Project
Illumina Body Map 2: averaged replicates
Navigation
Downloads
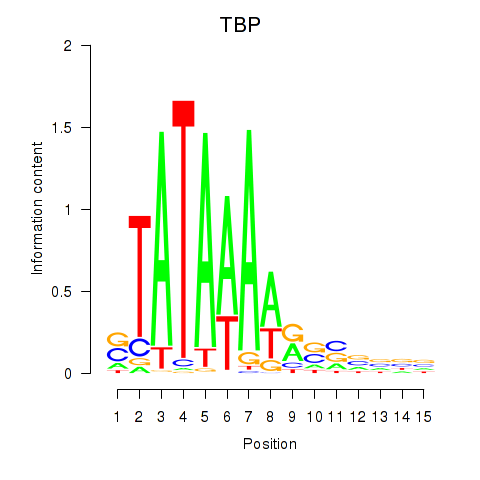
Results for TBP
Z-value: 4.63
Transcription factors associated with TBP
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
TBP
|
ENSG00000112592.8 | TATA-box binding protein |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| TBP | hg19_v2_chr6_+_170863421_170863484 | -0.18 | 3.3e-01 | Click! |
Activity profile of TBP motif
Sorted Z-values of TBP motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_-_59043166 | 11.92 |
ENST00000371225.2
|
TACSTD2
|
tumor-associated calcium signal transducer 2 |
| chr16_+_82068830 | 10.46 |
ENST00000199936.4
|
HSD17B2
|
hydroxysteroid (17-beta) dehydrogenase 2 |
| chr11_+_18287721 | 10.28 |
ENST00000356524.4
|
SAA1
|
serum amyloid A1 |
| chr11_+_18287801 | 10.25 |
ENST00000532858.1
ENST00000405158.2 |
SAA1
|
serum amyloid A1 |
| chr15_+_45722727 | 10.01 |
ENST00000396650.2
ENST00000558435.1 ENST00000344300.3 |
C15orf48
|
chromosome 15 open reading frame 48 |
| chr17_-_34345002 | 10.00 |
ENST00000293280.2
|
CCL23
|
chemokine (C-C motif) ligand 23 |
| chr1_+_159557607 | 9.55 |
ENST00000255040.2
|
APCS
|
amyloid P component, serum |
| chr7_+_142829162 | 9.48 |
ENST00000291009.3
|
PIP
|
prolactin-induced protein |
| chr19_+_46367518 | 8.42 |
ENST00000302177.2
|
FOXA3
|
forkhead box A3 |
| chr17_-_34344991 | 8.40 |
ENST00000591423.1
|
CCL23
|
chemokine (C-C motif) ligand 23 |
| chr21_-_43735628 | 7.93 |
ENST00000291525.10
ENST00000518498.1 |
TFF3
|
trefoil factor 3 (intestinal) |
| chr11_-_5248294 | 7.84 |
ENST00000335295.4
|
HBB
|
hemoglobin, beta |
| chr11_-_18270182 | 7.71 |
ENST00000528349.1
ENST00000526900.1 ENST00000529528.1 ENST00000414546.2 ENST00000256733.4 |
SAA2
|
serum amyloid A2 |
| chr3_-_165555200 | 7.57 |
ENST00000479451.1
ENST00000540653.1 ENST00000488954.1 ENST00000264381.3 |
BCHE
|
butyrylcholinesterase |
| chr21_-_43735446 | 7.29 |
ENST00000398431.2
|
TFF3
|
trefoil factor 3 (intestinal) |
| chr8_-_6735451 | 7.21 |
ENST00000297439.3
|
DEFB1
|
defensin, beta 1 |
| chr16_+_82068585 | 7.15 |
ENST00000563491.1
|
HSD17B2
|
hydroxysteroid (17-beta) dehydrogenase 2 |
| chr1_-_169703203 | 7.13 |
ENST00000333360.7
ENST00000367781.4 ENST00000367782.4 ENST00000367780.4 ENST00000367779.4 |
SELE
|
selectin E |
| chr11_+_43964055 | 6.86 |
ENST00000528572.1
|
C11orf96
|
chromosome 11 open reading frame 96 |
| chr16_+_82068873 | 6.78 |
ENST00000566213.1
|
HSD17B2
|
hydroxysteroid (17-beta) dehydrogenase 2 |
| chr3_+_148583043 | 6.76 |
ENST00000296046.3
|
CPA3
|
carboxypeptidase A3 (mast cell) |
| chr18_-_61329118 | 6.72 |
ENST00000332821.8
ENST00000283752.5 |
SERPINB3
|
serpin peptidase inhibitor, clade B (ovalbumin), member 3 |
| chr6_+_25727046 | 6.69 |
ENST00000274764.2
|
HIST1H2BA
|
histone cluster 1, H2ba |
| chr10_-_81320151 | 6.65 |
ENST00000372325.2
ENST00000372327.5 ENST00000417041.1 |
SFTPA2
|
surfactant protein A2 |
| chr12_+_13044787 | 6.65 |
ENST00000534831.1
|
GPRC5A
|
G protein-coupled receptor, family C, group 5, member A |
| chr17_-_39661849 | 6.63 |
ENST00000246635.3
ENST00000336861.3 ENST00000587544.1 ENST00000587435.1 |
KRT13
|
keratin 13 |
| chr9_+_117092149 | 6.59 |
ENST00000431067.2
ENST00000412657.1 |
ORM2
|
orosomucoid 2 |
| chr2_-_151344172 | 6.54 |
ENST00000375734.2
ENST00000263895.4 ENST00000454202.1 |
RND3
|
Rho family GTPase 3 |
| chr9_+_130911723 | 6.44 |
ENST00000277480.2
ENST00000373013.2 ENST00000540948.1 |
LCN2
|
lipocalin 2 |
| chr3_+_149192475 | 6.22 |
ENST00000465758.1
|
TM4SF4
|
transmembrane 4 L six family member 4 |
| chr17_-_39780634 | 6.10 |
ENST00000577817.2
|
KRT17
|
keratin 17 |
| chr19_+_11650709 | 6.06 |
ENST00000586059.1
|
CNN1
|
calponin 1, basic, smooth muscle |
| chr11_-_102651343 | 5.97 |
ENST00000279441.4
ENST00000539681.1 |
MMP10
|
matrix metallopeptidase 10 (stromelysin 2) |
| chr17_-_39780819 | 5.89 |
ENST00000311208.8
|
KRT17
|
keratin 17 |
| chr20_+_3052264 | 5.83 |
ENST00000217386.2
|
OXT
|
oxytocin/neurophysin I prepropeptide |
| chr5_-_42812143 | 5.81 |
ENST00000514985.1
|
SEPP1
|
selenoprotein P, plasma, 1 |
| chr13_+_53602894 | 5.74 |
ENST00000219022.2
|
OLFM4
|
olfactomedin 4 |
| chr16_+_2880369 | 5.74 |
ENST00000572863.1
|
ZG16B
|
zymogen granule protein 16B |
| chr17_-_48278983 | 5.62 |
ENST00000225964.5
|
COL1A1
|
collagen, type I, alpha 1 |
| chr17_+_32683456 | 5.59 |
ENST00000225844.2
|
CCL13
|
chemokine (C-C motif) ligand 13 |
| chr9_+_130911770 | 5.53 |
ENST00000372998.1
|
LCN2
|
lipocalin 2 |
| chr10_+_96698406 | 5.52 |
ENST00000260682.6
|
CYP2C9
|
cytochrome P450, family 2, subfamily C, polypeptide 9 |
| chr5_+_78407602 | 5.32 |
ENST00000274353.5
ENST00000524080.1 |
BHMT
|
betaine--homocysteine S-methyltransferase |
| chr15_-_62457480 | 5.31 |
ENST00000380392.3
|
C2CD4B
|
C2 calcium-dependent domain containing 4B |
| chr14_-_25045446 | 5.29 |
ENST00000216336.2
|
CTSG
|
cathepsin G |
| chr21_-_43786634 | 5.29 |
ENST00000291527.2
|
TFF1
|
trefoil factor 1 |
| chr1_-_168698433 | 5.26 |
ENST00000367817.3
|
DPT
|
dermatopontin |
| chrX_-_101771645 | 5.18 |
ENST00000289373.4
|
TMSB15A
|
thymosin beta 15a |
| chr11_-_5255861 | 4.92 |
ENST00000380299.3
|
HBD
|
hemoglobin, delta |
| chr20_+_31823792 | 4.88 |
ENST00000375413.4
ENST00000354297.4 ENST00000375422.2 |
BPIFA1
|
BPI fold containing family A, member 1 |
| chr12_-_11463353 | 4.85 |
ENST00000279575.1
ENST00000535904.1 ENST00000445719.2 |
PRB4
|
proline-rich protein BstNI subfamily 4 |
| chr20_-_43883197 | 4.81 |
ENST00000338380.2
|
SLPI
|
secretory leukocyte peptidase inhibitor |
| chr17_+_32612687 | 4.80 |
ENST00000305869.3
|
CCL11
|
chemokine (C-C motif) ligand 11 |
| chr11_-_102714534 | 4.79 |
ENST00000299855.5
|
MMP3
|
matrix metallopeptidase 3 (stromelysin 1, progelatinase) |
| chr11_-_5255696 | 4.76 |
ENST00000292901.3
ENST00000417377.1 |
HBD
|
hemoglobin, delta |
| chr12_-_11508520 | 4.72 |
ENST00000545626.1
ENST00000500254.2 |
PRB1
|
proline-rich protein BstNI subfamily 1 |
| chr3_+_193853927 | 4.63 |
ENST00000232424.3
|
HES1
|
hes family bHLH transcription factor 1 |
| chr17_-_39781054 | 4.57 |
ENST00000463128.1
|
KRT17
|
keratin 17 |
| chr17_-_48546232 | 4.53 |
ENST00000258969.4
|
CHAD
|
chondroadherin |
| chr6_-_25726781 | 4.48 |
ENST00000297012.3
|
HIST1H2AA
|
histone cluster 1, H2aa |
| chr5_-_42811986 | 4.47 |
ENST00000511224.1
ENST00000507920.1 ENST00000510965.1 |
SEPP1
|
selenoprotein P, plasma, 1 |
| chr17_-_48546324 | 4.46 |
ENST00000508540.1
|
CHAD
|
chondroadherin |
| chr20_+_30193083 | 4.41 |
ENST00000376112.3
ENST00000376105.3 |
ID1
|
inhibitor of DNA binding 1, dominant negative helix-loop-helix protein |
| chr16_+_78056412 | 4.09 |
ENST00000299642.4
ENST00000575655.1 |
CLEC3A
|
C-type lectin domain family 3, member A |
| chr4_-_74904398 | 4.09 |
ENST00000296026.4
|
CXCL3
|
chemokine (C-X-C motif) ligand 3 |
| chr11_-_116694009 | 4.07 |
ENST00000357780.3
|
APOA4
|
apolipoprotein A-IV |
| chr16_+_2014941 | 3.88 |
ENST00000531523.1
|
SNHG9
|
small nucleolar RNA host gene 9 (non-protein coding) |
| chr11_-_2950642 | 3.86 |
ENST00000314222.4
|
PHLDA2
|
pleckstrin homology-like domain, family A, member 2 |
| chr19_-_36247910 | 3.80 |
ENST00000587965.1
ENST00000004982.3 |
HSPB6
|
heat shock protein, alpha-crystallin-related, B6 |
| chr12_+_50355647 | 3.77 |
ENST00000293599.6
|
AQP5
|
aquaporin 5 |
| chr12_-_15038779 | 3.73 |
ENST00000228938.5
ENST00000539261.1 |
MGP
|
matrix Gla protein |
| chr1_-_159684371 | 3.72 |
ENST00000255030.5
ENST00000437342.1 ENST00000368112.1 ENST00000368111.1 ENST00000368110.1 ENST00000343919.2 |
CRP
|
C-reactive protein, pentraxin-related |
| chr16_+_2880157 | 3.71 |
ENST00000382280.3
|
ZG16B
|
zymogen granule protein 16B |
| chr1_+_86046433 | 3.65 |
ENST00000451137.2
|
CYR61
|
cysteine-rich, angiogenic inducer, 61 |
| chr1_+_161228517 | 3.64 |
ENST00000504449.1
|
PCP4L1
|
Purkinje cell protein 4 like 1 |
| chr17_-_39661947 | 3.64 |
ENST00000590425.1
|
KRT13
|
keratin 13 |
| chr9_-_33447584 | 3.61 |
ENST00000297991.4
|
AQP3
|
aquaporin 3 (Gill blood group) |
| chr1_-_161193349 | 3.57 |
ENST00000469730.2
ENST00000463273.1 ENST00000464492.1 ENST00000367990.3 ENST00000470459.2 ENST00000468465.1 ENST00000463812.1 |
APOA2
|
apolipoprotein A-II |
| chr15_-_41166414 | 3.54 |
ENST00000220507.4
|
RHOV
|
ras homolog family member V |
| chr16_+_57406368 | 3.52 |
ENST00000006053.6
ENST00000563383.1 |
CX3CL1
|
chemokine (C-X3-C motif) ligand 1 |
| chr1_+_152881014 | 3.49 |
ENST00000368764.3
ENST00000392667.2 |
IVL
|
involucrin |
| chr11_+_68451943 | 3.47 |
ENST00000265643.3
|
GAL
|
galanin/GMAP prepropeptide |
| chr15_-_50558223 | 3.42 |
ENST00000267845.3
|
HDC
|
histidine decarboxylase |
| chr8_+_66955648 | 3.39 |
ENST00000522619.1
|
DNAJC5B
|
DnaJ (Hsp40) homolog, subfamily C, member 5 beta |
| chr2_-_85895295 | 3.36 |
ENST00000428225.1
ENST00000519937.2 |
SFTPB
|
surfactant protein B |
| chr6_+_71104588 | 3.34 |
ENST00000418403.1
|
RP11-462G2.1
|
RP11-462G2.1 |
| chr1_-_117753540 | 3.33 |
ENST00000328189.3
ENST00000369458.3 |
VTCN1
|
V-set domain containing T cell activation inhibitor 1 |
| chr19_-_6720686 | 3.32 |
ENST00000245907.6
|
C3
|
complement component 3 |
| chr12_-_91539918 | 3.29 |
ENST00000548218.1
|
DCN
|
decorin |
| chr12_-_11036844 | 3.27 |
ENST00000428168.2
|
PRH1
|
proline-rich protein HaeIII subfamily 1 |
| chr12_+_11081828 | 3.25 |
ENST00000381847.3
ENST00000396400.3 |
PRH2
|
proline-rich protein HaeIII subfamily 2 |
| chrX_-_45629661 | 3.24 |
ENST00000602507.1
ENST00000602461.1 |
RP6-99M1.2
|
RP6-99M1.2 |
| chr4_+_74702214 | 3.22 |
ENST00000226317.5
ENST00000515050.1 |
CXCL6
|
chemokine (C-X-C motif) ligand 6 |
| chr2_-_89521942 | 3.19 |
ENST00000482769.1
|
IGKV2-28
|
immunoglobulin kappa variable 2-28 |
| chr22_+_23029188 | 3.15 |
ENST00000390305.2
|
IGLV3-25
|
immunoglobulin lambda variable 3-25 |
| chr3_+_142342228 | 3.13 |
ENST00000337777.3
|
PLS1
|
plastin 1 |
| chrX_-_105282712 | 3.12 |
ENST00000372563.1
|
SERPINA7
|
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 7 |
| chr5_+_40909354 | 3.10 |
ENST00000313164.9
|
C7
|
complement component 7 |
| chr2_-_89292422 | 3.07 |
ENST00000495489.1
|
IGKV1-8
|
immunoglobulin kappa variable 1-8 |
| chr11_-_2160611 | 3.07 |
ENST00000416167.2
|
IGF2
|
insulin-like growth factor 2 (somatomedin A) |
| chr9_-_34691201 | 3.05 |
ENST00000378800.3
ENST00000311925.2 |
CCL19
|
chemokine (C-C motif) ligand 19 |
| chr10_-_90712520 | 3.03 |
ENST00000224784.6
|
ACTA2
|
actin, alpha 2, smooth muscle, aorta |
| chr10_+_7745232 | 3.00 |
ENST00000358415.4
|
ITIH2
|
inter-alpha-trypsin inhibitor heavy chain 2 |
| chr12_-_21757774 | 2.99 |
ENST00000261195.2
|
GYS2
|
glycogen synthase 2 (liver) |
| chr2_+_135596106 | 2.96 |
ENST00000356140.5
|
ACMSD
|
aminocarboxymuconate semialdehyde decarboxylase |
| chr17_-_38545799 | 2.96 |
ENST00000577541.1
|
TOP2A
|
topoisomerase (DNA) II alpha 170kDa |
| chr18_+_56887381 | 2.94 |
ENST00000256857.2
ENST00000529320.2 ENST00000420468.2 |
GRP
|
gastrin-releasing peptide |
| chr20_+_36932521 | 2.92 |
ENST00000262865.4
|
BPI
|
bactericidal/permeability-increasing protein |
| chr16_+_56685796 | 2.91 |
ENST00000334346.2
ENST00000562399.1 |
MT1B
|
metallothionein 1B |
| chr1_-_209825674 | 2.91 |
ENST00000367030.3
ENST00000356082.4 |
LAMB3
|
laminin, beta 3 |
| chr2_-_89327228 | 2.91 |
ENST00000483158.1
|
IGKV3-11
|
immunoglobulin kappa variable 3-11 |
| chr4_-_110723134 | 2.89 |
ENST00000510800.1
ENST00000512148.1 |
CFI
|
complement factor I |
| chr4_-_110723194 | 2.89 |
ENST00000394635.3
|
CFI
|
complement factor I |
| chr11_-_2160180 | 2.89 |
ENST00000381406.4
|
IGF2
|
insulin-like growth factor 2 (somatomedin A) |
| chr1_+_31885963 | 2.88 |
ENST00000373709.3
|
SERINC2
|
serine incorporator 2 |
| chr16_+_57406437 | 2.87 |
ENST00000564948.1
|
CX3CL1
|
chemokine (C-X3-C motif) ligand 1 |
| chr1_+_192544857 | 2.86 |
ENST00000367459.3
ENST00000469578.2 |
RGS1
|
regulator of G-protein signaling 1 |
| chr6_-_133035185 | 2.84 |
ENST00000367928.4
|
VNN1
|
vanin 1 |
| chr10_+_124320156 | 2.83 |
ENST00000338354.3
ENST00000344338.3 ENST00000330163.4 ENST00000368909.3 ENST00000368955.3 ENST00000368956.2 |
DMBT1
|
deleted in malignant brain tumors 1 |
| chr6_-_26235206 | 2.78 |
ENST00000244534.5
|
HIST1H1D
|
histone cluster 1, H1d |
| chr8_-_23021533 | 2.77 |
ENST00000312584.3
|
TNFRSF10D
|
tumor necrosis factor receptor superfamily, member 10d, decoy with truncated death domain |
| chr22_+_39916558 | 2.76 |
ENST00000337304.2
ENST00000396680.1 |
ATF4
|
activating transcription factor 4 |
| chr2_-_190044480 | 2.75 |
ENST00000374866.3
|
COL5A2
|
collagen, type V, alpha 2 |
| chr19_+_840963 | 2.75 |
ENST00000234347.5
|
PRTN3
|
proteinase 3 |
| chr14_-_106478603 | 2.73 |
ENST00000390596.2
|
IGHV4-4
|
immunoglobulin heavy variable 4-4 |
| chr17_-_38574169 | 2.72 |
ENST00000423485.1
|
TOP2A
|
topoisomerase (DNA) II alpha 170kDa |
| chr3_-_170587974 | 2.71 |
ENST00000463836.1
|
RPL22L1
|
ribosomal protein L22-like 1 |
| chr15_-_50557863 | 2.69 |
ENST00000543581.1
|
HDC
|
histidine decarboxylase |
| chr4_-_110723335 | 2.66 |
ENST00000394634.2
|
CFI
|
complement factor I |
| chr2_-_238305397 | 2.64 |
ENST00000409809.1
|
COL6A3
|
collagen, type VI, alpha 3 |
| chr16_-_15950868 | 2.61 |
ENST00000396324.3
ENST00000452625.2 ENST00000576790.2 ENST00000300036.5 |
MYH11
|
myosin, heavy chain 11, smooth muscle |
| chr10_+_7745303 | 2.61 |
ENST00000429820.1
ENST00000379587.4 |
ITIH2
|
inter-alpha-trypsin inhibitor heavy chain 2 |
| chr3_-_170588163 | 2.60 |
ENST00000295830.8
|
RPL22L1
|
ribosomal protein L22-like 1 |
| chr4_-_100140331 | 2.56 |
ENST00000407820.2
ENST00000394897.1 ENST00000508558.1 ENST00000394899.2 |
ADH6
|
alcohol dehydrogenase 6 (class V) |
| chr1_-_8086343 | 2.54 |
ENST00000474874.1
ENST00000469499.1 ENST00000377482.5 |
ERRFI1
|
ERBB receptor feedback inhibitor 1 |
| chr8_-_6914251 | 2.54 |
ENST00000330590.2
|
DEFA5
|
defensin, alpha 5, Paneth cell-specific |
| chr10_-_99771079 | 2.52 |
ENST00000309155.3
|
CRTAC1
|
cartilage acidic protein 1 |
| chr19_+_10381769 | 2.52 |
ENST00000423829.2
ENST00000588645.1 |
ICAM1
|
intercellular adhesion molecule 1 |
| chr7_-_94285472 | 2.51 |
ENST00000437425.2
ENST00000447873.1 ENST00000415788.2 |
SGCE
|
sarcoglycan, epsilon |
| chr17_-_39041479 | 2.49 |
ENST00000167588.3
|
KRT20
|
keratin 20 |
| chr2_+_90211643 | 2.49 |
ENST00000390277.2
|
IGKV3D-11
|
immunoglobulin kappa variable 3D-11 |
| chr10_+_96522361 | 2.48 |
ENST00000371321.3
|
CYP2C19
|
cytochrome P450, family 2, subfamily C, polypeptide 19 |
| chr3_-_100712352 | 2.47 |
ENST00000471714.1
ENST00000284322.5 |
ABI3BP
|
ABI family, member 3 (NESH) binding protein |
| chr7_-_94285511 | 2.47 |
ENST00000265735.7
|
SGCE
|
sarcoglycan, epsilon |
| chr11_-_102826434 | 2.46 |
ENST00000340273.4
ENST00000260302.3 |
MMP13
|
matrix metallopeptidase 13 (collagenase 3) |
| chr3_-_170587815 | 2.44 |
ENST00000466674.1
|
RPL22L1
|
ribosomal protein L22-like 1 |
| chr12_-_120765565 | 2.42 |
ENST00000423423.3
ENST00000308366.4 |
PLA2G1B
|
phospholipase A2, group IB (pancreas) |
| chr15_-_63674034 | 2.41 |
ENST00000344366.3
ENST00000422263.2 |
CA12
|
carbonic anhydrase XII |
| chr2_-_163008903 | 2.41 |
ENST00000418842.2
ENST00000375497.3 |
GCG
|
glucagon |
| chr1_-_156675535 | 2.40 |
ENST00000368221.1
|
CRABP2
|
cellular retinoic acid binding protein 2 |
| chr11_+_1074870 | 2.40 |
ENST00000441003.2
ENST00000359061.5 |
MUC2
|
mucin 2, oligomeric mucus/gel-forming |
| chr11_-_5276008 | 2.40 |
ENST00000336906.4
|
HBG2
|
hemoglobin, gamma G |
| chr22_-_39715600 | 2.38 |
ENST00000427905.1
ENST00000402527.1 ENST00000216146.4 |
RPL3
|
ribosomal protein L3 |
| chr19_+_49468558 | 2.37 |
ENST00000331825.6
|
FTL
|
ferritin, light polypeptide |
| chr2_-_75426183 | 2.37 |
ENST00000409848.3
|
TACR1
|
tachykinin receptor 1 |
| chr17_+_77021702 | 2.36 |
ENST00000392445.2
ENST00000354124.3 |
C1QTNF1
|
C1q and tumor necrosis factor related protein 1 |
| chr8_+_99956759 | 2.36 |
ENST00000522510.1
ENST00000457907.2 |
OSR2
|
odd-skipped related transciption factor 2 |
| chr21_+_45905448 | 2.35 |
ENST00000449713.1
|
AP001065.15
|
AP001065.15 |
| chr14_-_55369525 | 2.35 |
ENST00000543643.2
ENST00000536224.2 ENST00000395514.1 ENST00000491895.2 |
GCH1
|
GTP cyclohydrolase 1 |
| chr12_+_21284118 | 2.35 |
ENST00000256958.2
|
SLCO1B1
|
solute carrier organic anion transporter family, member 1B1 |
| chr7_-_41742697 | 2.33 |
ENST00000242208.4
|
INHBA
|
inhibin, beta A |
| chr7_+_75931861 | 2.32 |
ENST00000248553.6
|
HSPB1
|
heat shock 27kDa protein 1 |
| chr14_-_47120956 | 2.31 |
ENST00000298283.3
|
RPL10L
|
ribosomal protein L10-like |
| chr3_-_194090460 | 2.31 |
ENST00000428839.1
ENST00000347624.3 |
LRRC15
|
leucine rich repeat containing 15 |
| chr16_+_2880254 | 2.29 |
ENST00000570670.1
|
ZG16B
|
zymogen granule protein 16B |
| chr12_-_52867569 | 2.27 |
ENST00000252250.6
|
KRT6C
|
keratin 6C |
| chr1_-_153518270 | 2.26 |
ENST00000354332.4
ENST00000368716.4 |
S100A4
|
S100 calcium binding protein A4 |
| chr2_+_135596180 | 2.25 |
ENST00000283054.4
ENST00000392928.1 |
ACMSD
|
aminocarboxymuconate semialdehyde decarboxylase |
| chr5_-_169816638 | 2.24 |
ENST00000521859.1
ENST00000274629.4 |
KCNMB1
|
potassium large conductance calcium-activated channel, subfamily M, beta member 1 |
| chr1_+_37940153 | 2.23 |
ENST00000373087.6
|
ZC3H12A
|
zinc finger CCCH-type containing 12A |
| chr10_+_5135981 | 2.21 |
ENST00000380554.3
|
AKR1C3
|
aldo-keto reductase family 1, member C3 |
| chr17_+_56315787 | 2.21 |
ENST00000262290.4
ENST00000421678.2 |
LPO
|
lactoperoxidase |
| chr7_-_7575477 | 2.20 |
ENST00000399429.3
|
COL28A1
|
collagen, type XXVIII, alpha 1 |
| chr12_-_92539614 | 2.17 |
ENST00000256015.3
|
BTG1
|
B-cell translocation gene 1, anti-proliferative |
| chr16_+_202686 | 2.16 |
ENST00000252951.2
|
HBZ
|
hemoglobin, zeta |
| chr6_+_33239787 | 2.16 |
ENST00000439602.2
ENST00000474973.1 |
RPS18
|
ribosomal protein S18 |
| chr4_+_129730779 | 2.15 |
ENST00000226319.6
|
PHF17
|
jade family PHD finger 1 |
| chr12_-_53074182 | 2.15 |
ENST00000252244.3
|
KRT1
|
keratin 1 |
| chr11_-_2182388 | 2.13 |
ENST00000421783.1
ENST00000397262.1 ENST00000250971.3 ENST00000381330.4 ENST00000397270.1 |
INS
INS-IGF2
|
insulin INS-IGF2 readthrough |
| chr16_+_71392616 | 2.10 |
ENST00000349553.5
ENST00000302628.4 ENST00000562305.1 |
CALB2
|
calbindin 2 |
| chr1_+_55446465 | 2.10 |
ENST00000371268.3
|
TMEM61
|
transmembrane protein 61 |
| chr10_+_22634384 | 2.10 |
ENST00000376624.3
ENST00000376603.2 ENST00000376601.1 ENST00000538630.1 ENST00000456231.2 ENST00000313311.6 ENST00000435326.1 |
SPAG6
|
sperm associated antigen 6 |
| chr1_-_935361 | 2.08 |
ENST00000484667.2
|
HES4
|
hes family bHLH transcription factor 4 |
| chr17_+_32646055 | 2.08 |
ENST00000394620.1
|
CCL8
|
chemokine (C-C motif) ligand 8 |
| chr10_+_54074033 | 2.07 |
ENST00000373970.3
|
DKK1
|
dickkopf WNT signaling pathway inhibitor 1 |
| chr16_+_56659687 | 2.07 |
ENST00000568293.1
ENST00000330439.6 |
MT1E
|
metallothionein 1E |
| chr10_+_124320195 | 2.06 |
ENST00000359586.6
|
DMBT1
|
deleted in malignant brain tumors 1 |
| chr19_-_14628645 | 2.05 |
ENST00000598235.1
|
DNAJB1
|
DnaJ (Hsp40) homolog, subfamily B, member 1 |
| chr7_-_94285402 | 2.05 |
ENST00000428696.2
ENST00000445866.2 |
SGCE
|
sarcoglycan, epsilon |
| chr17_-_38859996 | 2.05 |
ENST00000264651.2
|
KRT24
|
keratin 24 |
| chr12_+_56435637 | 2.01 |
ENST00000356464.5
ENST00000552361.1 |
RPS26
|
ribosomal protein S26 |
| chr6_-_27114577 | 1.99 |
ENST00000356950.1
ENST00000396891.4 |
HIST1H2BK
|
histone cluster 1, H2bk |
| chr9_-_104198042 | 1.97 |
ENST00000374855.4
|
ALDOB
|
aldolase B, fructose-bisphosphate |
| chr12_+_50344516 | 1.97 |
ENST00000199280.3
ENST00000550862.1 |
AQP2
|
aquaporin 2 (collecting duct) |
| chr5_-_153857819 | 1.96 |
ENST00000231121.2
|
HAND1
|
heart and neural crest derivatives expressed 1 |
| chr12_-_11548496 | 1.94 |
ENST00000389362.4
ENST00000565533.1 ENST00000546254.1 |
PRB2
PRB1
|
proline-rich protein BstNI subfamily 2 proline-rich protein BstNI subfamily 1 |
| chr9_+_136501478 | 1.94 |
ENST00000393056.2
ENST00000263611.2 |
DBH
|
dopamine beta-hydroxylase (dopamine beta-monooxygenase) |
| chr17_-_39674668 | 1.92 |
ENST00000393981.3
|
KRT15
|
keratin 15 |
| chr16_+_67840986 | 1.92 |
ENST00000561639.1
ENST00000567852.1 ENST00000565148.1 ENST00000388833.3 ENST00000561654.1 ENST00000431934.2 |
TSNAXIP1
|
translin-associated factor X interacting protein 1 |
| chr12_-_52845910 | 1.91 |
ENST00000252252.3
|
KRT6B
|
keratin 6B |
Network of associatons between targets according to the STRING database.
First level regulatory network of TBP
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.5 | 7.6 | GO:0014016 | neuroblast differentiation(GO:0014016) |
| 2.4 | 12.0 | GO:0015688 | iron chelate transport(GO:0015688) siderophore transport(GO:0015891) |
| 2.2 | 9.0 | GO:1900154 | regulation of bone trabecula formation(GO:1900154) negative regulation of bone trabecula formation(GO:1900155) |
| 2.1 | 6.4 | GO:0051040 | regulation of calcium-independent cell-cell adhesion(GO:0051040) |
| 2.0 | 6.1 | GO:0001694 | histamine biosynthetic process(GO:0001694) |
| 2.0 | 7.8 | GO:0030185 | nitric oxide transport(GO:0030185) |
| 1.9 | 5.8 | GO:0002125 | maternal aggressive behavior(GO:0002125) positive regulation of female receptivity(GO:0045925) |
| 1.9 | 5.6 | GO:0090265 | positive regulation of immune complex clearance by monocytes and macrophages(GO:0090265) |
| 1.8 | 5.3 | GO:0006579 | amino-acid betaine catabolic process(GO:0006579) |
| 1.7 | 6.7 | GO:0035425 | autocrine signaling(GO:0035425) |
| 1.5 | 4.6 | GO:0060164 | auditory receptor cell fate determination(GO:0042668) negative regulation of auditory receptor cell differentiation(GO:0045608) regulation of timing of neuron differentiation(GO:0060164) negative regulation of pro-B cell differentiation(GO:2000974) |
| 1.5 | 11.9 | GO:0090191 | negative regulation of branching involved in ureteric bud morphogenesis(GO:0090191) |
| 1.5 | 4.4 | GO:0010621 | negative regulation of transcription by transcription factor localization(GO:0010621) |
| 1.4 | 7.2 | GO:0060474 | positive regulation of sperm motility involved in capacitation(GO:0060474) |
| 1.4 | 5.6 | GO:0044691 | tooth eruption(GO:0044691) |
| 1.4 | 9.6 | GO:0052199 | negative regulation of catalytic activity in other organism involved in symbiotic interaction(GO:0052199) |
| 1.3 | 5.2 | GO:0010903 | negative regulation of very-low-density lipoprotein particle remodeling(GO:0010903) |
| 1.3 | 3.8 | GO:0015670 | carbon dioxide transport(GO:0015670) |
| 1.2 | 8.6 | GO:0016098 | monoterpenoid metabolic process(GO:0016098) |
| 1.2 | 4.9 | GO:1900228 | biofilm formation(GO:0042710) single-species biofilm formation(GO:0044010) single-species biofilm formation in or on host organism(GO:0044407) regulation of single-species biofilm formation(GO:1900190) negative regulation of single-species biofilm formation(GO:1900191) regulation of single-species biofilm formation in or on host organism(GO:1900228) negative regulation of single-species biofilm formation in or on host organism(GO:1900229) |
| 1.2 | 3.6 | GO:1903413 | cellular response to bile acid(GO:1903413) |
| 1.2 | 6.0 | GO:0044240 | multicellular organism lipid catabolic process(GO:0044240) |
| 1.2 | 3.5 | GO:0018969 | thiocyanate metabolic process(GO:0018969) |
| 1.2 | 3.5 | GO:0051795 | positive regulation of catagen(GO:0051795) |
| 1.1 | 3.3 | GO:0001798 | positive regulation of type IIa hypersensitivity(GO:0001798) positive regulation of type II hypersensitivity(GO:0002894) |
| 1.0 | 2.1 | GO:0003129 | heart induction(GO:0003129) negative regulation of Wnt signaling pathway involved in heart development(GO:0003308) regulation of heart induction(GO:0090381) regulation of heart induction by canonical Wnt signaling pathway(GO:0100012) |
| 1.0 | 3.0 | GO:2000547 | regulation of dendritic cell dendrite assembly(GO:2000547) |
| 1.0 | 2.0 | GO:0071288 | cellular response to mercury ion(GO:0071288) |
| 1.0 | 3.9 | GO:0060723 | spongiotrophoblast cell proliferation(GO:0060720) regulation of spongiotrophoblast cell proliferation(GO:0060721) cell proliferation involved in embryonic placenta development(GO:0060722) regulation of cell proliferation involved in embryonic placenta development(GO:0060723) |
| 1.0 | 4.8 | GO:0035962 | response to interleukin-13(GO:0035962) |
| 0.9 | 22.7 | GO:0006703 | estrogen biosynthetic process(GO:0006703) |
| 0.9 | 3.7 | GO:0010193 | response to ozone(GO:0010193) |
| 0.9 | 2.8 | GO:0061394 | regulation of transcription from RNA polymerase II promoter in response to arsenic-containing substance(GO:0061394) |
| 0.9 | 2.8 | GO:1903225 | negative regulation of endodermal cell differentiation(GO:1903225) |
| 0.9 | 3.6 | GO:0070295 | renal water absorption(GO:0070295) |
| 0.8 | 0.8 | GO:0003051 | angiotensin-mediated drinking behavior(GO:0003051) |
| 0.8 | 2.4 | GO:0036023 | limb joint morphogenesis(GO:0036022) embryonic skeletal limb joint morphogenesis(GO:0036023) |
| 0.8 | 2.3 | GO:0060279 | positive regulation of ovulation(GO:0060279) |
| 0.8 | 5.3 | GO:0070942 | neutrophil mediated cytotoxicity(GO:0070942) neutrophil mediated killing of symbiont cell(GO:0070943) neutrophil mediated killing of bacterium(GO:0070944) |
| 0.7 | 5.2 | GO:0019805 | quinolinate biosynthetic process(GO:0019805) |
| 0.7 | 2.2 | GO:2000625 | regulation of miRNA catabolic process(GO:2000625) positive regulation of miRNA catabolic process(GO:2000627) |
| 0.7 | 2.2 | GO:2000224 | sesquiterpenoid metabolic process(GO:0006714) sesquiterpenoid catabolic process(GO:0016107) farnesol metabolic process(GO:0016487) farnesol catabolic process(GO:0016488) regulation of testosterone biosynthetic process(GO:2000224) |
| 0.7 | 5.7 | GO:0072672 | neutrophil extravasation(GO:0072672) regulation of neutrophil extravasation(GO:2000389) |
| 0.7 | 2.1 | GO:0033861 | negative regulation of NAD(P)H oxidase activity(GO:0033861) |
| 0.7 | 3.5 | GO:0018262 | isopeptide cross-linking via N6-(L-isoglutamyl)-L-lysine(GO:0018153) isopeptide cross-linking(GO:0018262) |
| 0.7 | 2.7 | GO:0097029 | mature conventional dendritic cell differentiation(GO:0097029) |
| 0.7 | 6.7 | GO:0035093 | spermatogenesis, exchange of chromosomal proteins(GO:0035093) |
| 0.7 | 2.0 | GO:0061033 | secretion by lung epithelial cell involved in lung growth(GO:0061033) |
| 0.7 | 2.0 | GO:0003218 | cardiac left ventricle formation(GO:0003218) |
| 0.6 | 10.4 | GO:0043587 | tongue morphogenesis(GO:0043587) |
| 0.6 | 42.1 | GO:0048247 | lymphocyte chemotaxis(GO:0048247) |
| 0.6 | 16.6 | GO:0051798 | positive regulation of hair follicle development(GO:0051798) |
| 0.6 | 15.7 | GO:0015669 | gas transport(GO:0015669) oxygen transport(GO:0015671) |
| 0.6 | 1.9 | GO:1902299 | pre-replicative complex assembly involved in nuclear cell cycle DNA replication(GO:0006267) pre-replicative complex assembly(GO:0036388) pre-replicative complex assembly involved in cell cycle DNA replication(GO:1902299) |
| 0.6 | 1.2 | GO:0002874 | regulation of chronic inflammatory response to antigenic stimulus(GO:0002874) |
| 0.6 | 6.6 | GO:1904469 | positive regulation of tumor necrosis factor secretion(GO:1904469) |
| 0.6 | 2.3 | GO:0051066 | dihydrobiopterin metabolic process(GO:0051066) |
| 0.6 | 2.3 | GO:0099640 | axo-dendritic protein transport(GO:0099640) |
| 0.6 | 3.5 | GO:0045903 | positive regulation of translational fidelity(GO:0045903) |
| 0.6 | 5.7 | GO:0030263 | apoptotic chromosome condensation(GO:0030263) |
| 0.6 | 4.5 | GO:0098532 | histone H3-K27 trimethylation(GO:0098532) |
| 0.6 | 1.7 | GO:0042489 | negative regulation of odontogenesis of dentin-containing tooth(GO:0042489) |
| 0.6 | 3.3 | GO:1900042 | positive regulation of interleukin-2 secretion(GO:1900042) |
| 0.5 | 6.0 | GO:0038028 | insulin receptor signaling pathway via phosphatidylinositol 3-kinase(GO:0038028) positive regulation of glycogen (starch) synthase activity(GO:2000467) |
| 0.5 | 3.7 | GO:0003278 | apoptotic process involved in heart morphogenesis(GO:0003278) |
| 0.5 | 1.5 | GO:0045726 | positive regulation of integrin biosynthetic process(GO:0045726) |
| 0.5 | 3.0 | GO:0090131 | mesenchyme migration(GO:0090131) |
| 0.5 | 8.9 | GO:0002523 | leukocyte migration involved in inflammatory response(GO:0002523) |
| 0.5 | 4.8 | GO:0046813 | receptor-mediated virion attachment to host cell(GO:0046813) |
| 0.5 | 2.4 | GO:0035948 | positive regulation of gluconeogenesis by positive regulation of transcription from RNA polymerase II promoter(GO:0035948) |
| 0.5 | 2.9 | GO:1904222 | regulation of CDP-diacylglycerol-serine O-phosphatidyltransferase activity(GO:1904217) positive regulation of CDP-diacylglycerol-serine O-phosphatidyltransferase activity(GO:1904219) positive regulation of serine C-palmitoyltransferase activity(GO:1904222) |
| 0.5 | 4.3 | GO:0055064 | chloride ion homeostasis(GO:0055064) |
| 0.5 | 1.8 | GO:2000510 | positive regulation of dendritic cell chemotaxis(GO:2000510) |
| 0.4 | 1.8 | GO:1900738 | positive regulation of phospholipase C-activating G-protein coupled receptor signaling pathway(GO:1900738) |
| 0.4 | 1.8 | GO:0060455 | negative regulation of gastric acid secretion(GO:0060455) |
| 0.4 | 3.0 | GO:1902896 | terminal web assembly(GO:1902896) |
| 0.4 | 5.5 | GO:0070327 | thyroid hormone transport(GO:0070327) |
| 0.4 | 9.5 | GO:0090023 | positive regulation of neutrophil chemotaxis(GO:0090023) positive regulation of neutrophil migration(GO:1902624) |
| 0.4 | 2.9 | GO:0061737 | leukotriene signaling pathway(GO:0061737) |
| 0.4 | 9.2 | GO:0007175 | negative regulation of epidermal growth factor-activated receptor activity(GO:0007175) |
| 0.4 | 1.6 | GO:0018277 | protein deamination(GO:0018277) |
| 0.4 | 1.2 | GO:0034164 | negative regulation of toll-like receptor 9 signaling pathway(GO:0034164) |
| 0.4 | 1.2 | GO:0001207 | histone displacement(GO:0001207) positive regulation of transcription involved in meiotic cell cycle(GO:0051039) |
| 0.4 | 2.6 | GO:0030241 | skeletal muscle myosin thick filament assembly(GO:0030241) |
| 0.4 | 6.8 | GO:0002003 | angiotensin maturation(GO:0002003) |
| 0.3 | 4.9 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 0.3 | 12.2 | GO:0071294 | cellular response to zinc ion(GO:0071294) |
| 0.3 | 4.8 | GO:0010727 | negative regulation of hydrogen peroxide metabolic process(GO:0010727) |
| 0.3 | 1.3 | GO:0044027 | hypermethylation of CpG island(GO:0044027) |
| 0.3 | 2.0 | GO:0006001 | fructose catabolic process(GO:0006001) fructose catabolic process to hydroxyacetone phosphate and glyceraldehyde-3-phosphate(GO:0061624) |
| 0.3 | 0.7 | GO:0070434 | positive regulation of nucleotide-binding oligomerization domain containing signaling pathway(GO:0070426) positive regulation of nucleotide-binding oligomerization domain containing 2 signaling pathway(GO:0070434) |
| 0.3 | 1.9 | GO:0042421 | norepinephrine biosynthetic process(GO:0042421) |
| 0.3 | 1.3 | GO:0018101 | protein citrullination(GO:0018101) histone citrullination(GO:0036414) |
| 0.3 | 1.5 | GO:1900005 | positive regulation of serine-type endopeptidase activity(GO:1900005) positive regulation of serine-type peptidase activity(GO:1902573) |
| 0.3 | 3.6 | GO:2000623 | regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000622) negative regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000623) |
| 0.3 | 1.8 | GO:0071638 | negative regulation of monocyte chemotactic protein-1 production(GO:0071638) |
| 0.3 | 4.5 | GO:0048672 | positive regulation of collateral sprouting(GO:0048672) |
| 0.3 | 2.4 | GO:2000860 | positive regulation of mineralocorticoid secretion(GO:2000857) positive regulation of aldosterone secretion(GO:2000860) |
| 0.3 | 8.5 | GO:0030277 | maintenance of gastrointestinal epithelium(GO:0030277) |
| 0.3 | 1.2 | GO:0008050 | female courtship behavior(GO:0008050) |
| 0.3 | 8.2 | GO:0070233 | negative regulation of T cell apoptotic process(GO:0070233) |
| 0.3 | 6.1 | GO:1904706 | negative regulation of vascular smooth muscle cell proliferation(GO:1904706) |
| 0.3 | 1.4 | GO:0035993 | subthalamic nucleus development(GO:0021763) deltoid tuberosity development(GO:0035993) prolactin secreting cell differentiation(GO:0060127) superior vena cava morphogenesis(GO:0060578) cell proliferation involved in outflow tract morphogenesis(GO:0061325) |
| 0.3 | 1.4 | GO:0003430 | growth plate cartilage chondrocyte growth(GO:0003430) Harderian gland development(GO:0070384) |
| 0.3 | 1.1 | GO:0006566 | threonine metabolic process(GO:0006566) |
| 0.3 | 0.8 | GO:0060734 | regulation of endoplasmic reticulum stress-induced eIF2 alpha phosphorylation(GO:0060734) |
| 0.3 | 0.8 | GO:1903991 | positive regulation of iron ion transport(GO:0034758) positive regulation of iron ion transmembrane transport(GO:0034761) regulation of iron ion import(GO:1900390) regulation of ferrous iron import into cell(GO:1903989) positive regulation of ferrous iron import into cell(GO:1903991) regulation of ferrous iron binding(GO:1904432) positive regulation of ferrous iron binding(GO:1904434) regulation of transferrin receptor binding(GO:1904435) positive regulation of transferrin receptor binding(GO:1904437) regulation of ferrous iron import across plasma membrane(GO:1904438) positive regulation of ferrous iron import across plasma membrane(GO:1904440) |
| 0.3 | 1.5 | GO:0044800 | fusion of virus membrane with host plasma membrane(GO:0019064) membrane fusion involved in viral entry into host cell(GO:0039663) multi-organism membrane fusion(GO:0044800) |
| 0.3 | 0.5 | GO:0030718 | germ-line stem cell population maintenance(GO:0030718) |
| 0.2 | 0.9 | GO:0021524 | visceral motor neuron differentiation(GO:0021524) |
| 0.2 | 2.0 | GO:0042436 | tryptophan catabolic process(GO:0006569) indole-containing compound catabolic process(GO:0042436) indolalkylamine catabolic process(GO:0046218) |
| 0.2 | 1.8 | GO:1902527 | positive regulation of protein monoubiquitination(GO:1902527) |
| 0.2 | 0.7 | GO:0030474 | spindle pole body duplication(GO:0030474) spindle pole body organization(GO:0051300) spindle pole body localization(GO:0070631) establishment of spindle pole body localization(GO:0070632) spindle pole body localization to nuclear envelope(GO:0071789) establishment of spindle pole body localization to nuclear envelope(GO:0071790) |
| 0.2 | 0.7 | GO:0021986 | epithalamus development(GO:0021538) habenula development(GO:0021986) |
| 0.2 | 3.3 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 0.2 | 0.6 | GO:0002541 | activation of plasma proteins involved in acute inflammatory response(GO:0002541) |
| 0.2 | 3.4 | GO:0032930 | positive regulation of superoxide anion generation(GO:0032930) |
| 0.2 | 0.6 | GO:2000110 | negative regulation of macrophage apoptotic process(GO:2000110) |
| 0.2 | 1.6 | GO:0014807 | regulation of somitogenesis(GO:0014807) |
| 0.2 | 0.6 | GO:0045083 | negative regulation of interleukin-12 biosynthetic process(GO:0045083) |
| 0.2 | 1.6 | GO:1905232 | cellular response to L-glutamate(GO:1905232) |
| 0.2 | 6.0 | GO:0019731 | antibacterial humoral response(GO:0019731) |
| 0.2 | 2.1 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 0.2 | 0.8 | GO:0010430 | fatty acid omega-oxidation(GO:0010430) |
| 0.2 | 1.5 | GO:0006642 | triglyceride mobilization(GO:0006642) |
| 0.2 | 2.0 | GO:0050957 | equilibrioception(GO:0050957) |
| 0.2 | 2.5 | GO:1900121 | negative regulation of receptor binding(GO:1900121) |
| 0.2 | 1.8 | GO:0034465 | response to carbon monoxide(GO:0034465) |
| 0.2 | 0.5 | GO:1904798 | regulation of core promoter binding(GO:1904796) positive regulation of core promoter binding(GO:1904798) |
| 0.2 | 28.4 | GO:0006958 | complement activation, classical pathway(GO:0006958) |
| 0.2 | 2.6 | GO:0006069 | ethanol oxidation(GO:0006069) |
| 0.2 | 0.8 | GO:0038030 | non-canonical Wnt signaling pathway via MAPK cascade(GO:0038030) non-canonical Wnt signaling pathway via JNK cascade(GO:0038031) |
| 0.2 | 7.0 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 0.2 | 1.2 | GO:0038172 | interleukin-33-mediated signaling pathway(GO:0038172) |
| 0.2 | 2.8 | GO:0015939 | pantothenate metabolic process(GO:0015939) |
| 0.2 | 0.9 | GO:1900133 | regulation of renin secretion into blood stream(GO:1900133) |
| 0.2 | 2.9 | GO:0043031 | negative regulation of macrophage activation(GO:0043031) |
| 0.2 | 1.1 | GO:0035879 | plasma membrane lactate transport(GO:0035879) |
| 0.1 | 2.2 | GO:2000271 | positive regulation of fibroblast apoptotic process(GO:2000271) |
| 0.1 | 1.4 | GO:0061000 | negative regulation of dendritic spine development(GO:0061000) |
| 0.1 | 11.4 | GO:0001895 | retina homeostasis(GO:0001895) |
| 0.1 | 1.1 | GO:0048251 | elastic fiber assembly(GO:0048251) |
| 0.1 | 0.5 | GO:0032667 | interleukin-23 production(GO:0032627) regulation of interleukin-23 production(GO:0032667) positive regulation of interleukin-23 production(GO:0032747) |
| 0.1 | 0.8 | GO:1903378 | positive regulation of oxidative stress-induced neuron intrinsic apoptotic signaling pathway(GO:1903378) |
| 0.1 | 0.8 | GO:0001878 | response to yeast(GO:0001878) |
| 0.1 | 0.8 | GO:0043988 | histone H3-S28 phosphorylation(GO:0043988) |
| 0.1 | 0.5 | GO:0002215 | defense response to nematode(GO:0002215) |
| 0.1 | 3.2 | GO:0034587 | piRNA metabolic process(GO:0034587) |
| 0.1 | 1.9 | GO:0045945 | positive regulation of transcription from RNA polymerase III promoter(GO:0045945) cellular response to lithium ion(GO:0071285) |
| 0.1 | 2.1 | GO:0070389 | chaperone cofactor-dependent protein refolding(GO:0070389) |
| 0.1 | 2.5 | GO:0003417 | growth plate cartilage development(GO:0003417) |
| 0.1 | 3.7 | GO:0001502 | cartilage condensation(GO:0001502) |
| 0.1 | 0.6 | GO:2000969 | positive regulation of alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionate selective glutamate receptor activity(GO:2000969) |
| 0.1 | 7.2 | GO:0006953 | acute-phase response(GO:0006953) |
| 0.1 | 0.7 | GO:0060686 | regulation of prostatic bud formation(GO:0060685) negative regulation of prostatic bud formation(GO:0060686) |
| 0.1 | 0.5 | GO:0044210 | 'de novo' CTP biosynthetic process(GO:0044210) |
| 0.1 | 0.9 | GO:0071169 | establishment of protein localization to chromatin(GO:0071169) |
| 0.1 | 0.3 | GO:0021526 | medial motor column neuron differentiation(GO:0021526) |
| 0.1 | 0.4 | GO:0044314 | protein K27-linked ubiquitination(GO:0044314) |
| 0.1 | 3.3 | GO:0030728 | ovulation(GO:0030728) |
| 0.1 | 4.5 | GO:0050873 | brown fat cell differentiation(GO:0050873) |
| 0.1 | 2.9 | GO:0030199 | collagen fibril organization(GO:0030199) |
| 0.1 | 0.8 | GO:0060699 | regulation of endoribonuclease activity(GO:0060699) |
| 0.1 | 0.7 | GO:0046125 | thymidine metabolic process(GO:0046104) pyrimidine deoxyribonucleoside metabolic process(GO:0046125) |
| 0.1 | 15.2 | GO:0070268 | cornification(GO:0070268) |
| 0.1 | 8.9 | GO:0030574 | collagen catabolic process(GO:0030574) |
| 0.1 | 1.3 | GO:0050774 | negative regulation of dendrite morphogenesis(GO:0050774) |
| 0.1 | 0.8 | GO:0070934 | CRD-mediated mRNA stabilization(GO:0070934) |
| 0.1 | 1.4 | GO:1902101 | positive regulation of mitotic metaphase/anaphase transition(GO:0045842) positive regulation of mitotic sister chromatid separation(GO:1901970) positive regulation of metaphase/anaphase transition of cell cycle(GO:1902101) |
| 0.1 | 3.2 | GO:0010667 | negative regulation of cardiac muscle cell apoptotic process(GO:0010667) |
| 0.1 | 0.2 | GO:0035106 | operant conditioning(GO:0035106) |
| 0.1 | 8.0 | GO:0006614 | SRP-dependent cotranslational protein targeting to membrane(GO:0006614) |
| 0.1 | 0.9 | GO:0014067 | negative regulation of phosphatidylinositol 3-kinase signaling(GO:0014067) |
| 0.1 | 1.5 | GO:0090331 | negative regulation of platelet aggregation(GO:0090331) |
| 0.1 | 1.8 | GO:1900745 | positive regulation of p38MAPK cascade(GO:1900745) |
| 0.1 | 7.8 | GO:0007585 | respiratory gaseous exchange(GO:0007585) |
| 0.1 | 3.1 | GO:0016339 | calcium-dependent cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0016339) |
| 0.1 | 0.9 | GO:0031284 | positive regulation of guanylate cyclase activity(GO:0031284) |
| 0.1 | 1.5 | GO:0030903 | notochord development(GO:0030903) |
| 0.1 | 1.1 | GO:0006776 | vitamin A metabolic process(GO:0006776) |
| 0.1 | 2.0 | GO:0046597 | negative regulation of viral entry into host cell(GO:0046597) |
| 0.1 | 1.2 | GO:0006032 | chitin metabolic process(GO:0006030) chitin catabolic process(GO:0006032) |
| 0.1 | 0.7 | GO:1904948 | midbrain dopaminergic neuron differentiation(GO:1904948) |
| 0.1 | 8.6 | GO:0002181 | cytoplasmic translation(GO:0002181) |
| 0.1 | 2.5 | GO:0019730 | antimicrobial humoral response(GO:0019730) |
| 0.1 | 4.1 | GO:0030212 | hyaluronan metabolic process(GO:0030212) |
| 0.1 | 0.7 | GO:0006686 | sphingomyelin biosynthetic process(GO:0006686) |
| 0.1 | 0.8 | GO:0043568 | positive regulation of insulin-like growth factor receptor signaling pathway(GO:0043568) |
| 0.1 | 1.0 | GO:0035414 | negative regulation of catenin import into nucleus(GO:0035414) |
| 0.1 | 0.4 | GO:0060356 | leucine import(GO:0060356) |
| 0.1 | 2.3 | GO:0030261 | chromosome condensation(GO:0030261) |
| 0.1 | 3.8 | GO:0042246 | tissue regeneration(GO:0042246) |
| 0.0 | 0.4 | GO:0098912 | membrane depolarization during atrial cardiac muscle cell action potential(GO:0098912) |
| 0.0 | 0.1 | GO:0048611 | ectodermal digestive tract development(GO:0007439) embryonic ectodermal digestive tract development(GO:0048611) |
| 0.0 | 1.3 | GO:0051290 | protein heterotetramerization(GO:0051290) |
| 0.0 | 0.4 | GO:0097646 | calcitonin family receptor signaling pathway(GO:0097646) amylin receptor signaling pathway(GO:0097647) |
| 0.0 | 5.3 | GO:0006342 | chromatin silencing(GO:0006342) |
| 0.0 | 0.3 | GO:0006335 | DNA replication-dependent nucleosome assembly(GO:0006335) DNA replication-dependent nucleosome organization(GO:0034723) |
| 0.0 | 1.1 | GO:0008210 | estrogen metabolic process(GO:0008210) |
| 0.0 | 1.0 | GO:0045671 | negative regulation of osteoclast differentiation(GO:0045671) |
| 0.0 | 3.7 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.0 | 0.5 | GO:0019079 | viral genome replication(GO:0019079) |
| 0.0 | 0.3 | GO:0097577 | intracellular sequestering of iron ion(GO:0006880) sequestering of iron ion(GO:0097577) |
| 0.0 | 0.5 | GO:0030238 | male sex determination(GO:0030238) Leydig cell differentiation(GO:0033327) |
| 0.0 | 4.8 | GO:0060333 | interferon-gamma-mediated signaling pathway(GO:0060333) |
| 0.0 | 5.6 | GO:0007586 | digestion(GO:0007586) |
| 0.0 | 0.9 | GO:0042755 | eating behavior(GO:0042755) |
| 0.0 | 2.6 | GO:0009250 | glycogen biosynthetic process(GO:0005978) glucan biosynthetic process(GO:0009250) |
| 0.0 | 7.4 | GO:0001678 | cellular glucose homeostasis(GO:0001678) |
| 0.0 | 0.9 | GO:0045943 | positive regulation of transcription from RNA polymerase I promoter(GO:0045943) |
| 0.0 | 0.3 | GO:0035562 | negative regulation of chromatin binding(GO:0035562) snRNA modification(GO:0040031) |
| 0.0 | 1.6 | GO:0007339 | binding of sperm to zona pellucida(GO:0007339) |
| 0.0 | 0.5 | GO:0046784 | viral mRNA export from host cell nucleus(GO:0046784) |
| 0.0 | 0.9 | GO:0071364 | cellular response to epidermal growth factor stimulus(GO:0071364) |
| 0.0 | 0.1 | GO:0042756 | drinking behavior(GO:0042756) |
| 0.0 | 0.6 | GO:0071786 | endoplasmic reticulum tubular network organization(GO:0071786) |
| 0.0 | 0.3 | GO:1904936 | cerebral cortex GABAergic interneuron migration(GO:0021853) interneuron migration(GO:1904936) |
| 0.0 | 1.2 | GO:1901222 | regulation of NIK/NF-kappaB signaling(GO:1901222) |
| 0.0 | 0.7 | GO:0034694 | response to prostaglandin(GO:0034694) |
| 0.0 | 0.4 | GO:0031069 | hair follicle morphogenesis(GO:0031069) epidermis morphogenesis(GO:0048730) |
| 0.0 | 1.8 | GO:0033260 | nuclear DNA replication(GO:0033260) |
| 0.0 | 0.7 | GO:0016032 | viral process(GO:0016032) multi-organism cellular process(GO:0044764) |
| 0.0 | 4.2 | GO:0010811 | positive regulation of cell-substrate adhesion(GO:0010811) |
| 0.0 | 0.8 | GO:0050974 | detection of mechanical stimulus involved in sensory perception(GO:0050974) |
| 0.0 | 1.8 | GO:0042633 | molting cycle(GO:0042303) hair cycle(GO:0042633) |
| 0.0 | 5.5 | GO:0007160 | cell-matrix adhesion(GO:0007160) |
| 0.0 | 0.2 | GO:0035036 | sperm-egg recognition(GO:0035036) |
| 0.0 | 1.4 | GO:0000381 | regulation of alternative mRNA splicing, via spliceosome(GO:0000381) |
| 0.0 | 2.3 | GO:0001837 | epithelial to mesenchymal transition(GO:0001837) |
| 0.0 | 0.5 | GO:2000272 | negative regulation of receptor activity(GO:2000272) |
| 0.0 | 0.1 | GO:0070213 | protein auto-ADP-ribosylation(GO:0070213) |
| 0.0 | 0.3 | GO:0046069 | cGMP catabolic process(GO:0046069) |
| 0.0 | 0.3 | GO:0035855 | megakaryocyte development(GO:0035855) |
| 0.0 | 0.7 | GO:0051896 | regulation of protein kinase B signaling(GO:0051896) |
| 0.0 | 1.1 | GO:0006081 | cellular aldehyde metabolic process(GO:0006081) |
| 0.0 | 0.4 | GO:0036152 | phosphatidylethanolamine acyl-chain remodeling(GO:0036152) |
| 0.0 | 0.5 | GO:0010468 | regulation of gene expression(GO:0010468) |
| 0.0 | 1.4 | GO:0043154 | negative regulation of cysteine-type endopeptidase activity involved in apoptotic process(GO:0043154) |
| 0.0 | 0.8 | GO:0032781 | positive regulation of ATPase activity(GO:0032781) |
| 0.0 | 0.5 | GO:0007257 | activation of JUN kinase activity(GO:0007257) |
| 0.0 | 1.2 | GO:0042795 | snRNA transcription(GO:0009301) snRNA transcription from RNA polymerase II promoter(GO:0042795) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.0 | 7.8 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 1.9 | 5.7 | GO:0009330 | DNA topoisomerase complex (ATP-hydrolyzing)(GO:0009330) |
| 1.9 | 5.6 | GO:0005584 | collagen type I trimer(GO:0005584) |
| 1.9 | 5.6 | GO:0044299 | C-fiber(GO:0044299) |
| 1.2 | 7.2 | GO:1990742 | microvesicle(GO:1990742) |
| 1.1 | 15.7 | GO:0005833 | hemoglobin complex(GO:0005833) |
| 0.9 | 2.8 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.9 | 25.5 | GO:0071682 | endocytic vesicle lumen(GO:0071682) |
| 0.8 | 2.4 | GO:0070703 | inner mucus layer(GO:0070702) outer mucus layer(GO:0070703) |
| 0.8 | 2.3 | GO:0043512 | inhibin complex(GO:0043511) inhibin A complex(GO:0043512) |
| 0.7 | 7.0 | GO:0016012 | sarcoglycan complex(GO:0016012) |
| 0.6 | 1.9 | GO:0036387 | nuclear pre-replicative complex(GO:0005656) pre-replicative complex(GO:0036387) |
| 0.5 | 8.2 | GO:0000788 | nuclear nucleosome(GO:0000788) |
| 0.5 | 3.0 | GO:0030485 | smooth muscle contractile fiber(GO:0030485) |
| 0.5 | 7.6 | GO:0005641 | nuclear envelope lumen(GO:0005641) |
| 0.5 | 2.8 | GO:1990037 | Lewy body core(GO:1990037) |
| 0.4 | 5.9 | GO:0098647 | collagen type VI trimer(GO:0005589) collagen beaded filament(GO:0098647) |
| 0.4 | 3.2 | GO:0070876 | SOSS complex(GO:0070876) |
| 0.4 | 3.0 | GO:1990357 | terminal web(GO:1990357) |
| 0.3 | 36.7 | GO:0035580 | specific granule lumen(GO:0035580) |
| 0.3 | 10.0 | GO:0042599 | lamellar body(GO:0042599) |
| 0.3 | 15.3 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.3 | 5.8 | GO:0042589 | zymogen granule membrane(GO:0042589) |
| 0.3 | 2.5 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 0.3 | 3.1 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.3 | 0.8 | GO:0070195 | growth hormone receptor complex(GO:0070195) |
| 0.3 | 2.9 | GO:0005610 | laminin-5 complex(GO:0005610) |
| 0.3 | 15.7 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.2 | 0.7 | GO:0070762 | nuclear pore transmembrane ring(GO:0070762) |
| 0.2 | 18.7 | GO:0045095 | keratin filament(GO:0045095) |
| 0.2 | 1.4 | GO:0005971 | ribonucleoside-diphosphate reductase complex(GO:0005971) |
| 0.2 | 3.7 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.2 | 20.5 | GO:0035578 | azurophil granule lumen(GO:0035578) |
| 0.2 | 9.3 | GO:0000786 | nucleosome(GO:0000786) |
| 0.1 | 17.2 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.1 | 1.9 | GO:0042584 | chromaffin granule membrane(GO:0042584) |
| 0.1 | 0.7 | GO:0043196 | varicosity(GO:0043196) |
| 0.1 | 14.5 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.1 | 4.2 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.1 | 10.4 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.1 | 3.6 | GO:1990124 | messenger ribonucleoprotein complex(GO:1990124) |
| 0.1 | 24.4 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.1 | 1.4 | GO:0045179 | apical cortex(GO:0045179) |
| 0.1 | 0.8 | GO:0032133 | chromosome passenger complex(GO:0032133) |
| 0.1 | 53.2 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.1 | 3.4 | GO:0031093 | platelet alpha granule lumen(GO:0031093) |
| 0.1 | 5.5 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.1 | 1.5 | GO:0071438 | invadopodium membrane(GO:0071438) |
| 0.1 | 138.0 | GO:0005615 | extracellular space(GO:0005615) |
| 0.1 | 0.8 | GO:0070937 | CRD-mediated mRNA stability complex(GO:0070937) |
| 0.1 | 0.5 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 0.1 | 0.8 | GO:0097136 | Bcl-2 family protein complex(GO:0097136) |
| 0.1 | 2.0 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.1 | 2.1 | GO:0005921 | gap junction(GO:0005921) |
| 0.1 | 0.5 | GO:0005853 | eukaryotic translation elongation factor 1 complex(GO:0005853) |
| 0.1 | 0.7 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 0.0 | 1.2 | GO:0098563 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.0 | 0.3 | GO:0043204 | perikaryon(GO:0043204) |
| 0.0 | 0.8 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.0 | 0.3 | GO:0043527 | tRNA methyltransferase complex(GO:0043527) |
| 0.0 | 1.7 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.0 | 1.8 | GO:0035577 | azurophil granule membrane(GO:0035577) |
| 0.0 | 0.9 | GO:0005686 | U2 snRNP(GO:0005686) |
| 0.0 | 0.8 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.0 | 0.7 | GO:0043220 | Schmidt-Lanterman incisure(GO:0043220) |
| 0.0 | 2.0 | GO:0032420 | stereocilium(GO:0032420) |
| 0.0 | 1.1 | GO:0031011 | Ino80 complex(GO:0031011) DNA helicase complex(GO:0033202) |
| 0.0 | 0.4 | GO:0097550 | transcriptional preinitiation complex(GO:0097550) |
| 0.0 | 0.7 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.0 | 3.0 | GO:0030864 | cortical actin cytoskeleton(GO:0030864) |
| 0.0 | 0.2 | GO:0036128 | CatSper complex(GO:0036128) |
| 0.0 | 1.6 | GO:0043198 | dendritic shaft(GO:0043198) |
| 0.0 | 0.4 | GO:1990454 | L-type voltage-gated calcium channel complex(GO:1990454) |
| 0.0 | 0.3 | GO:0005796 | Golgi lumen(GO:0005796) |
| 0.0 | 1.8 | GO:0030286 | dynein complex(GO:0030286) |
| 0.0 | 0.1 | GO:1990745 | EARP complex(GO:1990745) |
| 0.0 | 0.5 | GO:0000346 | transcription export complex(GO:0000346) |
| 0.0 | 0.4 | GO:0097225 | sperm midpiece(GO:0097225) |
| 0.0 | 0.6 | GO:0001669 | acrosomal vesicle(GO:0001669) |
| 0.0 | 1.8 | GO:0005901 | caveola(GO:0005901) |
| 0.0 | 1.8 | GO:0042470 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.0 | 0.7 | GO:0030666 | endocytic vesicle membrane(GO:0030666) |
| 0.0 | 1.6 | GO:0001750 | photoreceptor outer segment(GO:0001750) |
| 0.0 | 10.8 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.0 | 3.9 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.1 | 24.4 | GO:0047006 | 17-alpha,20-alpha-dihydroxypregn-4-en-3-one dehydrogenase activity(GO:0047006) |
| 2.6 | 18.5 | GO:0031726 | CCR1 chemokine receptor binding(GO:0031726) |
| 2.4 | 7.2 | GO:0031731 | CCR6 chemokine receptor binding(GO:0031731) |
| 2.3 | 11.3 | GO:0033265 | choline binding(GO:0033265) |
| 2.1 | 16.6 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 1.8 | 5.3 | GO:0047150 | betaine-homocysteine S-methyltransferase activity(GO:0047150) |
| 1.4 | 5.7 | GO:0003918 | DNA topoisomerase type II (ATP-hydrolyzing) activity(GO:0003918) DNA topoisomerase II activity(GO:0061505) |
| 1.3 | 5.2 | GO:0070653 | high-density lipoprotein particle receptor binding(GO:0070653) |
| 1.3 | 7.8 | GO:0030492 | hemoglobin binding(GO:0030492) |
| 1.3 | 11.4 | GO:0001849 | complement component C1q binding(GO:0001849) |
| 1.2 | 6.1 | GO:0034875 | oxidoreductase activity, acting on CH or CH2 groups, quinone or similar compound as acceptor(GO:0033695) caffeine oxidase activity(GO:0034875) |
| 1.2 | 3.5 | GO:0036393 | thiocyanate peroxidase activity(GO:0036393) |
| 1.2 | 5.8 | GO:0031728 | CCR3 chemokine receptor binding(GO:0031728) |
| 1.1 | 5.6 | GO:0031727 | CCR2 chemokine receptor binding(GO:0031727) |
| 0.8 | 3.9 | GO:0004995 | tachykinin receptor activity(GO:0004995) |
| 0.8 | 15.7 | GO:0005344 | oxygen transporter activity(GO:0005344) |
| 0.8 | 3.0 | GO:0031735 | CCR10 chemokine receptor binding(GO:0031735) |
| 0.7 | 2.2 | GO:0047017 | geranylgeranyl reductase activity(GO:0045550) prostaglandin-F synthase activity(GO:0047017) androsterone dehydrogenase activity(GO:0047023) indanol dehydrogenase activity(GO:0047718) |
| 0.7 | 3.6 | GO:0015254 | glycerol channel activity(GO:0015254) |
| 0.7 | 10.3 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.7 | 2.0 | GO:0015168 | glycerol transmembrane transporter activity(GO:0015168) |
| 0.6 | 2.3 | GO:0008426 | protein kinase C inhibitor activity(GO:0008426) |
| 0.5 | 28.4 | GO:0042056 | chemoattractant activity(GO:0042056) |
| 0.5 | 4.1 | GO:0060228 | phosphatidylcholine-sterol O-acyltransferase activator activity(GO:0060228) |
| 0.5 | 9.5 | GO:0019864 | IgG binding(GO:0019864) |
| 0.5 | 3.5 | GO:0004966 | galanin receptor activity(GO:0004966) |
| 0.5 | 2.8 | GO:0017159 | pantetheine hydrolase activity(GO:0017159) |
| 0.4 | 3.5 | GO:1990932 | 5.8S rRNA binding(GO:1990932) |
| 0.4 | 7.1 | GO:0033691 | sialic acid binding(GO:0033691) |
| 0.4 | 2.5 | GO:0045569 | TRAIL binding(GO:0045569) |
| 0.4 | 8.5 | GO:0008009 | chemokine activity(GO:0008009) |
| 0.4 | 2.3 | GO:0070699 | type II activin receptor binding(GO:0070699) |
| 0.4 | 3.8 | GO:0015250 | water channel activity(GO:0015250) |
| 0.4 | 3.3 | GO:0004024 | alcohol dehydrogenase activity, zinc-dependent(GO:0004024) |
| 0.4 | 5.6 | GO:0048407 | platelet-derived growth factor binding(GO:0048407) |
| 0.3 | 9.0 | GO:0019887 | protein kinase regulator activity(GO:0019887) |
| 0.3 | 5.4 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.3 | 5.6 | GO:0019841 | retinol binding(GO:0019841) |
| 0.3 | 0.8 | GO:0005148 | prolactin receptor binding(GO:0005148) |
| 0.3 | 3.7 | GO:0032395 | MHC class II receptor activity(GO:0032395) |
| 0.3 | 2.3 | GO:0015349 | thyroid hormone transmembrane transporter activity(GO:0015349) |
| 0.3 | 1.5 | GO:0030171 | voltage-gated proton channel activity(GO:0030171) |
| 0.3 | 1.3 | GO:0004668 | protein-arginine deiminase activity(GO:0004668) |
| 0.2 | 1.6 | GO:0008048 | calcium sensitive guanylate cyclase activator activity(GO:0008048) |
| 0.2 | 2.3 | GO:0019238 | cyclohydrolase activity(GO:0019238) |
| 0.2 | 3.0 | GO:0035251 | UDP-glucosyltransferase activity(GO:0035251) |
| 0.2 | 0.7 | GO:0035033 | histone deacetylase regulator activity(GO:0035033) |
| 0.2 | 1.9 | GO:0016715 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced ascorbate as one donor, and incorporation of one atom of oxygen(GO:0016715) |
| 0.2 | 2.4 | GO:0047498 | calcium-dependent phospholipase A2 activity(GO:0047498) |
| 0.2 | 1.2 | GO:0032564 | adenyl deoxyribonucleotide binding(GO:0032558) dATP binding(GO:0032564) |
| 0.2 | 3.0 | GO:0050786 | RAGE receptor binding(GO:0050786) |
| 0.2 | 1.8 | GO:0060072 | large conductance calcium-activated potassium channel activity(GO:0060072) |
| 0.2 | 1.5 | GO:0046790 | virion binding(GO:0046790) |
| 0.2 | 7.6 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.2 | 9.4 | GO:0005159 | insulin-like growth factor receptor binding(GO:0005159) |
| 0.2 | 2.1 | GO:0001730 | 2'-5'-oligoadenylate synthetase activity(GO:0001730) |
| 0.2 | 1.4 | GO:0016728 | ribonucleoside-diphosphate reductase activity, thioredoxin disulfide as acceptor(GO:0004748) oxidoreductase activity, acting on CH or CH2 groups, disulfide as acceptor(GO:0016728) ribonucleoside-diphosphate reductase activity(GO:0061731) |
| 0.2 | 1.6 | GO:0004720 | protein-lysine 6-oxidase activity(GO:0004720) |
| 0.2 | 0.5 | GO:0051787 | misfolded protein binding(GO:0051787) |
| 0.2 | 1.2 | GO:0002114 | interleukin-33 receptor activity(GO:0002114) |
| 0.2 | 2.0 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 0.2 | 2.9 | GO:0015194 | L-serine transmembrane transporter activity(GO:0015194) serine transmembrane transporter activity(GO:0022889) |
| 0.2 | 2.1 | GO:0048019 | receptor antagonist activity(GO:0048019) |
| 0.2 | 2.4 | GO:0008097 | 5S rRNA binding(GO:0008097) |
| 0.2 | 25.6 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.1 | 0.4 | GO:0045518 | interleukin-22 receptor binding(GO:0045518) |
| 0.1 | 13.1 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.1 | 1.1 | GO:0010997 | anaphase-promoting complex binding(GO:0010997) |
| 0.1 | 2.8 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.1 | 3.6 | GO:0008494 | translation activator activity(GO:0008494) |
| 0.1 | 4.4 | GO:0070628 | proteasome binding(GO:0070628) |
| 0.1 | 1.1 | GO:0030060 | L-malate dehydrogenase activity(GO:0030060) |
| 0.1 | 1.1 | GO:0004957 | prostaglandin E receptor activity(GO:0004957) |
| 0.1 | 0.5 | GO:0030550 | acetylcholine receptor inhibitor activity(GO:0030550) |
| 0.1 | 2.5 | GO:0008391 | arachidonic acid monooxygenase activity(GO:0008391) arachidonic acid epoxygenase activity(GO:0008392) |
| 0.1 | 1.4 | GO:0097157 | pre-mRNA intronic binding(GO:0097157) |
| 0.1 | 0.7 | GO:0004797 | thymidine kinase activity(GO:0004797) |
| 0.1 | 9.3 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.1 | 1.9 | GO:0043138 | 3'-5' DNA helicase activity(GO:0043138) |
| 0.1 | 0.7 | GO:0033188 | sphingomyelin synthase activity(GO:0033188) ceramide cholinephosphotransferase activity(GO:0047493) |
| 0.1 | 1.3 | GO:0051525 | NFAT protein binding(GO:0051525) |
| 0.1 | 0.5 | GO:0003883 | CTP synthase activity(GO:0003883) |
| 0.1 | 0.5 | GO:0035473 | lipase binding(GO:0035473) |
| 0.1 | 0.8 | GO:0051434 | BH3 domain binding(GO:0051434) |
| 0.1 | 34.0 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.1 | 2.2 | GO:0035613 | RNA stem-loop binding(GO:0035613) |
| 0.1 | 1.5 | GO:0046870 | cadmium ion binding(GO:0046870) |
| 0.1 | 1.0 | GO:0070097 | delta-catenin binding(GO:0070097) |
| 0.1 | 2.7 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.1 | 21.9 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.1 | 1.4 | GO:0043008 | ATP-dependent protein binding(GO:0043008) |
| 0.1 | 0.8 | GO:0035174 | histone serine kinase activity(GO:0035174) |
| 0.1 | 1.7 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.1 | 1.2 | GO:0035374 | chondroitin sulfate binding(GO:0035374) |
| 0.1 | 2.2 | GO:0015269 | calcium-activated potassium channel activity(GO:0015269) |
| 0.1 | 0.6 | GO:0031849 | olfactory receptor binding(GO:0031849) |
| 0.1 | 1.1 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.1 | 0.8 | GO:0072542 | protein phosphatase activator activity(GO:0072542) |
| 0.1 | 2.7 | GO:0001530 | lipopolysaccharide binding(GO:0001530) |
| 0.1 | 0.8 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 0.1 | 8.5 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.1 | 3.9 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.1 | 11.0 | GO:0002020 | protease binding(GO:0002020) |
| 0.1 | 1.5 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.1 | 1.4 | GO:0034450 | ubiquitin-ubiquitin ligase activity(GO:0034450) |
| 0.1 | 0.7 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 0.1 | 0.7 | GO:0055131 | C3HC4-type RING finger domain binding(GO:0055131) |
| 0.1 | 1.2 | GO:0004887 | thyroid hormone receptor activity(GO:0004887) |
| 0.1 | 4.4 | GO:0051879 | Hsp90 protein binding(GO:0051879) |
| 0.1 | 0.9 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.1 | 2.1 | GO:0046965 | retinoid X receptor binding(GO:0046965) |
| 0.1 | 1.8 | GO:0045505 | dynein intermediate chain binding(GO:0045505) |
| 0.1 | 0.5 | GO:0019958 | C-X-C chemokine binding(GO:0019958) |
| 0.1 | 6.2 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 0.1 | 1.8 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.1 | 0.2 | GO:0033754 | indoleamine 2,3-dioxygenase activity(GO:0033754) |
| 0.0 | 0.7 | GO:0042826 | histone deacetylase binding(GO:0042826) |
| 0.0 | 2.9 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.0 | 0.4 | GO:0086056 | voltage-gated calcium channel activity involved in AV node cell action potential(GO:0086056) |
| 0.0 | 6.4 | GO:0051087 | chaperone binding(GO:0051087) |
| 0.0 | 1.6 | GO:0070064 | proline-rich region binding(GO:0070064) |
| 0.0 | 17.9 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 0.0 | 0.9 | GO:0003707 | steroid hormone receptor activity(GO:0003707) |
| 0.0 | 0.8 | GO:0022833 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 0.0 | 1.7 | GO:0005035 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.0 | 0.5 | GO:1990825 | sequence-specific mRNA binding(GO:1990825) |
| 0.0 | 0.9 | GO:0048027 | mRNA 5'-UTR binding(GO:0048027) |
| 0.0 | 1.5 | GO:0005504 | fatty acid binding(GO:0005504) |
| 0.0 | 0.3 | GO:0004118 | cGMP-stimulated cyclic-nucleotide phosphodiesterase activity(GO:0004118) |
| 0.0 | 0.2 | GO:0000293 | ferric-chelate reductase activity(GO:0000293) |
| 0.0 | 2.9 | GO:0030170 | pyridoxal phosphate binding(GO:0030170) |
| 0.0 | 0.1 | GO:0032190 | acrosin binding(GO:0032190) |
| 0.0 | 0.8 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.0 | 2.2 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 0.6 | GO:0000979 | RNA polymerase II core promoter sequence-specific DNA binding(GO:0000979) |
| 0.0 | 2.0 | GO:0043425 | bHLH transcription factor binding(GO:0043425) |
| 0.0 | 1.3 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.0 | 0.5 | GO:0030957 | Tat protein binding(GO:0030957) |
| 0.0 | 1.0 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.0 | 0.1 | GO:0004461 | lactose synthase activity(GO:0004461) |
| 0.0 | 0.3 | GO:0045159 | myosin II binding(GO:0045159) |
| 0.0 | 1.9 | GO:0008138 | protein tyrosine/serine/threonine phosphatase activity(GO:0008138) |
| 0.0 | 0.9 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.0 | 3.5 | GO:0031490 | chromatin DNA binding(GO:0031490) |
| 0.0 | 0.4 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 0.0 | 1.1 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.0 | 2.8 | GO:0003823 | antigen binding(GO:0003823) |
| 0.0 | 0.8 | GO:0015175 | neutral amino acid transmembrane transporter activity(GO:0015175) |
| 0.0 | 0.7 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.0 | 0.3 | GO:0008199 | ferric iron binding(GO:0008199) |
| 0.0 | 3.2 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.1 | GO:0001083 | transcription factor activity, RNA polymerase II basal transcription factor binding(GO:0001083) RNA polymerase II transcription factor activity, TBP-class protein binding, involved in preinitiation complex assembly(GO:0001129) RNA polymerase II transcription factor activity, TBP-class protein binding(GO:0001132) |
| 0.0 | 4.6 | GO:0042393 | histone binding(GO:0042393) |
| 0.0 | 0.5 | GO:0000217 | DNA secondary structure binding(GO:0000217) |
| 0.0 | 0.3 | GO:0016423 | tRNA (guanine) methyltransferase activity(GO:0016423) |
| 0.0 | 2.0 | GO:0008201 | heparin binding(GO:0008201) |
| 0.0 | 2.0 | GO:0016811 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amides(GO:0016811) |
| 0.0 | 0.3 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.0 | 1.9 | GO:0008083 | growth factor activity(GO:0008083) |
| 0.0 | 1.5 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.0 | 0.2 | GO:0008526 | phosphatidylinositol transporter activity(GO:0008526) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 21.3 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.2 | 6.7 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.2 | 21.2 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.2 | 28.5 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.2 | 3.8 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.2 | 12.3 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.2 | 3.6 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.2 | 14.7 | PID ATF2 PATHWAY | ATF-2 transcription factor network |
| 0.2 | 2.9 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.2 | 12.5 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 0.1 | 6.7 | PID ALK1 PATHWAY | ALK1 signaling events |
| 0.1 | 7.5 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.1 | 7.8 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.1 | 16.7 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
| 0.1 | 2.8 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
| 0.1 | 7.0 | ST ADRENERGIC | Adrenergic Pathway |
| 0.1 | 7.6 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.1 | 5.5 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 7.0 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.1 | 2.5 | PID IL23 PATHWAY | IL23-mediated signaling events |
| 0.1 | 28.0 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.1 | 4.5 | PID IL6 7 PATHWAY | IL6-mediated signaling events |
| 0.1 | 2.9 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.1 | 30.4 | NABA SECRETED FACTORS | Genes encoding secreted soluble factors |
| 0.1 | 1.5 | PID A6B1 A6B4 INTEGRIN PATHWAY | a6b1 and a6b4 Integrin signaling |
| 0.1 | 2.5 | PID TRAIL PATHWAY | TRAIL signaling pathway |
| 0.1 | 5.3 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.1 | 2.3 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.1 | 1.4 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
| 0.0 | 2.8 | PID IL12 2PATHWAY | IL12-mediated signaling events |
| 0.0 | 3.1 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.0 | 0.9 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.0 | 12.0 | NABA CORE MATRISOME | Ensemble of genes encoding core extracellular matrix including ECM glycoproteins, collagens and proteoglycans |
| 0.0 | 1.2 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.0 | 2.4 | PID FCER1 PATHWAY | Fc-epsilon receptor I signaling in mast cells |
| 0.0 | 10.3 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 1.5 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.0 | 2.9 | PID CXCR4 PATHWAY | CXCR4-mediated signaling events |
| 0.0 | 0.3 | PID ERBB1 RECEPTOR PROXIMAL PATHWAY | EGF receptor (ErbB1) signaling pathway |
| 0.0 | 2.1 | PID E2F PATHWAY | E2F transcription factor network |
| 0.0 | 0.7 | PID EPHB FWD PATHWAY | EPHB forward signaling |
| 0.0 | 0.2 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.0 | 0.8 | PID BARD1 PATHWAY | BARD1 signaling events |
| 0.0 | 2.0 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.0 | 0.8 | PID BMP PATHWAY | BMP receptor signaling |
| 0.0 | 1.2 | PID PDGFRB PATHWAY | PDGFR-beta signaling pathway |
| 0.0 | 0.3 | ST G ALPHA I PATHWAY | G alpha i Pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 9.3 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.6 | 20.5 | REACTOME ADVANCED GLYCOSYLATION ENDPRODUCT RECEPTOR SIGNALING | Genes involved in Advanced glycosylation endproduct receptor signaling |
| 0.5 | 1.5 | REACTOME BINDING AND ENTRY OF HIV VIRION | Genes involved in Binding and entry of HIV virion |
| 0.5 | 8.6 | REACTOME XENOBIOTICS | Genes involved in Xenobiotics |
| 0.4 | 17.6 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.4 | 8.8 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.4 | 11.8 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.3 | 5.4 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.3 | 4.6 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.3 | 7.3 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.3 | 4.8 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.3 | 5.6 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.3 | 3.7 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 0.3 | 3.1 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.3 | 19.4 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.2 | 6.0 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.2 | 2.9 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GLP1 | Genes involved in Synthesis, Secretion, and Inactivation of Glucagon-like Peptide-1 (GLP-1) |
| 0.2 | 22.2 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.2 | 13.1 | REACTOME REGULATION OF BETA CELL DEVELOPMENT | Genes involved in Regulation of beta-cell development |
| 0.2 | 9.8 | REACTOME DEFENSINS | Genes involved in Defensins |
| 0.2 | 5.1 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.2 | 3.3 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.2 | 2.3 | REACTOME GLYCOPROTEIN HORMONES | Genes involved in Glycoprotein hormones |
| 0.1 | 2.4 | REACTOME ACYL CHAIN REMODELLING OF PS | Genes involved in Acyl chain remodelling of PS |
| 0.1 | 9.7 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.1 | 3.0 | REACTOME ACTIVATION OF THE MRNA UPON BINDING OF THE CAP BINDING COMPLEX AND EIFS AND SUBSEQUENT BINDING TO 43S | Genes involved in Activation of the mRNA upon binding of the cap-binding complex and eIFs, and subsequent binding to 43S |
| 0.1 | 3.3 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.1 | 3.3 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 0.1 | 2.3 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.1 | 2.3 | REACTOME TRANSPORT OF ORGANIC ANIONS | Genes involved in Transport of organic anions |
| 0.1 | 1.1 | REACTOME PROSTANOID LIGAND RECEPTORS | Genes involved in Prostanoid ligand receptors |
| 0.1 | 7.6 | REACTOME EXTRACELLULAR MATRIX ORGANIZATION | Genes involved in Extracellular matrix organization |
| 0.1 | 5.6 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.1 | 0.8 | REACTOME APC C CDH1 MEDIATED DEGRADATION OF CDC20 AND OTHER APC C CDH1 TARGETED PROTEINS IN LATE MITOSIS EARLY G1 | Genes involved in APC/C:Cdh1 mediated degradation of Cdc20 and other APC/C:Cdh1 targeted proteins in late mitosis/early G1 |
| 0.1 | 7.7 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.1 | 15.2 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.1 | 1.5 | REACTOME AMINE DERIVED HORMONES | Genes involved in Amine-derived hormones |
| 0.1 | 2.5 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |
| 0.1 | 1.1 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.0 | 6.8 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.0 | 4.6 | REACTOME INTERFERON GAMMA SIGNALING | Genes involved in Interferon gamma signaling |
| 0.0 | 1.4 | REACTOME PHOSPHORYLATION OF THE APC C | Genes involved in Phosphorylation of the APC/C |
| 0.0 | 1.1 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
| 0.0 | 0.6 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.0 | 0.7 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.0 | 1.0 | REACTOME FORMATION OF FIBRIN CLOT CLOTTING CASCADE | Genes involved in Formation of Fibrin Clot (Clotting Cascade) |
| 0.0 | 1.1 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.0 | 1.5 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.0 | 0.7 | REACTOME DEPOSITION OF NEW CENPA CONTAINING NUCLEOSOMES AT THE CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.0 | 0.7 | REACTOME PYRIMIDINE METABOLISM | Genes involved in Pyrimidine metabolism |
| 0.0 | 0.8 | REACTOME DOWNREGULATION OF TGF BETA RECEPTOR SIGNALING | Genes involved in Downregulation of TGF-beta receptor signaling |
| 0.0 | 0.7 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 2.2 | REACTOME G ALPHA I SIGNALLING EVENTS | Genes involved in G alpha (i) signalling events |
| 0.0 | 0.4 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.0 | 0.5 | REACTOME RNA POL II TRANSCRIPTION PRE INITIATION AND PROMOTER OPENING | Genes involved in RNA Polymerase II Transcription Pre-Initiation And Promoter Opening |
| 0.0 | 0.3 | REACTOME PLC BETA MEDIATED EVENTS | Genes involved in PLC beta mediated events |
| 0.0 | 1.0 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |