hypotonic salinity response(GO:0042539) cellular hypotonic salinity response(GO:0071477) |
| 3.0 | 9.0 | GO:0002522 | leukocyte migration involved in immune response(GO:0002522) |
| 2.4 | 21.6 | GO:0034128 | negative regulation of MyD88-independent toll-like receptor signaling pathway(GO:0034128) |
| 2.3 | 15.9 | GO:0035709 | memory T cell activation(GO:0035709) regulation of memory T cell activation(GO:2000567) positive regulation of memory T cell activation(GO:2000568) |
| 2.2 | 13.5 | GO:1900147 | Schwann cell migration(GO:0036135) regulation of Schwann cell migration(GO:1900147) |
| 2.2 | 17.7 | GO:0051902 | negative regulation of mitochondrial depolarization(GO:0051902) |
| 2.2 | 6.6 | GO:0036058 | filtration diaphragm assembly(GO:0036058) slit diaphragm assembly(GO:0036060) |
| 2.2 | 26.3 | GO:0048298 | positive regulation of isotype switching to IgA isotypes(GO:0048298) |
| 2.2 | 12.9 | GO:0010897 | negative regulation of triglyceride catabolic process(GO:0010897) |
| 2.0 | 10.2 | GO:0042631 | cellular response to water deprivation(GO:0042631) |
| 2.0 | 6.1 | GO:1901076 | positive regulation of engulfment of apoptotic cell(GO:1901076) |
| 2.0 | 56.3 | GO:0035589 | G-protein coupled purinergic nucleotide receptor signaling pathway(GO:0035589) |
| 2.0 | 12.1 | GO:0070945 | neutrophil mediated killing of gram-negative bacterium(GO:0070945) |
| 2.0 | 5.9 | GO:0072356 | chromosome passenger complex localization to kinetochore(GO:0072356) |
| 1.9 | 13.6 | GO:0061737 | leukotriene signaling pathway(GO:0061737) |
| 1.9 | 7.7 | GO:1905205 | positive regulation of connective tissue replacement(GO:1905205) |
| 1.9 | 41.9 | GO:0019371 | cyclooxygenase pathway(GO:0019371) |
| 1.9 | 20.7 | GO:0060753 | regulation of mast cell chemotaxis(GO:0060753) |
| 1.8 | 3.6 | GO:0045356 | positive regulation of interferon-alpha biosynthetic process(GO:0045356) |
| 1.8 | 23.5 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 1.8 | 9.0 | GO:0034154 | toll-like receptor 7 signaling pathway(GO:0034154) |
| 1.8 | 5.4 | GO:0045083 | negative regulation of interleukin-12 biosynthetic process(GO:0045083) |
| 1.8 | 23.0 | GO:0036066 | protein O-linked fucosylation(GO:0036066) |
| 1.7 | 6.8 | GO:1904058 | positive regulation of sensory perception of pain(GO:1904058) |
| 1.6 | 13.0 | GO:0033625 | positive regulation of integrin activation(GO:0033625) |
| 1.6 | 11.0 | GO:0014846 | esophagus smooth muscle contraction(GO:0014846) |
| 1.5 | 17.6 | GO:2000623 | regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000622) negative regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000623) |
| 1.4 | 9.5 | GO:0014738 | regulation of muscle hyperplasia(GO:0014738) |
| 1.3 | 9.0 | GO:1904154 | positive regulation of retrograde protein transport, ER to cytosol(GO:1904154) |
| 1.3 | 25.4 | GO:0002523 | leukocyte migration involved in inflammatory response(GO:0002523) |
| 1.3 | 10.0 | GO:2000660 | negative regulation of interleukin-1-mediated signaling pathway(GO:2000660) |
| 1.2 | 4.7 | GO:1990637 | response to prolactin(GO:1990637) |
| 1.1 | 4.3 | GO:1903028 | positive regulation of opsonization(GO:1903028) |
| 1.1 | 17.0 | GO:0001955 | blood vessel maturation(GO:0001955) |
| 1.1 | 10.6 | GO:2001300 | lipoxin metabolic process(GO:2001300) |
| 1.1 | 4.2 | GO:0071883 | activation of MAPK activity by adrenergic receptor signaling pathway(GO:0071883) |
| 1.0 | 14.5 | GO:0032071 | regulation of endodeoxyribonuclease activity(GO:0032071) |
| 1.0 | 6.2 | GO:0021993 | initiation of neural tube closure(GO:0021993) |
| 1.0 | 5.1 | GO:0052056 | modulation by virus of host molecular function(GO:0039506) suppression by virus of host molecular function(GO:0039507) suppression by virus of host catalytic activity(GO:0039513) modulation by virus of host catalytic activity(GO:0039516) suppression by virus of host cysteine-type endopeptidase activity involved in apoptotic process(GO:0039650) negative regulation by symbiont of host catalytic activity(GO:0052053) negative regulation by symbiont of host molecular function(GO:0052056) modulation by symbiont of host catalytic activity(GO:0052148) |
| 1.0 | 4.0 | GO:0042144 | vacuole fusion, non-autophagic(GO:0042144) |
| 1.0 | 19.9 | GO:0045741 | positive regulation of epidermal growth factor-activated receptor activity(GO:0045741) |
| 1.0 | 8.0 | GO:0070091 | glucagon secretion(GO:0070091) regulation of glucagon secretion(GO:0070092) |
| 0.9 | 11.3 | GO:0001915 | negative regulation of T cell mediated cytotoxicity(GO:0001915) |
| 0.9 | 8.3 | GO:1902748 | positive regulation of lens fiber cell differentiation(GO:1902748) |
| 0.9 | 8.9 | GO:0000821 | regulation of arginine metabolic process(GO:0000821) |
| 0.9 | 7.7 | GO:1902255 | positive regulation of intrinsic apoptotic signaling pathway by p53 class mediator(GO:1902255) |
| 0.8 | 21.1 | GO:2000251 | positive regulation of actin cytoskeleton reorganization(GO:2000251) |
| 0.8 | 21.9 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.8 | 10.8 | GO:0060022 | hard palate development(GO:0060022) |
| 0.8 | 2.4 | GO:0002361 | CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation(GO:0002361) |
| 0.8 | 6.1 | GO:0090038 | negative regulation of protein kinase C signaling(GO:0090038) |
| 0.8 | 19.7 | GO:0060670 | branching involved in labyrinthine layer morphogenesis(GO:0060670) |
| 0.8 | 5.3 | GO:0070383 | DNA cytosine deamination(GO:0070383) |
| 0.7 | 9.7 | GO:0002291 | T cell activation via T cell receptor contact with antigen bound to MHC molecule on antigen presenting cell(GO:0002291) |
| 0.7 | 13.3 | GO:0015939 | pantothenate metabolic process(GO:0015939) |
| 0.7 | 8.1 | GO:0071287 | cellular response to manganese ion(GO:0071287) |
| 0.7 | 21.3 | GO:0032060 | bleb assembly(GO:0032060) |
| 0.7 | 3.5 | GO:0060800 | regulation of cell differentiation involved in embryonic placenta development(GO:0060800) |
| 0.7 | 2.8 | GO:0071894 | histone H2B conserved C-terminal lysine ubiquitination(GO:0071894) |
| 0.6 | 5.8 | GO:0007207 | phospholipase C-activating G-protein coupled acetylcholine receptor signaling pathway(GO:0007207) |
| 0.6 | 11.4 | GO:0002430 | complement receptor mediated signaling pathway(GO:0002430) |
| 0.6 | 13.1 | GO:0007252 | I-kappaB phosphorylation(GO:0007252) |
| 0.6 | 4.3 | GO:0071486 | cellular response to high light intensity(GO:0071486) retinal rod cell apoptotic process(GO:0097473) |
| 0.6 | 2.4 | GO:0032700 | negative regulation of interleukin-17 production(GO:0032700) |
| 0.6 | 4.8 | GO:0061198 | fungiform papilla formation(GO:0061198) |
| 0.6 | 4.6 | GO:2000158 | positive regulation of ubiquitin-specific protease activity(GO:2000158) |
| 0.6 | 22.2 | GO:0045671 | negative regulation of osteoclast differentiation(GO:0045671) |
| 0.5 | 54.8 | GO:0032945 | negative regulation of mononuclear cell proliferation(GO:0032945) negative regulation of lymphocyte proliferation(GO:0050672) |
| 0.5 | 2.0 | GO:0015910 | peroxisomal long-chain fatty acid import(GO:0015910) |
| 0.5 | 7.8 | GO:0002315 | marginal zone B cell differentiation(GO:0002315) |
| 0.5 | 4.4 | GO:0032876 | negative regulation of DNA endoreduplication(GO:0032876) |
| 0.5 | 6.6 | GO:0001562 | response to protozoan(GO:0001562) defense response to protozoan(GO:0042832) |
| 0.5 | 18.4 | GO:0002479 | antigen processing and presentation of exogenous peptide antigen via MHC class I, TAP-dependent(GO:0002479) |
| 0.4 | 2.2 | GO:2001045 | negative regulation of integrin-mediated signaling pathway(GO:2001045) |
| 0.4 | 6.7 | GO:0043249 | erythrocyte maturation(GO:0043249) |
| 0.4 | 8.1 | GO:0050869 | negative regulation of B cell activation(GO:0050869) |
| 0.4 | 4.7 | GO:0060613 | fat pad development(GO:0060613) |
| 0.4 | 6.1 | GO:0039536 | negative regulation of RIG-I signaling pathway(GO:0039536) |
| 0.4 | 13.3 | GO:0045730 | respiratory burst(GO:0045730) |
| 0.4 | 2.0 | GO:0035655 | interleukin-18-mediated signaling pathway(GO:0035655) cellular response to interleukin-18(GO:0071351) |
| 0.4 | 2.0 | GO:1903615 | regulation of protein tyrosine phosphatase activity(GO:1903613) positive regulation of protein tyrosine phosphatase activity(GO:1903615) |
| 0.4 | 13.2 | GO:0010447 | response to acidic pH(GO:0010447) |
| 0.4 | 11.6 | GO:0045954 | positive regulation of natural killer cell mediated cytotoxicity(GO:0045954) |
| 0.4 | 2.6 | GO:0043314 | negative regulation of neutrophil degranulation(GO:0043314) |
| 0.4 | 3.6 | GO:0010735 | positive regulation of transcription via serum response element binding(GO:0010735) |
| 0.3 | 5.5 | GO:0000290 | deadenylation-dependent decapping of nuclear-transcribed mRNA(GO:0000290) |
| 0.3 | 19.6 | GO:0006376 | mRNA splice site selection(GO:0006376) |
| 0.3 | 3.7 | GO:0032264 | IMP salvage(GO:0032264) |
| 0.3 | 7.1 | GO:0071474 | cellular hyperosmotic response(GO:0071474) |
| 0.3 | 19.3 | GO:0042100 | B cell proliferation(GO:0042100) |
| 0.3 | 14.7 | GO:0043551 | regulation of phosphatidylinositol 3-kinase activity(GO:0043551) |
| 0.3 | 35.6 | GO:0038083 | peptidyl-tyrosine autophosphorylation(GO:0038083) |
| 0.3 | 2.2 | GO:1901668 | regulation of superoxide dismutase activity(GO:1901668) |
| 0.3 | 25.2 | GO:0006968 | cellular defense response(GO:0006968) |
| 0.3 | 5.4 | GO:0007213 | G-protein coupled acetylcholine receptor signaling pathway(GO:0007213) |
| 0.3 | 1.4 | GO:0060339 | negative regulation of type I interferon-mediated signaling pathway(GO:0060339) |
| 0.3 | 3.7 | GO:0035973 | aggrephagy(GO:0035973) |
| 0.3 | 3.0 | GO:0002457 | T cell antigen processing and presentation(GO:0002457) |
| 0.3 | 8.2 | GO:0032012 | regulation of ARF protein signal transduction(GO:0032012) |
| 0.3 | 4.7 | GO:0007416 | synapse assembly(GO:0007416) regulation of synapse assembly(GO:0051963) |
| 0.3 | 9.4 | GO:1900027 | regulation of ruffle assembly(GO:1900027) |
| 0.3 | 14.9 | GO:0046627 | negative regulation of insulin receptor signaling pathway(GO:0046627) |
| 0.3 | 53.7 | GO:0002223 | stimulatory C-type lectin receptor signaling pathway(GO:0002223) |
| 0.3 | 2.5 | GO:0090043 | regulation of tubulin deacetylation(GO:0090043) |
| 0.2 | 1.5 | GO:0001692 | histamine metabolic process(GO:0001692) |
| 0.2 | 3.1 | GO:0010818 | T cell chemotaxis(GO:0010818) |
| 0.2 | 2.4 | GO:0015691 | cadmium ion transport(GO:0015691) cadmium ion transmembrane transport(GO:0070574) |
| 0.2 | 4.3 | GO:0090168 | Golgi reassembly(GO:0090168) |
| 0.2 | 3.3 | GO:0033299 | secretion of lysosomal enzymes(GO:0033299) |
| 0.2 | 23.4 | GO:0002819 | regulation of adaptive immune response(GO:0002819) |
| 0.2 | 2.5 | GO:0018094 | protein polyglycylation(GO:0018094) |
| 0.2 | 3.7 | GO:0051574 | positive regulation of histone H3-K9 methylation(GO:0051574) |
| 0.2 | 4.9 | GO:0042789 | mRNA transcription from RNA polymerase II promoter(GO:0042789) |
| 0.2 | 0.8 | GO:0036229 | glutamine secretion(GO:0010585) L-glutamine import(GO:0036229) L-glutamine import into cell(GO:1903803) |
| 0.2 | 8.1 | GO:0043984 | histone H4-K16 acetylation(GO:0043984) |
| 0.2 | 23.4 | GO:0071277 | cellular response to calcium ion(GO:0071277) |
| 0.2 | 2.2 | GO:1903859 | regulation of dendrite extension(GO:1903859) positive regulation of dendrite extension(GO:1903861) |
| 0.2 | 8.0 | GO:0050868 | negative regulation of T cell activation(GO:0050868) |
| 0.2 | 9.2 | GO:0002260 | lymphocyte homeostasis(GO:0002260) |
| 0.2 | 0.8 | GO:0070828 | heterochromatin assembly(GO:0031507) heterochromatin organization(GO:0070828) |
| 0.2 | 9.4 | GO:0046039 | GTP metabolic process(GO:0046039) |
| 0.2 | 0.7 | GO:0001767 | establishment of lymphocyte polarity(GO:0001767) |
| 0.2 | 6.8 | GO:0030225 | macrophage differentiation(GO:0030225) |
| 0.2 | 0.7 | GO:0006610 | ribosomal protein import into nucleus(GO:0006610) |
| 0.2 | 0.9 | GO:0010847 | regulation of chromatin assembly(GO:0010847) |
| 0.2 | 11.9 | GO:0043507 | positive regulation of JUN kinase activity(GO:0043507) |
| 0.2 | 8.7 | GO:0006607 | NLS-bearing protein import into nucleus(GO:0006607) |
| 0.2 | 1.2 | GO:0019805 | quinolinate biosynthetic process(GO:0019805) |
| 0.2 | 0.7 | GO:0000354 | cis assembly of pre-catalytic spliceosome(GO:0000354) |
| 0.2 | 3.4 | GO:0006020 | inositol metabolic process(GO:0006020) |
| 0.1 | 12.6 | GO:0006904 | vesicle docking involved in exocytosis(GO:0006904) |
| 0.1 | 1.1 | GO:2000812 | regulation of barbed-end actin filament capping(GO:2000812) |
| 0.1 | 1.1 | GO:0032483 | regulation of Rab protein signal transduction(GO:0032483) |
| 0.1 | 0.4 | GO:2000742 | anterior head development(GO:0097065) regulation of anterior head development(GO:2000742) positive regulation of anterior head development(GO:2000744) |
| 0.1 | 1.6 | GO:0070862 | negative regulation of protein exit from endoplasmic reticulum(GO:0070862) negative regulation of retrograde protein transport, ER to cytosol(GO:1904153) |
| 0.1 | 1.7 | GO:0051152 | positive regulation of smooth muscle cell differentiation(GO:0051152) |
| 0.1 | 19.7 | GO:1990830 | response to leukemia inhibitory factor(GO:1990823) cellular response to leukemia inhibitory factor(GO:1990830) |
| 0.1 | 14.5 | GO:0030218 | erythrocyte differentiation(GO:0030218) |
| 0.1 | 18.4 | GO:0006903 | vesicle targeting(GO:0006903) |
| 0.1 | 0.7 | GO:0039019 | pronephric nephron development(GO:0039019) |
| 0.1 | 1.6 | GO:1904354 | negative regulation of telomere capping(GO:1904354) |
| 0.1 | 5.6 | GO:0030855 | epithelial cell differentiation(GO:0030855) |
| 0.1 | 0.4 | GO:0021797 | forebrain anterior/posterior pattern specification(GO:0021797) |
| 0.1 | 1.7 | GO:1901096 | regulation of autophagosome maturation(GO:1901096) |
| 0.1 | 2.3 | GO:0002115 | store-operated calcium entry(GO:0002115) |
| 0.1 | 0.5 | GO:0006127 | glycerophosphate shuttle(GO:0006127) |
| 0.1 | 9.6 | GO:0015914 | phospholipid transport(GO:0015914) |
| 0.1 | 12.7 | GO:0002250 | adaptive immune response(GO:0002250) |
| 0.1 | 3.2 | GO:0007035 | vacuolar acidification(GO:0007035) |
| 0.1 | 1.2 | GO:0090481 | pyrimidine nucleotide-sugar transmembrane transport(GO:0090481) |
| 0.1 | 1.3 | GO:0034244 | negative regulation of transcription elongation from RNA polymerase II promoter(GO:0034244) |
| 0.1 | 0.8 | GO:0030579 | ubiquitin-dependent SMAD protein catabolic process(GO:0030579) |
| 0.1 | 0.4 | GO:0007198 | adenylate cyclase-inhibiting serotonin receptor signaling pathway(GO:0007198) |
| 0.1 | 5.6 | GO:0035329 | hippo signaling(GO:0035329) |
| 0.1 | 14.9 | GO:0007127 | meiosis I(GO:0007127) |
| 0.1 | 5.1 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
| 0.1 | 93.1 | GO:0043547 | positive regulation of GTPase activity(GO:0043547) |
| 0.1 | 2.9 | GO:0006359 | regulation of transcription from RNA polymerase III promoter(GO:0006359) |
| 0.1 | 2.2 | GO:0033169 | histone H3-K9 demethylation(GO:0033169) |
| 0.1 | 1.4 | GO:0051256 | mitotic spindle midzone assembly(GO:0051256) |
| 0.1 | 0.9 | GO:0007172 | signal complex assembly(GO:0007172) |
| 0.1 | 11.7 | GO:0070268 | cornification(GO:0070268) |
| 0.1 | 4.0 | GO:0031572 | G2 DNA damage checkpoint(GO:0031572) |
| 0.1 | 0.5 | GO:0051342 | regulation of cyclic-nucleotide phosphodiesterase activity(GO:0051342) negative regulation of cyclic-nucleotide phosphodiesterase activity(GO:0051344) |
| 0.1 | 0.3 | GO:0043686 | co-translational protein modification(GO:0043686) |
| 0.1 | 44.9 | GO:0002283 | neutrophil activation involved in immune response(GO:0002283) neutrophil degranulation(GO:0043312) |
| 0.1 | 1.4 | GO:0035360 | positive regulation of peroxisome proliferator activated receptor signaling pathway(GO:0035360) |
| 0.1 | 2.3 | GO:0061641 | CENP-A containing nucleosome assembly(GO:0034080) CENP-A containing chromatin organization(GO:0061641) |
| 0.1 | 5.5 | GO:0016266 | O-glycan processing(GO:0016266) |
| 0.1 | 2.5 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 0.1 | 10.4 | GO:0034605 | cellular response to heat(GO:0034605) |
| 0.1 | 1.2 | GO:0000729 | DNA double-strand break processing(GO:0000729) |
| 0.1 | 3.2 | GO:0072348 | sulfur compound transport(GO:0072348) |
| 0.1 | 1.3 | GO:1903204 | negative regulation of peptidyl-cysteine S-nitrosylation(GO:1902083) negative regulation of oxidative stress-induced neuron death(GO:1903204) |
| 0.1 | 2.6 | GO:0046686 | response to cadmium ion(GO:0046686) |
| 0.1 | 0.5 | GO:0000463 | maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000463) |
| 0.1 | 3.1 | GO:0006953 | acute-phase response(GO:0006953) |
| 0.1 | 1.0 | GO:0090383 | phagosome acidification(GO:0090383) |
| 0.1 | 1.4 | GO:0031663 | lipopolysaccharide-mediated signaling pathway(GO:0031663) |
| 0.1 | 6.6 | GO:0006487 | protein N-linked glycosylation(GO:0006487) |
| 0.1 | 5.4 | GO:0043154 | negative regulation of cysteine-type endopeptidase activity involved in apoptotic process(GO:0043154) |
| 0.0 | 2.1 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.0 | 2.2 | GO:0035338 | long-chain fatty-acyl-CoA biosynthetic process(GO:0035338) |
| 0.0 | 3.8 | GO:0043647 | inositol phosphate metabolic process(GO:0043647) |
| 0.0 | 1.8 | GO:0000186 | activation of MAPKK activity(GO:0000186) |
| 0.0 | 0.7 | GO:0014894 | response to muscle inactivity(GO:0014870) response to muscle inactivity involved in regulation of muscle adaptation(GO:0014877) response to denervation involved in regulation of muscle adaptation(GO:0014894) |
| 0.0 | 5.8 | GO:0034968 | histone lysine methylation(GO:0034968) |
| 0.0 | 0.6 | GO:0001780 | neutrophil homeostasis(GO:0001780) |
| 0.0 | 5.1 | GO:0006334 | nucleosome assembly(GO:0006334) |
| 0.0 | 0.0 | GO:0031642 | negative regulation of myelination(GO:0031642) |
| 0.0 | 5.4 | GO:0045047 | protein targeting to ER(GO:0045047) |
| 0.0 | 0.8 | GO:0016024 | CDP-diacylglycerol biosynthetic process(GO:0016024) |
| 0.0 | 2.1 | GO:0070301 | cellular response to hydrogen peroxide(GO:0070301) |
| 0.0 | 3.3 | GO:0042795 | snRNA transcription(GO:0009301) snRNA transcription from RNA polymerase II promoter(GO:0042795) |
| 0.0 | 0.8 | GO:0032201 | DNA ligation(GO:0006266) nucleotide-excision repair, DNA gap filling(GO:0006297) telomere maintenance via semi-conservative replication(GO:0032201) |
| 0.0 | 0.5 | GO:0005513 | detection of calcium ion(GO:0005513) |
| 0.0 | 1.2 | GO:1901998 | toxin transport(GO:1901998) |
| 0.0 | 35.2 | GO:0006952 | defense response(GO:0006952) |
| 0.0 | 0.4 | GO:0015886 | heme transport(GO:0015886) |
| 0.0 | 3.9 | GO:0015992 | proton transport(GO:0015992) |
| 0.0 | 0.4 | GO:0010990 | regulation of SMAD protein complex assembly(GO:0010990) negative regulation of SMAD protein complex assembly(GO:0010991) |
| 0.0 | 2.1 | GO:0065004 | protein-DNA complex assembly(GO:0065004) |
| 0.0 | 1.2 | GO:0071377 | cellular response to glucagon stimulus(GO:0071377) |
| 0.0 | 0.3 | GO:1990118 | sodium ion import across plasma membrane(GO:0098719) sodium ion import into cell(GO:1990118) |
| 0.0 | 1.0 | GO:0008625 | extrinsic apoptotic signaling pathway via death domain receptors(GO:0008625) |
| 0.0 | 1.4 | GO:0006977 | DNA damage response, signal transduction by p53 class mediator resulting in cell cycle arrest(GO:0006977) |
| 0.0 | 2.0 | GO:0046777 | protein autophosphorylation(GO:0046777) |
| 0.0 | 8.3 | GO:0016032 | viral process(GO:0016032) |
| 0.0 | 0.6 | GO:0000245 | spliceosomal complex assembly(GO:0000245) |
| 0.0 | 0.7 | GO:0035116 | embryonic hindlimb morphogenesis(GO:0035116) |
| 0.0 | 1.2 | GO:0014066 | regulation of phosphatidylinositol 3-kinase signaling(GO:0014066) |
| 0.0 | 0.8 | GO:0006611 | protein export from nucleus(GO:0006611) |
| 0.0 | 3.2 | GO:0008654 | phospholipid biosynthetic process(GO:0008654) |
| 0.0 | 4.5 | GO:0098656 | anion transmembrane transport(GO:0098656) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 11.1 | 33.2 | GO:0030526 | granulocyte macrophage colony-stimulating factor receptor complex(GO:0030526) |
| 6.9 | 27.6 | GO:0005944 | phosphatidylinositol 3-kinase complex, class IB(GO:0005944) |
| 6.3 | 43.8 | GO:0019815 | B cell receptor complex(GO:0019815) |
| 3.2 | 19.0 | GO:0070436 | Grb2-EGFR complex(GO:0070436) |
| 3.2 | 25.4 | GO:0034688 | integrin alphaM-beta2 complex(GO:0034688) |
| 2.9 | 25.7 | GO:0046696 | lipopolysaccharide receptor complex(GO:0046696) |
| 2.2 | 28.7 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 1.8 | 17.6 | GO:0032010 | phagolysosome(GO:0032010) |
| 1.1 | 31.0 | GO:0031254 | uropod(GO:0001931) cell trailing edge(GO:0031254) |
| 1.1 | 3.2 | GO:0032002 | interleukin-28 receptor complex(GO:0032002) |
| 1.0 | 33.1 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.9 | 3.7 | GO:0097635 | extrinsic component of autophagosome membrane(GO:0097635) |
| 0.8 | 11.3 | GO:0000788 | nuclear nucleosome(GO:0000788) |
| 0.7 | 0.7 | GO:0071159 | NF-kappaB complex(GO:0071159) |
| 0.7 | 7.3 | GO:0005955 | calcineurin complex(GO:0005955) |
| 0.7 | 13.8 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.7 | 9.7 | GO:0042101 | T cell receptor complex(GO:0042101) |
| 0.7 | 4.7 | GO:0071256 | Sec61 translocon complex(GO:0005784) translocon complex(GO:0071256) |
| 0.7 | 8.1 | GO:0072487 | MSL complex(GO:0072487) |
| 0.6 | 10.9 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.5 | 33.1 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.5 | 3.6 | GO:0032009 | early phagosome(GO:0032009) |
| 0.5 | 29.7 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.5 | 5.4 | GO:0005785 | signal recognition particle receptor complex(GO:0005785) |
| 0.5 | 15.9 | GO:1990124 | messenger ribonucleoprotein complex(GO:1990124) |
| 0.5 | 4.7 | GO:0098554 | cytoplasmic side of endoplasmic reticulum membrane(GO:0098554) |
| 0.5 | 1.9 | GO:0071006 | U2-type catalytic step 1 spliceosome(GO:0071006) |
| 0.4 | 83.0 | GO:0035579 | specific granule membrane(GO:0035579) |
| 0.4 | 105.7 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |
| 0.4 | 5.4 | GO:0033256 | I-kappaB/NF-kappaB complex(GO:0033256) |
| 0.4 | 3.2 | GO:0000220 | vacuolar proton-transporting V-type ATPase, V0 domain(GO:0000220) |
| 0.3 | 2.2 | GO:1990130 | Iml1 complex(GO:1990130) |
| 0.3 | 1.6 | GO:0000333 | telomerase catalytic core complex(GO:0000333) |
| 0.3 | 30.2 | GO:0035580 | specific granule lumen(GO:0035580) |
| 0.3 | 36.2 | GO:0045335 | phagocytic vesicle(GO:0045335) |
| 0.3 | 59.5 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.3 | 4.5 | GO:0046930 | pore complex(GO:0046930) |
| 0.3 | 2.7 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 0.3 | 22.2 | GO:0035577 | azurophil granule membrane(GO:0035577) |
| 0.2 | 5.5 | GO:0016442 | RISC complex(GO:0016442) RNAi effector complex(GO:0031332) |
| 0.2 | 43.8 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.2 | 4.0 | GO:0030897 | HOPS complex(GO:0030897) |
| 0.2 | 1.3 | GO:0032044 | DSIF complex(GO:0032044) |
| 0.2 | 5.1 | GO:0031265 | CD95 death-inducing signaling complex(GO:0031265) ripoptosome(GO:0097342) |
| 0.2 | 7.7 | GO:0030686 | 90S preribosome(GO:0030686) |
| 0.2 | 2.2 | GO:0016600 | flotillin complex(GO:0016600) |
| 0.2 | 2.4 | GO:0031906 | late endosome lumen(GO:0031906) |
| 0.2 | 4.6 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.2 | 63.6 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.2 | 3.3 | GO:0030904 | retromer complex(GO:0030904) |
| 0.2 | 3.9 | GO:0005885 | Arp2/3 protein complex(GO:0005885) |
| 0.1 | 59.1 | GO:0098857 | membrane raft(GO:0045121) membrane microdomain(GO:0098857) |
| 0.1 | 1.4 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.1 | 0.6 | GO:0071012 | catalytic step 1 spliceosome(GO:0071012) |
| 0.1 | 0.8 | GO:0043625 | delta DNA polymerase complex(GO:0043625) |
| 0.1 | 4.8 | GO:0016580 | Sin3 complex(GO:0016580) |
| 0.1 | 11.0 | GO:0005795 | Golgi stack(GO:0005795) |
| 0.1 | 2.5 | GO:0000124 | SAGA complex(GO:0000124) STAGA complex(GO:0030914) |
| 0.1 | 32.2 | GO:0030667 | secretory granule membrane(GO:0030667) |
| 0.1 | 16.9 | GO:0031093 | platelet alpha granule lumen(GO:0031093) |
| 0.1 | 10.4 | GO:0005811 | lipid particle(GO:0005811) |
| 0.1 | 3.1 | GO:0031932 | TORC2 complex(GO:0031932) |
| 0.1 | 262.8 | GO:0005887 | integral component of plasma membrane(GO:0005887) |
| 0.1 | 5.2 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.1 | 3.7 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.1 | 1.7 | GO:0030008 | TRAPP complex(GO:0030008) |
| 0.1 | 5.1 | GO:0000786 | nucleosome(GO:0000786) |
| 0.1 | 1.4 | GO:0044666 | MLL3/4 complex(GO:0044666) |
| 0.1 | 4.8 | GO:0016592 | mediator complex(GO:0016592) |
| 0.1 | 1.4 | GO:0060091 | kinocilium(GO:0060091) |
| 0.1 | 0.5 | GO:0034751 | aryl hydrocarbon receptor complex(GO:0034751) |
| 0.1 | 5.3 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.1 | 12.1 | GO:0016605 | PML body(GO:0016605) |
| 0.1 | 3.4 | GO:0031519 | PcG protein complex(GO:0031519) |
| 0.1 | 1.0 | GO:0097225 | sperm midpiece(GO:0097225) |
| 0.1 | 0.7 | GO:0005787 | signal peptidase complex(GO:0005787) |
| 0.1 | 0.5 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 0.1 | 8.3 | GO:0043235 | receptor complex(GO:0043235) |
| 0.1 | 30.7 | GO:0000790 | nuclear chromatin(GO:0000790) |
| 0.1 | 0.8 | GO:0005677 | chromatin silencing complex(GO:0005677) |
| 0.1 | 1.1 | GO:0008290 | F-actin capping protein complex(GO:0008290) |
| 0.1 | 3.1 | GO:0030864 | cortical actin cytoskeleton(GO:0030864) |
| 0.1 | 321.0 | GO:0016021 | integral component of membrane(GO:0016021) |
| 0.1 | 3.7 | GO:0005793 | endoplasmic reticulum-Golgi intermediate compartment(GO:0005793) |
| 0.1 | 2.2 | GO:0098852 | lysosomal membrane(GO:0005765) lytic vacuole membrane(GO:0098852) |
| 0.1 | 4.7 | GO:0000118 | histone deacetylase complex(GO:0000118) |
| 0.1 | 0.7 | GO:0031527 | filopodium membrane(GO:0031527) contractile ring(GO:0070938) |
| 0.0 | 9.0 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 7.2 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.0 | 4.7 | GO:0031965 | nuclear membrane(GO:0031965) |
| 0.0 | 4.3 | GO:0042470 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.0 | 0.6 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.0 | 0.7 | GO:0005776 | autophagosome(GO:0005776) |
| 0.0 | 2.2 | GO:0098791 | Golgi subcompartment(GO:0098791) |
| 0.0 | 1.9 | GO:0016235 | aggresome(GO:0016235) |
| 0.0 | 1.2 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.0 | 0.6 | GO:0032045 | guanyl-nucleotide exchange factor complex(GO:0032045) |
| 0.0 | 1.9 | GO:0001750 | photoreceptor outer segment(GO:0001750) |
| 0.0 | 3.2 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.0 | 4.3 | GO:0000777 | condensed chromosome kinetochore(GO:0000777) |
| 0.0 | 2.6 | GO:0044309 | dendritic spine(GO:0043197) neuron spine(GO:0044309) |
| 0.0 | 1.7 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 0.7 | GO:0046540 | U4/U6 x U5 tri-snRNP complex(GO:0046540) |
| 0.0 | 1.5 | GO:0000775 | chromosome, centromeric region(GO:0000775) |
| 0.0 | 0.2 | GO:0016602 | CCAAT-binding factor complex(GO:0016602) |
| 0.0 | 0.2 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.0 | 1.1 | GO:0000139 | Golgi membrane(GO:0000139) |
| 0.0 | 4.2 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.0 | 4.7 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.0 | 0.5 | GO:0005902 | microvillus(GO:0005902) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 12.8 | 38.4 | GO:0005136 | interleukin-4 receptor binding(GO:0005136) |
| 12.6 | 50.5 | GO:0050528 | acyloxyacyl hydrolase activity(GO:0050528) |
| 10.0 | 39.9 | GO:0004796 | thromboxane-A synthase activity(GO:0004796) 12-hydroxyheptadecatrienoic acid synthase activity(GO:0036134) |
| 8.0 | 40.1 | GO:0004914 | interleukin-5 receptor activity(GO:0004914) |
| 4.2 | 21.2 | GO:0031727 | CCR2 chemokine receptor binding(GO:0031727) |
| 4.0 | 47.6 | GO:0005094 | Rho GDP-dissociation inhibitor activity(GO:0005094) |
| 3.5 | 10.6 | GO:0004464 | leukotriene-C4 synthase activity(GO:0004464) |
| 3.3 | 10.0 | GO:0005150 | interleukin-1, Type I receptor binding(GO:0005150) |
| 3.2 | 28.7 | GO:0019763 | immunoglobulin receptor activity(GO:0019763) |
| 2.8 | 11.3 | GO:0004917 | interleukin-7 receptor activity(GO:0004917) |
| 2.6 | 13.1 | GO:0036435 | IkappaB kinase activity(GO:0008384) K48-linked polyubiquitin binding(GO:0036435) |
| 2.6 | 33.7 | GO:0051525 | NFAT protein binding(GO:0051525) |
| 2.5 | 19.9 | GO:0016314 | phosphatidylinositol-3,4,5-trisphosphate 3-phosphatase activity(GO:0016314) |
| 2.3 | 13.6 | GO:0004974 | leukotriene receptor activity(GO:0004974) |
| 2.3 | 15.9 | GO:0023030 | MHC class Ib protein complex binding(GO:0023025) MHC class Ib protein binding, via antigen binding groove(GO:0023030) |
| 2.2 | 13.3 | GO:0017159 | pantetheine hydrolase activity(GO:0017159) |
| 2.2 | 15.3 | GO:0016019 | peptidoglycan receptor activity(GO:0016019) |
| 2.1 | 25.4 | GO:0030369 | ICAM-3 receptor activity(GO:0030369) |
| 2.0 | 56.3 | GO:0001608 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 1.8 | 10.9 | GO:0030144 | alpha-1,6-mannosylglycoprotein 6-beta-N-acetylglucosaminyltransferase activity(GO:0030144) |
| 1.7 | 10.4 | GO:0001875 | lipopolysaccharide receptor activity(GO:0001875) |
| 1.6 | 19.6 | GO:0019864 | IgG binding(GO:0019864) |
| 1.6 | 7.8 | GO:0033829 | O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase activity(GO:0033829) |
| 1.5 | 6.1 | GO:0090556 | apolipoprotein A-I receptor activity(GO:0034188) phosphatidylserine-translocating ATPase activity(GO:0090556) |
| 1.5 | 17.6 | GO:0005168 | neurotrophin TRKA receptor binding(GO:0005168) |
| 1.5 | 13.2 | GO:0016812 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in cyclic amides(GO:0016812) |
| 1.3 | 17.3 | GO:0022820 | potassium:chloride symporter activity(GO:0015379) potassium ion symporter activity(GO:0022820) |
| 1.3 | 10.1 | GO:0005157 | macrophage colony-stimulating factor receptor binding(GO:0005157) |
| 1.3 | 50.2 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 1.2 | 24.5 | GO:0004716 | receptor signaling protein tyrosine kinase activity(GO:0004716) |
| 1.2 | 12.9 | GO:0035005 | 1-phosphatidylinositol-4-phosphate 3-kinase activity(GO:0035005) |
| 1.1 | 11.0 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 1.1 | 18.5 | GO:0019957 | C-C chemokine binding(GO:0019957) |
| 1.1 | 4.3 | GO:0045322 | unmethylated CpG binding(GO:0045322) |
| 1.0 | 16.3 | GO:0042608 | T cell receptor binding(GO:0042608) |
| 1.0 | 6.1 | GO:0039552 | RIG-I binding(GO:0039552) |
| 1.0 | 10.1 | GO:0008525 | phosphatidylcholine transporter activity(GO:0008525) |
| 1.0 | 21.9 | GO:0005344 | oxygen transporter activity(GO:0005344) |
| 1.0 | 17.6 | GO:0016176 | superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 1.0 | 52.1 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.9 | 17.5 | GO:0032036 | myosin heavy chain binding(GO:0032036) |
| 0.9 | 14.7 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.8 | 29.3 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.8 | 27.2 | GO:0019992 | diacylglycerol binding(GO:0019992) |
| 0.8 | 7.8 | GO:0055104 | ligase inhibitor activity(GO:0055104) ubiquitin ligase inhibitor activity(GO:1990948) |
| 0.8 | 4.6 | GO:0035800 | deubiquitinase activator activity(GO:0035800) |
| 0.8 | 4.5 | GO:0015333 | peptide:proton symporter activity(GO:0015333) proton-dependent peptide secondary active transmembrane transporter activity(GO:0022897) |
| 0.7 | 9.0 | GO:0036312 | phosphatidylinositol 3-kinase regulatory subunit binding(GO:0036312) |
| 0.7 | 5.5 | GO:0004534 | 5'-3' exoribonuclease activity(GO:0004534) |
| 0.7 | 6.8 | GO:0004985 | opioid receptor activity(GO:0004985) |
| 0.7 | 70.4 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.7 | 17.6 | GO:0008494 | translation activator activity(GO:0008494) |
| 0.6 | 11.4 | GO:0004875 | complement receptor activity(GO:0004875) |
| 0.5 | 7.7 | GO:0033192 | calmodulin-dependent protein phosphatase activity(GO:0033192) |
| 0.5 | 6.5 | GO:0070290 | N-acylphosphatidylethanolamine-specific phospholipase D activity(GO:0070290) |
| 0.5 | 20.2 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.5 | 3.2 | GO:0004609 | phosphatidylserine decarboxylase activity(GO:0004609) |
| 0.4 | 22.9 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.4 | 13.0 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.4 | 3.4 | GO:0033857 | diphosphoinositol-pentakisphosphate kinase activity(GO:0033857) |
| 0.4 | 3.7 | GO:0047623 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 0.4 | 1.9 | GO:0004666 | prostaglandin-endoperoxide synthase activity(GO:0004666) |
| 0.4 | 8.9 | GO:0015174 | basic amino acid transmembrane transporter activity(GO:0015174) |
| 0.4 | 4.5 | GO:0051434 | BH3 domain binding(GO:0051434) |
| 0.4 | 41.8 | GO:0001618 | virus receptor activity(GO:0001618) |
| 0.3 | 2.4 | GO:0005143 | interleukin-12 receptor binding(GO:0005143) |
| 0.3 | 5.8 | GO:0005537 | mannose binding(GO:0005537) |
| 0.3 | 150.6 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 0.3 | 6.2 | GO:0051864 | histone demethylase activity (H3-K36 specific)(GO:0051864) |
| 0.3 | 3.2 | GO:0004920 | interleukin-10 receptor activity(GO:0004920) |
| 0.3 | 4.2 | GO:0005078 | MAP-kinase scaffold activity(GO:0005078) |
| 0.3 | 4.3 | GO:0004571 | mannosyl-oligosaccharide 1,2-alpha-mannosidase activity(GO:0004571) mannosyl-oligosaccharide mannosidase activity(GO:0015924) |
| 0.3 | 4.8 | GO:0050291 | sphingosine N-acyltransferase activity(GO:0050291) |
| 0.3 | 2.0 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 0.3 | 19.3 | GO:0036002 | pre-mRNA binding(GO:0036002) |
| 0.3 | 3.7 | GO:0003831 | beta-N-acetylglucosaminylglycopeptide beta-1,4-galactosyltransferase activity(GO:0003831) |
| 0.2 | 5.5 | GO:0005225 | volume-sensitive anion channel activity(GO:0005225) |
| 0.2 | 8.2 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.2 | 2.5 | GO:0070735 | protein-glycine ligase activity(GO:0070735) |
| 0.2 | 14.5 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
| 0.2 | 0.8 | GO:0032184 | SUMO polymer binding(GO:0032184) |
| 0.2 | 18.7 | GO:0016279 | protein-lysine N-methyltransferase activity(GO:0016279) |
| 0.2 | 6.4 | GO:0001104 | RNA polymerase II transcription cofactor activity(GO:0001104) |
| 0.2 | 10.1 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.2 | 92.3 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.2 | 5.1 | GO:0005328 | neurotransmitter:sodium symporter activity(GO:0005328) |
| 0.2 | 4.0 | GO:0001222 | transcription corepressor binding(GO:0001222) |
| 0.2 | 5.9 | GO:0032452 | histone demethylase activity(GO:0032452) |
| 0.2 | 2.4 | GO:0030297 | transmembrane receptor protein tyrosine kinase activator activity(GO:0030297) |
| 0.1 | 5.8 | GO:0031683 | G-protein beta/gamma-subunit complex binding(GO:0031683) |
| 0.1 | 6.6 | GO:0051959 | dynein light intermediate chain binding(GO:0051959) |
| 0.1 | 4.8 | GO:0032041 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.1 | 0.5 | GO:0052591 | sn-glycerol-3-phosphate:ubiquinone oxidoreductase activity(GO:0052590) sn-glycerol-3-phosphate:ubiquinone-8 oxidoreductase activity(GO:0052591) |
| 0.1 | 0.9 | GO:0001042 | RNA polymerase I core binding(GO:0001042) |
| 0.1 | 4.3 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.1 | 4.7 | GO:0042288 | MHC class I protein binding(GO:0042288) |
| 0.1 | 10.9 | GO:0004521 | endoribonuclease activity(GO:0004521) |
| 0.1 | 0.5 | GO:0016435 | rRNA (guanine) methyltransferase activity(GO:0016435) |
| 0.1 | 2.2 | GO:0045295 | gamma-catenin binding(GO:0045295) |
| 0.1 | 3.7 | GO:0046961 | proton-transporting ATPase activity, rotational mechanism(GO:0046961) |
| 0.1 | 51.5 | GO:0003924 | GTPase activity(GO:0003924) |
| 0.1 | 7.9 | GO:0019843 | rRNA binding(GO:0019843) |
| 0.1 | 3.0 | GO:0102391 | decanoate--CoA ligase activity(GO:0102391) |
| 0.1 | 11.4 | GO:0001190 | transcriptional activator activity, RNA polymerase II transcription factor binding(GO:0001190) transcriptional repressor activity, RNA polymerase II activating transcription factor binding(GO:0098811) |
| 0.1 | 5.3 | GO:0016814 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in cyclic amidines(GO:0016814) |
| 0.1 | 2.1 | GO:0001011 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.1 | 1.4 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.1 | 1.2 | GO:0005068 | transmembrane receptor protein tyrosine kinase adaptor activity(GO:0005068) |
| 0.1 | 8.1 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.1 | 1.9 | GO:0035613 | RNA stem-loop binding(GO:0035613) |
| 0.1 | 2.9 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.1 | 2.0 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 0.1 | 2.1 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.1 | 9.5 | GO:0097110 | scaffold protein binding(GO:0097110) |
| 0.1 | 44.4 | GO:0005085 | guanyl-nucleotide exchange factor activity(GO:0005085) |
| 0.1 | 1.2 | GO:0015165 | pyrimidine nucleotide-sugar transmembrane transporter activity(GO:0015165) |
| 0.1 | 3.1 | GO:0005041 | low-density lipoprotein receptor activity(GO:0005041) |
| 0.1 | 2.4 | GO:0043395 | heparan sulfate proteoglycan binding(GO:0043395) |
| 0.1 | 2.0 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.1 | 3.3 | GO:0000983 | transcription factor activity, RNA polymerase II core promoter sequence-specific(GO:0000983) |
| 0.1 | 1.5 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.1 | 7.4 | GO:0001085 | RNA polymerase II transcription factor binding(GO:0001085) |
| 0.1 | 4.3 | GO:0043531 | ADP binding(GO:0043531) |
| 0.1 | 1.1 | GO:0031435 | mitogen-activated protein kinase kinase kinase binding(GO:0031435) |
| 0.1 | 2.6 | GO:0035035 | histone acetyltransferase binding(GO:0035035) |
| 0.1 | 1.4 | GO:0019789 | SUMO transferase activity(GO:0019789) |
| 0.1 | 6.5 | GO:0070491 | repressing transcription factor binding(GO:0070491) |
| 0.1 | 3.2 | GO:0061650 | ubiquitin conjugating enzyme activity(GO:0061631) ubiquitin-like protein conjugating enzyme activity(GO:0061650) |
| 0.1 | 1.7 | GO:0004435 | phosphatidylinositol phospholipase C activity(GO:0004435) |
| 0.1 | 3.1 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.1 | 0.5 | GO:0017162 | aryl hydrocarbon receptor binding(GO:0017162) |
| 0.0 | 1.3 | GO:0031434 | mitogen-activated protein kinase kinase binding(GO:0031434) |
| 0.0 | 3.2 | GO:0048365 | Rac GTPase binding(GO:0048365) |
| 0.0 | 4.7 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.0 | 79.4 | GO:0060089 | receptor activity(GO:0004872) molecular transducer activity(GO:0060089) |
| 0.0 | 1.8 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 0.2 | GO:0001032 | RNA polymerase III type 3 promoter DNA binding(GO:0001032) |
| 0.0 | 1.2 | GO:0008198 | ferrous iron binding(GO:0008198) |
| 0.0 | 0.5 | GO:0001055 | RNA polymerase II activity(GO:0001055) |
| 0.0 | 4.7 | GO:0005179 | hormone activity(GO:0005179) |
| 0.0 | 1.2 | GO:0004869 | cysteine-type endopeptidase inhibitor activity(GO:0004869) |
| 0.0 | 3.0 | GO:0003725 | double-stranded RNA binding(GO:0003725) |
| 0.0 | 1.5 | GO:0008170 | N-methyltransferase activity(GO:0008170) |
| 0.0 | 0.7 | GO:0017166 | vinculin binding(GO:0017166) |
| 0.0 | 3.9 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.0 | 0.9 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.0 | 0.8 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.0 | 0.4 | GO:0015232 | heme transporter activity(GO:0015232) |
| 0.0 | 3.3 | GO:0019888 | protein phosphatase regulator activity(GO:0019888) |
| 0.0 | 0.8 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.0 | 0.3 | GO:0004579 | dolichyl-diphosphooligosaccharide-protein glycotransferase activity(GO:0004579) |
| 0.0 | 0.3 | GO:0015386 | potassium:proton antiporter activity(GO:0015386) |
| 0.0 | 0.4 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.0 | 7.1 | GO:0001077 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
| 0.0 | 2.7 | GO:0005525 | GTP binding(GO:0005525) |
| 0.0 | 1.9 | GO:0035257 | nuclear hormone receptor binding(GO:0035257) |
| 0.0 | 0.6 | GO:0030552 | cAMP binding(GO:0030552) |
| 0.0 | 0.3 | GO:0008656 | cysteine-type endopeptidase activator activity involved in apoptotic process(GO:0008656) |
| 0.0 | 0.5 | GO:0015269 | calcium-activated potassium channel activity(GO:0015269) |
| 0.0 | 1.4 | GO:0005261 | cation channel activity(GO:0005261) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.7 | 57.7 | PID IL5 PATHWAY | IL5-mediated signaling events |
| 1.0 | 141.3 | PID BCR 5PATHWAY | BCR signaling pathway |
| 0.9 | 62.7 | PID IL8 CXCR2 PATHWAY | IL8- and CXCR2-mediated signaling events |
| 0.9 | 26.2 | SA PTEN PATHWAY | PTEN is a tumor suppressor that dephosphorylates the lipid messenger phosphatidylinositol triphosphate. |
| 0.9 | 65.5 | PID CD8 TCR PATHWAY | TCR signaling in naïve CD8+ T cells |
| 0.8 | 46.5 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
| 0.8 | 66.9 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.7 | 45.3 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.7 | 25.7 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.5 | 16.6 | PID TCR PATHWAY | TCR signaling in naïve CD4+ T cells |
| 0.5 | 14.7 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.4 | 37.7 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
| 0.4 | 20.4 | PID FCER1 PATHWAY | Fc-epsilon receptor I signaling in mast cells |
| 0.4 | 53.7 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.4 | 4.7 | PID AR NONGENOMIC PATHWAY | Nongenotropic Androgen signaling |
| 0.3 | 14.9 | PID IL6 7 PATHWAY | IL6-mediated signaling events |
| 0.3 | 10.0 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.3 | 9.1 | PID IL3 PATHWAY | IL3-mediated signaling events |
| 0.2 | 13.6 | PID ENDOTHELIN PATHWAY | Endothelins |
| 0.2 | 4.8 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.2 | 21.6 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.2 | 60.5 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.2 | 5.1 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.2 | 7.4 | PID IL12 2PATHWAY | IL12-mediated signaling events |
| 0.1 | 1.5 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.1 | 3.3 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.1 | 8.7 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.1 | 3.4 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
| 0.1 | 2.4 | PID TXA2PATHWAY | Thromboxane A2 receptor signaling |
| 0.1 | 10.7 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.1 | 1.6 | PID LYSOPHOSPHOLIPID PATHWAY | LPA receptor mediated events |
| 0.1 | 24.0 | NABA SECRETED FACTORS | Genes encoding secreted soluble factors |
| 0.0 | 3.2 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 1.7 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.0 | 2.4 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.0 | 0.9 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.0 | 1.7 | PID PDGFRB PATHWAY | PDGFR-beta signaling pathway |
| 0.0 | 1.4 | PID CDC42 PATHWAY | CDC42 signaling events |
| 0.0 | 1.6 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.0 | 1.3 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.0 | 1.3 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.0 | 0.8 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.3 | 56.3 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 1.9 | 28.7 | REACTOME PECAM1 INTERACTIONS | Genes involved in PECAM1 interactions |
| 1.9 | 55.1 | REACTOME CD28 DEPENDENT VAV1 PATHWAY | Genes involved in CD28 dependent Vav1 pathway |
| 1.6 | 38.8 | REACTOME ACTIVATION OF IRF3 IRF7 MEDIATED BY TBK1 IKK EPSILON | Genes involved in Activation of IRF3/IRF7 mediated by TBK1/IKK epsilon |
| 1.3 | 86.5 | REACTOME GPVI MEDIATED ACTIVATION CASCADE | Genes involved in GPVI-mediated activation cascade |
| 1.0 | 40.1 | REACTOME IL RECEPTOR SHC SIGNALING | Genes involved in Interleukin receptor SHC signaling |
| 0.9 | 10.1 | REACTOME REGULATION OF SIGNALING BY CBL | Genes involved in Regulation of signaling by CBL |
| 0.9 | 7.8 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.8 | 29.1 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 0.8 | 17.3 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.8 | 11.3 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.7 | 15.1 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.7 | 13.6 | REACTOME EICOSANOID LIGAND BINDING RECEPTORS | Genes involved in Eicosanoid ligand-binding receptors |
| 0.7 | 41.7 | REACTOME RAP1 SIGNALLING | Genes involved in Rap1 signalling |
| 0.6 | 2.5 | REACTOME CROSS PRESENTATION OF SOLUBLE EXOGENOUS ANTIGENS ENDOSOMES | Genes involved in Cross-presentation of soluble exogenous antigens (endosomes) |
| 0.6 | 73.5 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.6 | 42.2 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.5 | 39.9 | REACTOME CYTOCHROME P450 ARRANGED BY SUBSTRATE TYPE | Genes involved in Cytochrome P450 - arranged by substrate type |
| 0.4 | 5.8 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.4 | 9.0 | REACTOME SIGNALING BY SCF KIT | Genes involved in Signaling by SCF-KIT |
| 0.4 | 9.7 | REACTOME GRB2 SOS PROVIDES LINKAGE TO MAPK SIGNALING FOR INTERGRINS | Genes involved in GRB2:SOS provides linkage to MAPK signaling for Intergrins |
| 0.4 | 106.7 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.4 | 13.5 | REACTOME SEMA4D IN SEMAPHORIN SIGNALING | Genes involved in Sema4D in semaphorin signaling |
| 0.4 | 8.5 | REACTOME SIGNAL REGULATORY PROTEIN SIRP FAMILY INTERACTIONS | Genes involved in Signal regulatory protein (SIRP) family interactions |
| 0.3 | 16.9 | REACTOME THE ROLE OF NEF IN HIV1 REPLICATION AND DISEASE PATHOGENESIS | Genes involved in The role of Nef in HIV-1 replication and disease pathogenesis |
| 0.3 | 37.3 | REACTOME CELL SURFACE INTERACTIONS AT THE VASCULAR WALL | Genes involved in Cell surface interactions at the vascular wall |
| 0.3 | 17.6 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.3 | 31.4 | REACTOME INTERFERON GAMMA SIGNALING | Genes involved in Interferon gamma signaling |
| 0.3 | 3.6 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION IN TLR7 8 OR 9 SIGNALING | Genes involved in TRAF6 mediated IRF7 activation in TLR7/8 or 9 signaling |
| 0.3 | 2.4 | REACTOME SHC1 EVENTS IN EGFR SIGNALING | Genes involved in SHC1 events in EGFR signaling |
| 0.3 | 4.7 | REACTOME HORMONE LIGAND BINDING RECEPTORS | Genes involved in Hormone ligand-binding receptors |
| 0.3 | 4.5 | REACTOME INFLAMMASOMES | Genes involved in Inflammasomes |
| 0.3 | 23.4 | REACTOME G ALPHA Z SIGNALLING EVENTS | Genes involved in G alpha (z) signalling events |
| 0.3 | 8.1 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.3 | 5.5 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |
| 0.2 | 5.2 | REACTOME CTLA4 INHIBITORY SIGNALING | Genes involved in CTLA4 inhibitory signaling |
| 0.2 | 9.1 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.2 | 4.6 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION | Genes involved in TRAF6 mediated IRF7 activation |
| 0.2 | 5.1 | REACTOME NFKB ACTIVATION THROUGH FADD RIP1 PATHWAY MEDIATED BY CASPASE 8 AND10 | Genes involved in NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 |
| 0.2 | 16.3 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.2 | 2.0 | REACTOME SPRY REGULATION OF FGF SIGNALING | Genes involved in Spry regulation of FGF signaling |
| 0.2 | 23.9 | REACTOME REGULATION OF MRNA STABILITY BY PROTEINS THAT BIND AU RICH ELEMENTS | Genes involved in Regulation of mRNA Stability by Proteins that Bind AU-rich Elements |
| 0.2 | 32.7 | REACTOME G ALPHA Q SIGNALLING EVENTS | Genes involved in G alpha (q) signalling events |
| 0.2 | 5.7 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.2 | 5.8 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.2 | 5.1 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.2 | 6.4 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.2 | 4.3 | REACTOME SIGNALING BY FGFR1 FUSION MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |
| 0.1 | 30.1 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.1 | 14.6 | REACTOME SIGNALING BY ERBB2 | Genes involved in Signaling by ERBB2 |
| 0.1 | 3.3 | REACTOME LYSOSOME VESICLE BIOGENESIS | Genes involved in Lysosome Vesicle Biogenesis |
| 0.1 | 21.9 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.1 | 3.7 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.1 | 9.5 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.1 | 2.4 | REACTOME ACYL CHAIN REMODELLING OF PS | Genes involved in Acyl chain remodelling of PS |
| 0.1 | 4.4 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.1 | 5.1 | REACTOME RNA POL I PROMOTER OPENING | Genes involved in RNA Polymerase I Promoter Opening |
| 0.1 | 4.8 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.1 | 10.5 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.1 | 4.3 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.1 | 6.5 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.1 | 9.5 | REACTOME TOLL RECEPTOR CASCADES | Genes involved in Toll Receptor Cascades |
| 0.1 | 6.3 | REACTOME 3 UTR MEDIATED TRANSLATIONAL REGULATION | Genes involved in 3' -UTR-mediated translational regulation |
| 0.1 | 2.2 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.1 | 5.5 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.1 | 1.9 | REACTOME PHASE1 FUNCTIONALIZATION OF COMPOUNDS | Genes involved in Phase 1 - Functionalization of compounds |
| 0.1 | 0.8 | REACTOME REMOVAL OF THE FLAP INTERMEDIATE FROM THE C STRAND | Genes involved in Removal of the Flap Intermediate from the C-strand |
| 0.1 | 0.7 | REACTOME ER PHAGOSOME PATHWAY | Genes involved in ER-Phagosome pathway |
| 0.0 | 4.8 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.0 | 6.4 | REACTOME GLYCEROPHOSPHOLIPID BIOSYNTHESIS | Genes involved in Glycerophospholipid biosynthesis |
| 0.0 | 0.7 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.0 | 1.3 | REACTOME ABORTIVE ELONGATION OF HIV1 TRANSCRIPT IN THE ABSENCE OF TAT | Genes involved in Abortive elongation of HIV-1 transcript in the absence of Tat |
| 0.0 | 0.4 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.0 | 0.7 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.0 | 1.1 | REACTOME RIG I MDA5 MEDIATED INDUCTION OF IFN ALPHA BETA PATHWAYS | Genes involved in RIG-I/MDA5 mediated induction of IFN-alpha/beta pathways |
| 0.0 | 0.7 | REACTOME INSULIN RECEPTOR RECYCLING | Genes involved in Insulin receptor recycling |
| 0.0 | 0.7 | REACTOME MRNA SPLICING MINOR PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |
| 0.0 | 0.6 | REACTOME RNA POL III TRANSCRIPTION INITIATION FROM TYPE 3 PROMOTER | Genes involved in RNA Polymerase III Transcription Initiation From Type 3 Promoter |
| 0.0 | 1.2 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.0 | 0.7 | REACTOME REGULATION OF MITOTIC CELL CYCLE | Genes involved in Regulation of mitotic cell cycle |
| 0.0 | 0.5 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |
| 0.0 | 0.4 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |